Week 11 HW: Bioproduction & Cloud Labs
Contribution to the Collective Bioart Project


I contributed to several designs during the experiment. My final contribution was trying to create a geometric pattern inspired by Islamic geometric art in the bottom-right corner of the pixel canvas. The design did not stay until the end because other participants kept modifying it, but it was interesting to see how the artwork kept changing with everyone’s input.

 What I Liked About the Project
What I Liked About the Project
I really liked the collaborative aspect of this project. It was fun to work with others at the same time, contribute to different designs, and watch them change in real time. The canvas was dynamic and creative, and it encouraged experimentation and shared participation.
Suggestions for Improvement
One improvement could be to limit each participant to only one or two pixels. This would encourage more collaboration, because people would need to work together to create designs instead of working alone on bigger parts. It could make the final artwork more coordinated and truly collaborative.
- Component Roles (20-Hour NMP–Ribose–Glucose System)
- E. coli Lysate
BL21 (DE3) Star Lysate: Provides the core cellular machinery required for gene expression, including ribosomes, tRNAs, aminoacyl-tRNA synthetases, and metabolic enzymes. The BL21 (DE3) strain also supplies T7 RNA polymerase, enabling strong transcription from T7 promoters.
- Salts / Buffer
Potassium Glutamate: Maintains proper ionic strength and mimics the natural intracellular environment, helping stabilize proteins and support enzyme activity.
HEPES-KOH (pH 7.5): Acts as a buffering agent to keep the pH stable, which is essential for maintaining enzyme function during long incubations.
Magnesium Glutamate: Provides Mg²⁺ ions, which are essential cofactors for ribosome stability, RNA polymerase activity, and interactions with nucleic acids.
Potassium Phosphate (monobasic/dibasic): Serves as a secondary buffer and provides inorganic phosphate needed for ATP regeneration and nucleotide metabolism.
- Energy / Nucleotide System
Ribose: Feeds into the pentose phosphate pathway to generate precursors (like PRPP) required for nucleotide synthesis.
Glucose: Acts as the main energy source, supporting ATP production through metabolic pathways such as glycolysis.
AMP, CMP, UMP: These nucleoside monophosphates (NMPs) are low-cost precursors that are enzymatically converted into NTPs (ATP, CTP, UTP) for RNA synthesis.
Guanine: Supplied as a nucleobase that is converted into GMP through salvage pathways, then further phosphorylated into GTP for transcription.
- Translation Mix (Amino Acids)
17 Amino Acid Mix + Tyrosine + Cysteine: Provide all amino acids required for protein synthesis. Tyrosine and cysteine are added separately because they are less stable or less soluble in standard mixtures.
- Additives / Backfill
Nicotinamide: Acts as a precursor for NAD⁺, an important cofactor in metabolic reactions that support long-term energy regeneration.
Nuclease-Free Water: Serves as the solvent to adjust final concentrations while preventing degradation of DNA or RNA by nucleases.
2. Differences Between 1-Hour PEP-NTP and 20-Hour NMP–Ribose–Glucose Systems
The 1-hour PEP-NTP system is designed for rapid protein production by providing ready-to-use NTPs and a high-energy phosphate donor (PEP), allowing fast transcription and translation but for a short duration due to quick depletion of resources. In contrast, the 20-hour NMP–Ribose–Glucose system uses cheaper precursors (NMPs, ribose, glucose) and relies on the lysate’s metabolic pathways to gradually regenerate NTPs and energy, enabling longer and more sustained protein production.
3. Bonus Question
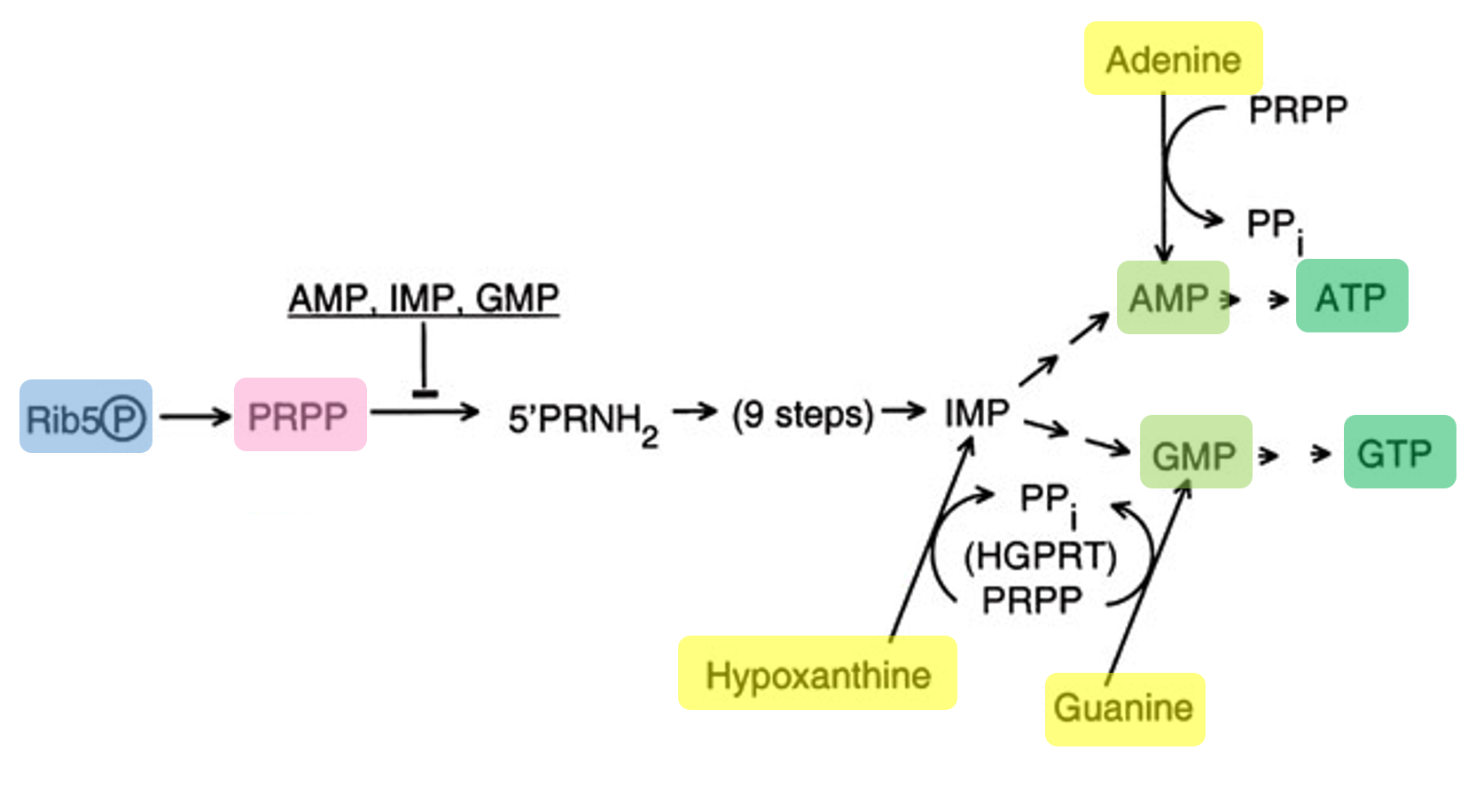
Transcription can still occur without GMP because the system includes guanine, which is converted into GMP through the salvage pathway. In this process, guanine is combined with PRPP (derived from ribose metabolism) to form GMP, which is then phosphorylated into GDP and GTP. The produced GTP is then used by T7 RNA polymerase for RNA synthesis.
1. Fluorescent Protein Properties Affecting Cell-Free Expression
The biophysical properties of fluorescent proteins (FPs), including folding efficiency, maturation time, pH sensitivity, oxygen dependence, and structural stability, play a critical role in determining their fluorescence output in cell-free systems, especially during extended incubations such as 36 hours.
This protein exhibits very fast folding and high structural stability, with efficient chromophore maturation that is oxygen-dependent. Its resistance to misfolding and aggregation allows it to maintain strong and consistent fluorescence over long incubation periods, making it a reliable reference protein in cell-free systems.
mRFP1 (Monomeric Red Fluorescent Protein 1)
It is characterized by slow maturation kinetics and incomplete chromophore formation, which delays the appearance of fluorescence. Additionally, it may form non-fluorescent intermediates, leading to lower overall signal intensity compared to more advanced red fluorescent proteins.
mKO2 (Monomeric Kusabira Orange 2)
This protein shows relatively fast maturation and high brightness, but its chromophore formation is strongly dependent on oxygen availability and can also be influenced by temperature. In conditions with limited oxygen or suboptimal temperature, its fluorescence intensity may be reduced.
mTurquoise2 It has a complex maturation mechanism and high quantum yield but is sensitive to environmental conditions such as pH and oxygen levels. Acidic conditions can reduce fluorescence, while insufficient oxygen may limit proper chromophore formation.
This protein is known for its very high brightness due to an excellent extinction coefficient and quantum yield. However, its performance depends on proper folding, and it can be sensitive to temperature or conditions that promote misfolding, which may reduce fluorescence output.
Electra2 is engineered for rapid maturation and improved performance under reducing conditions commonly found in cell-free systems. Its stability in such environments allows it to maintain fluorescence where other proteins may struggle, although its long-term stability or photostability may vary depending on conditions.
2. Hypothesis (Electra2 Optimization)
For Electra2, fluorescence output over a 36-hour incubation may be limited by the availability of nucleotides and the sustainability of transcription in the cell-free system. I hypothesize that increasing the concentrations of ribose and nucleoside monophosphates (AMP, CMP, UMP, and guanine) will enhance the regeneration of nucleoside triphosphates (NTPs) through the lysate’s metabolic pathways. Ribose can be converted into phosphoribosyl pyrophosphate (PRPP), which is required for nucleotide synthesis, while NMPs and guanine serve as precursors that are enzymatically converted into NTPs. By increasing these components, the system should maintain a continuous supply of NTPs, thereby sustaining transcription by T7 RNA polymerase and increasing mRNA production over time. As a result, this enhanced transcriptional activity is expected to support prolonged translation and lead to higher cumulative protein production and fluorescence intensity over the 36-hour period. This strategy is particularly suitable for Electra2, which is designed for rapid maturation and can efficiently convert increased protein synthesis into measurable fluorescence.
3. Master Mix Design: The Three-Well Strategy
To test this hypothesis, I designed three distinct reagent compositions to identify the “sweet spot” between fuel availability and metabolic stability.
- Mix 1: The “Maximized” Fuel Mix (Well Q4-H20)
Goal: To test the absolute capacity of the system by pushing precursors to the high end.
Key Adjustments: Ribose was increased to 19.0 g/L (+63.4 %) and NMPs (AMP/CMP/UMP) were increased by 60-100 %. Guanine was doubled to 0.313 mM to provide a surplus of base molecules for the salvage pathway.
- Mix 2: The “Intermediate” Mix (Well Q4-G21)
Goal: To establish a bridge between the standard mix and the maximum boost.
Key Adjustments: Ribose was set at 15.0 g/L (+29%) and NMPs/Guanine were increased by 20-33 %. This well helps determine if the “Max” mix is overkill or if a moderate increase is sufficient.
- Mix 3: The “Direct Supply” Mix (Well Q4-I21)
Goal: To test if bypassing the enzymatic salvage of Guanine improves initial speed.
Key Adjustments: While maintaining the Intermediate fuel levels, I added 0.500 mM of pure GMP. This tests whether providing a direct nucleotide (GMP) is more efficient for Electra2 than relying solely on Guanine-to-GMP conversion.
Final Concentration Comparison Table
| Component | Mix 1 (Max Fuel) | Mix 2 (Intermediate) | Mix 3 (Direct Boost) |
|---|---|---|---|
| Cell Lysate | 1X (6.00 µL) | 1X (6.00 µL) | 1X (6.00 µL) |
| DNA Template | 50 nM (2.00 µL) | 50 nM (2.00 µL) | 50 nM (2.00 µL) |
| Ribose | 19.000 g/L | 15.000 g/L | 15.000 g/L |
| AMP | 1.000 mM | 0.750 mM | 0.750 mM |
| CMP | 0.750 mM | 0.500 mM | 0.500 mM |
| UMP | 0.750 mM | 0.500 mM | 0.500 mM |
| GMP | 0.000 mM | 0.000 mM | 0.500 mM |
| Guanine | 0.313 mM | 0.188 mM | 0.156 mM |
| Potassium Glutamate | 312.563 mM | 312.563 mM | 312.563 mM |
| Magnesium Glutamate | 6.975 mM | 6.975 mM | 6.975 mM |
| HEPES-KOH (pH 7.5) | 45.000 mM | 45.000 mM | 45.000 mM |
| 17 Amino Acid Mix | 4.063 mM | 4.063 mM | 4.063 mM |
| Glucose | 1.250 g/L | 1.250 g/L | 1.250 g/L |
| Nicotinamide | 3.125 mM | 3.125 mM | 3.125 mM |
| Backfill (NF Water) | 0.175 µL | 1.225 µL | 1.150 µL |
4. Data Analysis Strategy
Once the 36-hour fluorescence data is returned, I will compare the slopes and peak intensities of these three wells.
- Validation: If Mix 1 > Mix 2 > Standard, the limiting factor was raw fuel.
- Metabolic Insights: If Mix 3 reaches a plateau faster than Mix 2, it proves the enzymatic conversion of Guanine was a kinetic bottleneck for Electra2 production.
Sources:
- Banks, A. M., Whitfield, C. J., Brown, S. R., Fulton, D. A., Goodchild, S. A., Grant, C., Love, J., Lendrem, D. W., Fieldsend, J. E., & Howard, T. P. (2022). Key reaction components affect the kinetics and performance robustness of cell-free protein synthesis reactions. Computational and Structural Biotechnology Journal, 20, 218–229. https://doi.org/10.1016/j.csbj.2021.12.013
- Burrington, L. R., Watts, K. R., & Oza, J. P. (2021). Characterizing and Improving Reaction Times for E. coli-Based Cell-Free Protein Synthesis. ACS Synthetic Biology, 10(8), 1821–1829. https://doi.org/10.1021/acssynbio.1c00195
- Deng, H., Callender, R., Schramm, V. L., & Grubmeyer, C. (2010). Pyrophosphate Activation in Hypoxanthine-Guanine Phosphoribosyltransferase with Transition State Analogue. Biochemistry, 49(12), 2705–2714. https://doi.org/10.1021/bi100012u
- Dopp, B. J. L., Tamiev, D. D., & Reuel, N. F. (2019). Cell-free supplement mixtures: Elucidating the history and biochemical utility of additives used to support in vitro protein synthesis in E. coli extract. Biotechnology Advances, 37(1), 246–258. https://doi.org/10.1016/j.biotechadv.2018.12.006
- Dudzinska, W., Lubkowska, A., Dolegowska, B., Safranow, K., & Jakubowska, K. (2010). Adenine, guanine and pyridine nucleotides in blood during physical exercise and restitution in healthy subjects. European Journal of Applied Physiology, 110(6), 1155–1162. https://doi.org/10.1007/s00421-010-1611-7
- Gregorio, N. E., Levine, M. Z., & Oza, J. P. (2019). A User’s Guide to Cell-Free Protein Synthesis. Methods and Protocols, 2(1), 24. https://doi.org/10.3390/mps2010024
- Hashimura, H., Nakagawa, H., & Sawai, S. (2025). Use of blue fluorescent protein Electra2 for live-cell imaging in Dictyostelium discoideum. microPublication Biology. https://doi.org/10.17912/micropub.biology.001774
- Hove-Jensen, B., Andersen, K. R., Kilstrup, M., Martinussen, J., Switzer, R. L., & Willemoës, M. (2016). Phosphoribosyl Diphosphate (PRPP): Biosynthesis, Enzymology, Utilization, and Metabolic Significance. Microbiology and Molecular Biology Reviews : MMBR, 81(1), e00040-16. https://doi.org/10.1128/MMBR.00040-16
- Jiang, L., Zhao, J., Lian, J., & Xu, Z. (2018). Cell-free protein synthesis enabled rapid prototyping for metabolic engineering and synthetic biology. Synthetic and Systems Biotechnology, 3(2), 90–96. https://doi.org/10.1016/j.synbio.2018.02.003
- Jiang, N., Ding, X., & Lu, Y. (2021). Development of a robust Escherichia coli-based cell-free protein synthesis application platform. Biochemical Engineering Journal, 165, 107830. https://doi.org/10.1016/j.bej.2020.107830
- Krinsky, N., Kaduri, M., Shainsky-Roitman, J., Goldfeder, M., Ivanir, E., Benhar, I., Shoham, Y., & Schroeder, A. (2016). A Simple and Rapid Method for Preparing a Cell-Free Bacterial Lysate for Protein Synthesis. PLOS ONE, 11(10), e0165137. https://doi.org/10.1371/journal.pone.0165137
- Vengut-Climent, E., Peñalver, P., Lucas, R., Gómez-Pinto, I., Aviñó, A., Muro-Pastor, A. M., Galbis, E., de Paz, M. V., Fonseca Guerra, C., Bickelhaupt, F. M., Eritja, R., González, C., & Morales, J. C. (2018). Glucose-nucleobase pairs within DNA: Impact of hydrophobicity, alternative linking unit and DNA polymerase nucleotide insertion studies †Electronic supplementary information (ESI) available. See DOI: 10.1039/c7sc04850e. Chemical Science, 9(14), 3544–3554. https://doi.org/10.1039/c7sc04850e
- Zhang, Y., Huang, Q., Deng, Z., Xu, Y., & Liu, T. (2018). Enhancing the efficiency of cell-free protein synthesis system by systematic titration of transcription and translation components. Biochemical Engineering Journal, 138, 47–53. https://doi.org/10.1016/j.bej.2018.07.001