Week 07 HW: genetic circuits part-II


Traditional genetic circuits work using Boolean logic, where the output is binary (either ON or OFF). The output depends on whether the input signals pass a fixed threshold. For example, in a genetic AND gate, a protein is only produced when both transcription factors A and B are present above a certain level. If one of them is missing or below the threshold, the output is zero. This type of system is useful for simple decisions, but it has important limitations because real biological signals are usually continuous, variable, and noisy, not strictly ON or OFF.
Intracellular Artificial Neural Networks (IANNs) solve these limitations by mimicking artificial neural networks inside the cell. Instead of treating inputs as binary, IANNs assign a continuous weight to each input. These weighted inputs are then summed, and the result is passed through a biological activation mechanism (such as a riboswitch or a protease-regulated system) to generate a graded output.
IANN approach provides several important advantages:
Continuous output resolution: Unlike Boolean circuits that only produce ON or OFF outputs, IANNs generate different levels of expression depending on the strength of the inputs. This allows cells to respond in a more precise and dose-dependent way, which is important for applications like metabolic regulation or controlled therapeutic delivery.
Weighted signal integration: Each input does not contribute equally. Instead, every signal has a specific weight that determines how much it influences the final output. This allows the system to prioritize certain signals over others, which is not possible in traditional AND/OR gates where all inputs are treated equally.
Robustness to biological noise: Cellular environments are naturally noisy, and signals can vary between cells. Because IANNs work with continuous values rather than strict thresholds, they are more tolerant to noise and variability, making them more reliable in real biological conditions.
Greater computational power: A multilayer IANN can act as a universal function approximator, meaning it can represent very complex relationships between inputs and outputs. In contrast, Boolean circuits are limited to simple logical combinations, which restricts the complexity of decisions they can perform.
Rational tunability: The weights and biases in an IANN can be adjusted through DNA design (for example, by modifying promoters or regulatory elements) or improved through directed evolution. This makes it possible to “train” the system to recognize complex patterns, such as a specific combination of biomarkers, with much higher precision than traditional Boolean circuits.
Application of an IANN: Smart Lactase-Producing Probiotic System
As a highly relevant and practical application of an Intracellular Artificial Neural Network (IANN) I chose to apply it in the engineering of a probiotic bacterium capable of context-aware lactase production for the management of lactose intolerance. Unlike conventional synthetic circuits that respond to a single input in a binary manner, this system integrates multiple physiological signals from the gastrointestinal environment to produce a graded and condition-dependent enzymatic response.
-Input Layer: Multidimensional Environmental Sensing
The system incorporates multiple biologically relevant input signals, each representing a distinct physiological parameter of the gastrointestinal environment:
X1: Lactose concentration This serves as the primary input signal, directly reflecting the presence and abundance of the substrate requiring enzymatic degradation.
X2: pH level This input provides spatial context by distinguishing between different regions of the gastrointestinal tract. The acidic pH of the stomach versus the near-neutral pH of the intestine allows the system to restrict activation to physiologically appropriate locations, thereby preventing premature or energetically wasteful enzyme production.
X3: Inflammatory biomarkers Molecules such as nitric oxide, reactive oxygen species or cytokine-associated metabolites act as indicators of intestinal stress or dysbiosis. This input enables modulation of the system’s response based on host physiological state, allowing adaptive tuning of output under pathological conditions.
Lactose sensitivity can be increased using a strong promoter or high-affinity regulator, corresponding to a positive weight. pH sensitivity may be implemented through a regulatory element that suppresses output under acidic conditions, corresponding to a negative or inhibitory weight. Inflammatory signals could be integrated via modulatory promoters or regulatory RNAs that amplify output under stress conditions, acting as an adjustable positive or negative weight depending on the desired response.
At the molecular level, each input is transduced into regulatory signals (e.g., transcription factors, small RNAs, or protease-mediated regulators). These signals are then integrated through combinatorial gene regulation, where promoter strengths, ribosome binding site efficiencies, and degradation dynamics collectively encode the effective weights.
The aggregated signal undergoes a transformation through a biological activation function, which may be implemented via nonlinear regulatory elements such as riboswitches, cooperative transcriptional regulators, or proteolytic cascades. This step introduces thresholding and saturation effects analogous to activation functions in artificial neural networks, thereby enabling continuous and nonlinear input–output relationships.
Output Layer: Graded Lactase Expression
The final output of the system is the expression of the lactase enzyme, with expression levels determined by the integrated and nonlinearly transformed input signal
This enables a spectrum of responses:
- Sub-threshold activation: (e.g., low lactose concentration or inhibitory pH conditions) result in negligible or no enzyme production.
- Intermediate activation: moderate enzyme expression
- High activation: (e.g., high lactose concentration under optimal pH conditions, potentially combined with inflammatory signals) drive maximal enzyme production.
Functional Behavior and Decision-Making Capability
The system effectively implements a context-dependent decision-making process, wherein output is not determined by a single condition but by the weighted combination of multiple environmental cues. For example:
- The presence of lactose alone is insufficient to trigger activation under acidic conditions, thereby preventing inappropriate expression in the stomach.
- Under intestinal pH, lactose induces activation in a concentration-dependent manner.
- In the presence of both high lactose and inflammatory signals, the system can upregulate lactase production, potentially enhancing digestive efficiency under stress conditions.
Limitations and Practical Constraints
Despite its conceptual advantages, the implementation of such an IANN-based system faces several challenges:
- Stochastic gene expression: Intrinsic and extrinsic noise can introduce variability in circuit performance across individual cells. Parameter tuning complexity: Precise calibration of weights and activation thresholds through genetic elements (e.g., promoters, RBSs) remains experimentally demanding.
- Kinetic limitations: Transcriptional and translational processes impose temporal delays, limiting the speed of system response.
- Regulatory crosstalk: Interactions between synthetic and endogenous pathways may lead to unintended behaviors.
- Metabolic burden: The expression of complex regulatory networks can reduce host fitness and stability.
- Environmental variability: Dynamic and heterogeneous gut conditions may challenge the robustness and predictability of the system.
Implementation of a Multilayer Perceptron Using Endoribonucleases
To implement a multilayer perceptron in a biological system, the output of one computational layer must regulate the activity of the next. This can be achieved using a cascade of endoribonucleases, where each layer processes inputs and produces a regulatory molecule that serves as the input for the subsequent layer.
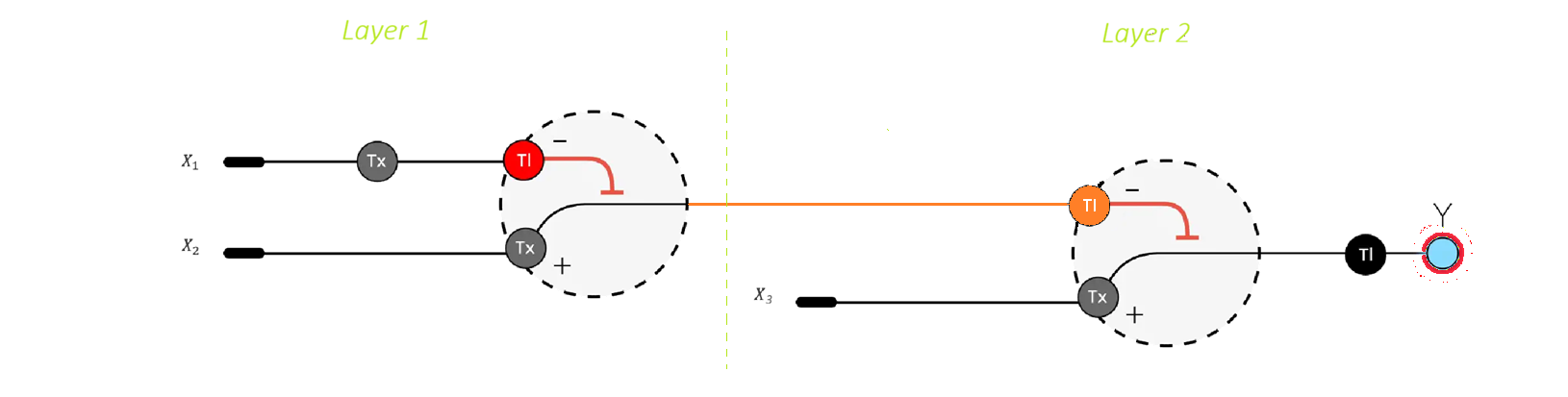

- Input Representation
The system integrates multiple biological inputs represented as molecular signals:
X1: Csy4 endoribonuclease (constitutively or inducibly expressed)
X2: an additional regulatory signal (e.g., inducible promoter or transcriptional activator)
X3: environmental or metabolic signal (e.g., pH, or inflammatory markers such as nitric oxide)
These inputs are converted into regulatory effects at the gene expression level, analogous to numerical inputs in an artificial neural network.
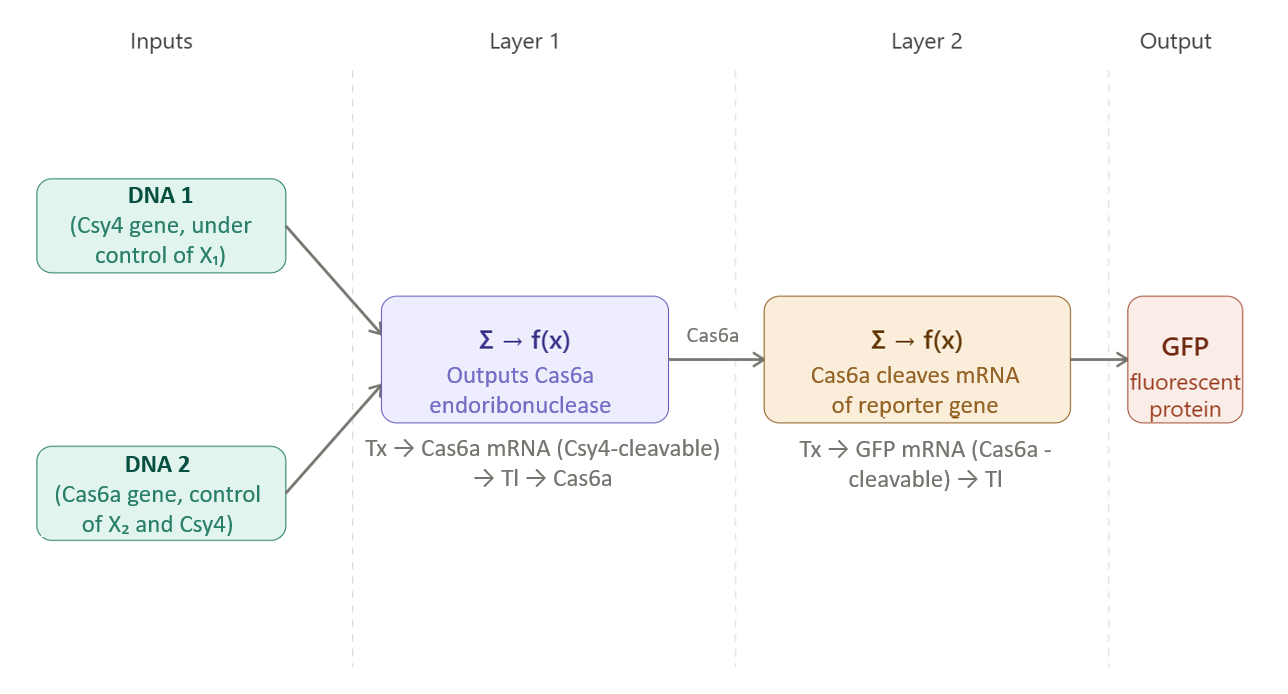

- Layer 1: Intermediate Processing In the first layer, the inputs jointly regulate the expression of an intermediate endoribonuclease (e.g., Cas6a). The mRNA encoding this enzyme is engineered to contain specific recognition sites for Csy4. As a result:
–> The presence of Csy4 (X1) induces cleavage of the mRNA, leading to repression of Cas6a expression
–> The second input (X2) can act as an activator, promoting transcription of the Cas6a gene
Thus, Layer 1 integrates activating and inhibitory signals. The resulting expression level of Cas6a reflects a balance between these opposing regulatory effects, analogous to a weighted sum followed by a nonlinear activation function in a perceptron.
- Layer 2: Output Generation The output of Layer 1 (Cas6a protein) serves as the regulatory input for Layer 2.
The mRNA encoding a reporter protein (e.g., GFP) is engineered to contain Cas6a recognition sites. Consequently:
–> High levels of Cas6a lead to cleavage of GFP mRNA and repression of fluorescence
–> Low levels of Cas6a allow GFP expression
This establishes a second computational layer in which the input is not external, but derived from the processed output of the first layer.
- System-Level Behavior
This cascading architecture enables hierarchical signal processing within the cell. –> When Csy4 levels are high, Cas6a production is suppressed, allowing GFP expression
–> When Csy4 levels are low and activation dominates, Cas6a is produced and represses GFP
Therefore, the final output depends on both the original inputs and the intermediate computation performed in Layer 1.
Fungal Materials and Their Uses
Fungi, especially their root-like networks called mycelium, are becoming a surprisingly powerful source of sustainable materials. Unlike plastics or leather, which require heavy manufacturing and chemicals, mycelium-based materials are grown. The fungi take agricultural waste—things like sawdust, rice husks, or hemp—and weave it into solid, structured materials. It’s almost like nature is doing the 3D printing for us.
One of the most familiar uses today is in packaging. Mycelium can form lightweight, shock-absorbing foams that replace Styrofoam or plastic inserts. Fragile items like electronics, furniture, or delicate goods can be safely packed in these eco-friendly alternatives. Companies like Ecovative and IKEA have already begun experimenting with this approach, showing that sustainable materials don’t have to compromise practicality.
image_ref
Fungi are also stepping into the fashion world through myco-leather. By processing mats of mycelium into flexible sheets, it’s possible to make shoes, bags, and even clothing without harming animals. Myco-leather is fully biodegradable, reduces chemical waste, and offers a much lower environmental footprint than traditional leather production. It’s a great example of how biology can meet design.
image_ref
In construction, fungal materials are finding their place as well. Mycelium boards can provide thermal and acoustic insulation, or serve as lightweight panels for ceilings and walls. They are naturally fire-resistant, resistant to pests like termites, and completely biodegradable. This means that even in building applications, mycelium offers both functionality and sustainability.
The versatility doesn’t stop there. Designers and researchers are exploring fungal foams, textiles, and even furniture, taking advantage of mycelium’s ability to grow into complex shapes. People are also experimenting with specialty applications, like wearable electronics, wound dressings, filters, and acoustic panels. Fungi aren’t just materials—they’re living factories that can be shaped, molded, and sometimes even programmed to do more.
image_ref


Advantages and Disadvantages of Fungal Materials
| Advantages | Disadvantages |
|---|---|
| Made from renewable agricultural waste and fully biodegradable | Lower mechanical strength compared to plastics, metals, or treated leather |
| “Grown” in controlled conditions with minimal energy and no toxic chemicals | Sensitive to moisture; can deform or degrade if untreated |
| Naturally fire-resistant and termite-resistant | Slower production—growing a material takes days or weeks |
| Lightweight with good strength-to-weight ratio | Batch-to-batch variability due to biological growth |
| Can return nutrients to the soil after disposal | Limited durability under extreme conditions without extra treatment |
Genetic Engineering and Synthetic Biology in Fungi
Fungi are not just fascinating organisms—they are also incredibly versatile tools for engineering. If I were to genetically engineer fungi, I would aim to enhance the properties that currently limit their use while maximizing their natural strengths. For instance, one limitation of mycelium-based materials is their mechanical strength, which can make them less competitive compared to plastics or synthetic foams. I would focus on modifying the cell wall composition or growth patterns to produce stronger, more durable materials, making fungi a realistic alternative for packaging, textiles, and construction.
Another area I would target is environmental resilience. Fungal materials are naturally biodegradable, which is a huge advantage, but they can degrade too quickly in humid or wet environments. By engineering fungi to better tolerate moisture or extreme temperatures, it would be possible to create materials that maintain their structure and functionality in a wider range of conditions, expanding their practical applications.
Beyond materials, fungi can also be engineered for functional enhancements. I would consider adding traits like pigmentation for natural coloring, antimicrobial properties to extend shelf life, or even self-healing abilities so that minor damage doesn’t ruin the product. These modifications could transform mycelium into “smart materials” that are not only sustainable but also highly functional.
Why Use Fungi Instead of Bacteria?
Fungi offer several important advantages over bacteria when it comes to synthetic biology. First, as eukaryotic organisms, they have more advanced cellular machinery. This allows them to properly fold and modify complex proteins through processes like glycosylation, which is essential for many pharmaceuticals and functional biomolecules.
Another major advantage is their filamentous growth. Many fungi grow as long branching structures (hyphae), which makes them very efficient at secreting enzymes and other products into their environment. This simplifies downstream processing because the desired product is often already outside the cell.
Fungi also have a much richer and more diverse metabolism compared to most bacteria. They naturally produce a wide range of secondary metabolites, which means they can be engineered to generate a broader variety of useful compounds, from drugs to pigments to bioactive molecules. In addition, fungi are generally more robust in industrial settings. They can grow on cheap, low-quality substrates like agricultural waste and tolerate harsher conditions than many bacteria, making them more practical for large-scale, sustainable production.
That said, working with fungi can be more complex. They tend to grow more slowly than bacteria, and genetic engineering tools are less standardized. However, despite these challenges, their unique capabilities make them extremely valuable for applications where bacteria fall short.
for this part, I have developed three potential ideas for my final project and would greatly appreciate your feedback to help refine my direction. While I am still open to suggestions, I currently find myself most aligned with my second idea, as it feels both biologically intuitive and well-matched to the techniques we have learned throughout the course.
The idea I am leaning toward is focused on engineering an E. coli reporter system to monitor protein aging during heterologous expression using a fluorescent timer protein. I am particularly drawn to this concept because it allows me to integrate multiple core synthetic biology tools, including DNA construct design, protein engineering, and computational structure prediction, while also remaining experimentally feasible within the scope of the course. In addition, the system is mechanistically clear, which makes it easier to design, test, and interpret.
I have further refined this idea into a more specific and functional design: a time-dependent protein quality control system in which a fluorescent timer regulates the exposure of a degron, leading to the selective degradation of aged proteins. In this system, a protein of interest is fused to a fluorescent timer and a C-terminal degron. As the protein matures and the timer shifts from its “young” to “old” fluorescence state, conformational or structural changes are expected to increase the accessibility of the degron. This, in turn, allows recognition by the host proteolytic machinery, enabling targeted degradation of older protein populations.
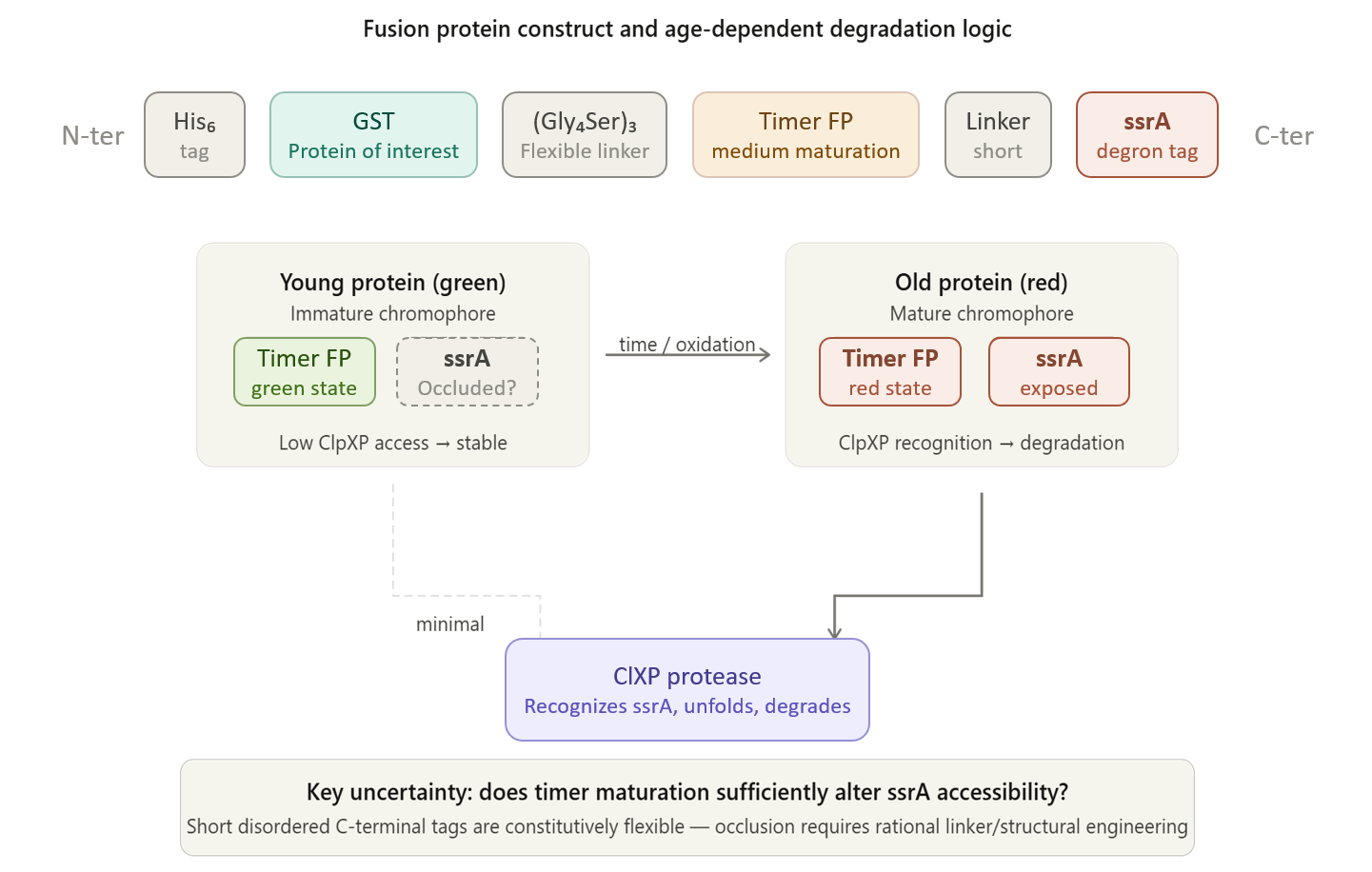
 The key modification from the original idea is the addition of a functional outcome—degradation—rather than only monitoring protein age. This transforms the system from a passive reporter into an active quality control mechanism. The purpose of this change is to address a limitation in current heterologous expression systems, where proteins can accumulate in misfolded or non-functional states over time. By selectively degrading older or potentially damaged proteins, this system could improve overall protein quality and stability.
The key modification from the original idea is the addition of a functional outcome—degradation—rather than only monitoring protein age. This transforms the system from a passive reporter into an active quality control mechanism. The purpose of this change is to address a limitation in current heterologous expression systems, where proteins can accumulate in misfolded or non-functional states over time. By selectively degrading older or potentially damaged proteins, this system could improve overall protein quality and stability.
The broader gap I am attempting to address is the lack of dynamic, time-resolved control over protein lifespan in bacterial systems. Most current approaches either measure protein expression statically or rely on constitutive degradation signals that do not account for protein age. This project introduces a strategy to link protein function, age, and degradation in a single genetically encoded system.
At this stage, I would greatly value any feedback on the conceptual design, feasibility, or potential improvements. In particular, I would appreciate input on whether the proposed mechanism for degron exposure is realistic, and whether there are alternative design strategies that could strengthen the system. Any suggestions on experimental design, protein choice, or construct optimization would also be extremely helpful.
Please feel free to share feedback through any preferred channel, including email or whatsApp. Thank you for your time and guidance.
Designing the isert sequence in Benchling:
for this idea i designed the genetic construct in Benchling that encodes a fusion protein consisting of GST as the protein of interest, followed by a flexible linker, a fluorescent timer protein, a second short linker, and a C-terminal ssrA degron whose sequences are represented in the following table:
| Genetic Element | Function | DNA Sequence (5' → 3') |
|---|---|---|
| Start Codon | Initiates translation | |
| Protein of Interest (GST - Schistosoma japonicum) | Reporter protein for studying protein aging and degradation | |
| Linker 1 ((Gly₄Ser)₃) | Flexible linker between GST and timer | |
| Fluorescent Timer (Medium FT) | Reports protein age (green to red transition) | |
| Linker 2 (GGGGS) | Provides flexibility and enables degron exposure | |
| Degron (ssrA tag) | Targets protein for degradation by ClpXP | |
| Stop Codon | Terminates translation | |
This design enables time-dependent exposure of the degron, allowing selective degradation of aged proteins by the host proteolytic system.
The designed insert will be cloned into a pET28 expression vector for protein expression in Escherichia coli BL21(DE3). This vector provides a T7 promoter, ribosome binding site, transcription terminator, and an N-terminal His₆ tag for protein purification. Therefore, only the coding sequence of the fusion protein was designed in Benchling.
the direct link to Benchling: GST_Timer_Degron_Insert
In this homework, ChatGPT helped me structure and write the answers and descriptions clearly, while Cloud AI generated the diagrams comparing Boolean genetic circuits and INNAs, the example illustrating a multilayer perceptron application, and the diagram describing my final project idea proposal.
Sources:
- Brophy, J. A. N., & Voigt, C. A. (2014). Principles of Genetic Circuit Design. Nature Methods, 11(5), 508–520. https://doi.org/10.1038/nmeth.2926 Differences Between SLP and MLP | PDF | Theoretical Computer Science | Machine Learning. (n.d.). Retrieved March 30, 2026, from https://fr.scribd.com/document/858039220/Single-Layer-Perceptron-and-Multilayer-Perceptron
- Gandia, A., van den Brandhof, J. G., Appels, F. V. W., & Jones, M. P. (2021). Flexible Fungal Materials: Shaping the Future. Trends in Biotechnology, 39(12), 1321–1331. https://doi.org/10.1016/j.tibtech.2021.03.002
- Halužan Vasle, A., & Moškon, M. (2024). Synthetic biological neural networks: From current implementations to future perspectives. BioSystems, 237, 105164. https://doi.org/10.1016/j.biosystems.2024.105164
- Hinneburg, H., Gu, S., & Naseri, G. (2025). Fungal Innovations—Advancing Sustainable Materials, Genetics, and Applications for Industry. Journal of Fungi, 11(10). https://doi.org/10.3390/jof11100721
- Lim, H. G., Jang, S., Jang, S., Seo, S. W., & Jung, G. Y. (2018). Design and optimization of genetically encoded biosensors for high-throughput screening of chemicals. Current Opinion in Biotechnology, Analytical Biotechnology, 54, 18–25. https://doi.org/10.1016/j.copbio.2018.01.011
- Mattern, D. J., Valiante, V., Unkles, S. E., & Brakhage, A. A. (2015). Synthetic biology of fungal natural products. Frontiers in Microbiology, 6, 775. https://doi.org/10.3389/fmicb.2015.00775
- Moorman, A., Samaniego, C. C., Maley, C., & Weiss, R. (2019). A Dynamical Biomolecular Neural Network. 2019 IEEE 58th Conference on Decision and Control (CDC), 1797–1802. https://doi.org/10.1109/CDC40024.2019.9030122
- Parhizi, Z., Dearnaley, J., Kauter, K., Mikkelsen, D., Pal, P., Shelley, T., & Burey, P. (Polly). (2025). The Fungus Among Us: Innovations and Applications of Mycelium-Based Composites. Journal of Fungi, 11(8), 549. https://doi.org/10.3390/jof11080549
- Seak, L. C. U., Lo, O. L. I., Suen, W. C.-W., & Wu, M.-T. (2021). Next-generation biocomputing: Mimicking artificial neural network with genetic circuits (p. 2021.03.12.435120). bioRxiv. https://doi.org/10.1101/2021.03.12.435120
- Secret fungi in everyday life | Kew. (n.d.). Retrieved March 30, 2026, from https://www.kew.org/read-and-watch/everyday-fungi-food-medicine Stock, C. H., Harvey, S. E., Ocko, S. A., & Ganguli, S. (2022). Synaptic balancing: A biologically plausible local learning rule that provably increases neural network noise robustness without sacrificing task performance. PLoS Computational Biology, 18(9), e1010418. https://doi.org/10.1371/journal.pcbi.1010418
- Undecided with Matt Ferrell. (2021, June 22). Is Mycelium Fungus the Plastic of the Future? [Video recording]. https://www.youtube.com/watch?v=cApVVuuqLFY
- van der Linden, A. J., Pieters, P. A., Bartelds, M. W., Nathalia, B. L., Yin, P., Huck, W. T. S., Kim, J., & de Greef, T. F. A. (2022). DNA Input Classification by a Riboregulator-Based Cell-Free Perceptron. ACS Synthetic Biology, 11(4), 1510–1520. https://doi.org/10.1021/acssynbio.1c00596
- Wang, X., Chen, Y.-Z., Qiu, X.-D., Chen, L., Teng, Y.-M., Ding, C., Huang, Y.-T., Wang, S.-Y., Liu, S.-Y., Ding, B., Laborda, P., & Zhu, S.-Q. (2026). Bioactivity and mechanisms of Ewingella americana for the control of Alternaria leaf spot on peanut. Physiological and Molecular Plant Pathology, 142, 103088. https://doi.org/10.1016/j.pmpp.2025.103088
- Yang, P., Condrich, A., Lu, L., Scranton, S., Hebner, C., Sheykhhasan, M., & Ali, M. A. (2024). Genetic Engineering in Bacteria, Fungi, and Oomycetes, Taking Advantage of CRISPR. DNA, 4(4), 427–454. https://doi.org/10.3390/dna4040030