Week 2 HW: DNA Reading
I was thinking of choosing between two proteins
- Titin (also known as connectin) which is the largest known protein encoded by the TTN gene. In humans it accounts for 0.5kg of body weight! Titin is important in muscle cells, acting as a molecular spring. It is the third most abundant protein in muscles, giving them their elasticity, structural integrity, and stability.
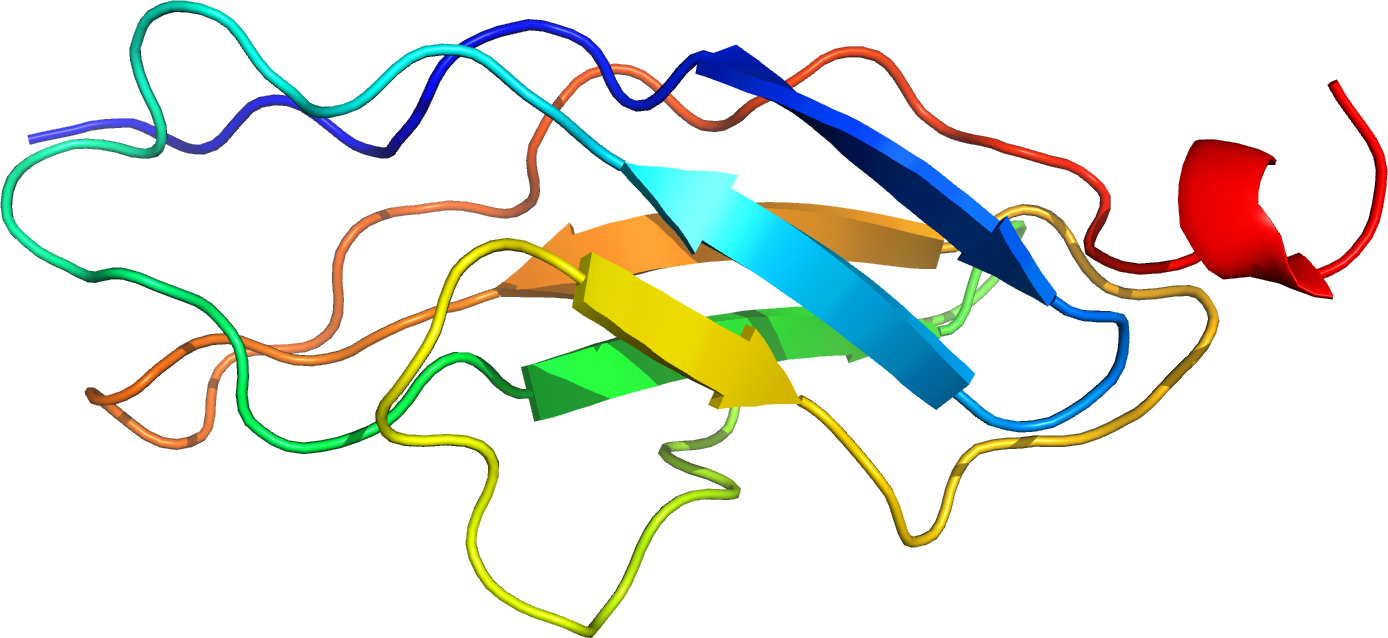

- Green fluorescent protein or GFP which is found in the crystal jelly or hydromedusa (Aequorea Victoria), as well as various species of coral, sea anemones, and crustaceans. GFP is often used as a reporter gene in cell as well as molecular biology.
Scientists have created many organisms which can express GFP which is thusly a proof of concept that a gene can be expressed by a given organism. This protein has been introduced and expressed by many species, maintained in their genome, and even passed on to their offspring; such organisms include bacteria, yeast, fungi, fish, and mammalian cells, including those of humans.
Interestingly, the winners of the 2008 Nobel Prize in Chemistry: Roger Y. Tsien, Osamu Shimomura, and Martin Chalfie, were awarded such due to their discovery and development of GFP.
I decided to choose GFP as it is extremely abundant and familiar to scientists due to its in vivo and in vitro applications, but also because of my interest in creating a bioluminescent biosensor for radiation.
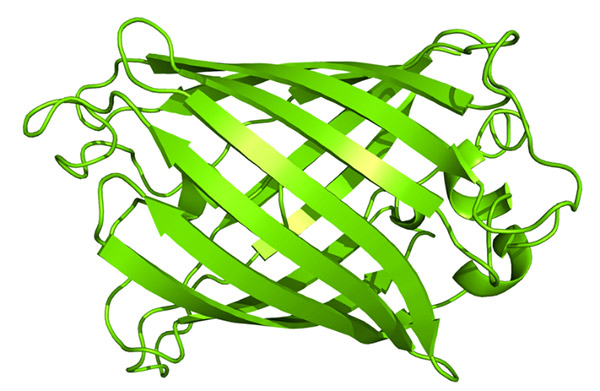
I obtained the protein sequence for such from NCBI:
- https://www.ncbi.nlm.nih.gov/protein/AMQ45836.1
- https://www.ncbi.nlm.nih.gov/protein/AMQ45836.1?report=fasta
Click to view GFP Protein Sequence
MSKGEELFTG VVPILVELDG DVNGHKFSVS GEGEGDATYG KLTLKFICTT GKLPVPWPTL VTTFSYGVQC FSRYPDHMKQ HDFFKSAMPE GYVQERTIFF KDDGNYKTRA EVKFEGDTLV NRIELKGIDF KEDGNILGHK LEYNYNSHNV YIMADKQKNG IKVNFKIRHN IEDGSVQLAD HYQQNTPIGD GPVLLPDNHY LSTQSALSKD PNEKRDHMVL LEFVTAAGIT HGMDELYK
I obtained the DNA (nucleotide) sequence from NCBI also:
- https://www.ncbi.nlm.nih.gov/nuccore/L29345.1
- https://www.ncbi.nlm.nih.gov/nuccore/L29345.1?report=fasta
Click to view GFP DNA Sequence
TACACACGAATAAAAGATAACAAAGATGAGTAAAGGAGAAGAACTTTTCACTGGAGTTGTCCCAATTCTT GTTGAATTAGATGGCGATGTTAATGGGCAAAAATTCTCTGTCAGTGGAGAGGGTGAAGGTGATGCAACAT ACGGAAAACTTACCCTTAAATTTATTTGCACTACTGGGAAGCTACCTGTTCCATGGCCAACACTTGTCAC TACTTTCTCTTATGGTGTTCAATGCTTTTCAAGATACCCAGATCATATGAAACAGCATGACTTTTTCAAG AGTGCCATGCCCGAAGGTTATGTACAGGAAAGAACTATATTTTACAAAGATGACGGGAACTACAAGACAC GTGCTGAAGTCAAGTTTGAAGGTGATACCCTTGTTAATAGAATCGAGTTAAAAGGTATTGATTTTAAAGA AGATGGAAACATTCTTGGACACAAAATGGAATACAACTATAACTCACATAATGTATACATCATGGCAGAC AAACCAAAGAATGGAATCAAAGTTAACTTCAAAATTAGACACAACATTAAAGATGGAAGCGTTCAATTAG CAGACCATTATCAACAAAATACTCCAATTGGCGATGGCCCTGTCCTTTTACCAGACAACCATTACCTGTC CACACAATCTGCCCTTTCCAAAGATCCCAACGAAAAGAGAGATCACATGATCCTTCTTGAGTTTGTAACA GCTGCTGGGATTACACATGGCATGGATGAACTATACAAATAAATGTCCAGACTTCCAATTGACACTAAAG TGTCCGAACAATTACTAAATTCTCAGGGTTCCTGGTTAAATTCAGGCTGAGACTTTATTTATATATTTAT AGATTCATTAAAATTTTATGAATAATTTATTGATGTTATTAATAGGGGCTATTTTCTTATTAAATAGGCT ACTGGAGTGTAT
. . . I then looked for the CDS
Click to view GFP DNA Sequence (CDS)
ATGAGTAAAGGAGAAGAACTTTTCACTGGAGTTGTCCCAATTCTTGTTGAATTAGATGGCGATGTTAATGGGCAAAAATTCTCTGTCAGTGGAGAGGGTGAAGGTGATGCAACATACGGAAAACTTACCCTTAAATTTATTTGCACTACTGGGAAGCTACCTGTTCCATGGCCAACACTTGTCACTACTTTCTCTTATGGTGTTCAATGCTTTTCAAGATACCCAGATCATATGAAACAGCATGACTTTTTCAAGAGTGCCATGCCCGAAGGTTATGTACAGGAAAGAACTATATTTTACAAAGATGACGGGAACTACAAGACACGTGCTGAAGTCAAGTTTGAAGGTGATACCCTTGTTAATAGAATCGAGTTAAAAGGTATTGATTTTAAAGAAGATGGAAACATTCTTGGACACAAAATGGAATACAACTATAACTCACATAATGTATACATCATGGCAGACAAACCAAAGAATGGAATCAAAGTTAACTTCAAAATTAGACACAACATTAAAGATGGAAGCGTTCAATTAGCAGACCATTATCAACAAAATACTCCAATTGGCGATGGCCCTGTCCTTTTACCAGACAACCATTACCTGTCCACACAATCTGCCCTTTCCAAAGATCCCAACGAAAAGAGAGATCACATGATCCTTCTTGAGTTTGTAACAGCTGCTGGGATTACACATGGCATGGATGAACTATACAAATAA
The genetic code is degenerate, meaning that multiple codons can code for the same amino acid. However, certain codons are preferred over others in difference organisms. It is for that reason that we must optimise codon usage depending on the host organism (e.g. human or jellyfish genes in E.coli).
Replacing rare coding with more frequently used ones has the effects of increasing translation speed, improving mRNA stability, and preventing premature translation termination.
I have opted to use E.coli as the host organism for which the codon sequence is optimised for. This is due to the fact that E.coli is abundantly used due to its fast reproduction, low cost, and well charactrised genetics. It is for these reasons that E.coli is an ideal host for producing proteins, espeicially those found in prokaryotes, such as the jellyfish from which GFP is isolated.
I used IDT DNA to optimise my codon sequence
Click to view E. coli Optimised GFP DNA Sequence
ATG TCC AAA GGT GAG GAA CTG TTT ACA GGT GTT GTG CCT ATC CTG GTT GAA CTG GAT GGG GAC GTT AAC GGG CAG AAG TTT AGT GTA TCA GGC GAG GGG GAG GGG GAT GCT ACA TAT GGC AAA CTT ACT TTG AAA TTC ATC TGT ACT ACT GGC AAG TTA CCG GTT CCC TGG CCA ACA TTG GTC ACA ACG TTT TCT TAT GGG GTC CAG TGT TTT TCC AGA TAC CCG GAC CAC ATG AAG CAG CAC GAC TTT TTC AAG TCT GCT ATG CCC GAA GGT TAT GTG CAA GAA CGT ACT ATT TTC TAT AAG GAC GAC GGC AAT TAC AAA ACC AGA GCC GAA GTA AAA TTC GAA GGG GAT ACC TTG GTT AAT CGT ATC GAG CTG AAA GGT ATC GAT TTT AAA GAG GAC GGT AAT ATT TTG GGC CAC AAA ATG GAG TAT AAT TAT AAT TCC CAC AAT GTA TAT ATC ATG GCG GAC AAA CCC AAA AAT GGC ATC AAA GTC AAT TTT AAA ATA CGC CAT AAC ATT AAA GAC GGC TCG GTG CAG CTT GCG GAT CAT TAT CAG CAG AAT ACC CCC ATA GGC GAC GGT CCT GTA CTG CTG CCT GAC AAT CAC TAT CTT TCA ACA CAA TCA GCC CTG TCG AAA GAC CCG AAT GAA AAA CGC GAC CAT ATG ATA CTG CTT GAA TTC GTA ACT GCT GCC GGA ATA ACA CAC GGT ATG GAC GAG TTA TAC AAA TAA
Cell-dependent method (using E.coli)
In order to produce this protein from our DNA we can use the host cells of E.coli. This method is the most common for producing high protein yields.
1. Transcription
We would clone the optimised DNA sequence into a plasmic vector with a strong promoter. Inside the E.coli cell, the RNA polymerase would bind to the promoter and transcribe the DNA sequence into mRNA.
2. Translation
Ribosomes within the E.coli cell bind to the mRNA and commence translation by reading the codon sequence and assembling the corresponding amino acids. The optimised codon sequence would prevent translation from stalling thus folding the amino acids into the functional GFP protein.
3. Induction
The E.coli would then produce GFP. IPTG is a common reagent used to induce protein expression in bacteria.
Cell independent method
This method does not require the use of living cells through using extracts with the necessary machinery.
1. Mixing
You combine your DNA sequence, sources of energy (ATP, GTP), amino acids, and translation machinery (ribosomes, tRNA, enzymes) derived from bacterial lysates in a test tube.
2. Transcription and Translation
The processes of transcription and translation occur simultaneously in this open system producing the GFP protein without such limitation as maintaining the viability of cells.
An advantage of this method if that you can produce proteins that are toxic to living bacterial (i.e. E.coli) cells.
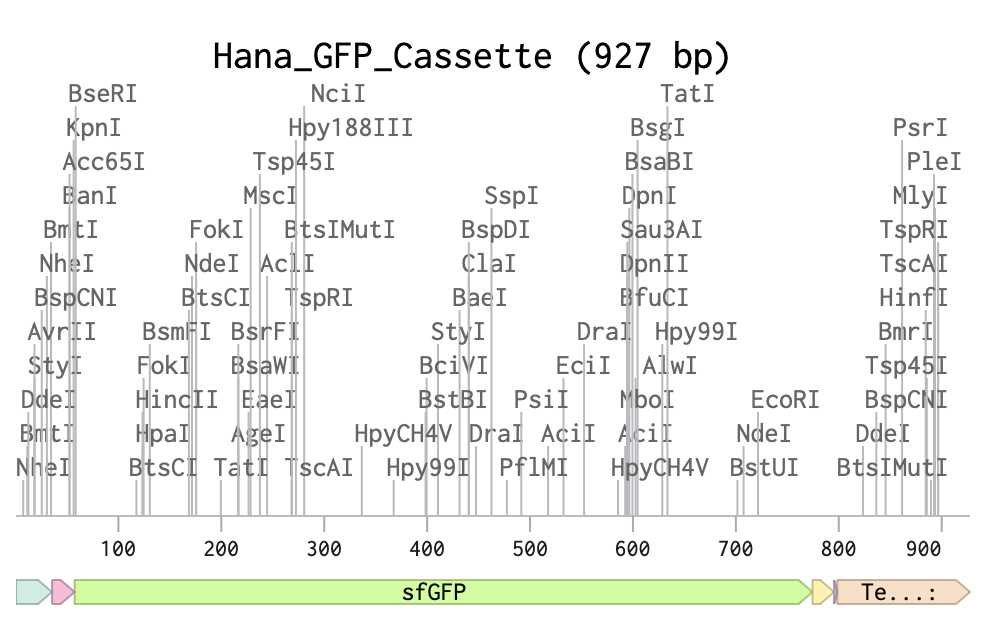
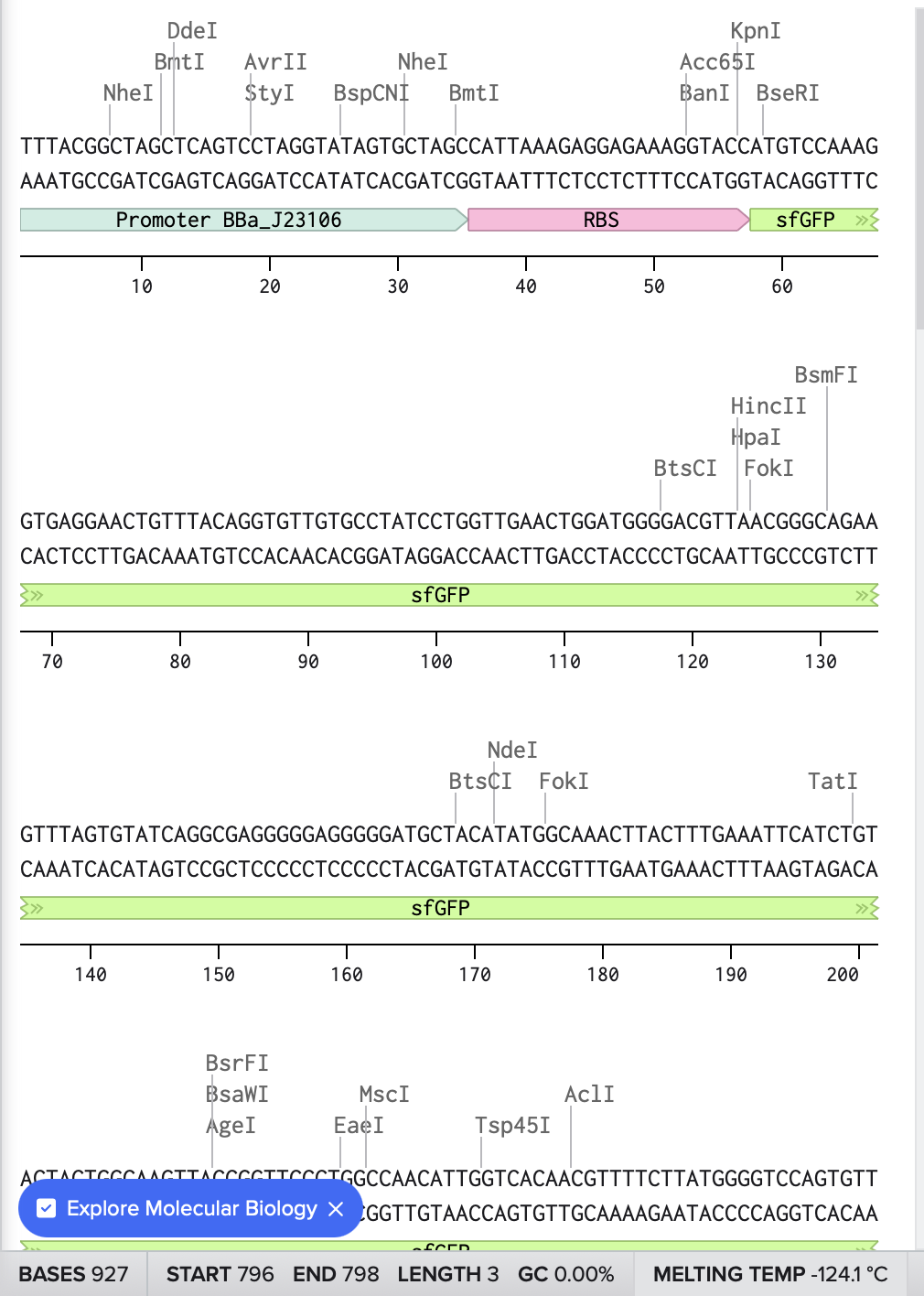
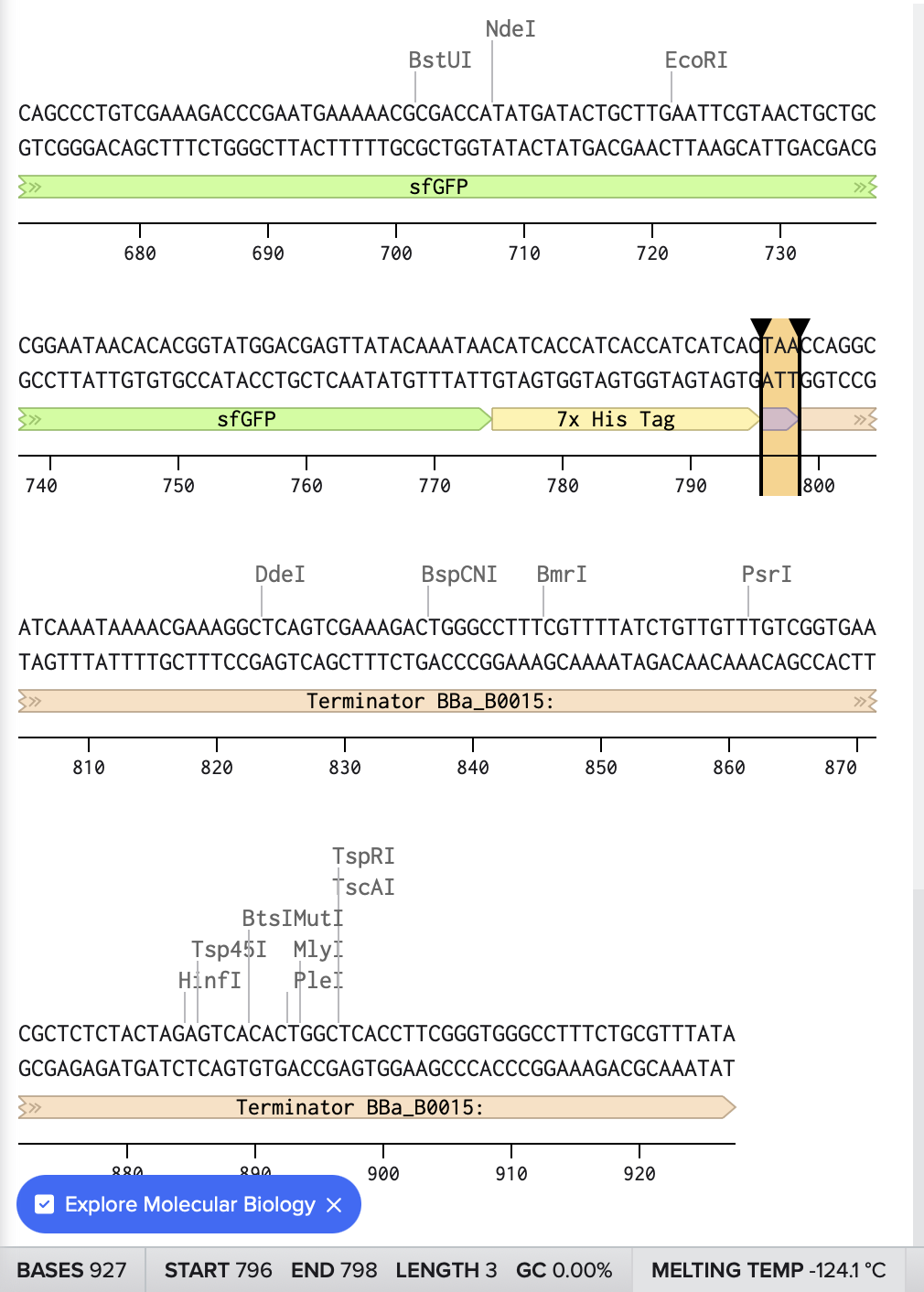

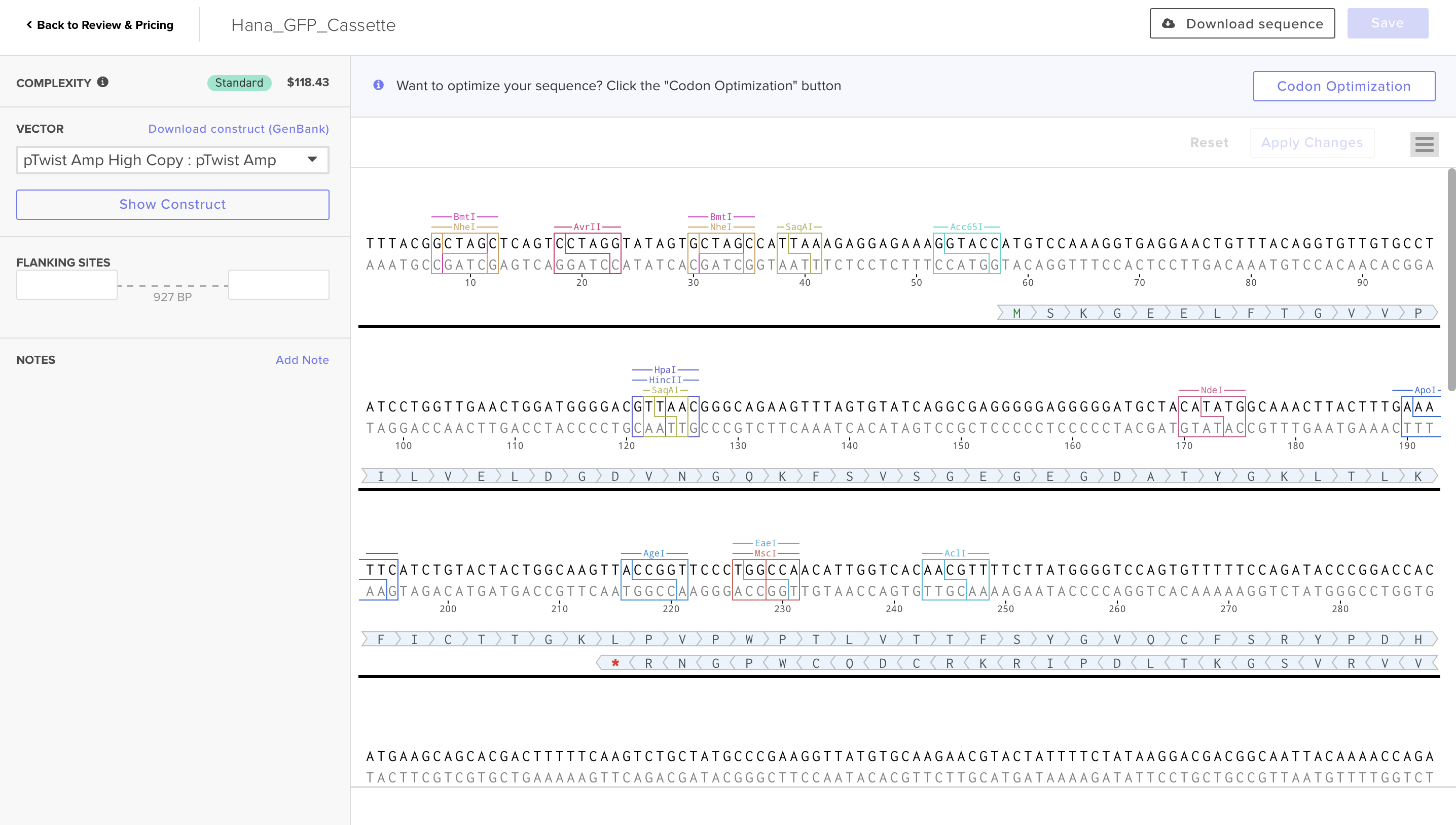
Here is the link for my final sequence:
https://benchling.com/s/seq-P12tVvT9nqlExDjD5hkS?m=slm-DFTTw8aWcdihybyBVM8P
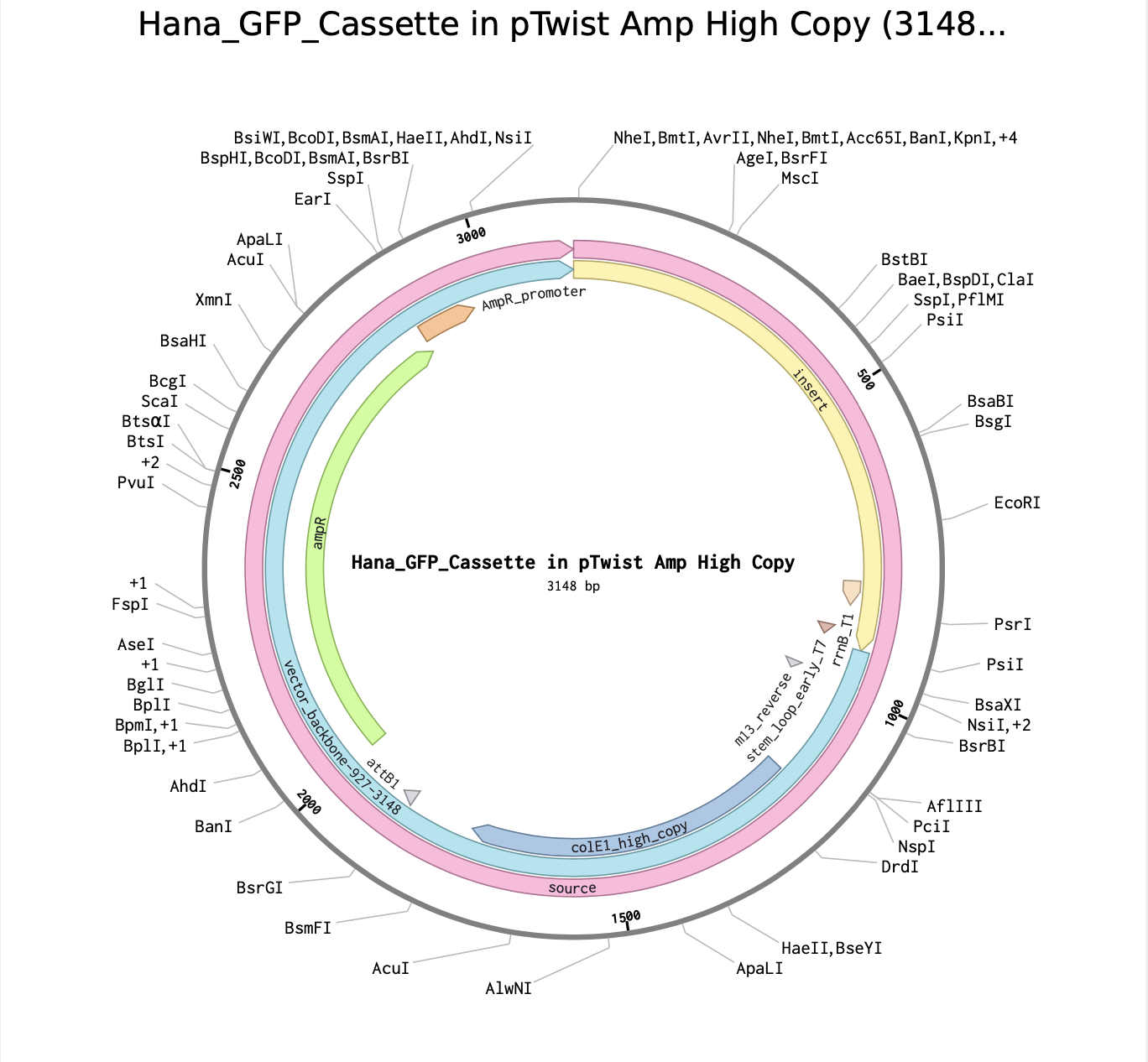
(i) I would like to sequence the DNA of sunflowers, Conan the Bacterium, bowhead whales, and yeast.
- Sunflowers have been used for the purposes of phytoremediation (removing radioactive contaminants in soil), for exmaple in the aftermath of the disasters in Chernobyl (1986) and Fukushima (2011). This is due to the fact that sunflowers are able to act as radiation sponges, thus classifying them as “hyperaccumulators.” They are able soak up Pb (lead) and Cd (cadmium), for example, and store such in their leaves and shoots (Alaboudi et al 2018), (Al-Jabori et al 2019).
They also absorb radioactive isotopes because these toxins resemble essential nutrients due to chemical mimicry. Specifically, Cesium-137 acts like potassium, and Strontium-90 acts like calcium. Their extensive root systems mean that they are able to soak up contaminants from large volumes of soil. Moreover, their capacity to grow quickly and produce large amounts of biomass meand that they are a high-capacity storage system for contaminants, thereby preventing damage to the surrounding ecosystem. It should be notes that sunflowers do not neutralise radiation but make it easier to clean up.
https://link.springer.com/article/10.1007/s11270-009-0128-3 (Adesodun et al. 2010)
https://www.sciencedirect.com/science/article/pii/S0570178318300174 (Alaboudi et al 2018)
https://www.proquest.com/openview/7e4eea8c2ab721e20eb9481b4a2a96b0/1?pq-origsite=gscholar&cbl=54977 (Al-Jabori et al 2019)
I would like to sequence the DNA of sunflowers to better understand how they don’t die under radiation stress - what about their DNA makes them so resilient? I am interested in creating a radiation biosensor. Sequencing the DNA of sunflowers could be useful for human, animal, and plant health, environmental monitoring, AND potentially data storage.
- Conan the Bacterium otherwise known as Deinococcus radiodurans is a bacterium and the most radiation resistant organism known to man. Not only can it withstand radiation, but also dehydration, vacuum, and highly acidic condition, therefore qualifying it the classification of a polyextremophile. It can withstand up to 5 kGy (kiloGrays) of gamma radiation and 1,000 J/m² of UV radiation without significant loss of viability. It has even been recorded as having survived on the exterior of the ISS for 3 years.
Deinococcus radiodurans is able to do this employing specialised DNA repair mechanisms and creating proteome shields against oxidation: namely PprI and DdrC. These proteins are able to repair hundreds of DNA double-strand breaks (DSBs) caused by ionising radiation or desiccation within hours. It achieves this through a two-phase process: rapid Extended Synthesis-Dependent Strand Annealing (ESDSA) and then RecA-dependent homologous recombination.
I would sequence the DNA of this organism to understand what makes it so hardy. It could have applications for human, animal, and plant health in the context of genotoxic stress inducing conditions. They could also be used for the creation of radiation biosensors/useful biomaterials. I care less about tech bro space applications.
- Bowhead whales have incredible resistance to cancer. This is despite the fact that they have many times more cells (greater likelihood of mutation) than other smaller animals such as humans and mice for instance, - a phenomena known as Peto’s Paradox.
So what is the reason for their remarkable cancer resistance? Whilst some large mammals such as elephants have extra TP53 tumour suppressor genes, bonehead whales have highly efficient DNA repair systems. They have high levels of CIRBP ( cold inducible RNA-binding protein) - a stress response protein which minimises genotoxic stress. The production of this protein is induced by such factors as low temperatures, UV light, and hypoxia (low oxygen levels). It is located in the nucleus of a cell where it regulates alternative splicing, and the cytoplasm where it stabilises mRNA. It this influences cell survivial, circadian rhythms, and inflammation. (Zhong and Huang 2017)
Simply put, this protein aids in the repair of double-strand DNA breaks, thus minimising tumour cell growth and proliferation, and maintaining genomic stability.
- https://pmc.ncbi.nlm.nih.gov/articles/PMC5674272/ (Zhong and Huang 2017)
Bowhead what CIRPB (bwCIRBP) is extremely stable while human CIRPB (hCIRBP) is relatively unstable. The whale version has five unique amino acids. I want to sequence bowhead whale DNA for the purposes of human, animal, and plant health, particualarly for oncological research and therapy.
- Yeast have incredible prion suppression abilities. Saccharomyces cerevisiae - yes the yeast used for baking and wine-making. Whilst mammals, humans included, are unable to “cure” prions (misfolded proteins), yeast have developed advanced mechanisms for eliminating such.
For instance, they possess the protein Hsp104 which works alongside the proteins Hsp70 and Hsp40. Hsp104 breaks apart amyloid filaments in cells by resolubilising aggregated proteins. Hsp70 and Hsp40 work together in what is known as a “molecular chaperone system.” Hsp40 delivers misfolded proteins to Hsp70 which remodels such. Humans do not have homologs for Hsp70 though we have the Hsp70/Hsp40/Hsp110 system. Unfortunately our system is not as affective at disaggregating existing clumps of amyloid hence the prevalence of prion diseases.
The gene for Hsp104 in Saccharomyces cerevisiae is the HSP104 gene (also known as YLL026W). I would sequence the DNA of yeast to better understand the mechanisms of prion suppression to help combat neurodegenerative disorders.
(ii) I would use a variety of DNA sequencing technologies for the DNA of different organisms that I wish to sequence.
For sunflowers
Technology and Generation
- I would like to use a combination of ONT (Oxford Nanopore Technology) long read sequencing + Illumina short read sequencing
- The goal is to identify novel repair genes, structural variants and “hardy” regulatory genes ∴ hybrid sequencing approach prevents gaps in research
- Long reads with ONT are essential for the Sunflower genome, which is notoriously large (approx. 3.6 Gb/base pairs) and filled with repetitive “junk” DNA that hides the hyperaccumulator genes + over 78% to 80% of the sunflower genome is made of repetitive sequences
- Short read technologies such as Illumina have greater accuracies and can be used to “polish” long-read data
- ONT: third generation, Illumina: second generation
Input
- Input: High Molecular Weight (HMW) Genomic DNA extracted from sunflower leaf or root tissue
- Extraction: Using a specialized method (like CTAB or magnetic beads) to get ultra-pure, long DNA strands
- End-Repair: We use enzymes to “blunt” the ends of the DNA strands so they aren’t ragged
- Adapter Ligation: We attach specialized sequencing adapters to both ends of the DNA to help unzip 👖 double helix
Steps
- Translocation: the motor protein zips the DNA strand through a protein nanopore embedded in an electrically resistant membrane.
- Ionic Current Blockage: a constant flow of ions passes through the pore. As the DNA bases (A, T, C, G) pass through, they physically block the pore
- Base Calling: each of the four bases has a different shape and size. Therefore, each base (or more accurately, a “k-mer” or group of bases) causes a unique disruption in the electrical current
- Decoding electric signals with AI
- Output
- The output of this process is a FASTQ file
For Conan the Bacterium
Technology and Generation
- I would use Illumina once again as this short read technology is accurate and sufficient for sequencing the genome of Conan at only 3.2 Mb/bp
- Extreme accuracy is required to identify the specific mutations in the PprI and DdrC repair genes
- The Goal: High-fidelity “polishing” to ensure every single base of those rapid-repair enzymes is mapped correctly for biosensor
Input
- Input: Genomic DNA (gDNA) extracted from a culture of D. radiodurans
- Fragmentation (cut DNA into consistent chunks)
- A-Tailing (prevent sticking)
- Adapter Ligation (We attach Y-shaped DNA adapters for sample identification)
- PCR Amplification
- Steps
- Illumina uses fluorescent light
- Cluster Generation: grow colonies of identical strands
- Reversible Terminator Dance: machine floods the flow cell with four types of fluorescently labeled nucleotides (A, T, C, G)
- Competition: Only the correct matching base can bind to the DNA strand. When it binds, it emits a specific color of light (e.g., Green for G, Red for T)
- Action!: high-resolution camera takes a photo of the entire flow cell; color of the “dot” tells the computer which base was just added
- Cleave and Repeat: The “terminator” (the chemical block) is washed away, and the next base is added - happens hundreds of times
- Output
- The output of this process is a FASTQ file
- To build a radiation biosensor using D. radiodurans we must isolate the Promoters—the “DNA switches”—that activate the RecA and PprI proteins the moment radiation is detected
- Short-read sequencing is the gold standard for finding these precise regulatory sequences
For Bowhead whales
- Technology and Generation
- The goal: looking for the unique amino acid swaps in the bwCIRBP protein and the duplication of DNA repair genes
- ∴ We need a sequencing method that combines the length of the sunflower strategy with the accuracy of the bacterial strategy
- Solution: PacBio HiFi Sequencing, third generation
- Unlike standard Nanopore (which can be “noisy”) or Illumina (which is too short), HiFi reads are both long (up to 20,000 bp) and ultra-accurate
- Input
- Input: HMW (High Molecular Weight) DNA, usually taken from a skin biopsy or blood sample of the whale 🐋
- Shearing: The DNA is sheared into large fragments of approximately 15–20 kb
- Exonuclease Digestion: We clean up any stray single-stranded DNA to ensure only high-quality double-stranded templates remain
- Hairpin Adapter Ligation: This is the “secret sauce” of PacBio. We attach hairpin adapters to both ends of the DNA fragment, turning it into a closed circle (called a SMRTbell)
- Polymerase Binding: A highly engineered DNA Polymerase is attached to the adapter, ready to start “looping” around the circle
- Steps
Decoding: Circular Consensus Sequencing (CCS) - this is where the name HiFi comes from
- The ZMW (Zero-Mode Waveguide): The SMRTbell is dropped into a tiny well called a ZMW. There are millions of these on a single chip.
- The “Rolling Circle”: The polymerase starts unzipping and copying the DNA. Because the DNA is a circle, the polymerase doesn’t stop at the end—it just keeps going around and around.
- Fluorescence Detection: As each base (A, T, C, G) is added, it lets off a flash of light. A camera at the bottom of the ZMW records these flashes.
- Base Calling by Consensus: The machine reads the same strand of DNA 10, 20, or 30 times. If there was a random error in the first pass, it gets corrected by the subsequent passes.
- Result: You get a single, long “Consensus” read that is as accurate as Illumina but long enough to capture a whole gene! 🧬
- Output
- The output is a BAM file (which contains the raw signal) that is converted into a FASTQ file
- You can align the whale’s CIRBP sequence directly against the human hCIRBP sequence
- Because the data is so clean, those five unique amino acids you mentioned will stand out clearly, allowing you to design synthetic versions for oncological therapy 🥳
If we can understand how the whale stabilises its mRNA via CIRBP, we might be able to replicate that stability in human cells undergoing chemotherapy.
For regular degular yeast
- Technology and Generation Ok admittedly this is a little curveball answer but instead of sequencing the DNA of yeast, I would sequence the RNA of yeast. I am seeking to understand the mechanism of how Hsp104 interacts with Hsp70 and Hsp40. Just knowing the DNA sequence is not sufficient - instead I need to know how yeast regulates these genes in a protein-folding crisis.
- The goal: To map the “Chaperone Network” and identify the regulatory triggers that allow yeast to resolubilise amyloid filaments
Therefore I would use Illumina for transcriptonomic purposes (RNA sequencing) as it is
- Cost effective
- Highly quantitative: doesn’t just tell you the sequence, but also how many copies of the Hsp104 mRNA are present
- Input
- RNA extracted from yeast cells that have been “stressed” (e.g., heat-shocked or exposed to misfolded proteins)
- Poly-A Selection: Use magnetic beads to grab only the mRNA (the instructions for proteins), which have a “Poly-A tail”
- Reverse Transcription: Illumina cannot sequence RNA directly so we must use reverse transcriptase to turn the unstable RNA back into stable cDNA (complementary DNA)
- Fragmentation & Adapters: Just like with Conan the Bacterium, we shear the cDNA into small bits and attach Illumina adapters (barcodes and anchors)
- PCR: We amplify the library to ensure we can detect even the rarest “stress-response” instructions
- Steps
We use the same technique we used for Conan:
- Fluorescence: Each time a base pair is added to the cDNA strand, it flashes a specific colour
- High throughput: We can sequence millions of RNA transcripts simultaneously
- Output
- The output is a FASTQ file, but the way we use it is different
- Instead of building a map, we count
- If we see 5,000 reads for HSP104 in stressed yeast but only 10 reads in healthy yeast, we know exactly how much the “prion-curing” machinery has been ramped up
- Alternative Splicing: You can see if the yeast is “editing” its mRNA to create different versions of the Hsp104 protein for different types of protein clumps
| Organism | Primary Research Goal | Best Technology | Data Output Type |
|---|---|---|---|
| Sunflower | Find metal-transporter clusters | Oxford Nanopore | Long-Read Map |
| Conan (Bacterium) | Map rapid-repair genes (PprI) | Illumina (SBS) | High-Accuracy Map |
| Bowhead Whale | Compare CIRBP amino acids | PacBio HiFi | Ultra-Clean Gene Sequence |
| Yeast | Watch Hsp104 “curing” prions | RNA-Seq (Illumina) | Gene Expression Levels |
I would synthesise a Chimeric Genetic Circuit that combines the “sensing” power of the sunflowers and Conan with the “repair” power of bowhead whales and yeast. It would have these specific compenents:
- Switch (Promoter): The DNA sequence for the PprI promoter from D. radiodurans. This acts as a high-sensitivity “on-switch” that only activates in the presence of ionising radiation.
- Shield (Protective Protein): The DNA sequence for the Bowhead Whale CIRBP (with those five unique amino acids). This protein would be produced the moment the PprI switch is flipped.
- Cleaner: The DNA sequence for Yeast Hsp104, modified with human-compatible codons (through codon optimisation) to ensure it works efficiently in mammalian cells to prevent protein clumping under radiation stress
By synthesizing this custom circuit, we can create a prophylactic genetic shield. This could be used to create radiation-proof crops 🌾 for phytoremediation or specialized cell therapies for organisms living in high radiation areas, or even cancer patients undergoing high-dose radiotherapy.
In order to do this, I would use Phosphoramidite Synthesis (Oligonucleotide Synthesis) to create primers and concise units of DNA, which is what happens at Twist as Emily LeProust explained. I would also use Enzymatic DNA Synthesis because this enables the synthesis of long strands of DNA. And finally, I would make the circuit by joining together all of the concise units of DNA and necessary primers by using Gibson assembly.
Maybe I would edit stem cells . . . not sure. Have to think about the risks and ethical implications. But if I were going to I would use CRISPR because it’s really good at targeting DNA for editing with great precision.