Week 6 HW: Genetic Circuits Part I
1. What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
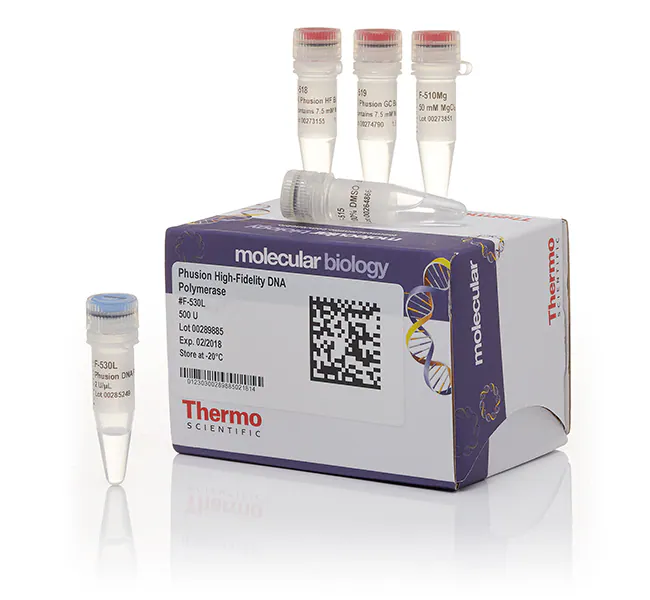

Phusion-High Fidelity PCR Master Mix contains
Phusion DNA Polymerase
→ High fidelity, thermostable enzyme for fast, robust, and accurate DNA amplification in PCR; it is used particualrly for cloning and sequencing
“Phusion is one of the most accurate thermostable polymerases available.” (New England Biolabs Website 2026)
Deoxynucleotide Triphosphates (dNTPs)
→ The essential building blocks of DNA; they store the genetic information of an organism
Optimised reaction buffer including MgCl
→ Premixed solution that is designed to serve as the ideal chemical environment for DNA polymerase to function at maximum efficiency; it acts as a ready to use foundation for PCR experiments, thus eliminating tedious optimisation of key components. It does this by maintaining the maintains the optimal pH and ionic strength for the enzyme to function. It often contains additives that stabilize the enzyme during the high temperatures.
→ MgCl is an essential cofactor for the enzyme as it enables its catalyctic ability. This is due to the fact that the magnesium ions coordinate with the phosphate groups of the dNTPs as well as the active site of the enzyme, thereby facilitation the formation of the phosphodiester bond. It also helps primers anneal to the template DNA by reducing electrostatic repulsion.
“All that is required is the addition of template, primers and water." (Fisher Scientific Website 2026)
Sources:
New England Biolabs Website
Fisher Scientific Website
https://www.fishersci.fi/shop/products/phusion-high-fidelity-pcr-master-mix-gc-buffer/10523288
2. What are some factors that determine primer annealing temperature during PCR?
- Primer’s melting temperature
- Primer length
- Proportion of GC relative to AT (GC content)
- Primer concentration
- Ionic strength of the buffer
3. There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
PCR (Polymerase Chain Reaction)
What it does: Amplifies specific DNA sequences, basically a biological photocopier ADDITIVE - synthesises DNA. Lots of copies of one fragment
How: Uses thermal cycling (95°C→Variable→72°C) and DNA polymerase to exponentially increase the amount of copies of a particular DNA sequence
With: Requires primers and dNTPs
Why: Finding and amplifying
When: You need to isolate some DNA out of the whole genome, and also you don’t have a lot of DNA to start with i.e. a trace amount of DNA such as a cheek swab or ancient DNA!
Restriction Enzyme Digest
What it does: Cuts specific DNA sequences, basically biological scissors SUBTRACTIVE - subtracts DNA. Several fragments of varying sizes
How: Uses restriction endonucleases which recognise short, specific DNA sequences, and cuts the sugar-phosphate backbone at those sites, kept at a constant (usually 37°C)
With: Requires Specific recognition sites (GAATTC, etc.)
Why: Cutting and checking
When: You want to
- Manipulate or verify existing high-concentration DNA (i.e. plasmids - you can cut them open and check if desired gene is actually inside by using gel electrophoresis),
- Do Cloning/Ligation (opening up and inserting gene into a plasmid)
- Do Genomic mapping - before modern sequencing Restriction Fragment Length Polymorphism (RFLP) was used to compare DNA samples by looking at unique cut patterns
Here is that same information tabulated for the sake of easier comparison:
Comparison of PCR and Restriction Enzyme Digests
| Feature | PCR (Polymerase Chain Reaction) | Restriction Enzyme Digest |
|---|---|---|
| Action | Additive: Synthesizes and amplifies DNA. | Subtractive: Cleaves and fragments DNA |
| Analogy | Biological Photocopier | Biological Scissors |
| The “How” | Thermal Cycling: Uses DNA polymerase; cycles through 95°C - 72°C | Isothermal: Uses endonucleases; kept at a constant temperature (usually 37°C) |
| Requirements | Specific Primers and dNTPs. | Specific Recognition Sites (e.g., GAATTC) |
| Output | Millions of copies of one specific fragment | Several fragments of varying sizes |
| Primary Goal | Finding and Amplifying | Cutting and Checking |
| Ideal Scenario | Trace amounts of DNA: Isolating a single gene from a whole genome (e.g., cheek swab or ancient DNA) | High-concentration DNA: Manipulating or verifying plasmids (e.g., cloning/ligation or genomic mapping) |
| Verification | Used to “zoom in” on a needle in a haystack | Used in RFLP to compare DNA samples via unique “cut patterns” on a gel |
Key Takeaways for Lab Application
PCR is your primary tool for creation It allows you to build a massive amount of DNA from a microscopic starting point by repeatedly synthesizing the target sequence.
Restriction Digests are your primary tool for modification. They allow you to “cut and paste” DNA or verify that a plasmid contains the correct insert by checking fragment sizes via gel electrophoresis.
4. How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
- Remove enzymes/buffers using PCR clean up because they might interfere with the Gibson Master Mix
- Run gel electrophoresis to check if you have the correct DNA fragment/gene
- Have at least 20 bp overlap
- Design PCR primers with appropriate tails
5. How does the plasmid DNA enter the E. coli cells during transformation?
Heat Shock
This is the process of generating pores in the bacterial cell wall through abrupt temperature changes
Electroporation
This is the process of generating pores in the bacterial cell wall using a high electrical voltage
Both of these methods causes the cell membrane to “open up” thus allowing the plasmid to enter the cells via diffusion. After being shocked with either heat or electricity, the E.coli cells are incubated in a nutrient-rich liquid broth such as Super Optimal Broth (SOB) or Lysogeny Broth (LB) at 37°C for about an hour. This period of time allows the cells to recover and begin multiplication. some of these cells will have the plasmid inside. After this process, the transformed cells are placed on an agar plate with antibiotics. Only the sucessful recombinant cells which have received the plasmid will survive as they contain a gene for antibiotic resistance. Colonies of the E.coli with the plasmid will grow.
After a day or two, you will see the colours expressed by the inserted gene if the plasmid containsn GFP for example.
6. Describe another assembly method in detail (such as Golden Gate Assembly)
I did some research on Golden Gate Assembly. It is a goated assembly method which was invented in 2008 in Köln by Carola Engler, Sylvestre Marillonnet, Romy Kandzia, and colleagues but has its origins in 1996. I erroneously thought it was named so because it was invented in San Francisco but no - it is called Golden Gate Assembly as it can seamlessly combine multiple DNA fragmanets, acting as a “golden gate” which allows desired fragments to pass through while restricting unwanted or empty products.This one-pot, one-step cloning method was developed to allow efficient and seamless assembly of multiple DNA fragments using Type IIS restriction enzymes (like BsaI).
It is a scarless method (does not leave any unwanted sequences) that can be used to assemble multiple fragments, even up to 50+ (Thermo Fisher Scientific Website).
Here is the pioneering 2008 paper “A One Pot, One Step, Precision Cloning Method with High Throughput Capability” (Engler et al. 2008):
https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0003647
I watched this video to help me understand it better:
https://www.youtube.com/watch?v=NzQdLQ44I7w
Background info:
Golden Gate Assembly is a versatile cloning technique which allows for the seamless assembly of multiple fragments in a single cloning reaction. It leverages upon the features of Type IIS restriction enzymes which makes them unsuitable for standard restriction enzyme cloning:
- Restriction site is non-palindromic (Type IIS enzymes recognise and bind to asymmetric DNA sequences)
- In general sites range from 4-7 nucleotides
- DNA is cleaved outside of the recognition site!
Golden Gate Assemblies are done using a destination vector and one or more fragment of interest - these can come from multiple oraganisms. A lot of vectors are available through commerical sources or Addgene. But you can adapt a vector for Golden Gate Assembly in your lab also by making sure:
- The destination vector has type IIS restriction sites which flank desired insert region
- The enzymes flank a selectable or screenable gene
- Different Type IIS enzymes will cleave DNA to produce different length overhangs
- 4 base overhangs are often referred to as “fusion sites”
- Type IIS restriction enzymes most commonly used for Golden Gate Assembly are Bsa1, BSM, B1, BBSB1 (all of these create 4 nucleotide overhangs)
The orientation of type IIS enzymes as well as the recognition and fusion sites is critical to the success of Golden Gate Assembly!
In destination vector, the sites must face away from each other or be outward facing and flank the region being excised. This means that after cleavage, the recognition sequences are removed from the remaining plasmid backbone, thus making it resistant to further digestion by the restriction enzyme.
The resulting overhangs are not compatible with each other so the plasmid cannot recircularise!
Preparation:
You may use any double stranded DNA fragment for Golden Gate Assembly including plasmids or PCR products. Thankfully a lot of Golden Gate Assembly kits come with plasmids with sequence-verified inserts. PCR makes it simple to convert or accommodate any sequence for the assembly. You must design PCR primers with flanking bases, type IIS recognition sites, and an overhang sequence. This way you can introduce the required recognition site at each end of the plasmid/PCR product.
Our fragment of interest will have type IIS sies that are inward facing. This means that cuts will occure between two recognition sites. This way the insert fragment will not contain a type IIS recognition after clevage, ensuring a cease of futher digestion.
Reaction time baby!
You combine all your components into a single reaction tube:
- Destination Vector: The backbone plasmid.
- Inserts: The DNA fragments you want to assemble.
- Type IIS Restriction Enzyme: (e.g., BsaI or BsmBI) to cut the DNA
- T4 DNA Ligase: To join the fragments together
- Reaction Buffer: Containing ATP (required for the ligase)
A thermocycler rotates through different reaction conditions/temperatures
Digestion (37°C): The optimal temperature for the restriction enzyme to cut the DNA and reveal the 4-bp overhangs.
Ligation (16°C): The optimal temperature for the T4 Ligase to stabilize the short overhangs and seal the phosphodiester backbone.
Annealing (Generally between 50°C and 65°C the specific temperature is determined by your primers): Chemical sweet spot which is cool enough to allow the DNA strands to come together but warm enough to prevent the primers from sticking to the wrong places.
During the enzyme digestion the type IIS restriction enzyme exposes single stranded ends which are no longer associated with the type IIS recognition sequence. Complementary single stranded overhangs anneal creating the “Golden Gate.” Voila! The annealed overhangs are then covalently closed by DNA ligase. Cleavage with a Type IIS enzyme eliminates any unmodified destination vector that survived through the reaction.
Finally, at 80°C remaining enzyme activity is inactivated, the plasmid product is now ready for transformation into an appropriate host!
7. Explain the other method in 5 - 7 sentences plus diagrams (either handmade or online).
Golden Gate Assembly TLDR:
- Design primers. Choose your DNA fragments. Make sure they are “clean” and have type IIS restriction sites which flank desired insert region.
- Put all of your components into a reaction tube. This includes the vector, the inserts, the Type IIS enzyme, T4 ligase, and reaction buffer.
- The Type IIS enzyme cuts outside its recognition sequence, creating unique 4-bp sticky ends that dictate the exact order of assembly.
- The thermocycler should alternate between different reaction cycles to enable digestion, ligation, and annealing.
- As correct assemblies form, they lose the restriction sites and become “immune” to further cutting, while original vectors are repeatedly re-cut.
- The mix is heated to 80°C to stop all enzyme activity. This temperature will denature the Type IIS restriction enzyme and the ligase. This prevents the Type IIS restriction enzyme from cutting up the DNA or the ligase from sticking to the DNA.
- Plasmid can now be tranformed into desired host.
8. Model this assembly method with Benchling or Asimov Kernel!