TobUbi.U4 proximal promoter:
The 263 bp proximal promoter region of the Ubi.U4 gene from Nicotiana tabacum was obtained based on the study by Genschik et al., (1994)This region corresponds to the sequence spanning −263 to −1 relative to the transcription start site (TSS) and contains key cis-regulatory elements involved in transcriptional regulation.
The transcription start site (TSS, +1) was not directly annotated in the GenBank entry. Therefore, it was determined based on the promoter analysis presented in the original publication by Genschik et al. (1994), where the TSS was experimentally identified and illustrated in Figure 3.
The nucleotide sequence was retrieved from the GenBank database (accession: X77456.1), corresponding to positions 575–837 of the N. tabacum Ubi.U4 gene.
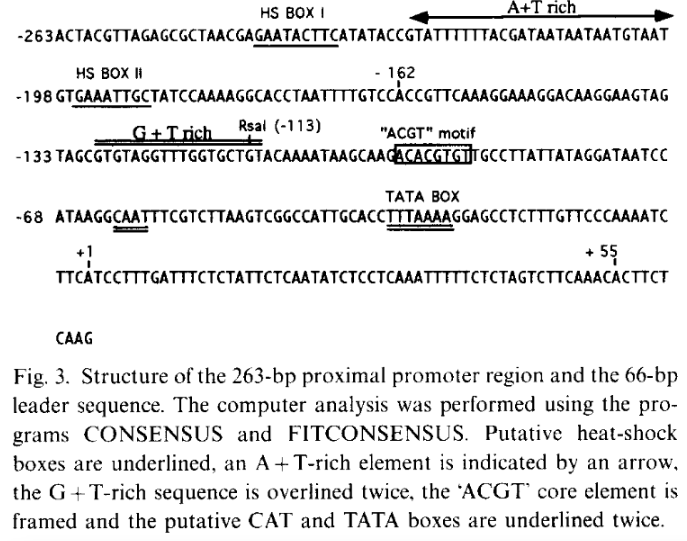


> emb|X77456.1 :575-837 N.tabacum Ubi.U4 gene
ACTACGTTAGAGCGCTAACGAGAATACTTCATATACCGTATTTTTTACGATAATAATAATGTAATGTGAAATTGCTATCCAAAAGGCACCTAATTTTGTCCACCGTTCAAAGGAAAGGACAAGGAAGTAGTAGCGTGTAGGTTTGGTGCTGTACAAAATAAGCAAGACACGTGTTGCCTTATTATAGGATAATCCATAAGGCAATTTCGTCTTAAGTCGGCCATTGCACCTTTAAAAGGAGCCTCTTTGTTCCCAAAATCTTC
D100 chimeric promoter (Dahlia mosaic virus - DaMV):
The D100 promoter is a synthetic construct derived from the Dahlia mosaic virus (DaMV) genome, as described by (Khadanga et al., 2021)based on the work of (Sahoo et al., 2015). It is designed by combining an upstream activation sequence with a core promoter region to enhance transcriptional activity.
- DaMV14UAS (−203 to −33): an upstream activation sequence acting as a transcriptional enhancer
- A short linker sequence (CCCGAC)
- DaMV4CP (−474 to +82): a core promoter region required for basal transcription
The source promoter region corresponds to a 706 bp fragment (6579–7280) of the DaMV genome (GenBank: JX272320.1), with the transcription start site (TSS, +1) located at position 7053 based on coordinate mapping.
The following sequences were extracted based on coordinate mapping:
DaMV14UAS (−203 to −33):
> gb|JX272320.1|:6850-7020 Dahlia mosaic virus clone pDaMV-p2, complete genome
TCTGCAAACAGCTCAAAAAGCTACTGGCCGACAATCATAATTGCTCGGCATGTGCAGGTGGGGCCTCCACTAGCAATAATACAAGCTTTACAGCTTGCAGTGACTCATCCTCCAATAATGAGGAAAAAGACGTCAGCAGTGACGAACAAGGGCCTGAAGACTTGCCTATAT
DaMV4CP (−474 to +82):
> gb|JX272320.1|:6579-7134 Dahlia mosaic virus clone pDaMV-p2, complete genome
GAATTCAATCCTCCTCAGGAAATGAAGGATTCAGGAGATCTTCTCTATCAACTTGCTCAAGTAAGGACAAACGGGTTCACCCGGATCCTCCAGAAGACCCAGTCTATCAACGGAGAAACAAAGATAAAAATCAATTACTCACATGAAAGAGTATTGATCACGAGTCACTATGGAGCGACAATCTCCAGACAGGATGTCAGCATCTTATCTTCCTTTGAAGAAAGCATCATCAATAACGATGTAATGGTGGGGACATCCACTAAGTTATTGCTCTGCAAACAGCTCAAAAAGCTACTGGCCGACAATCATAATTGCTCGGCATGTGCAGGTGGGGCCTCCACTAGCAATAATACAAGCTTTACAGCTTGCAGTGACTCATCCTCCAATAATGAGGAAAAAGACGTCAGCAGTGACGAACAAGGGCCTGAAGACTTGCCTATATAATGGCATTCACCCCTCAGTTGAAGAGCATCAGGAGTTTCAGCATAGAAACTTTCTCTTTAACAAATCTATCTTTTCTTTAAAGCATGTGTGAGTAGAAACCCATATAGGGTTA
Initially, the promoter sequence was reconstructed using GenBank coordinates. However, slight discrepancies were observed when compared to the promoter structure illustrated in the published figure.
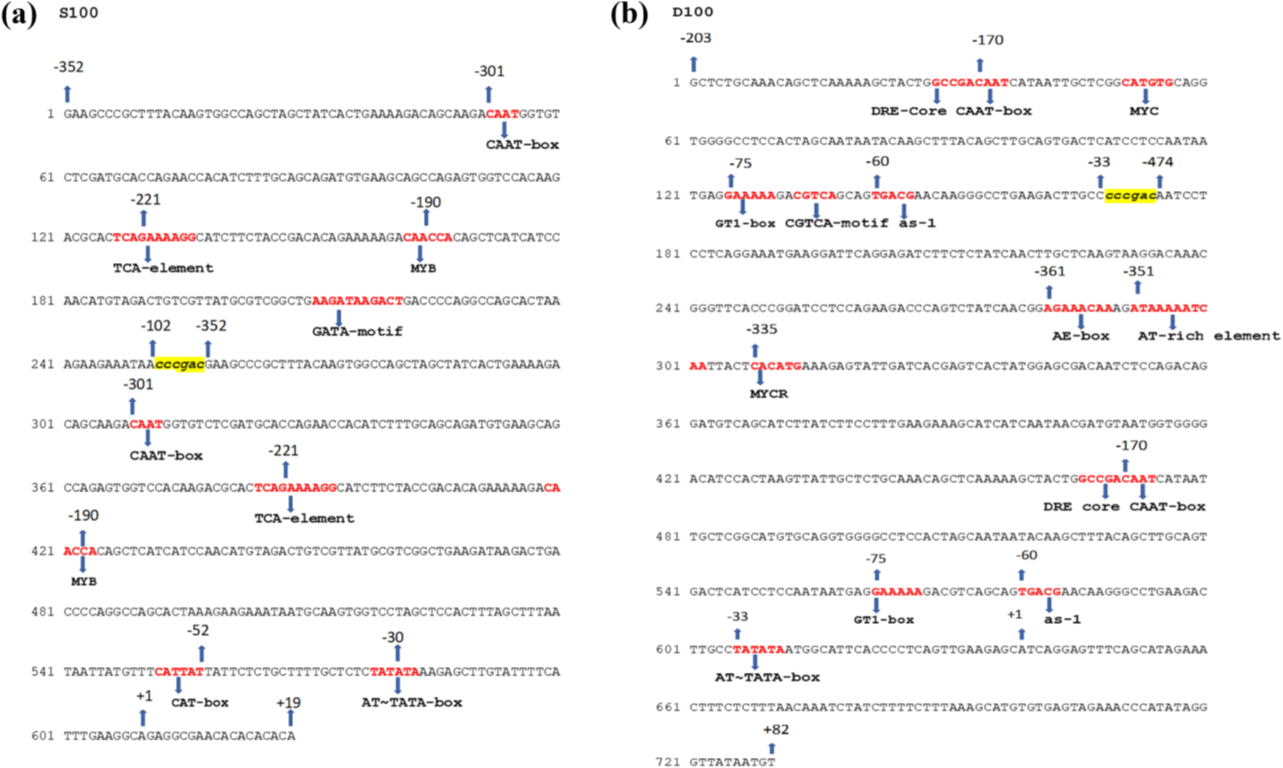

 Therefore, the final D100 promoter sequence was generated using an Gemini AI tool based on the figure from Khadanga et al. (2021), as it accurately reflects the reported experimental construct:
Therefore, the final D100 promoter sequence was generated using an Gemini AI tool based on the figure from Khadanga et al. (2021), as it accurately reflects the reported experimental construct:
GCTCTGCAAACAGCTCAAAAAGCTACTGGCCGACAATCATAATTGCTCGGCATGTGCAGGTGGGGCCTCCACTAGCAATAATACAAGCTTTACAGCTTGCAGTGACTCATCCTCCAATAATGAGGAAAAAGACGTCAGCAGTGACGAACAAGGGCCTGAAGACTTGCCcccgacAATCCTCCTCAGGAAATGAAGGATTCAGGAGATCTTCTCTATCAACTTGCTCAAGTAAGGACAAACGGGTTCACCCGGATCCTCCAGAAGACCCAGTCTATCAACGGAGAAACAAAGATAAAAATCAATTACTCACATGAAAGAGTATTGATCACGAGTCACTATGGAGCGACAATCTCCAGACAGGATGTCAGCATCTTATCTTCCTTTGAAGAAAGCATCATCAATAACGATGTAATGGTGGGGACATCCACTAAGTTATTGCTCTGCAAACAGCTCAAAAAGCTACTGGCCGACAATCATAATTGCTCGGCATGTGCAGGTGGGGCCTCCACTAGCAATAATACAAGCTTTACAGCTTGCAGTGACTCATCCTCCAATAATGAGGAAAAAGACGTCAGCAGTGACGAACAAGGGCCTGAAGACTTGCCTATATAATGGCATTCACCCCTCAGTTGAAGAGCATCAGGAGTTTCAGCATAGAAACTTTCTCTTTAACAAATCTATCTTTTCTTTAAAGCATGTGTGAGTAGAAACCCATATAGGGTTATAATGT
S100 chimeric promoter (Soybean vein clearing virus, SVBV):
The S100 promoter is a synthetic chimeric construct derived from the Soybean vein clearing virus (SVBV), as described by Khadanga et al., (2021)based on Pattanaik et al., (2004). It is designed by combining an upstream activation sequence with a core promoter region to enhance transcriptional activity.
- SV10UAS (250 bp) (-352 to -102): This is the Upstream Activation Sequence that contains major regulatory elements contributing to transcriptional enhancement.
2.2. The Linker: CCCGAC sequence: A synthetic 6 bp linker (CCCGAC) inserted between the enhancer and core promoter, similar to the design used in the D100 promoter.
- SV10CP (371 bp) (-352 to +19): The core promoter fragment (also referred to as SVBVFLt10) containing the TATA box (around −30) and the transcription start site (TSS, +1) required for transcription initiation.
The S100 promoter sequence was directly extracted from Figure 1 of Pattanaik et al. (2004), where the nucleotide sequence is explicitly provided in text format, and assembled in this order [SV10UAS] + [CCCGAC linker] + [SV10CP]:
GAAGCCCGCTTTACAAGTGGCCAGCTAGCTATCACTGAAAAGACAGCAAGACAATGGTGTCTCGATGCACCAGAACCACATCTTTGCAGCAGATGTGAAGCAGCCAGAGTGGTCCACAAGACGCACTCAGAAAAGGCATCTTCTACCGACACAGAAAAAGACAACCACAGCTCATCATCCAACATGTAGACTGTCGTTATGCGTCGGCTGAAGATAAGACTGACCCCAGGCCAGCACTAAAGAAGAAATAAcccgacGAAGCCCGCTTTACAAGTGGCCAGCTAGCTATCACTGAAAAGACAGCAAGACAATGGTGTCTCGATGCACCAGAACCACATCTTTGCAGCAGATGTGAAGCAGCCAGAGTGGTCCACAAGACGCACTCAGAAAAGGCATCTTCTACCGACACAGAAAAAGACAACCACAGCTCATCATCCAACATGTAGACTGTCGTTATGCGTCGGCTGAAGATAAGACTGACCCCAGGCCAGCACTAAAGAAGAAATAATGCAAGTGGTCCTAGCTCCACTTTAGCTTTAATAATTATGTTTCATTATTATTCTCTGCTTTTGCTCTCTATATAAAGAGCTTGTATTTTCATTTGAAGGCAGAGGCGAACACACACACA
DaMVFLt4 promoter (556 pb):
The DaMV4CP fragment corresponds to a natural promoter region derived from the Dahlia mosaic virus (DaMV). It consists of a 556 bp sequence spanning positions −474 to +82 relative to the transcription start site (TSS) according to Sahoo et al., (2014) study.
This fragment was directly extracted from the DaMV genome available in the GenBank database (accession: JX272320.1), corresponding to genomic coordinates 6579–7134.
> gb|JX272320.1|:6579-7134 Dahlia mosaic virus clone pDaMV-p2, complete genome
GAATTCAATCCTCCTCAGGAAATGAAGGATTCAGGAGATCTTCTCTATCAACTTGCTCAAGTAAGGACAAACGGGTTCACCCGGATCCTCCAGAAGACCCAGTCTATCAACGGAGAAACAAAGATAAAAATCAATTACTCACATGAAAGAGTATTGATCACGAGTCACTATGGAGCGACAATCTCCAGACAGGATGTCAGCATCTTATCTTCCTTTGAAGAAAGCATCATCAATAACGATGTAATGGTGGGGACATCCACTAAGTTATTGCTCTGCAAACAGCTCAAAAAGCTACTGGCCGACAATCATAATTGCTCGGCATGTGCAGGTGGGGCCTCCACTAGCAATAATACAAGCTTTACAGCTTGCAGTGACTCATCCTCCAATAATGAGGAAAAAGACGTCAGCAGTGACGAACAAGGGCCTGAAGACTTGCCTATATAATGGCATTCACCCCTCAGTTGAAGAGCATCAGGAGTTTCAGCATAGAAACTTTCTCTTTAACAAATCTATCTTTTCTTTAAAGCATGTGTGAGTAGAAACCCATATAGGGTTA
SM chimeric hybrid promoter (SUAS + MUAS fusion):
The SM promoter is a synthetic chimeric hybrid promoter constructed by combining regulatory elements from two plant viruses, as described by Kumari et al., (2024). It integrates an upstream activation sequence from Sugarcane bacilliform virus with an enhancer domain from Mirabilis mosaic virus to enhance transcriptional activity.
- SUAS ( SCBV Upstream Activation Sequence): This fragment corresponds to the Upstream Activation Sequence (UAS) derived from Sugarcane bacilliform virus (SCBV), as described by Davies et al., (2014). The selected region spans −434 bp to −153 bp relative to the transcription start site (TSS), resulting in a fragment of 282 bp. This region functions as a transcriptional enhancer.
- MUAS (MMV Upstream Activation Sequence): This fragment corresponds to the transcriptional enhancer domain derived from the full-length transcript (FLt) promoter of Mirabilis mosaic virus (MMV), as reported by Dey & Maiti, (1999).The sequence spans −297 to −38 relative to the TSS, with a total length of 259 bp, and contributes strong enhancer activity.
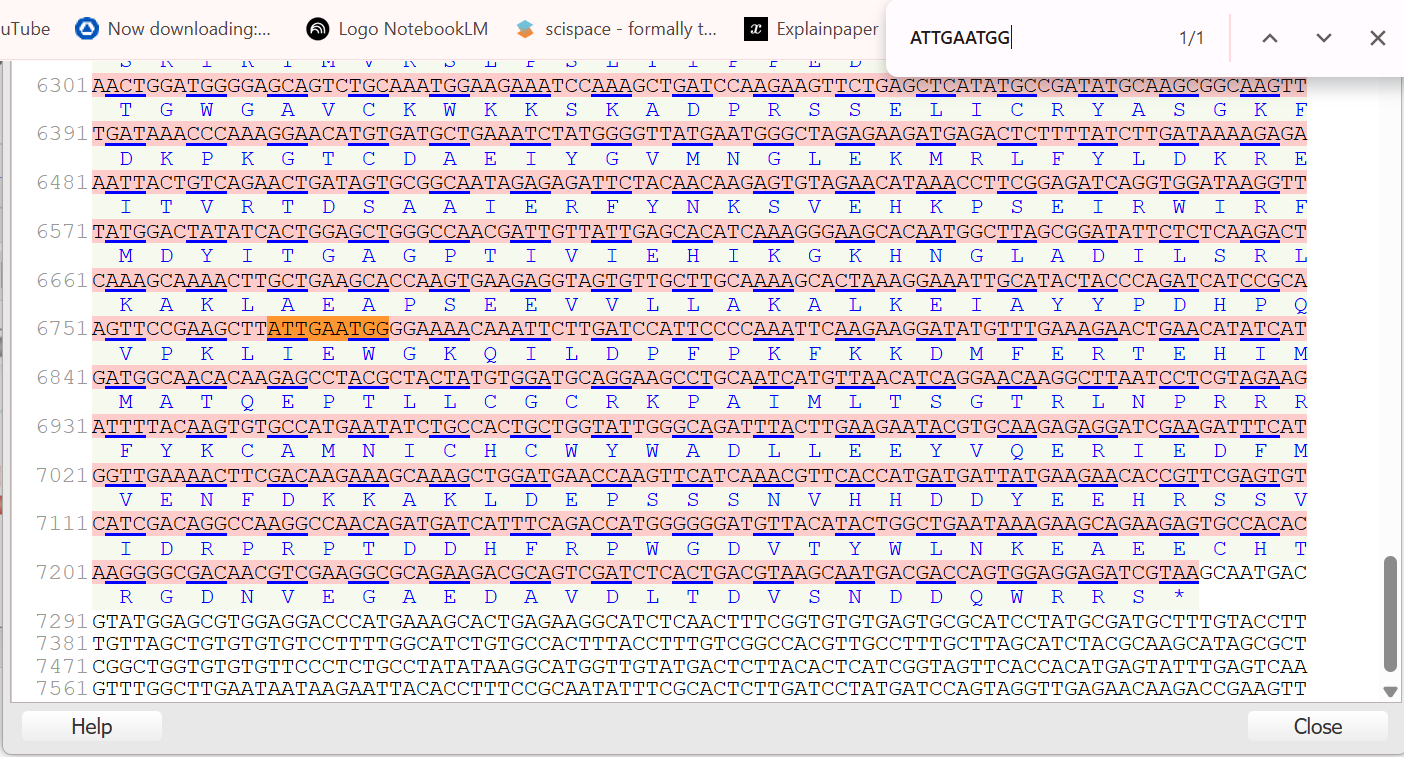
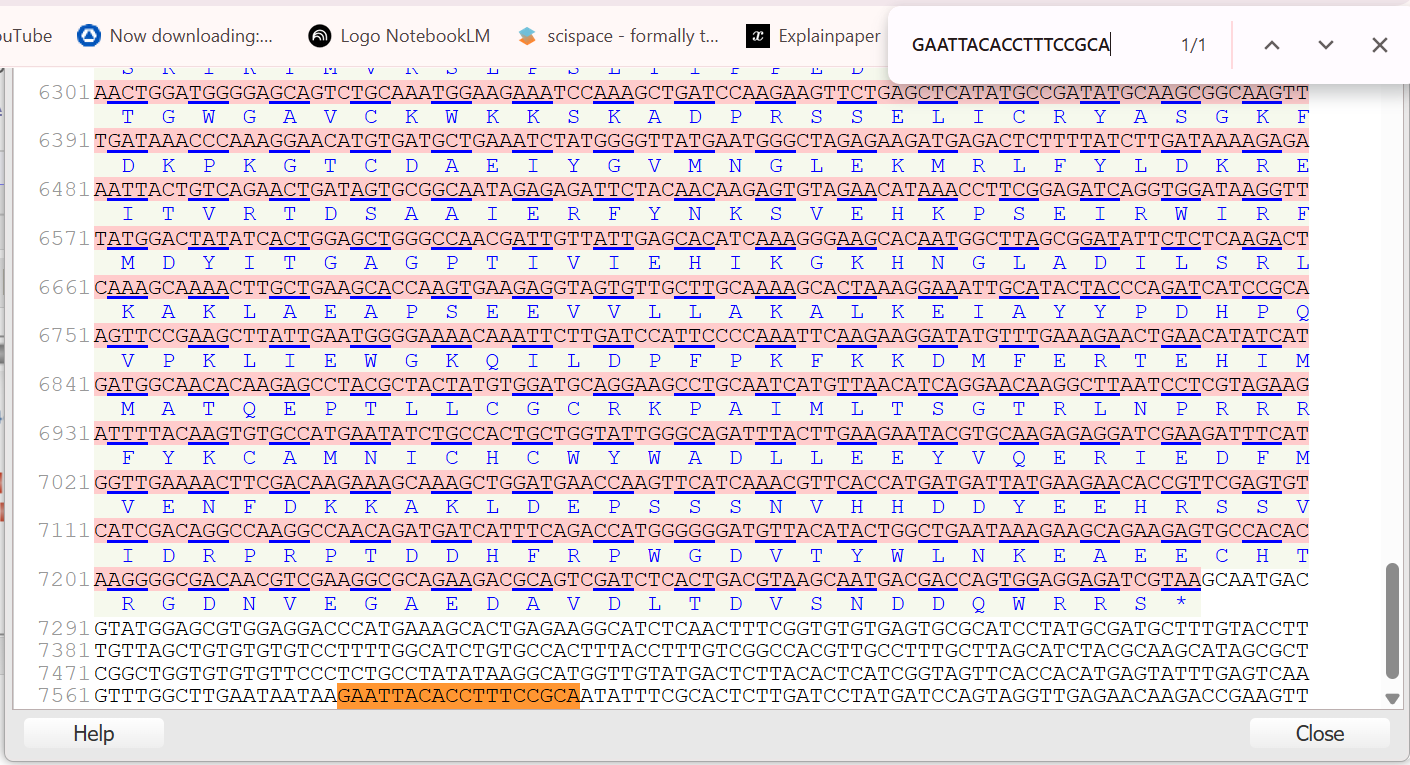
To find the first fragment SUAS, I first mapped both boundaries of the 839 bp SCBV promoter using the SCBV-F primer anchor (ATTGAATGG) and the complement of the SCBV-R primer (GAATTACACCTTTCCGCA) against the Sugarcane bacilliform virus (SCBV) Ireng Maleng isolate sequence (accession AJ277091). This allowed me to confirm the full span of the mother fragment from relative coordinate −770 to +69

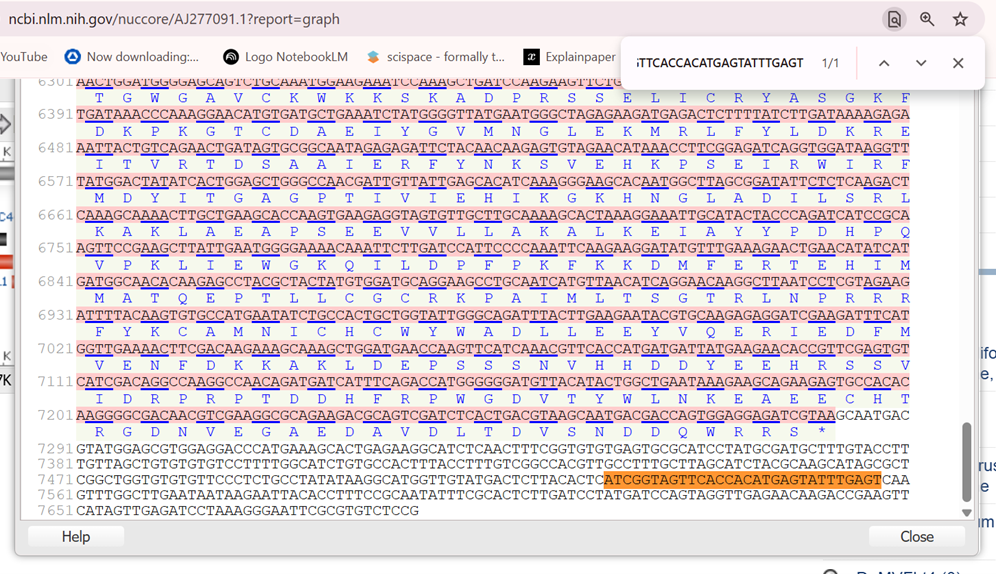
 Next, I identified the Transcription Start Site (TSS) based on the underlined leader sequence reported in the Figure 2 from the Davies (2014) study. I could identify the TSS (+1) as the 7528th nucleotide in the Sugarcane bacilliform virus (SCBV) Ireng Maleng isolate sequence:
7528 ATC GGTAGTTCAC CACATGAGTA TTTGAGTCAA 7560
Next, I identified the Transcription Start Site (TSS) based on the underlined leader sequence reported in the Figure 2 from the Davies (2014) study. I could identify the TSS (+1) as the 7528th nucleotide in the Sugarcane bacilliform virus (SCBV) Ireng Maleng isolate sequence:
7528 ATC GGTAGTTCAC CACATGAGTA TTTGAGTCAA 7560
 To isolate the specific SUAS domain for the SM promoter, which the sources define as the segment from relative coordinates −434 to −153, I calculated the internal absolute indices within the 839 bp mother fragment. By mapping these relative coordinates back from the TSS, I determined the exact 282 bp enhancer sequence required to be joined directly to the MMV core promoter to build the chimeric SM promoter:
To isolate the specific SUAS domain for the SM promoter, which the sources define as the segment from relative coordinates −434 to −153, I calculated the internal absolute indices within the 839 bp mother fragment. By mapping these relative coordinates back from the TSS, I determined the exact 282 bp enhancer sequence required to be joined directly to the MMV core promoter to build the chimeric SM promoter:
> emb|AJ277091.1|:7094-7375 Sugarcane bacilliform IM virus complete genome, isolate Ireng Maleng
GAACACCGTTCGAGTGTCATCGACAGGCCAAGGCCAACAGATGATCATTTCAGACCATGGGGGGATGTTACATACTGGCTGAATAAAGAAGCAGAAGAGTGCCACACAAGGGGCGACAACGTCGAAGGCGCAGAAGACGCAGTCGATCTCACTGACGTAAGCAATGACGACCAGTGGAGGAGATCGTAAGCAATGACGTATGGAGCGTGGAGGACCCATGAAAGCACTGAGAAGGCATCTCAACTTTCGGTGTGTGAGTGCGCATCCTATGCGATGCTTTGT
To find the second fragment MUAS, I first identified the source as the Mirabilis mosaic virus (MMV) full-length transcript (FLt) promoter from the Dey and Maiti (1999) article. Because the original study provided the literal nucleotide sequence in Figure 1 rather than a GenBank accession number, I used the printed sequence obtained from Gemini AI tool as my primary reference.

 I then established the Transcription Start Site (TSS or +1) as the anchor point, which the researchers mapped via primer extension to a guanidine (G) residue located 24 nucleotides downstream of the TATATAA box. To isolate the specific MUAS fragment, which spans the relative coordinates −297 to −38, I counted upstream from the TSS to locate the nucleotide at position −297 and extracted the sequence through to the nucleotide at position −38. This process provided the 259 bp enhancer domain required for the construction of the SM and BM chimeric promoters:
I then established the Transcription Start Site (TSS or +1) as the anchor point, which the researchers mapped via primer extension to a guanidine (G) residue located 24 nucleotides downstream of the TATATAA box. To isolate the specific MUAS fragment, which spans the relative coordinates −297 to −38, I counted upstream from the TSS to locate the nucleotide at position −297 and extracted the sequence through to the nucleotide at position −38. This process provided the 259 bp enhancer domain required for the construction of the SM and BM chimeric promoters:
TTCGTCCACAGACATCAACATCTTATCGTCCTTTGAAGATAAGATAATAATGTTGAAGATAAGAGTGGGAGCCACCACTAAAACATTGCTTTGTCAAAAGCTAAAAAAGATGATGCCCGACAGCCACTTGTGTGAAGCATGTGAAGCCGGTCCCTCCACTAAGAAAATTAGTGAAGCATCTTCCAGTGGTCCCTCCACTCACAGCTCAATCAGTGAGCAACAGGACGAAGGAAATGACGTAAGCCATGACGTCTAATCCC
The SM promoter was generated by directly fusing the SUAS fragment upstream of the MUAS enhancer sequence, as described by (Kumari et al., 2024a) based on the source sequence described in Dey & Maiti, (1999) study:
GAACACCGTTCGAGTGTCATCGACAGGCCAAGGCCAACAGATGATCATTTCAGACCATGGGGGGATGTTACATACTGGCTGAATAAAGAAGCAGAAGAGTGCCACACAAGGGGCGACAACGTCGAAGGCGCAGAAGACGCAGTCGATCTCACTGACGTAAGCAATGACGACCAGTGGAGGAGATCGTAAGCAATGACGTATGGAGCGTGGAGGACCCATGAAAGCACTGAGAAGGCATCTCAACTTTCGGTGTGTGAGTGCGCATCCTATGCGATGCTTTGTTTCGTCCACAGACATCAACATCTTATCGTCCTTTGAAGATAAGATAATAATGTTGAAGATAAGAGTGGGAGCCACCACTAAAACATTGCTTTGTCAAAAGCTAAAAAAGATGATGCCCGACAGCCACTTGTGTGAAGCATGTGAAGCCGGTCCCTCCACTAAGAAAATTAGTGAAGCATCTTCCAGTGGTCCCTCCACTCACAGCTCAATCAGTGAGCAACAGGACGAAGGAAATGACGTAAGCCATGACGTCTAATCCC
BM chimeric hybrid promoter (BUAS + MUAS fusion):
The BM promoter is a synthetic chimeric hybrid promoter constructed by the fusion of two regulatory elements, as described by (Kumari et al., 2024a). It combines an upstream activation sequence from Banana streak virus with an enhancer domain from Mirabilis mosaic virus to enhance transcriptional efficiency.
- BUAS (BSV Upstream Activation Sequence) : This fragment corresponds to the Upstream Activation Sequence (UAS) derived from Banana streak virus (BSV), as reported by Remans et al., (2005). The selected region spans −1150 bp to −33 bp relative to the transcription start site (TSS), resulting in an expected length of approximately 1117 bp. This region functions as a strong transcriptional enhancer.
- MUAS (MMV Upstream Activation Sequence): This sequence corresponds to the transcriptional enhancer domain derived from the full-length transcript (FLt) promoter of Mirabilis mosaic virus (MMV). It is identical to the MUAS element used in the SM promoter and contributes additional transcriptional activation capacity.
To find the first fragment BUAS, I first identified the source as the Banana streak virus (BSV) Cavendish isolate, which corresponds to GenBank accession AF215815. Although the current database entry for this accession may show a length of 1,287 bp, I noted that the sources utilize a 1,304 bp synthesized version of this isolate spanning from relative coordinates −1,150 to +154.
Next, I used the BSV-F primer anchor sequence (GGTTGCATGGAAGG) to locate the beginning of the promoter region within the GenBank file. By finding this exact sequence at the very start of the file, I established that Nucleotide 1 of the GenBank entry corresponds to the relative coordinate −1,150.
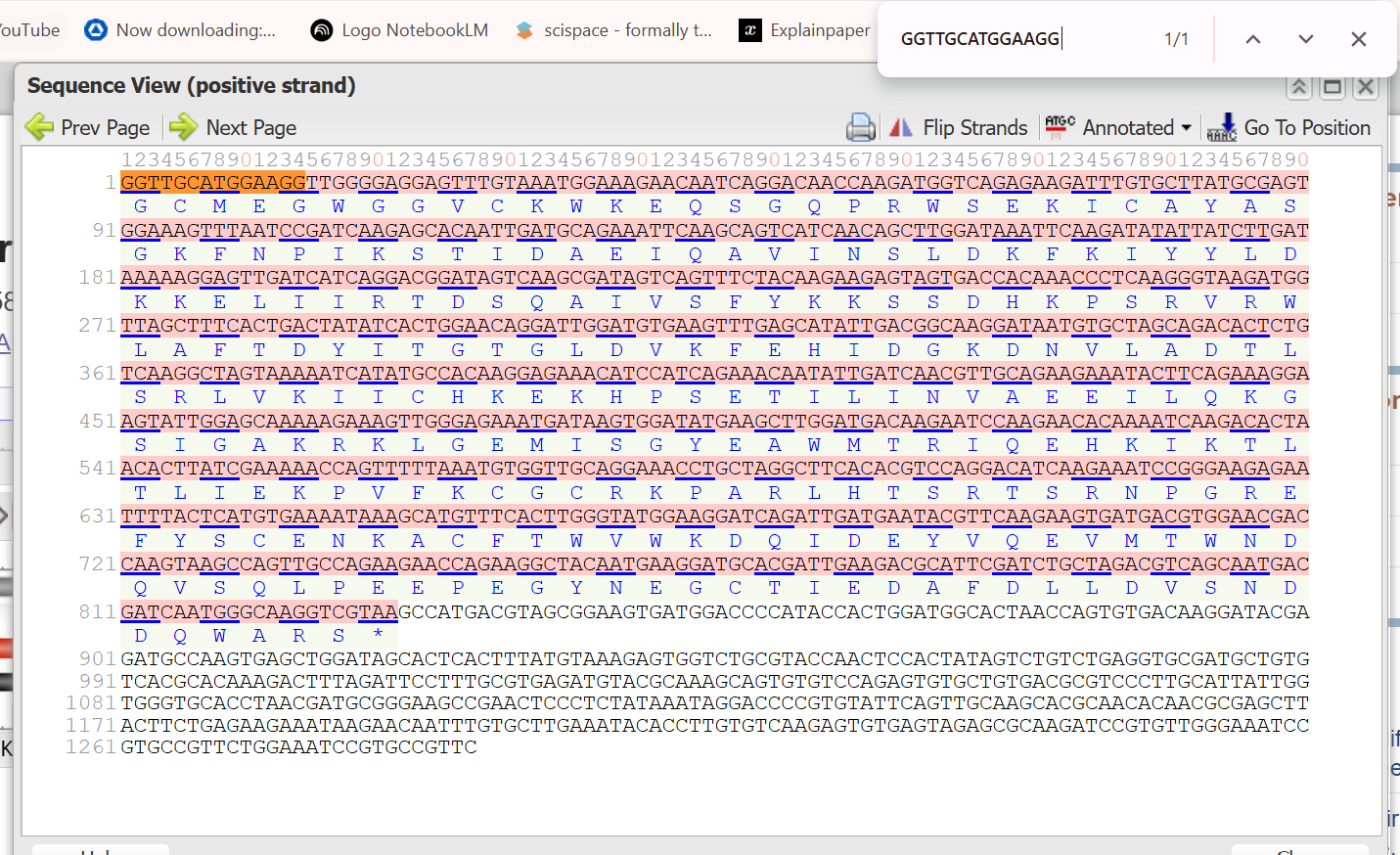
 I then determined the Transcription Start Site (TSS or +1) by mapping the relative coordinates to the absolute indices of the 1,304 bp sequence. Since there are 1,150 bases upstream of the start site, the TSS is located at Nucleotide 1151. To isolate the specific BUAS domain, which the sources define as the segment from −1,150 to −33, I calculated the end index by subtracting 33 from the TSS (1151−33=1118).
Finally, I extracted the sequence from Nucleotide 1 to Nucleotide 1118, which provided the approximately 1,117 bp (mathematically 1,118 bp) enhancer fragment required to construct the BM chimeric promoter:
I then determined the Transcription Start Site (TSS or +1) by mapping the relative coordinates to the absolute indices of the 1,304 bp sequence. Since there are 1,150 bases upstream of the start site, the TSS is located at Nucleotide 1151. To isolate the specific BUAS domain, which the sources define as the segment from −1,150 to −33, I calculated the end index by subtracting 33 from the TSS (1151−33=1118).
Finally, I extracted the sequence from Nucleotide 1 to Nucleotide 1118, which provided the approximately 1,117 bp (mathematically 1,118 bp) enhancer fragment required to construct the BM chimeric promoter:
> gb|AF215815.1|:1-1118 Banana streak virus ORF III polyprotein gene, partial cds
GGTTGCATGGAAGGTTGGGGAGGAGTTTGTAAATGGAAAGAACAATCAGGACAACCAAGATGGTCAGAGAAGATTTGTGCTTATGCGAGTGGAAAGTTTAATCCGATCAAGAGCACAATTGATGCAGAAATTCAAGCAGTCATCAACAGCTTGGATAAATTCAAGATATATTATCTTGATAAAAAGGAGTTGATCATCAGGACGGATAGTCAAGCGATAGTCAGTTTCTACAAGAAGAGTAGTGACCACAAACCCTCAAGGGTAAGATGGTTAGCTTTCACTGACTATATCACTGGAACAGGATTGGATGTGAAGTTTGAGCATATTGACGGCAAGGATAATGTGCTAGCAGACACTCTGTCAAGGCTAGTAAAAATCATATGCCACAAGGAGAAACATCCATCAGAAACAATATTGATCAACGTTGCAGAAGAAATACTTCAGAAAGGAAGTATTGGAGCAAAAAGAAAGTTGGGAGAAATGATAAGTGGATATGAAGCTTGGATGACAAGAATCCAAGAACACAAAATCAAGACACTAACACTTATCGAAAAACCAGTTTTTAAATGTGGTTGCAGGAAACCTGCTAGGCTTCACACGTCCAGGACATCAAGAAATCCGGGAAGAGAATTTTACTCATGTGAAAATAAAGCATGTTTCACTTGGGTATGGAAGGATCAGATTGATGAATACGTTCAAGAAGTGATGACGTGGAACGACCAAGTAAGCCAGTTGCCAGAAGAACCAGAAGGCTACAATGAAGGATGCACGATTGAAGACGCATTCGATCTGCTAGACGTCAGCAATGACGATCAATGGGCAAGGTCGTAAGCCATGACGTAGCGGAAGTGATGGACCCCATACCACTGGATGGCACTAACCAGTGTGACAAGGATACGAGATGCCAAGTGAGCTGGATAGCACTCACTTTATGTAAAGAGTGGTCTGCGTACCAACTCCACTATAGTCTGTCTGAGGTGCGATGCTGTGTCACGCACAAAGACTTTAGATTCCTTTGCGTGAGATGTACGCAAAGCAGTGTGTCCAGAGTGTGCTGTGACGCGTCCCTTGCATTATTGGTGGGTGCACCTAACGATGCGGGAAGCCGAACTCCCTCT
The BM promoter was generated by directly fusing the BUAS fragment upstream of the MUAS enhancer sequence, as described by Kumari et al., (2024):
GGTTGCATGGAAGGTTGGGGAGGAGTTTGTAAATGGAAAGAACAATCAGGACAACCAAGATGGTCAGAGAAGATTTGTGCTTATGCGAGTGGAAAGTTTAATCCGATCAAGAGCACAATTGATGCAGAAATTCAAGCAGTCATCAACAGCTTGGATAAATTCAAGATATATTATCTTGATAAAAAGGAGTTGATCATCAGGACGGATAGTCAAGCGATAGTCAGTTTCTACAAGAAGAGTAGTGACCACAAACCCTCAAGGGTAAGATGGTTAGCTTTCACTGACTATATCACTGGAACAGGATTGGATGTGAAGTTTGAGCATATTGACGGCAAGGATAATGTGCTAGCAGACACTCTGTCAAGGCTAGTAAAAATCATATGCCACAAGGAGAAACATCCATCAGAAACAATATTGATCAACGTTGCAGAAGAAATACTTCAGAAAGGAAGTATTGGAGCAAAAAGAAAGTTGGGAGAAATGATAAGTGGATATGAAGCTTGGATGACAAGAATCCAAGAACACAAAATCAAGACACTAACACTTATCGAAAAACCAGTTTTTAAATGTGGTTGCAGGAAACCTGCTAGGCTTCACACGTCCAGGACATCAAGAAATCCGGGAAGAGAATTTTACTCATGTGAAAATAAAGCATGTTTCACTTGGGTATGGAAGGATCAGATTGATGAATACGTTCAAGAAGTGATGACGTGGAACGACCAAGTAAGCCAGTTGCCAGAAGAACCAGAAGGCTACAATGAAGGATGCACGATTGAAGACGCATTCGATCTGCTAGACGTCAGCAATGACGATCAATGGGCAAGGTCGTAAGCCATGACGTAGCGGAAGTGATGGACCCCATACCACTGGATGGCACTAACCAGTGTGACAAGGATACGAGATGCCAAGTGAGCTGGATAGCACTCACTTTATGTAAAGAGTGGTCTGCGTACCAACTCCACTATAGTCTGTCTGAGGTGCGATGCTGTGTCACGCACAAAGACTTTAGATTCCTTTGCGTGAGATGTACGCAAAGCAGTGTGTCCAGAGTGTGCTGTGACGCGTCCCTTGCATTATTGGTGGGTGCACCTAACGATGCGGGAAGCCGAACTCCCTCTTTCGTCCACAGACATCAACATCTTATCGTCCTTTGAAGATAAGATAATAATGTTGAAGATAAGAGTGGGAGCCACCACTAAAACATTGCTTTGTCAAAAGCTAAAAAAGATGATGCCCGACAGCCACTTGTGTGAAGCATGTGAAGCCGGTCCCTCCACTAAGAAAATTAGTGAAGCATCTTCCAGTGGTCCCTCCACTCACAGCTCAATCAGTGAGCAACAGGACGAAGGAAATGACGTAAGCCATGACGTCTAATCCC
MSD3 chimeric deletion-hybrid promoter (MUAS + SD3):
The MSD3 promoter is a “deletion-hybrid” construct composed of the following two fragments joined directly together as described in the study of (Kumari et al., 2024b):
- MUAS (MMV Upstream Activation Sequence): This is the same sequence of the transcriptional enhancer domain isolated from the Mirabilis mosaic virus (MMV) full-length transcript (FLt) promoter, as used in SM and BM promoters.
- SD3 (SCBV Deletion Fragment 3): This fragment is a truncated promoter region derived from the Sugarcane bacilliform virus (SCBV), as described by Davies et al., 2014. The SD3 sequence corresponds to the region spanning −340 bp to +69 bp relative to the transcription start site, resulting in a fragment of 409 bp. This region retains essential core promoter elements required for basal transcription.
The SD3 fragment was extracted from the SCBV genome (GenBank accession: AJ277091.1, positions 7188–7597):
emb|AJ277091.1|:7188-7597 Sugarcane bacilliform IM virus complete genome, isolate Ireng Maleng
AAGAGTGCCACACAAGGGGCGACAACGTCGAAGGCGCAGAAGACGCAGTCGATCTCACTGACGTAAGCAATGACGACCAGTGGAGGAGATCGTAAGCAATGACGTATGGAGCGTGGAGGACCCATGAAAGCACTGAGAAGGCATCTCAACTTTCGGTGTGTGAGTGCGCATCCTATGCGATGCTTTGTACCTTTGTTAGCTGTGTGTGTCCTTTTGGCATCTGTGCCACTTTACCTTTGTCGGCCACGTTGCCTTTGCTTAGCATCTACGCAAGCATAGCGCTCGGCTGGTGTGTGTTCCCTCTGCCTATATAAGGCATGGTTGTATGACTCTTACACTCATCGGTAGTTCACCACATGAGTATTTGAGTCAAGTTTGGCTTGAATAATAAGAATTACACCTTTCCGCAA
The final MSD3 promoter was obtained by direct assembly of the MUAS enhancer upstream of the SD3 core promoter fragment:
TTCGTCCACAGACATCAACATCTTATCGTCCTTTGAAGATAAGATAATAATGTTGAAGATAAGAGTGGGAGCCACCACTAAAACATTGCTTTGTCAAAAGCTAAAAAAGATGATGCCCGACAGCCACTTGTGTGAAGCATGTGAAGCCGGTCCCTCCACTAAGAAAATTAGTGAAGCATCTTCCAGTGGTCCCTCCACTCACAGCTCAATCAGTGAGCAACAGGACGAAGGAAATGACGTAAGCCATGACGTCTAATCCCAAGAGTGCCACACAAGGGGCGACAACGTCGAAGGCGCAGAAGACGCAGTCGATCTCACTGACGTAAGCAATGACGACCAGTGGAGGAGATCGTAAGCAATGACGTATGGAGCGTGGAGGACCCATGAAAGCACTGAGAAGGCATCTCAACTTTCGGTGTGTGAGTGCGCATCCTATGCGATGCTTTGTACCTTTGTTAGCTGTGTGTGTCCTTTTGGCATCTGTGCCACTTTACCTTTGTCGGCCACGTTGCCTTTGCTTAGCATCTACGCAAGCATAGCGCTCGGCTGGTGTGTGTTCCCTCTGCCTATATAAGGCATGGTTGTATGACTCTTACACTCATCGGTAGTTCACCACATGAGTATTTGAGTCAAGTTTGGCTTGAATAATAAGAATTACACCTTTCCGCAA
M24 synthetic promoter (MMV-derived):
The M24 promoter is a synthetic high-expression promoter derived from the Mirabilis mosaic virus (MMV), as described by (Sahoo et al., 2014). It was engineered to enhance transcriptional activity in plant systems. Based on the full-length transcript (FLt) promoter of MMV, the promoter was enhanced by duplication of upstream enhancer domains, leading to a significant increase in transcriptional strength.
The M24 promoter sequence was retrieved from the binary vector pSiM24 available in GenBank (accession: KF032933.1). The promoter corresponds to the region spanning positions 235–860 of the vector sequence.
KF032933.1:235-860 Binary vector pSiM24, complete sequence
TTCGTCCACAGACATCAACATCTTATCGTCCTTTGAAGATAAGATAATAATGTTGAAGATAAGAGTGGGAGCCACCACTAAAACATTGCTTTGTCAAAAGCTAAAAAAGATGATGCCCGACAGCCACTTGTGTGAAGCATGTGAAGCCGGTCCCTCCACTAAGAAAATTAGTGAAGCATCTTCCAGTGGTCCCTCCACTCACAGCTCAATCAGTGAGCAACAGGACGAAGGAAATGACGTAAGCCATGACGTCTAATCCCCCCAACTTCGTCCACAGACATCAACATCTTATCGTCCTTTGAAGATAAGATAATAATGTTGAAGATAAGAGTGGGAGCCACCACTAAAACATTGCTTTGTCAAAAGCTAAAAAAGATGATGCCCGACAGCCACTTGTGTGAAGCATGTGAAGCCGGTCCCTCCACTAAGAAAATTAGTGAAGCATCTTCCAGTGGTCCCTCCACTCACAGCTCAATCAGTGAGCAACAGGACGAAGGAAATGACGTAAGCCATGACGTCTAATCCCACAAGAATTTCCTTATATAAGGAACACAAATCAGAAGGAAGAGATCAATCGAAATCAAAATCGGAATCGAAATCAAAATCGGAATCGAAATCTCTCATCT
PClSV FLt promoter (Peanut chlorotic streak caulimovirus):
The PClSV FLt promoter is a constitutive plant promoter derived from the Peanut chlorotic streak caulimovirus. It is composed of a basic full-length transcript (FLt) promoter region and upstream enhancer elements, which can be arranged in single or duplicated configurations to modulate transcriptional strength.
The promoter elements were identified from the PClSV genome (GenBank accession: U13988.1) as follows:
- Basic FLt promoter (core region):
Spans positions 5852–6101 (~250 bp) and contains essential elements required for transcription initiation
> gb|U13988.1|PCU13988:5852-6101 Peanut chlorotic streak caulimovirus, complete genome
GAGATCTTGAGCCAATCAAAGAGGAGTGATGTAGACCTAAAGCAATAATGGAGCCATGACGTAAGGGCTTACGCCATTACGAAATAATTAAAGGCTGATGTGACCTGTCGGTCTCTCAGAACCTTTACTTTTTATATTTGGCGTGTATTTTTAAATTTCCACGGCAATGACGATGTGACCTGTGCATCCGCTTTGCCTATAAATAAGTTTTAGTTTGTATTGATCGACACGATCGAGAAGACACGGCCAT
- Enhancer element:
A 178 bp upstream regulatory sequence (5852–6029) responsible for increasing transcriptional activity
> gb|U13988.1|PCU13988:5852-6029 Peanut chlorotic streak caulimovirus, complete genome
GAGATCTTGAGCCAATCAAAGAGGAGTGATGTAGACCTAAAGCAATAATGGAGCCATGACGTAAGGGCTTACGCCATTACGAAATAATTAAAGGCTGATGTGACCTGTCGGTCTCTCAGAACCTTTACTTTTTATATTTGGCGTGTATTTTTAAATTTCCACGGCAATGACGATGTGA
The assembled PClSV FLt promoter [Enhancer] + [Core promoter] sequence:
GAGATCTTGAGCCAATCAAAGAGGAGTGATGTAGACCTAAAGCAATAATGGAGCCATGACGTAAGGGCTTACGCCATTACGAAATAATTAAAGGCTGATGTGACCTGTCGGTCTCTCAGAACCTTTACTTTTTATATTTGGCGTGTATTTTTAAATTTCCACGGCAATGACGATGTGAGAGATCTTGAGCCAATCAAAGAGGAGTGATGTAGACCTAAAGCAATAATGGAGCCATGACGTAAGGGCTTACGCCATTACGAAATAATTAAAGGCTGATGTGACCTGTCGGTCTCTCAGAACCTTTACTTTTTATATTTGGCGTGTATTTTTAAATTTCCACGGCAATGACGATGTGACCTGTGCATCCGCTTTGCCTATAAATAAGTTTTAGTTTGTATTGATCGACACGATCGAGAAGACACGGCCAT
Double enhancer PCisV FLt promoter:
Based on (Maiti & Shepherd, 1998), the double enhancer configuration was constructed by duplicating the enhancer region upstream of the core promoter: [Enhancer] + [Enhancer] + [Core promoter] (~428 bp)
The PClSV FLt promoter sequence was reconstructed from GenBank (U13988.1) and assembled in a double enhancer configuration based on the design described by Maiti & Shepherd (1998):
GAGATCTTGAGCCAATCAAAGAGGAGTGATGTAGACCTAAAGCAATAATGGAGCCATGACGTAAGGGCTTACGCCATTACGAAATAATTAAAGGCTGATGTGACCTGTCGGTCTCTCAGAACCTTTACTTTTTATATTTGGCGTGTATTTTTAAATTTCCACGGCAATGACGATGTGAGAGATCTTGAGCCAATCAAAGAGGAGTGATGTAGACCTAAAGCAATAATGGAGCCATGACGTAAGGGCTTACGCCATTACGAAATAATTAAAGGCTGATGTGACCTGTCGGTCTCTCAGAACCTTTACTTTTTATATTTGGCGTGTATTTTTAAATTTCCACGGCAATGACGATGTGAGAGATCTTGAGCCAATCAAAGAGGAGTGATGTAGACCTAAAGCAATAATGGAGCCATGACGTAAGGGCTTACGCCATTACGAAATAATTAAAGGCTGATGTGACCTGTCGGTCTCTCAGAACCTTTACTTTTTATATTTGGCGTGTATTTTTAAATTTCCACGGCAATGACGATGTGACCTGTGCATCCGCTTTGCCTATAAATAAGTTTTAGTTTGTATTGATCGACACGATCGAGAAGACACGGCCAT
The double enhancer configuration of the PClSV FLt promoter results in an approximately threefold increase in transcriptional activity compared to the single enhancer version. Overall, this promoter exhibits strong constitutive expression in transgenic plants, with activity levels reported to be comparable to the FLt promoter of the Figwort mosaic virus and functionally similar to the widely used CaMV 35S promoter, making it a robust alternative for high-level gene expression in plant systems.
CVP1 and CVP2 promoters (Cassava vein mosaic virus, CsVMV):
The CVP1 and CVP2 promoters are constitutive plant promoters derived from the Cassava vein mosaic virus (CsVMV), as described by Verdaguer et al., (1996) and Verdaguer et al., (1998) based on the reference genome reported by Calvert et al., (1995). These promoters correspond to two fragments of different lengths within the viral genome and differ in their regulatory strength.
- CVP1 (short fragment): corresponds to a 388 bp fragment spanning nucleotides 7235 to 7623, which maps to the region −368 to +20 relative to the transcription start site (TSS).
- CVP2 (long fragment): represents a longer 511 bp fragment extending from nucleotides 7160 to 7675, corresponding to positions −443 to +72 relative to the TSS.
Both fragments contain core promoter elements, including the TATA box and upstream regulatory motifs, with CVP2 retaining additional upstream sequences that enhance transcriptional activity.
The sequences were directly retrieved from the CsVMV reference genome (GenBank accession: U20341.1) using the genomic coordinates reported in the original studies:
CPV 1 :
>gb|U20341.1|CVU20341:7235-7623 Cassava vein mosaic virus, complete genome
GCTCAGCAAGAAGCAGATCAATATGCGGCACATATGCAACCTATGTTCAAAAATGAAGAATGTACAGATACAAGATCCTATACTGCCAGAATACGAAGAAGAATACGTAGAAATTGAAAAAGAAGAACCAGGCGAAGAAAAGAATCTTGAAGACGTAAGCACTGACGACAACAATGAAAAGAAGAAGATAAGGTCGGTGATTGTGAAAGAGACATAGAGGACACATGTAAGGTGGAAAATGTAAGGGCGGAAAGTAACCTTATCACAAAGGAATCTTATCCCCCACTACTTATCCTTTTATATTTTTCCGTGTCATTTTTGCCCTTGAGTTTTCCTATATAAGGAACCAAGTTCGGCATTTGTGAAAACAAGAAAAAATTTGGTGTAAG
CPV 2 :
>gb|U20341.1|CVU20341:7160-7675 Cassava vein mosaic virus, complete genome
TCCAGAAGGTAATTATCCAAGATGTAGCATCAAGAATCCAATGTTTACGGGAAAAACTATGGAAGTATTATGTGAGCTCAGCAAGAAGCAGATCAATATGCGGCACATATGCAACCTATGTTCAAAAATGAAGAATGTACAGATACAAGATCCTATACTGCCAGAATACGAAGAAGAATACGTAGAAATTGAAAAAGAAGAACCAGGCGAAGAAAAGAATCTTGAAGACGTAAGCACTGACGACAACAATGAAAAGAAGAAGATAAGGTCGGTGATTGTGAAAGAGACATAGAGGACACATGTAAGGTGGAAAATGTAAGGGCGGAAAGTAACCTTATCACAAAGGAATCTTATCCCCCACTACTTATCCTTTTATATTTTTCCGTGTCATTTTTGCCCTTGAGTTTTCCTATATAAGGAACCAAGTTCGGCATTTGTGAAAACAAGAAAAAATTTGGTGTAAGCTATTTTCTTTGAAGTACTGAGGATACAACTTCAGAGAAATTTGTAAGTTTG
Functional analyses have demonstrated that CVP2 exhibits expression levels comparable to the enhanced CaMV 35S promoter (e35S), whereas CVP1 shows approximately half of this activity, indicating that CVP2 is about twofold more active than CVP1. These results highlight the importance of additional upstream regulatory sequences in driving stronger gene expression in plant systems.
FMV Sgt (34S) promoter (Figwort mosaic virus):
The Sgt (34S) promoter is a subgenomic promoter derived from the Figwort mosaic virus (FMV). It is located between ORF V and ORF VI and is responsible for driving the expression of ORF VI via a subgenomic transcript.
According to (Bhattacharyya et al., 2002) , a 301 bp fragment spanning −270 to +31 relative to the transcription start site (TSS) provides maximal promoter activity.
The promoter sequence was extracted from the published figure using an AI tool (Gemini), as it was only available in image format:
.png?width=500px)
.png?width=500px)
TTTACAGTAAGAACTGATAACAAAAATTTTACTTATTTCCTTAGAATTAATCTTAAAGGTGATAGTAAACAAGGACGATTAGTCCGTTGGCAAAATTGGTTCAGCAAGTATCAATTTGATGTCGAACATCTTGAAGGTGTAAAAAACGTTTTAGCAGATTGCCTCACGAGAGATTTTAATGCTTAAAAACGTAAGCGCTGACGTATGATTTCAAAAAACGCAGCTATAAAAGAAGCCCTCCAGCTTCAAAGTTTTCATCAACACAAATTCTAAAAACAAAATTTTTAGAGAGGGGGAGTG
PTSB1 promoter (Arabidopsis thaliana):
The PTSB1 promoter is a constitutive plant promoter I derived from the Arabidopsis thaliana tryptophan synthase β-subunit gene (TSB1). I identified this as a powerful alternative to the CaMV 35S promoter for high-level gene expression in tobacco (Shirasawa-Seo et al. 2002).
I retrieved this promoter from GenBank accession M23872, corresponding to a 1.5 kb fragement. I defined the exact boundaries of this fragment by mapping the reported PCR primers directly onto the reference sequence (Shirasawa-Seo et al. 2002):
- 5’ Border (Forward primer): GAATTCTTTCATATCTCCTGCAAAGT
- 3’ Border (Reverse primer): TCAGAGAGAGATTCATTCAGTA (This is the reverse complement of the primer sequence TACTGAATGAATCTCTCTCTGA listed in the sources.)
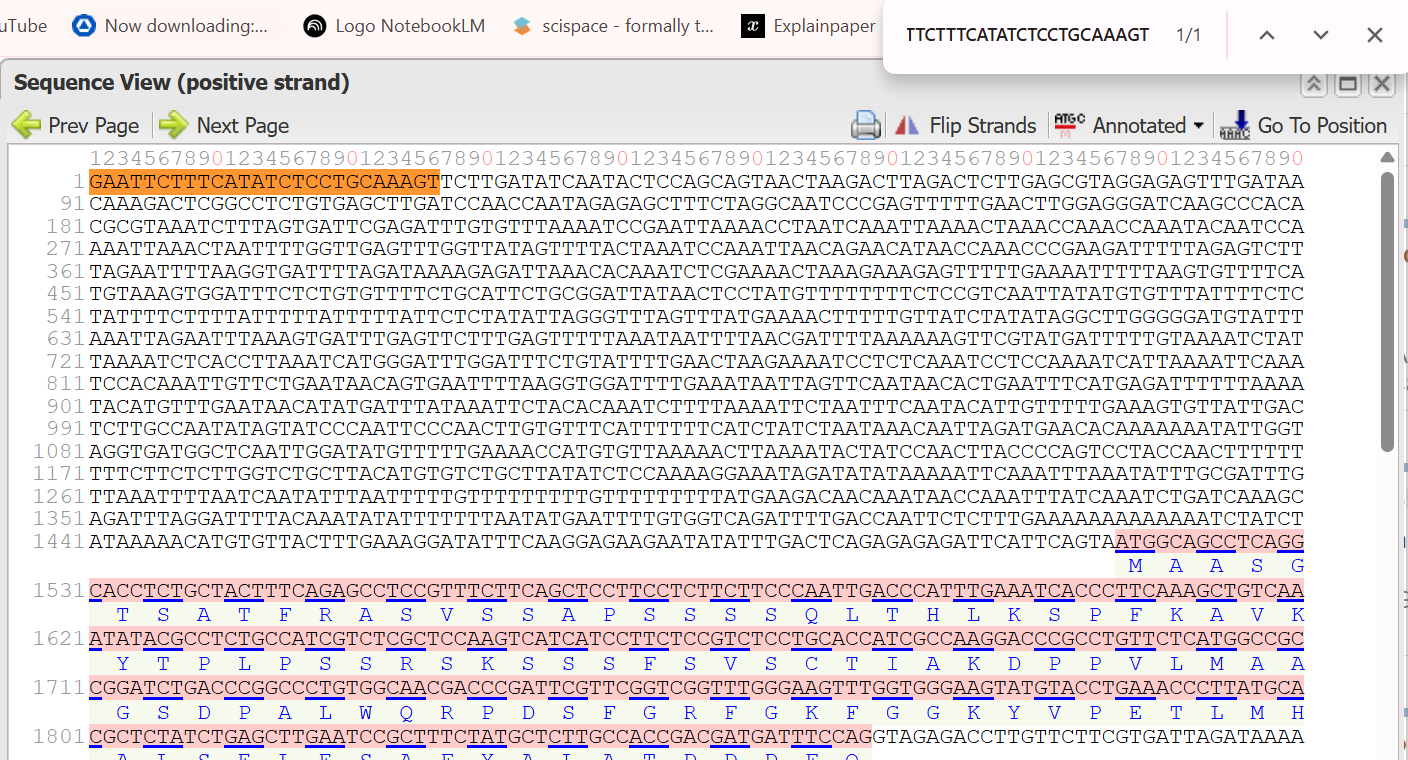

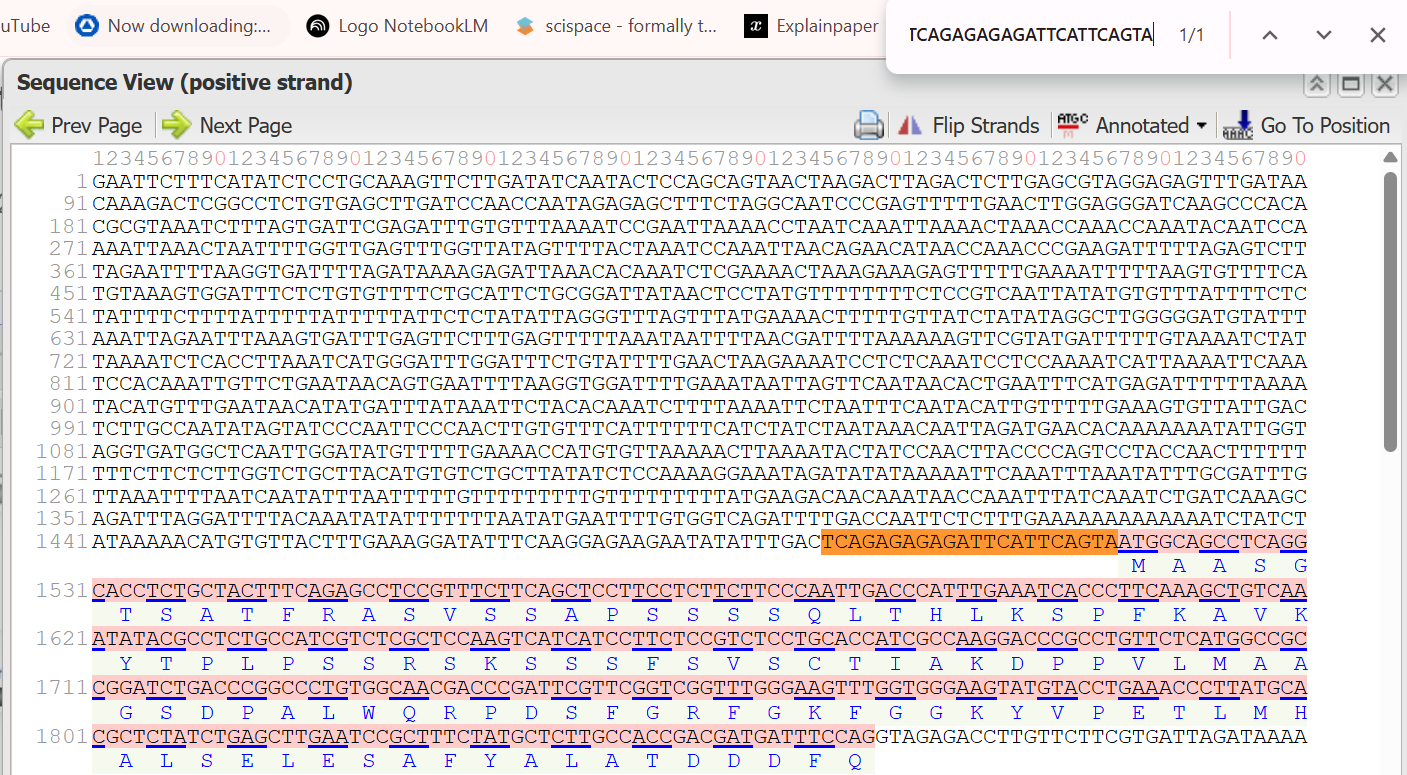
 The resulted extracted sequence of PTSB1 promoter:
The resulted extracted sequence of PTSB1 promoter:
GAATTCTTTCATATCTCCTGCAAAGTTCTTGATATCAATACTCCAGCAGTAACTAAGACTTAGACTCTTGAGCGTAGGAGAGTTTGATAACAAAGACTCGGCCTCTGTGAGCTTGATCCAACCAATAGAGAGCTTTCTAGGCAATCCCGAGTTTTTGAACTTGGAGGGATCAAGCCCACACGCGTAAATCTTTAGTGATTCGAGATTTGTGTTTAAAATCCGAATTAAAACCTAATCAAATTAAAACTAAACCAAACCAAATACAATCCAAAATTAAACTAATTTTGGTTGAGTTTGGTTATAGTTTTACTAAATCCAAATTAACAGAACATAACCAAACCCGAAGATTTTTAGAGTCTTTAGAATTTTAAGGTGATTTTAGATAAAAGAGATTAAACACAAATCTCGAAAACTAAAGAAAGAGTTTTTGAAAATTTTTAAGTGTTTTCATGTAAAGTGGATTTCTCTGTGTTTTCTGCATTCTGCGGATTATAACTCCTATGTTTTTTTTCTCCGTCAATTATATGTGTTTATTTTCTCTATTTTCTTTTATTTTTATTTTTATTCTCTATATTAGGGTTTAGTTTATGAAAACTTTTTGTTATCTATATAGGCTTGGGGGATGTATTTAAATTAGAATTTAAAGTGATTTGAGTTCTTTGAGTTTTTAAATAATTTTAACGATTTTAAAAAAGTTCGTATGATTTTTGTAAAATCTATTAAAATCTCACCTTAAATCATGGGATTTGGATTTCTGTATTTTGAACTAAGAAAATCCTCTCAAATCCTCCAAAATCATTAAAATTCAAATCCACAAATTGTTCTGAATAACAGTGAATTTTAAGGTGGATTTTGAAATAATTAGTTCAATAACACTGAATTTCATGAGATTTTTTAAAATACATGTTTGAATAACATATGATTTATAAATTCTACACAAATCTTTTAAAATTCTAATTTCAATACATTGTTTTTGAAAGTGTTATTGACTCTTGCCAATATAGTATCCCAATTCCCAACTTGTGTTTCATTTTTTCATCTATCTAATAAACAATTAGATGAACACAAAAAAATATTGGTAGGTGATGGCTCAATTGGATATGTTTTTGAAAACCATGTGTTAAAAACTTAAAATACTATCCAACTTACCCCAGTCCTACCAACTTTTTTTTTCTTCTCTTGGTCTGCTTACATGTGTCTGCTTATATCTCCAAAAGGAAATAGATATATAAAAATTCAAATTTAAATATTTGCGATTTGTTAAATTTTAATCAATATTTAATTTTTGTTTTTTTTTGTTTTTTTTTATGAAGACAACAAATAACCAAATTTATCAAATCTGATCAAAGCAGATTTAGGATTTTACAAATATATTTTTTTAATATGAATTTTGTGGTCAGATTTTGACCAATTCTCTTTGAAAAAAAAAAAAATCTATCTATAAAAACATGTGTTACTTTGAAAGGATATTTCAAGGAGAAGAATATATTTGACTCAGAGAGAGATTCATTCAGTA
This region contains the core promoter and upstream regulatory elements responsible for its strong constitutive activity.
This promoter exhibited approximately 2.4-fold higher expression than the CaMV 35S promoter in mature tobacco leaves, with activity increasing in lower leaf positions (Shirasawa-Seo et al. 2002).
PPHYB promoter (Arabidopsis thaliana):
The PPHYB promoter is a constitutive promoter derived from the Arabidopsis thaliana phytochrome B (PHYB) gene (Goosey et al. 1997; Shirasawa-Seo et al. 2002).
I retrieved this sequence from GenBank accession L09262, which corresponds to a 2.3 kb fragment. The promoter boundaries were defined by mapping the experimentally reported primers onto the sequence (Shirasawa-Seo et al. 2002):
- 5’ Border (Forward primer): GTCGACTTGTGCACCACCGTCT
- 3’ Border (Reverse primer): CGGAGAAGAAGAACCGTCGTCA (This is the reverse complement of the primer sequence TGACGACGGTTCTTCTTCTCCG listed in the sources.)
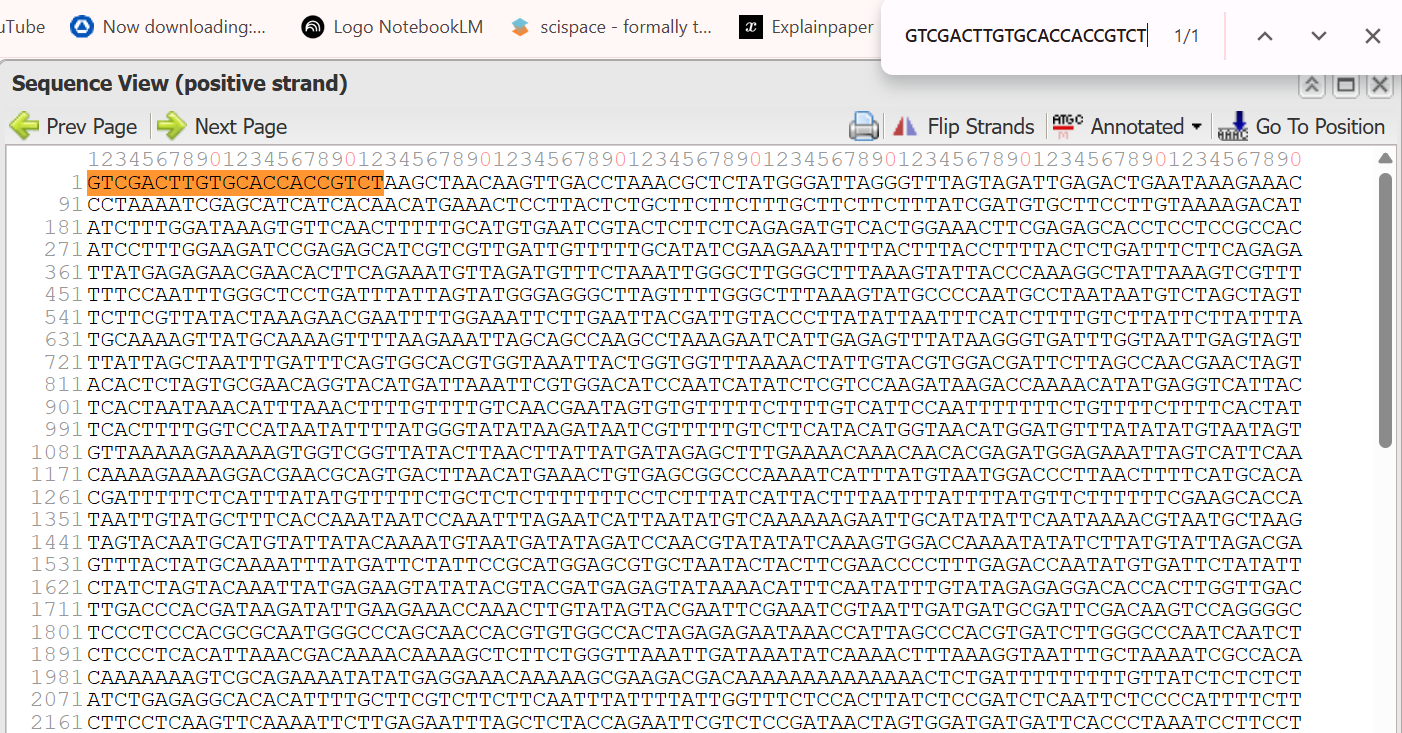

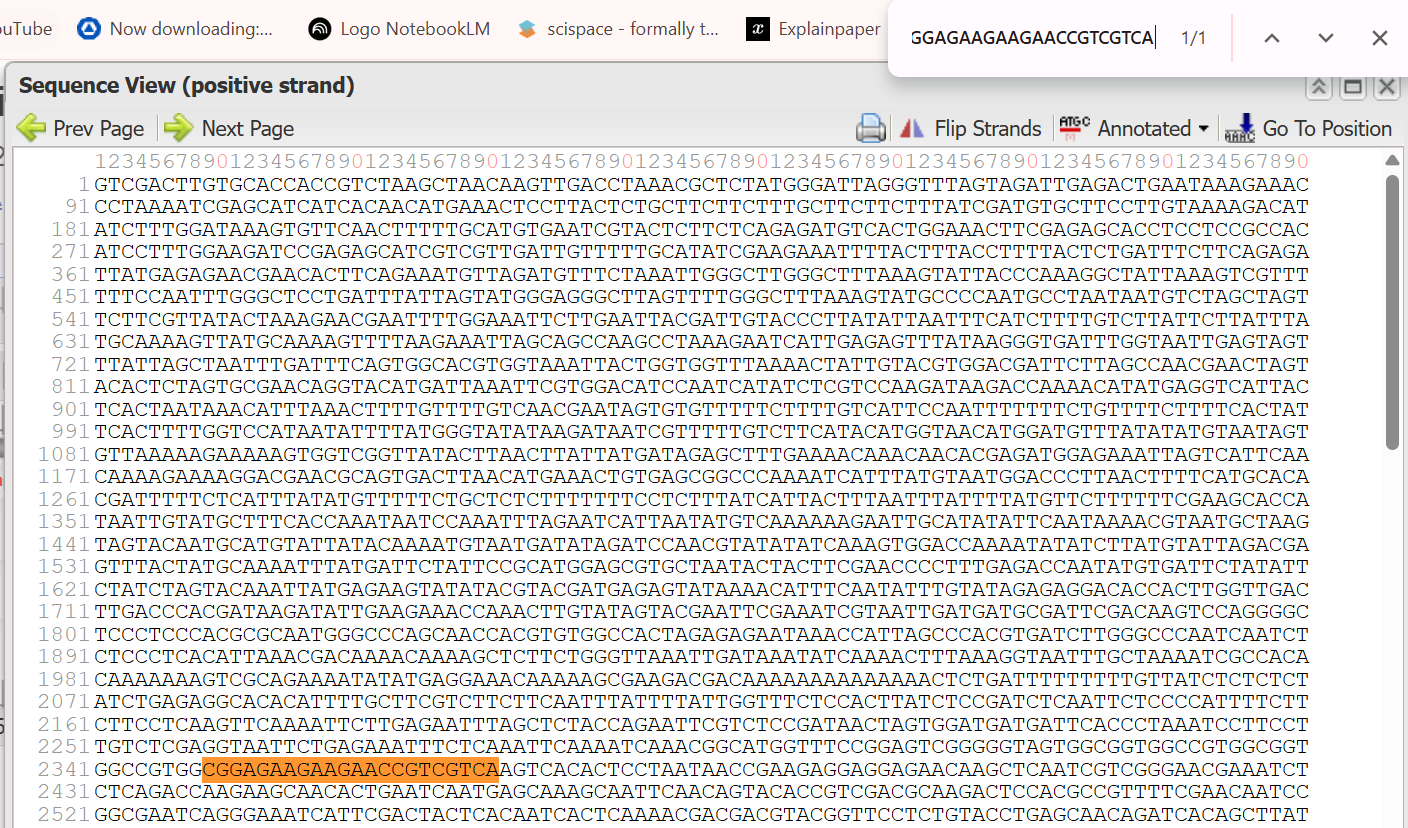
 The resulted extracted sequence of PPHYB promoter:
The resulted extracted sequence of PPHYB promoter:
GTCGACTTGTGCACCACCGTCTAAGCTAACAAGTTGACCTAAACGCTCTATGGGATTAGGGTTTAGTAGATTGAGACTGAATAAAGAAACCCTAAAATCGAGCATCATCACAACATGAAACTCCTTACTCTGCTTCTTCTTTGCTTCTTCTTTATCGATGTGCTTCCTTGTAAAAGACATATCTTTGGATAAAGTGTTCAACTTTTTGCATGTGAATCGTACTCTTCTCAGAGATGTCACTGGAAACTTCGAGAGCACCTCCTCCGCCACATCCTTTGGAAGATCCGAGAGCATCGTCGTTGATTGTTTTTGCATATCGAAGAAATTTTACTTTACCTTTTACTCTGATTTCTTCAGAGATTATGAGAGAACGAACACTTCAGAAATGTTAGATGTTTCTAAATTGGGCTTGGGCTTTAAAGTATTACCCAAAGGCTATTAAAGTCGTTTTTTCCAATTTGGGCTCCTGATTTATTAGTATGGGAGGGCTTAGTTTTGGGCTTTAAAGTATGCCCCAATGCCTAATAATGTCTAGCTAGTTCTTCGTTATACTAAAGAACGAATTTTGGAAATTCTTGAATTACGATTGTACCCTTATATTAATTTCATCTTTTGTCTTATTCTTATTTATGCAAAAGTTATGCAAAAGTTTTAAGAAATTAGCAGCCAAGCCTAAAGAATCATTGAGAGTTTATAAGGGTGATTTGGTAATTGAGTAGTTTATTAGCTAATTTGATTTCAGTGGCACGTGGTAAATTACTGGTGGTTTAAAACTATTGTACGTGGACGATTCTTAGCCAACGAACTAGTACACTCTAGTGCGAACAGGTACATGATTAAATTCGTGGACATCCAATCATATCTCGTCCAAGATAAGACCAAAACATATGAGGTCATTACTCACTAATAAACATTTAAACTTTTGTTTTGTCAACGAATAGTGTGTTTTTCTTTTGTCATTCCAATTTTTTTCTGTTTTCTTTTCACTATTCACTTTTGGTCCATAATATTTTATGGGTATATAAGATAATCGTTTTTGTCTTCATACATGGTAACATGGATGTTTATATATGTAATAGTGTTAAAAAGAAAAAGTGGTCGGTTATACTTAACTTATTATGATAGAGCTTTGAAAACAAACAACACGAGATGGAGAAATTAGTCATTCAACAAAAGAAAAGGACGAACGCAGTGACTTAACATGAAACTGTGAGCGGCCCAAAATCATTTATGTAATGGACCCTTAACTTTTCATGCACACGATTTTTCTCATTTATATGTTTTTCTGCTCTCTTTTTTTCCTCTTTATCATTACTTTAATTTATTTTATGTTCTTTTTTCGAAGCACCATAATTGTATGCTTTCACCAAATAATCCAAATTTAGAATCATTAATATGTCAAAAAAGAATTGCATATATTCAATAAAACGTAATGCTAAGTAGTACAATGCATGTATTATACAAAATGTAATGATATAGATCCAACGTATATATCAAAGTGGACCAAAATATATCTTATGTATTAGACGAGTTTACTATGCAAAATTTATGATTCTATTCCGCATGGAGCGTGCTAATACTACTTCGAACCCCTTTGAGACCAATATGTGATTCTATATTCTATCTAGTACAAATTATGAGAAGTATATACGTACGATGAGAGTATAAAACATTTCAATATTTGTATAGAGAGGACACCACTTGGTTGACTTGACCCACGATAAGATATTGAAGAAACCAAACTTGTATAGTACGAATTCGAAATCGTAATTGATGATGCGATTCGACAAGTCCAGGGGCTCCCTCCCACGCGCAATGGGCCCAGCAACCACGTGTGGCCACTAGAGAGAATAAACCATTAGCCCACGTGATCTTGGGCCCAATCAATCTCTCCCTCACATTAAACGACAAAACAAAAGCTCTTCTGGGTTAAATTGATAAATATCAAAACTTTAAAGGTAATTTGCTAAAATCGCCACACAAAAAAAGTCGCAGAAAATATATGAGGAAACAAAAAGCGAAGACGACAAAAAAAAAAAAAACTCTGATTTTTTTTTGTTATCTCTCTCTATCTGAGAGGCACACATTTTGCTTCGTCTTCTTCAATTTATTTTATTGGTTTCTCCACTTATCTCCGATCTCAATTCTCCCCATTTTCTTCTTCCTCAAGTTCAAAATTCTTGAGAATTTAGCTCTACCAGAATTCGTCTCCGATAACTAGTGGATGATGATTCACCCTAAATCCTTCCTTGTCTCGAGGTAATTCTGAGAAATTTCTCAAATTCAAAATCAAACGGCATGGTTTCCGGAGTCGGGGGTAGTGGCGGTGGCCGTGGCGGTGGCCGTGGCGGAGAAGAAGAACCGTCGTCA
This fragment includes the core promoter and regulatory regions required for stable expression.
Functionally, PPHYB provides approximately 1.5-fold higher expression than the CaMV 35S promoter in mature tobacco leaves, with a more uniform expression pattern across leaf positions compared to PTSB1 (Shirasawa-Seo et al. 2002).
PNCR promoter (Soybean chlorotic mottle virus):
The PNCR promoter is a viral-derived constitutive promoter isolated from the large noncoding region of the Soybean chlorotic mottle virus (Conci et al. 1993).
Based on the reported genome size (~8,175 bp), I identified the corresponding genomic sequence and retrieved it from GenBank accession X15828.2.
I then defined the functional ~486 bp promoter fragment by mapping the reported PCR primers onto the genome (Conci et al. 1993):
- 5’ Border (Forward primer): ATGTAGGACATGCCAGCTGTAA
- 3’ Border (Reverse primer): CAAGCACAAGAGAAAAGAAAGG (Note: This is the reverse complement of the primer sequence CCGGATCCTTTCTTTTCTCTTGTGCTTG provided in the source, after removing the restriction enzyme site.):
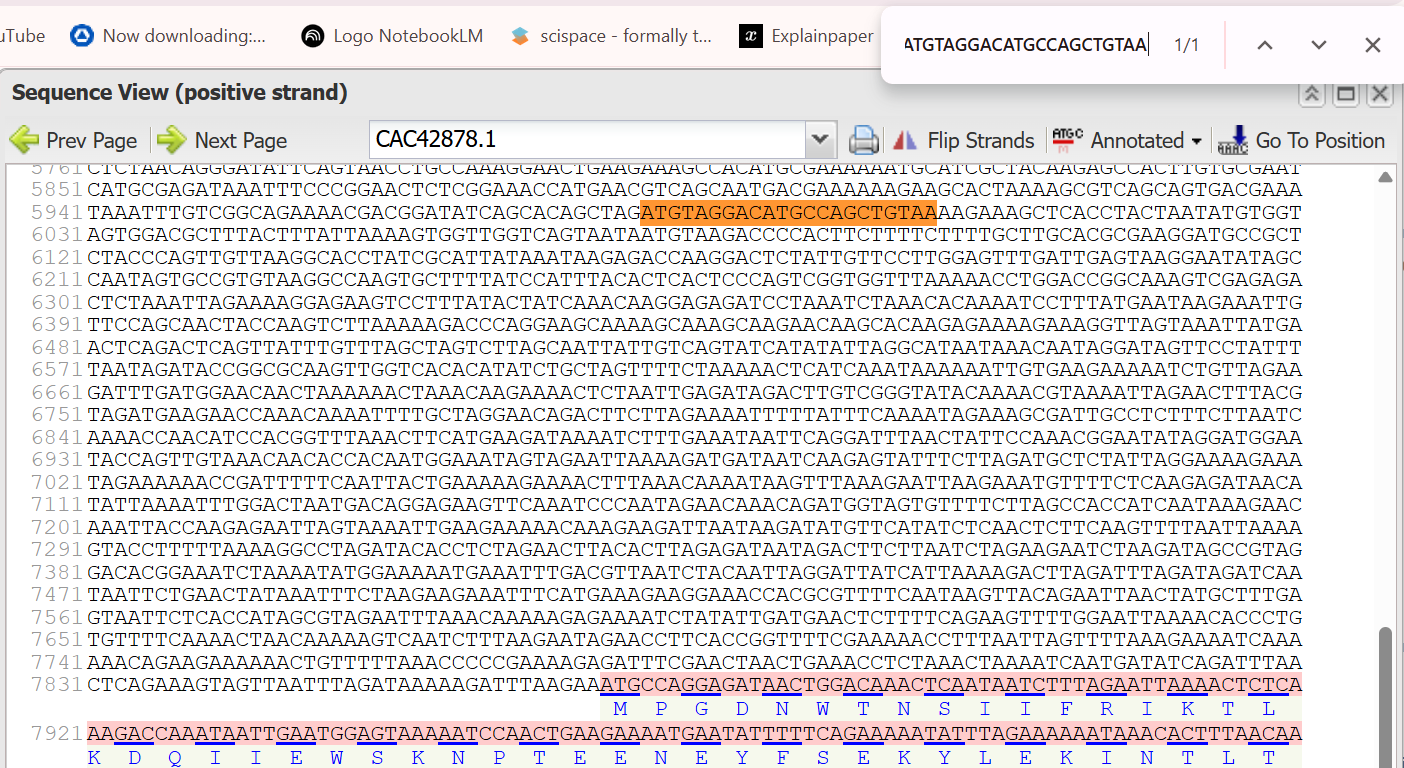

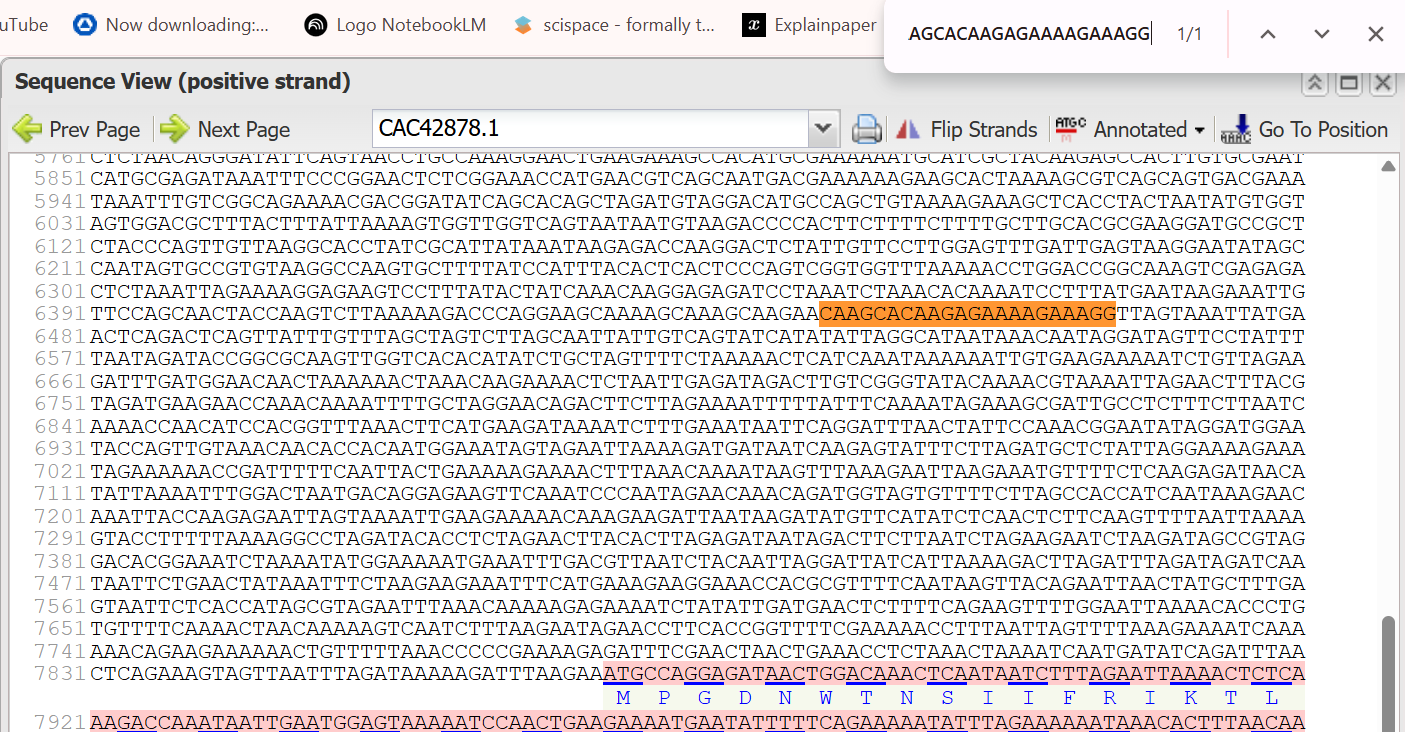

The extracted sequence of PNCR promoter:
ATGTAGGACATGCCAGCTGTAAAAGAAAGCTCACCTACTAATATGTGGTAGTGGACGCTTTACTTTATTAAAAGTGGTTGGTCAGTAATAATGTAAGACCCCACTTCTTTTCTTTTGCTTGCACGCGAAGGATGCCGCTCTACCCAGTTGTTAAGGCACCTATCGCATTATAAATAAGAGACCAAGGACTCTATTGTTCCTTGGAGTTTGATTGAGTAAGGAATATAGCCAATAGTGCCGTGTAAGGCCAAGTGCTTTTATCCATTTACACTCACTCCCAGTCGGTGGTTTAAAAACCTGGACCGGCAAAGTCGAGAGACTCTAAATTAGAAAAGGAGAAGTCCTTTATACTATCAAACAAGGAGAGATCCTAAATCTAAACACAAAATCCTTTATGAATAAGAAATTGTTCCAGCAACTACCAAGTCTTAAAAAGACCCAGGAAGCAAAAGCAAAGCAAGAACAAGCACAAGAGAAAAGAAAGG
This region contains key regulatory features including a TATA box, CAAT-like motifs, and multiple enhancer-related elements.
Functionally, this promoter exhibits approximately five-fold higher expression than the CaMV 35S promoter in tobacco protoplasts (Conci et al. 1993), while showing moderate constitutive activity (~67% of P35S) in mature leaves (Shirasawa-Seo et al. 2002).
FMV promoter (Figwort mosaic virus):
The FMV promoter is a constitutive viral promoter derived from the Figwort mosaic virus genome. In this work, I used the promoter sequence obtained directly from the supplementary Benchling file provided in (Shakhova et al., 2022):
tcatcaaaatatttagcagcattccagattgggttcaatcaacaaggtacgagccatatcactttattcaaattggtatcgccaaaaccaagaaggaactcccatcctcaaaggtttgtaaggaagaattctcagtccaaagcctcaacaaggtcagggtacagagtctccaaaccattagccAaaagctacaggagatcaatgaagaatcttcaatcaaagtaaactactgttccagcacatgcatcatggtcagtaagtttcagaaaaagacatccaccgaGgacttaaagttagtgggcatctttgaaagtaatcttgtcaacatcgagcagctggcttgtggggaccagacaaaaaaggaatggtgcagaattgttaggcgcacctaccaaaagcatctttgcctttattgcaaagataaagcagattcctctagtacaagtggggaacaaaataacgtggaaaagagctgtcctgacagcccactcactaatgcgtatgacgaacgcagtgacgaccacaaaagaattccctctatataagaaggcattcattcccatttgaaggatcatcagatactGaaccaatatttctc
To verify its genomic origin, I performed a BLAST analysis using the NCBI nblast, and obtained a 100% sequence match corresponding to coordinates 6358 to 6955 of the reference genome (GenBank accession NC_003554.1), confirming the exact location of the promoter fragment within the FMV genome.
According to (Shakhova et al., 2022), the FMV promoter exhibited lower activity compared to the CaMV 35S promoter under their experimental conditions, indicating that while it remains a functional constitutive promoter, it is not as strong as p35S in this specific system.
p35S (CAMV 35S promoter):
The p35S promoter is a canonical constitutive promoter derived from the Cauliflower mosaic virus and is one of the most widely used regulatory elements in plant biotechnology.
In my study, I used the specific p35S sequence provided in the supplementary Benchling file of (Shakhova et al., 2022):
tgagacttttcaacaaaggataatttcgggaaacctcctcggattccattgcccagctatctgtcacttcatcgaaaggacagtagaaaaggaaggtggctcctacaaatgccatcattgcgataaaggaaaggctatcattcaagatctctctgccgacagtggtcccaaagatggacccccacccacgaggagcatcgtggaaaaagaagaggttccaaccacgtctacaaagcaagtggattgatgtgacatctccactgacgtaagggatgacgcacaatcccactatccttcgcaagacccttcctctatataaggaagttcatttcatttggagaggaca
pAtUBQ10 promoter (Arabidopsis thaliana):
The pAtUBQ10 promoter (version 0.8) is a strong constitutive plant promoter derived from the Arabidopsis thaliana ubiquitin-10 gene (At4g05320).
In this work, I used the exact ~800 bp upstream fragment as characterized in (Shakhova et al., 2022).
I obtained the sequence directly from the supplementary Benchling file provided in the study, ensuring that the construct corresponds precisely to the experimentally validated version used for expression analysis:
tgggacccacggttcaattattgccaattttcagctccaccgtatatttaaaaaataaaacgataatgctaaaaaaatataaatcgtaacgatcgttaaatctcaacggctggatcttatgacgaccgttagaaattgtggttgtcgacgagtcagtaataaacggcgtcaaagtggttgcagccggcacacacgagtcgtgtttatcaactcaaagcacaaatacttttcctcaacctaaaaataaggcaattagccaaaaacaactttgcgtgtaaacaacgctcaatacacgtgtcattttattattagctattgcttcaccgccttagctttctcgtgacctagtcgtcctcgtcttttcttcttcttcttctataaaacaatacccaaagagctcttcttcttcacaattcagatttcaatttctcaaaatcttaaaaactttctctcaattctctctaccgtgatcaaggtaaatttctgtgttccttattctctcaaaatcttcgattttgttttcgttcgatcccaatttcgtatatgttctttggtttagattctgttaatcttagatcgaagtcgattttctgggtttgatcgttagatatcatcttaattctcgattagggtttcatagatatcatccgatttgttcaaataatttgagttttgtcgaataattactcttcgatttgtgatttctatctagatctggtgttagtttctagtttgtgcgatcgaatttgtcgattaatctgagtttttctgattaaca
This fragment represents the regulatory region immediately upstream of the translation start site and includes key cis-regulatory elements responsible for its constitutive activity.
Functionally, in Nicotiana systems, this promoter provides high and stable expression levels, outperforming several endogenous plant promoters such as pAtAct2, pAtTCTP, and pAtPD7 (Shakhova et al., 2022). Although its activity is lower than the viral Cauliflower mosaic virus 35S promoter, it shows comparable expression strength to other viral promoters such as Figwort mosaic virus (FMV) and Cotton leaf curl Multan virus (CmYLCV), making it a reliable and predictable option for high-level gene expression in both Nicotiana benthamiana leaves and tobacco BY-2 cell packs.
pAtAct2 promoter (Arabidopsis thaliana):
The pAtAct2 promoter is a constitutive plant promoter derived from the Arabidopsis thaliana actin 2 gene (AT3G18780). In this work, I used the specific version characterized in (Shakhova et al., 2022).
I obtained the sequence directly from the supplementary Benchling file provided in the study, ensuring that the construct corresponds exactly to the experimentally tested version. In this configuration, the native promoter was fused to the 5′UTR omega sequence of the Tobacco mosaic virus (TMV), a common modification used to enhance translation efficiency in Nicotiana expression systems:
tcgacaaaatttagaacgaacttaattatgatctcaaatacattgatacatatctcatctagatctaggttatcattatgtaagaaagttttgacgaatatggcacgacaaaatggctagactcgatgtaattggtatctcaactcaacattatacttataccaaacattagttagacaaaatttaaacaactattttttatgtatgcaagagtcagcatatgtataattgattcagaatcgttttgacgagttcggatgtagtagtagccattatttaatgtacatactaatcgtgaatagtgaatatgatgaaacattgtatcttattgtataaatatccataaacacatcatgaaagacactttctttcacggtctgaattaattatgatacaattctaatagaaaacgaattaaattacgttgaattgtatgaaatctaattgaacaagccaaccacgacgacgactaacgttgcctggattgactcggtttaagttaaccactaaaaaaacggagctgtcatgtaacacgcggatcgagcaggtcacagtcatgaagccatcaaagcaaaagaactaatccaagggctgagatgattaattagtttaaaaattagttaacacgagggaaaaggctgtctgacagccaggtcacgttatctttacctgtggtcgaaatgattcgtgtctgtcgattttaattatttttttgaaaggccgaaaataaagttgtaagagataaacccgcctatataaattcatatattttcctctccgctttgaatactgtatttttac
Functionally, although pAtAct2 is historically described as a strong constitutive promoter in Arabidopsis, the results of (Shakhova et al., 2022) show that it exhibits relatively low activity in tobacco systems. When compared to the 0.4 kb version of the Cauliflower mosaic virus 35S promoter (p35S) used as the reference in this study, pAtAct2 ranks among the weakest promoters in the tested set. This indicates that, despite its native strength in Arabidopsis, pAtAct2 behaves as a moderate-to-low strength promoter in Nicotiana, even after optimization via the TMV omega 5′UTR fusion.
NOS promoter (Agrobacterium tumefaciens nopaline synthase):
The NOS promoter is a constitutive plant promoter derived from the nopaline synthase (nos) gene of Agrobacterium tumefaciens, and is widely used in plant transformation vectors for moderate gene expression.
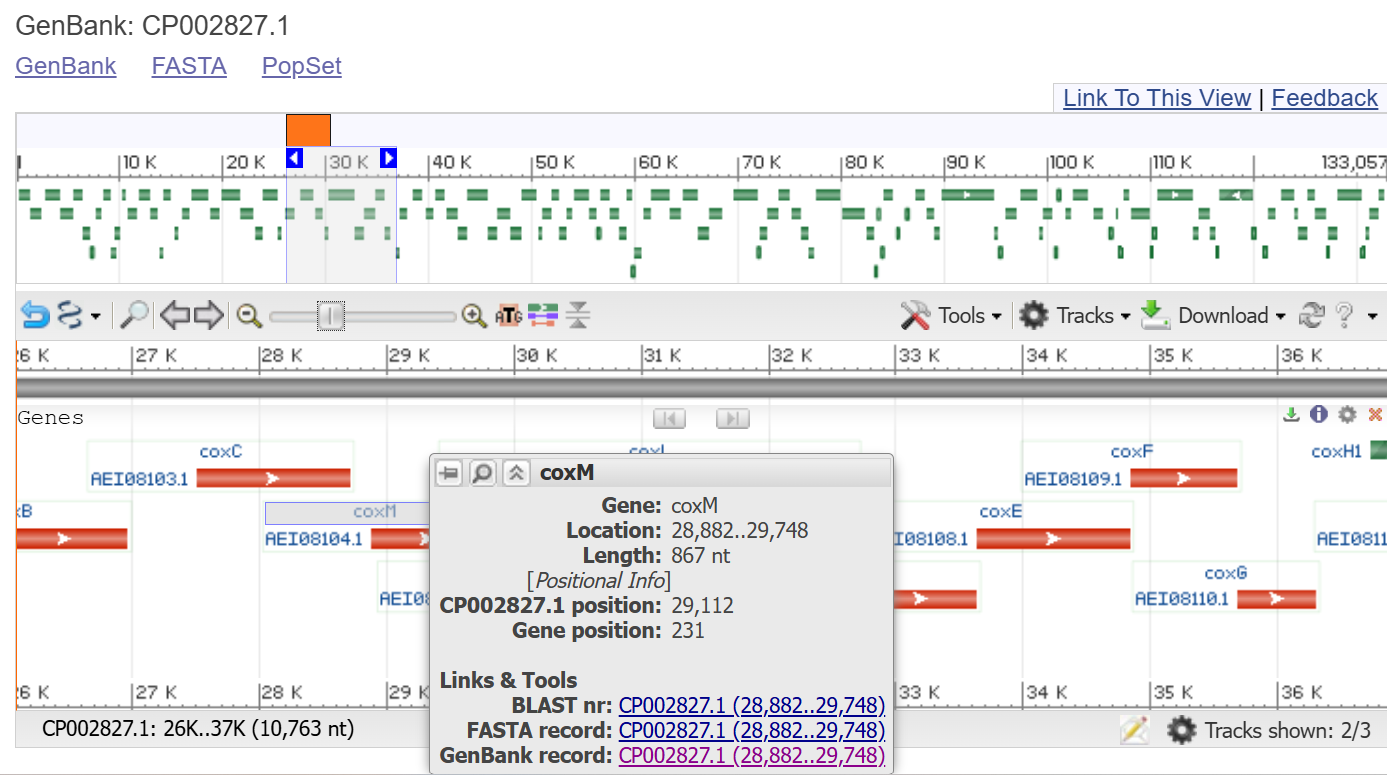
In this work, I retrieved the NOS promoter sequence from GenBank entry AF485783.1, corresponding to the binary vector pBI121, using the coordinates 2519 to 2825. This fragment represents the regulatory region upstream of the nos gene as commonly implemented in plant expression constructs.
The sequence was directly extracted from the annotated GenBank record, ensuring consistency with a well-established and experimentally validated vector backbone frequently used in plant biotechnology.
>AF485783.1:7727-7979 Binary vector pBI121, complete sequence
GATCGTTCAAACATTTGGCAATAAAGTTTCTTAAGATTGAATCCTGTTGCCGGTCTTGCGATGATTATCATATAATTTCTGTTGAATTACGTTAAGCATGTAATAATTAACATGTAATGCATGACGTTATTTATGAGATGGGTTTTTATGATTAGAGTCCCGCAATTATACATTTAATACGCGATAGAAAACAAAATATAGCGCGCAAACTAGGATAAATTATCGCGCGCGGTGTCATCTATGTTACTAGATC
Functionally, the NOS promoter is considered a moderate-to low strength constitutive promoter, typically weaker than strong viral promoters such as the Cauliflower mosaic virus 35S promoter, but valued for its stable and reliable expression across different plant tissues.










.png?width=500px)