Week 09 HW: Cell Free Systems


Homework Part A: General and Lecturer-Specific Questions
- Main Advantages of Cell-Free Protein Synthesis (CFPS) Over Traditional In Vivo Methods
Cell-free protein synthesis removes the constraints of using living cells. You are working in a test tube, which gives you direct control over the reaction environment without worrying about cell viability.
image ref


Flexibility and Control:
- Direct manipulation: You can easily change pH, salt concentration, redox potential, or add detergents, chaperones, or unnatural amino acids at any time. In living cells, these changes would kill the cells or fail to enter.
- No cell walls or membranes: You add DNA directly to the extract. There is no need for transformation, selection, or cell lysis steps. This saves hours or days.
- Toxic protein production: You can synthesize proteins that would kill living cells (e.g., membrane proteins, proteases, toxins).
- Speed and efficiency: Protein production takes 2–4 hours instead of days. All energy goes into making your target protein, not cell growth. Two cases where CFPS is more beneficial than cell production:
High-throughput screening of enzyme variants or genetic circuits – Because reactions are fast and can be done in 96- or 384-well plates, you can test hundreds of conditions or mutants in a single afternoon.
Production of toxic membrane proteins (e.g., GPCRs, viral ion channels) – These proteins kill E. coli or insect cells when produced in vivo. In CFPS, you can add detergents or nanodiscs directly to the reaction to keep the protein soluble and stable.
Main Components of a Cell-Free Expression System and Their Roles
A cell-free system combines cellular machinery with necessary nutrients and energy. Below are the key components and what each does.
| Component | Role in the System |
|---|---|
| Cell extract (lysate) | Derived from broken cells (e.g., E. coli, wheat germ, rabbit reticulocytes). Provides ribosomes, tRNAs, aminoacyl-tRNA synthetases, initiation/elongation/termination factors, and native enzymes needed for transcription and translation. |
| Genetic template (DNA or mRNA) | The instruction manual. DNA (plasmid or linear PCR product) is transcribed into mRNA, then translated into protein. If you add mRNA directly, translation starts immediately without transcription. |
| Amino acids | The 20 building blocks that ribosomes link together to form the protein chain. |
| Energy source (ATP, GTP) | Provides the chemical energy needed for bond formation during translation, transcription, and tRNA charging. |
| Energy regeneration system | Converts spent ADP back to ATP. Without this, the reaction stops within 10–20 minutes. Common systems include creatine phosphate/creatine kinase or phosphoenolpyruvate (PEP)/pyruvate kinase. |
| RNA polymerase (e.g., T7 RNA polymerase) | If using a DNA template with a T7 promoter, you add this enzyme separately to transcribe DNA into mRNA efficiently. |
| Buffer solution (salts and cofactors) | Maintains optimal pH (usually 7.4–8.0) and ionic conditions. Magnesium (Mg²⁺) and potassium (K⁺) concentrations are critical – too little and ribosomes fall apart, too much and they stop working. |
| RNase and protease inhibitors | Protect your mRNA and protein from degradation by native enzymes present in the cell extract. |
These components are combined either as a crude extract (fast and cheap) or a PURE system (reconstituted from purified components, cleaner but more expensive).
- Why Energy Provision Regeneration Is Critical and a Method to Ensure Continuous ATP Supply
- Why it is critical?
Cell-free systems lack the metabolic networks of living cells that continuously generate ATP. Translation consumes ATP rapidly – each peptide bond uses 2 ATP equivalents. Without regeneration, ATP drops to zero within 10–20 minutes, and protein synthesis stops. To produce protein for 2–6 hours, you need a way to keep making ATP from ADP.
Method for continuous ATP supply: Phosphoenolpyruvate (PEP) / Pyruvate Kinase system
- What you add: Phosphoenolpyruvate (PEP) and the enzyme pyruvate kinase.
- How it works: Pyruvate kinase transfers a high-energy phosphate group from PEP to ADP, regenerating ATP and producing pyruvate as a byproduct.
- Why it works well: PEP has a higher phosphate transfer potential than ATP, so the reaction favors ATP formation. It is reliable and commonly used in E. coli systems.
Alternative methods (if PEP causes problems):
If the PEP system presents limitations, other options can be used:
- Creatine phosphate / creatine kinase: Converts ADP + creatine phosphate → ATP + creatine. Very common and stable.
- Glucose / hexokinase or maltodextrin – cheaper but can cause pH drops.
- Comparison of Prokaryotic vs. Eukaryotic Cell-Free Expression Systems
Cell-free expression systems can be broadly divided into prokaryotic and eukaryotic platforms, and the choice between them mainly depends on the complexity of the target protein.
- Prokaryotic system (e.g., Escherichia coli)
These systems are typically derived from E. coli and are widely used because they are fast, cost-effective, and produce high protein yields in a short time. However, they lack the machinery needed for post-translational modifications such as glycosylation, and they often have difficulty forming correct disulfide bonds and folding complex proteins properly.
- Eukaryotic system (e.g., rabbit reticulocyte lysate, wheat germ extract)
These systems provide a more suitable environment for protein folding. They contain molecular chaperones and can support disulfide bond formation and, in some cases, post-translational modifications. However, they are generally more expensive, slower, and produce lower yields compared to prokaryotic systems.
- Choosing proteins for each system
–> For prokaryotic systems:
The general rule is to choose proteins that are simple, relatively small, and do not require post-translational modifications or complex folding. These proteins should be able to fold easily in the cytoplasm. Based on these criteria, bacterial luciferase is a suitable choice. This enzyme produces a measurable light signal, making it very useful as a reporter protein. It does not require glycosylation and can be efficiently expressed and folded in E. coli, allowing easy detection through luminescence assays.
–> For eukaryotic systems:
The selection criteria are different. Proteins are usually more complex, may contain multiple domains, require disulfide bonds, or need chaperones for correct folding. Some are also membrane proteins and need a suitable environment to function. Membrane proteins, such as G protein-coupled receptors (GPCRs), are good examples. These proteins have complex structures with multiple transmembrane domains and require proper folding machinery and membrane-like conditions. Such requirements cannot be met by prokaryotic systems, while eukaryotic systems can support their correct folding and functionality
| Feature | Prokaryotic (e.g., E. coli lysate) | Eukaryotic (e.g., Wheat germ, Rabbit reticulocyte, Insect cell) |
|---|---|---|
| Yield | High (up to 1–2 mg/mL) | Low to moderate (µg/mL range) |
| Speed | Fast – 2 to 4 hours | Slower – 4 to 12 hours |
| Cost | Low | High |
| Folding machinery | Limited chaperones; no natural membrane structures | Better chaperones; some systems contain microsomes (ER vesicles) |
| Post-translational modifications (PTMs) | None (no glycosylation, limited disulfide bonds) | Can perform glycosylation, phosphorylation, and efficient disulfide bonds (if microsomes present) |
| Best for | Simple cytoplasmic proteins, enzymes, high-throughput screening | Complex human proteins, antibodies, secreted proteins, membrane proteins requiring PTMs |
- Designing a Cell-Free Experiment to Optimize Membrane Protein Expression
Optimizing the expression of a membrane protein in a cell-free system requires careful consideration of the protein’s complexity, folding requirements, and membrane integration. Membrane proteins are challenging to produce because of their hydrophobic transmembrane domains, tendency to aggregate, and need for a membrane-like environment and proper chaperones.
- Choosing the right Expression System
The choice of a cell-free system depends on the nature and complexity of the membrane protein:
Prokaryotic system (e.g., Escherichia coli): Suitable for simpler membrane proteins with few transmembrane domains that do not require complex folding or post-translational modifications. Advantages include fast expression, high yield, and low cost. However, proper folding must be supported using membrane mimics such as liposomes, nanodiscs, or mild detergents.
Eukaryotic system (e.g., rabbit reticulocyte lysate, wheat germ extract): Preferable for complex membrane proteins with multiple transmembrane domains or disulfide bonds. These systems contain molecular chaperones and provide a more natural folding environment, reducing aggregation and increasing the chance of functional protein production. Limitations include higher cost, slower expression, and lower yields.
Providing a Membrane-Like Environment
Membrane proteins require an environment that mimics a lipid bilayer. In both prokaryotic and eukaryotic systems, this can be achieved by:
- Adding liposomes or nanodiscs
- Using mild detergents carefully optimized to prevent aggregation
This ensures proper insertion of the protein into a membrane-like environment, which is critical for correct folding and functionality.
- Optimizing Folding and Expression
To further improve expression and functionality:
- Add chaperones if the protein is prone to misfolding
- Adjust reaction conditions such as temperature, Mg²⁺ concentration, and DNA template concentration
- Use a continuous ATP regeneration system (e.g., PEP/pyruvate kinase) to sustain protein synthesis
- Employ a Continuous Exchange Cell-Free (CECF) setup to extend reaction time up to 24 hours. This setup constantly provides fresh energy (ATP/GTP) and removes inhibitory byproducts, which significantly improves protein yield and folding efficiency
Challenges and how to address them:
| Challenge | Why it happens | Solution |
|---|---|---|
| Protein aggregation | Membrane proteins are hydrophobic and clump together in water. | Add liposomes or nanodiscs from the start. Test different detergents (0.1–1% DDM, Brij-35, or LMNG). |
| Low yield | Detergents can inhibit ribosomes. | Titrate detergent concentration – start low, increase until protein is soluble but yield remains acceptable. |
| Ribosome stalling | The hydrophobic nascent chain sticks to the ribosome exit tunnel. | Optimize the N-terminal sequence. Use a fusion tag like Mistic (from Bacillus subtilis) that helps membrane proteins fold. |
| No activity (misfolding) | Protein inserted incorrectly or in wrong lipid environment. | Test different lipid compositions (e.g., POPC, POPG, or E. coli polar lipids). Add chaperones (GroEL/GroES). |
| Short reaction time | Energy runs out or inhibitors accumulate. | Use CECF (dialysis) format. Double the energy regeneration components. |
Optimization checklist:
- Titrate magnesium (8–16 mM) – critical for ribosome function.
- Test temperatures (20°C, 25°C, 30°C, 37°C).
- Try 2–3 different detergents or lipid preparations.
- Run a small-scale (10 µL) screening reaction before scaling up.
- Low Yield of Target Protein – Three Possible Reasons and Troubleshooting
If your cell-free reaction produces very low yield protein, check these common issues:
Reason 1: Low quality of DNA template
The DNA may contain inhibitors (salts, ethanol, phenol, agarose) or be degraded by nucleases. Without a good template, no mRNA is made.
- Troubleshooting:
✅ Purify DNA using a spin column kit (not just alcohol precipitation).
✅ Avoid using DNA cut from agarose gels – re-extract if necessary.
✅ Check DNA concentration and run an agarose gel to see if it is intact.
✅ Use 10–20 µg of plasmid or 5–10 µg of linear PCR product per 1 mL reaction.
Reason 2: Codon bias (rare codons in the target gene)
If your gene contains many codons that are rare in the host (e.g., human gene expressed in E. coli extract), ribosomes stall or terminate early. This produces truncated or no protein.
- Troubleshooting:
✅ Re-synthesize the gene with codons optimized for your extract (E. coli or wheat germ). Many online tools and services do this.
✅ Use an extract from a strain that supplies extra rare tRNAs (e.g., E. coli Rosetta or BL21 CodonPlus).
✅ Switch to a PURE system, which is less sensitive to codon bias.
Reason 3: Rapid energy depletion
ATP runs out after 30–60 minutes because the energy regeneration system is weak or missing. The reaction stops while plenty of template and amino acids remain.
- Troubleshooting:
✅ Switch to a Continuous Exchange Cell-Free (CECF) format (dialysis membrane or two-chamber system). This constantly supplies fresh energy and removes waste.
✅ Increase the concentration of your energy regeneration components (e.g., double creatine phosphate from 50 mM to 100 mM).
✅ Use a more efficient energy source: PEP/pyruvate kinase or a maltodextrin-based system.
✅ Check the pH after the reaction – if it dropped below 7.0, your energy system may be producing acid. Switch to creatine phosphate (less pH drop).
Additional common reasons (if the above don’t help):
- Protein aggregation: Lower temperature to 20–25°C. Add 0.5% detergent or 1 mM DTT.
- RNase contamination: Use nuclease-free tubes, add RNase inhibitor (e.g., murine RNase inhibitor at 1 U/µL), and wear gloves.
- Wrong magnesium concentration: Test a range from 8 to 16 mM Mg²⁺. Too low and ribosomes dissociate; too high and they lock up.
- Pick a function and describe it.
a. What would your synthetic cell do? What is the input and what is the output?
My synthetic minimal cell (SMC) is a “killer biosensor” that detects the presence of Staphylococcus aureus and responds by producing and secreting lysostaphin, a specific anti-staphylococcal enzyme.
- Input: AIP-1 (autoinducing peptide-1), a quorum sensing molecule secreted by S. aureus (Group I strains) when it reaches high cell density.
- Output: Lysostaphin (27 kDa zinc metalloprotease from Simulans staphylolyticus), which specifically cleaves the pentaglycine cross-bridges in the S. aureus cell wall, causing bacterial lysis.
Overall function: The SMC acts as a sentinel that detects S. aureus quorum signaling and releases a targeted killer, preventing infection, biofilm formation, and the spread of antibiotic-resistant strains.
b. Could this function be realized by cell-free Tx/Tl alone, without encapsulation?
No. Without encapsulation, the cell-free reaction would produce lysostaphin immediately and continuously, regardless of whether AIP-1 is present. The SMC would release its output constitutively, wasting the enzyme and providing no sensing function. Encapsulation creates a barrier that allows the system to wait for the input signal before producing the output. Additionally, without a membrane: The membrane-bound receptor AgrC could not be properly inserted and oriented and Lysostaphin would diffuse away uncontrollably instead of being released only after detecting S. aureus.
c. Could this function be realized by genetically modified natural cell?
Yes, in principle, but with significant drawbacks compared to a synthetic minimal cell (SMC). Natural GMOs can grow, divide, and potentially spread in the environment, and they may transfer genes to other bacteria through horizontal gene transfer. They can also mutate over time and lose their function, and the produced antibacterial molecule (e.g., lysostaphin) might harm the host cell itself. In contrast, SMCs do not replicate, cannot transfer genes, and do not evolve, making them safer and more stable. Additionally, their activity is more controlled, since the toxic compound is produced only when needed and released outward, which makes SMCs more suitable for applications such as medical treatments or topical use.
d. Describe the desired outcome of your synthetic cell operation.
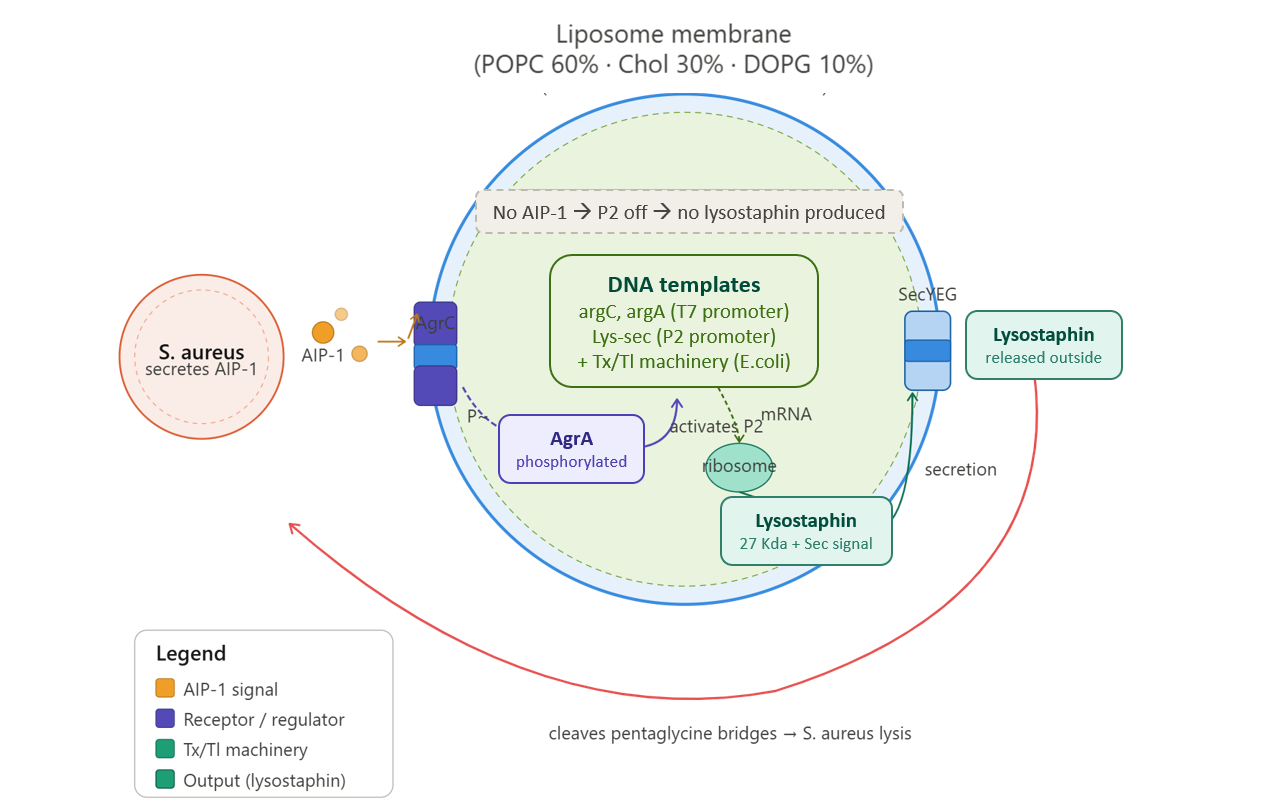
In the presence of S. aureus (which secretes AIP-1), the synthetic cell detects AIP-1 via the membrane-bound AgrC receptor. This triggers a phosphorylation cascade that activates AgrA, which then binds the P2 promoter and drives transcription of the lysostaphin gene. Lysostaphin is produced inside the vesicle and secreted into the environment. The released lysostaphin specifically cleaves the pentaglycine bridges in the S. aureus cell wall, causing bacterial lysis and death.
In the absence of AIP-1 (no S. aureus), the synthetic cell remains inactive. The P2 promoter is “off” (no leak), and no lysostaphin is produced. This ensures the toxin is only made when and where it is needed.
- Design all components that would need to be part of your synthetic cell.
a. What would the membrane be made of?
The membrane needs to be stable but also allow the AgrC histidine kinase (a transmembrane protein) to insert properly. A suitable choice is liposomes composed of:
- POPC (1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine) – 60 mol% → Main structural lipid of the membrane
- Cholesterol – 30 mol% → Increases membrane stability and reduces leakage
- DOPG (1,2-dioleoyl-sn-glycero-3-phospho-(1’-rac-glycerol)) – 10 mol% → Adds negative charge, which helps the insertion and function of membrane proteins like AgrC
b. What would you encapsulate inside? Enzymes, small molecules.
Inside the synthetic cell, i would encapsulate the basic components needed for protein production and function. First, a cell-free transcription–translation system from Escherichia coli is included, which contains all the machinery such as ribosomes, tRNAs, enzymes, and T7 RNA polymerase to make proteins.
I also add the DNA templates: agrC and agrA genes (from Staphylococcus aureus) under a constitutive promoter to sense the signal and activate the response, and the lysostaphin (lys) gene (from Staphylococcus simulans) under the P2 promoter to produce the antibacterial protein. A secretion signal is fused to the lys gene so the protein can be exported outside the cell.
In addition, small molecules like:
- ATP, GTP, CTP, UTP (nucleotide triphosphates for transcription)
- 20 amino acids (building blocks for protein synthesis)
- Creatine phosphate + creatine kinase (energy regeneration system)
- Magnesium acetate (10–14 mM) – critical for ribosome function
- Potassium glutamate (100–150 mM) – maintains ionic strength
- DTT (1–2 mM) – maintains reducing environment
- RNase inhibitors – protect mRNA from degradation
c. Which organism will your Tx/Tl system come from?
The Tx/Tl system will come from a bacterial source, specifically an Escherichia coli extract. This is because the AgrC/AgrA system is naturally bacterial and works well in an E. coli cell-free system, where AgrC can insert into liposomes properly. In addition, lysostaphin is a bacterial enzyme that does not require complex modifications, so it can be produced efficiently in this system. Finally, using a bacterial extract is simpler, faster, and cheaper than using a mammalian system, which is not needed in this case.
d. How will your synthetic cell communicate with the environment?
This synthetic cell communicates with its environment in a simple and efficient way using natural bacterial mechanisms:
For input, the signaling molecule AIP-1 does not need to enter the cell; instead, it binds directly to the external part of AgrC, a membrane protein embedded in the liposome. This means the sensor is already on the surface, so no channels are needed.
For output, lysostaphin (a relatively large protein, about 27 kDa) cannot pass through the membrane by diffusion. To solve this, a secretion signal peptide is added to lysostaphin, which directs it to the membrane during its synthesis. The protein is then transported across the membrane through the SecYEG translocon, a natural protein channel present in the Escherichia coli extract. This allows the protein to be released outside the synthetic cell in a controlled and efficient way, without needing artificial pores.
- Experimental details
a. List all lipids and genes (specific names).
- Lipids:
| Lipid | Full name | mol% |
|---|---|---|
| POPC | 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine | 60% |
| Cholesterol | Cholesterol | 30% |
| DOPG | 1,2-dioleoyl-sn-glycero-3-phospho-(1'-rac-glycerol) | 10% |
- Genes:
| Gene | Source | Promoter | Function |
|---|---|---|---|
| agrC | S. aureus (GI, for example: strain RN6390) | Constitutive (T7) | Membrane histidine kinase that binds AIP-1 on the extracellular side |
| agrA | S. aureus (same strain) | Constitutive (T7) | Response regulator; when phosphorylated by AgrC, activates P2 promoter |
| lys (lysostaphin) Fused to sec-secretion signal | Simulans staphylolyticus | P2 promoter (from S. aureus agr operon) | Zinc metalloprotease that kills S. aureus, directed to be secreted across the membrane via the sec-secretion signal |
Cell-free Tx/Tl system: All machinery for transcription and translation from E. coli extract (ribosomes, tRNAs, aminoacyl-tRNA synthetases, initiation/elongation/termination factors, T7 RNA polymerase).
b. How will you measure the function of your system?
- Measurement 1: AIP-1 sensing (dose–response)
The synthetic cells can be exposed to different concentrations of AIP-1. After a few hours, lysostaphin production is measured using methods like ELISA, Western blot, or an enzyme activity assay. If the system works properly, higher AIP-1 levels should lead to higher lysostaphin production.
- Measurement 2: Lysostaphin production (fluorescent reporter)
The lysostaphin gene can be replaced with GFP (green fluorescent protein) under the same promoter. The synthetic cells can then be monitored over time using a plate reader to measure fluorescence. Higher fluorescence indicates stronger gene expression.
- Measurement 3: Killing of Staphylococcus aureus (functional assay)
The synthetic cells can be incubated with live bacteria in culture medium. After several hours, bacterial growth can be measured using OD600, colony counting (CFU), or live/dead staining. Reduced growth shows that the system is effective.
- Measurement 4: Secretion efficiency
The synthetic cells can be centrifuged to separate them from the surrounding liquid. Lysostaphin activity is then measured both in the supernatant (outside) and inside the cells. A good system will show most of the protein in the supernatant.
- Measurement 5: Promoter leakiness (control test)
The synthetic cells can be tested without adding AIP-1 to check background expression. Ideally, very little lysostaphin should be produced. If significant production is observed, the promoter may be leaky and require optimization.

- One-sentence pitch
A wall paint containing synthetic minimal cells that detect toxic mold signals in damp walls and produce enzymes to neutralize mycotoxins and inhibit mold growth.
- How will the idea work?
The paint is embedded with microcapsules containing freeze-dried synthetic minimal cells (SMCs). When the wall becomes damp, the SMCs are activated by chemical signals released by mold, such as those from Stachybotrys chartarum. Once triggered, the SMCs produce enzymes or antimicrobial proteins that either degrade mycotoxins or prevent further mold growth. This creates a self-protecting coating that actively reduces mold and mycotoxin levels in real-time, improving indoor air safety.
- What societal challenge or market need does this address?
Toxic wall moisture is a serious indoor environmental problem. Persistent dampness encourages growth of black mold, which releases mycotoxins harmful to human health, causing respiratory issues, chronic fatigue, and neurological problems. Current paints only act as passive barriers and do not remove toxins. This smart paint provides active protection, reducing health risks and the need for costly remediation.
- How will you address limitations of cell-free systems?
The SMCs are freeze-dried within protective microcapsules, remaining inactive until moisture activates them. Microcapsules shield the system during storage and paint application. Activation only occurs when mold is present, ensuring efficient use. The one-time-use limitation is addressed by applying fresh paint layers during regular maintenance, keeping the wall continuously protected.
- One-sentence summary pitc
We will use a freeze-dried cell-free system to test how microgravity affects protein production using a GFP reporter, providing insight into reduced collagen synthesis in space.
- How the idea works
Freeze-dried cell-free reactions containing a GFP reporter gene will be prepared in sealed chambers. In space, they will be rehydrated and incubated under microgravity conditions. GFP fluorescence will act as a direct indicator of protein synthesis efficiency. By comparing fluorescence levels between space and Earth conditions, we can determine whether microgravity directly affects the molecular machinery responsible for producing proteins such as collagen.
- Societal challenge / market need
Long-duration space missions lead to bone loss and tissue weakening in astronauts, partly due to reduced production of structural proteins like collagen. Understanding whether this reduction is caused by fundamental limits in protein synthesis will help develop countermeasures for bone loss, injury prevention, and tissue regeneration, improving astronaut health during missions to Mars and beyond.
- Limitation of cell-free reactions and how to address them
Cell-free reactions are single-use and require activation by water. To overcome this, we will freeze-dry the reactions in sealed chambers, ensuring long-term stability. The experiment will be activated by rehydration in space, allowing controlled and efficient protein production measurements under microgravity conditions.
- Molecular or genetic target
Green Fluorescent Protein (GFP) gene used as a reporter to measure protein synthesis efficiency linked to collagen-related biological processes.
- How the target relates to the space biology challenge
Collagen is essential for maintaining bone and tissue structure, but its production decreases in microgravity. Instead of directly expressing collagen, which is complex, GFP is used as a reporter to measure overall protein synthesis efficiency. If microgravity reduces GFP production, it suggests that the basic machinery needed to produce proteins like collagen is affected. This helps determine whether tissue weakening in space is caused by direct physical effects on protein production or by cellular regulation, providing clearer insight into astronaut health challenges.
- Hypothesis or research goal
We hypothesize that microgravity reduces protein synthesis efficiency, which contributes to decreased production of structural proteins such as collagen in astronauts. The goal is to measure GFP production in a cell-free system under microgravity and Earth conditions. Since cell-free systems isolate transcription and translation from cellular signaling, any observed decrease in GFP fluorescence would indicate that physical factors—such as altered diffusion, molecular interactions, or protein folding—directly impact protein synthesis. This would suggest that microgravity imposes fundamental constraints on biological processes, helping explain tissue weakening. The results could guide the development of targeted countermeasures to maintain astronaut health during long-duration missions.
- Experimental plan
Freeze-dried BioBits® reactions containing GFP DNA will be used. Samples include: (1) microgravity test reactions, (2) Earth-based positive controls, and (3) negative controls without DNA. Reactions will be rehydrated and incubated using the miniPCR®. GFP fluorescence will be measured with the P51 Molecular Fluorescence Viewer. Fluorescence intensity will be compared between conditions to determine whether microgravity reduces protein synthesis efficiency.
- For this homework, I used DeepSeek and Google as sources of information. ChatGPT was used to improve the structure and clarity of the writing, while Cloud AI was used to generate the illustration of the synthetic minimal cell function.
Sources:
- A Comparative Guide: Prokaryotic vs. Eukaryotic Cell-Free Expression Systems for Eukaryotic Proteins—CD Biosynsis. (n.d.). Retrieved April 5, 2026, from https://www.biosynsis.com/a-comparative-guide-prokaryotic-vs-eukaryotic-cell-free-expression-systems-for-eukaryotic-proteins.html
- Advantages of Cell-Free Protein Expression. (n.d.). Retrieved April 5, 2026, from https://www.biocompare.com/Editorial-Articles/594727-Advantages-of-Cell-Free-Protein-Expression/
- Biology Goal. (2023, November 29). Autoinducing Peptides (AIP) in Staphylococcus Quorum Sensing [Video recording]. https://www.youtube.com/watch?v=nXXVLU7Dh-w
- Black Mold: Health Risks, Costs & Identification Guide. (n.d.). Mold Busters. Retrieved April 5, 2026, from https://www.bustmold.com/resources/about-mold/types-of-mold/black-mold/
- Brookwell, A., Oza, J. P., & Caschera, F. (2021). Biotechnology Applications of Cell-Free Expression Systems. Life, 11(12), 1367. https://doi.org/10.3390/life11121367
- Cell-Free Expression Support—Troubleshooting—DZ. (n.d.). Retrieved April 5, 2026, from https://www.thermofisher.com/ng/en/home/technical-resources/technical-reference-library/protein-expression-support-center/cell-free-expression-support/cell-free-expression-support-troubleshooting.html
- Cell-Free Protein Expression—DZ. (n.d.). Retrieved April 5, 2026, from https://www.thermofisher.com/ng/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/pierce-protein-methods/cell-free-protein-expression.html
- Cell-Free Systems for Protein Production: Advantages Over Living Cells. (n.d.). Cytion. Retrieved April 5, 2026, from https://www.cytion.com/ca/About-Cytion/Knowledge-Hub/Blog/Cell-Free-Systems-for-Protein-Production-Advantages-Over-Living-Cells/
- Chong, S. (2014). Overview of Cell-Free Protein Synthesis: Historic Landmarks, Commercial Systems, and Expanding Applications. Current Protocols in Molecular Biology / Edited by Frederick M. Ausubel … [et Al.], 108, 16.30.1-16.30.11. https://doi.org/10.1002/0471142727.mb1630s108
- Contact us via LiveChat! (n.d.). Retrieved April 5, 2026, from https://secure.livechatinc.com/customer/action/open_chat?license_id=7496091&group=0&embedded=1&widget_version=3&unique_groups=0&organization_id=eef41f83-49a8-4805-8843-da337d811599&use_parent_storage=1&x-region=us-south1
- Don’t Ignore Mould In The Home | EnviroVent. (n.d.). Retrieved April 5, 2026, from https://www.envirovent.com/blog/dont-ignore-mould-in-the-home/ Jiang, S., Çelen, G., Glatter, T., Niederholtmeyer, H., & Yuan, J. (2024). A cell-free system for functional studies of small membrane proteins. The Journal of Biological Chemistry, 300(11), 107850. https://doi.org/10.1016/j.jbc.2024.107850
- Jiang, Y., Geng, M., & Bai, L. (2020). Targeting Biofilms Therapy: Current Research Strategies and Development Hurdles. Microorganisms, 8(8), 1222. https://doi.org/10.3390/microorganisms8081222
- Khambhati, K., Bhattacharjee, G., Gohil, N., Braddick, D., Kulkarni, V., & Singh, V. (2019). Exploring the Potential of Cell-Free Protein Synthesis for Extending the Abilities of Biological Systems. Frontiers in Bioengineering and Biotechnology, 7, 248. https://doi.org/10.3389/fbioe.2019.00248
- Lysostaphin—An overview | ScienceDirect Topics. (n.d.). Retrieved April 5, 2026, from https://www.sciencedirect.com/topics/neuroscience/lysostaphin Meyer, C., Arizzi, A., Henson, T., Aviran, S., Longo, M. L., Wang, A., & Tan, C. (2025). Designer artificial environments for membrane protein synthesis. Nature Communications, 16(1), 4363. https://doi.org/10.1038/s41467-025-59471-1
- NASA OSDR: Open Science for Life in Space. (n.d.). Retrieved April 6, 2026, from https://osdr.nasa.gov/bio/repo/search?q=&data_source=cgene,alsda,esa&data_type=study
- Optimization of Energy Regeneration in Cell-free Protein Synthesis. (n.d.). PatSnap Eureka. Retrieved April 5, 2026, from https://eureka.patsnap.com/report-optimization-of-energy-regeneration-in-cell-free-protein-synthesis
- Seki, E., Matsuda, N., Yokoyama, S., & Kigawa, T. (2008). Cell-free protein synthesis system from Escherichia coli cells cultured at decreased temperatures improves productivity by decreasing DNA template degradation. Analytical Biochemistry, 377(2), 156–161. https://doi.org/10.1016/j.ab.2008.03.001
- Sitaraman, K., Esposito, D., Klarmann, G., Le Grice, S. F., Hartley, J. L., & Chatterjee, D. K. (2004). A novel cell-free protein synthesis system. Journal of Biotechnology, 110(3), 257–263. https://doi.org/10.1016/j.jbiotec.2004.02.014
- Szaflarski, W., & Nierhaus, K. H. (2007). Question 7: Optimized Energy Consumption for Protein Synthesis. Origins of Life and Evolution of Biospheres, 37(4), 423–428. https://doi.org/10.1007/s11084-007-9091-4
- Troubleshooting Guide for NEBExpressTM Cell-free E. coli Protein Synthesis System (NEB #E5360) | NEB. (n.d.). Retrieved April 5, 2026, from https://www.neb.com/en/tools-and-resources/troubleshooting-guides/troubleshooting-guide-for-nebexpress-cell-free-e-coli-protein-synthesis-system-neb-e5360
- Williams, P., Hill, P., Bonev, B., & Chan, W. C. (2023). Quorum-sensing, intra- and inter-species competition in the staphylococci. Microbiology, 169(8), 001381. https://doi.org/10.1099/mic.0.001381
- Wu, J. A., Kusuma, C., Mond, J. J., & Kokai-Kun, J. F. (2003). Lysostaphin disrupts Staphylococcus aureus and Staphylococcus epidermidis biofilms on artificial surfaces. Antimicrobial Agents and Chemotherapy, 47(11), 3407–3414. https://doi.org/10.1128/AAC.47.11.3407-3414.2003

