Week 9 HW: Cell Free Systems
1. Explain the main advantages of cell-free protein synthesis over traditional in vivo methods, specifically in terms of flexibility and control over experimental variables. Name at least two cases where cell-free expression is more beneficial than cell production.
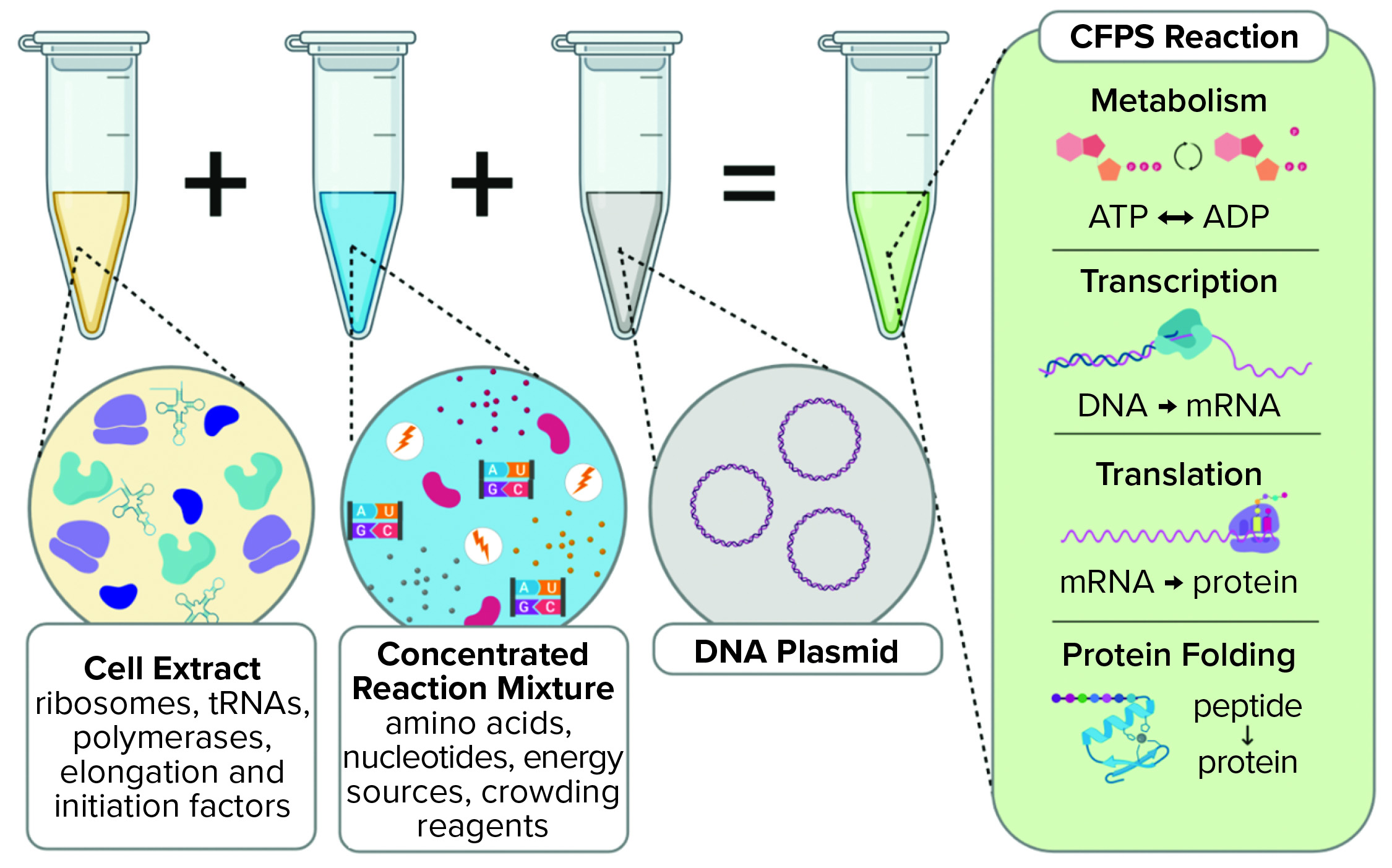
Cell-free protein synthesis leverages biology as an engineering tool (Kate). Living cells require a lot of resources such as the correct amounts of
- Water
- Energy
- Gases
- Pressure
- Heat
- Equipment to sustain
- Less need for cold chain distribution
and produce a lot of waste (Peter). With cell-free, you are able to freeze dry systems for upto a year. It has the advantages of
- Transportability
- Therapeutics on demand → just add water!
- Rapid manufacturing
- Greater biosafety: don’t have to worry about living cells getting about
Cell free is more beneficial in
- Space
- Places where supply chains are not as strong/resources are scarce i.e. in developing countries
The military was also mentioned . . .
Applications of cell-free
- Synthetic Biology: Designing and testing biological circuits or pathways without cellular constraints
- Protein Engineering: Rapid protein production and screening, especially for proteins that are toxic or hard to express in cells
- Metabolic Engineering: Production of high-value chemicals, biofuels, and pharmaceuticals via synthetic pathways
- Biosensing: Creating diagnostic tools that are portable and easy to use, like paper-based biosensors
- Gene Editing Research: Testing CRISPR-based systems or genome editing tools in a controlled environment
2. Describe the main components of a cell-free expression system and explain the role of each component.
Whole cell extract including:
- Ribosomes
- tRNA
- aminoacyl-tRNA synthetases
- translation factors
- RNA polymerase*
The cell extract provides the necessary machinery, as it were, for translation.
DNA Template such as a plasmid or a linear PCR product
This functions as the blueprint for the desired protein that we will be producing as it contains the coding sequence for such. Before, this sequence is a strong promoter such as a T7 promoter which the RNA polymerase will recognise in order to begin transcription.Source of energy i.e. ATP and GTP and also phosphoenolpyruvate and pyruvate kinase (for regeneration of ADP to ATP)
The process of protein synthesis is very energy-intensive; and as there is not a living cell metabolism to make use of, energy must be supplied in an external form.
Amino acids
These function as the fundamental building blocks of the protein. The ribosomes inside the whole cell extract will link these together according to the DNA template!
Nucleotides (NTPs)
These will serve as the fundamental building blocks for the mRNA strand.
Salts and organic molecules
- Magnesium is necessary for stabilising the ribosome structure and the polymerase activity
- Potassium is necessary for maintaining the right ionic strength which allows protein folding and enzyme activity
- Buffer such as HEPES, is necessary for maintaining a stable pH so that the enzymes do not denature
Chaperones and Protease inhibitors
The chaperones help the complex proteins to fold properly, while the protease inhibitors prevent the breakdown of the protein as it is being made.3. Why is energy provision regeneration critical in cell-free systems? Describe a method you could use to ensure continuous ATP supply in your cell-free experiment.
Energy provision regeneration is critical in cell-free systems because there is no living cell metabolism. There are no mitochondria so you don’t have the normal mechanisms of energy production.
One of the most common methods of enusring a continous ATP supply is by using PEP (phosphoenolpyruvate) and PK (pyruvate kinase). PEP is a high energy phosphate, whilst PK is an enzyme which catalyses the transfer of the phosphate group from PEP to ADP - the products are which are pyruvate and ATP.
Mechanisms of such:
- You add a high-energy phosphate donor (PEP) and an enzyme (pyruvate kinase) to your reaction mix
- The ribosomes in the whole cell extract use ATP, converting it to ADP
- The pyruvate kinase enzyme takes a phosphate group from the PEP and sticks it back onto the ADP
- This recreates ATP, allowing the cycle to continue until the PEP supply is exhausted
Click to view Tangent I went on . . .
I wondered why they didn’t just put mitochondria into the whole cell extract. I found out that it is because mitochondria are delicate. But I was confused because I read that there are free floating mitochondria in human blood.
But in blood they are wrapped in cell membranes or extracellular vesicles which act like protective bubbles, thus shielding them from the harsh plasma environment and keeping their internal chemistry stable. Also most of these free floating mitochondria come from platelets as a stress response or to help other cells. So they are quite delicate, in that they require intact membranes to maintain the proton gradient (electrical charge) across their inner membrane which causes ATP synthesis. With any membrane damage, they lose their function.
Also the mitochondria need oxygen to function and this can be used up quickly in a microcentrifuge tube. Ok they are quite precious I see.
I also asked Juan Diego and he told me that if you were to introduce mitochondria into cell-free you would just be complicating matters because as endosymbiotic theory posits, mitochondria are the descendants of free-living bacteria. Hence, you would kind of be introducing bacterial DNA into cell-free. You should just stick to what is strictly necessary essentially.
I remember that mitochondria posess their own DNA which is inherited from the mother. Because of this scientists came up with the concept of “mitochondrial Eve” and indeed found one some years ago, or so I saw.
4. Compare prokaryotic versus eukaryotic cell-free expression systems. Choose a protein to produce in each system and explain why.
Prokaryotic cell-free systems
- Use E.coli
Protein synthesis is
- High yield
- Rapid
- Low-cost
Ideal for simple protein synthesis and high throughput screening
Eukaryotic cell-free systems
- Use Rabbit reticulocytes (immature red blood cells), Wheat germ, HeLa (Henrietta Lack’s cervical cancer cells), Chinese Hamster Ovary cells
Protein synthesis is
- Lower yield
- Slower
- Higher-cost
Ideal for complex folding, post-translational modifications, production of membrane proteins
Source: Cell-Free Protein Synthesis: Pros and Cons of Prokaryotic and Eukaryotic Systems (Zemella et al. 2015)
In a prokaryotic cell-free system I would synthesise GFP as it is a robust, single domain protein with a quarternary structure constisting of 238 amino acids, which does not require post-translational modifications i.e. the addition of sugars, in order to function.
In a eukaryotic cell-free system I would synthesise human erythropoietin (EPO). It is a glycoprotein hormone that is mainly produced in the kidneys. Though it is composed or relatively fewer amino acids at 165, it must undergo glycosylation - that is the addition of sugar chains in order for it to be biologically active and stable in the human body. Around 40% of its total weight consists of sugar chains/carbohydrates (Jelkmann 2013). A prokatyotic cell-free system using E. coli for example cannot perform glycosylation - it must be engineered to do such. Without this engineering, it was produce a protein without the sugar chains, rendering it unusable in a medical context. Eukaryotic cell-free systems take advantage of the endoplasmic reticulum vesicles and enzymes to add these sugar groups, thereby making sure that EPO is folded correctly into its complex teritiary shape.
In case you are wondering . . . Yes GFP has a quarternary strucutre whilst EPO has a tertiary one, but as I explained, the latter requires post-translational modifications.
5. How would you design a cell-free experiment to optimize the expression of a membrane protein? Discuss the challenges and how you would address them in your setup.
The hydrophoic element of membrane proteins poses one of the greatest challenged to designing an experiment with them. In a cell-free extract they may misfold or aggregate as there is no lipid membrane for them to be in. The phospholipid bilayer stabilises their transmembrane domains.
To overcome this we would have to add a synthetic membrane in the cell-free system to mimic the normal interaction of membrane proteins. In addition to this we can add nanodiscs which help keep the protein soluble, and liposomes which will essentially “house” the membrane proteins.
In addition to this we can add lot of protein chaperones which assist newly synthesised proteins to fold into their correct functional 3D shape.
6. Imagine you observe a low yield of your target protein in a cell-free system. Describe three possible reasons for this and suggest a troubleshooting strategy for each.
There are a few reasons as to why you may experience a low yield of your target protein. These include:
Energy/Nutrient Depletion which causes Phosphate Byproduct Accumulation Cell-free protein synthesis requires large amounts of the energy molecules ATP and GTP. In addition to this, standard systems also often rely on secondary energy sources such as phosphoenolpyruvate (PEP) or creatine phosphate. As these are used up, inorganic phosphate builds up. This can inhibit the reaction and change the pH which thereby stalls translation.
Troubleshooting Strategy: Secondary Energy Source Shift to a secondary energy source which does not produce byproducts which can inhibit reactions. Good examples inclue pyruvate or glucose which regenerate ATP more “cleanly.”
Troubleshooting Strategy: Dialysis Run the reaction in a dialysis cassette submerged in a large reservoir of “feeding buffer.” this will enable the “fresh” substrates to diffuse in and the toxic phosphate to diffuse out, thereby achieving continous exchange.
Template Issues: You may have a template with a suboptimal purity level which inhibits reactions from occuring, or you may encounter degredation by endonucleases/exonucleases in the lysate. This is particularly common in cell free systems derived from E. coli or wheat germ. What happens is that endogenous nucleases essentially “chew up” your DNA or mRNA template before the ribosomes have have the time to complete protein synthesis (translation).
Troubleshooting Strategy: Use a circular template It would be best to use a plasmid as opposed to linear PRC products as this will protect against the action of nucleases.
Troubleshooting Strategy: Add RNase inhibitors or small molecule stabilisers. This will protect the mRNA.
Troubleshooting Strategy: Chemical modification or linear DNA This can be achived by adding a GamS protein (which inhibits RecBCD) or by having phosphorothioate bonds at the ends of your primers to make the DNA “indigestible” to the nucleases.
Protein Folding and Solubility Issues may arise. Low yields may not be due to a lack of synthesis action but rather becase the protein may aggrefate or form inclusion bodies after leaving the ribosome. Cell-free extracts often lack the chaperones or the specific redox environment (for disulfide bonds) found in a living cell. Otherwise you may be using codons poorly represented in the extract (e.g., using a eukaryotic protein gene in a bacterial lysate)..
Troubleshooting Strategy: Adjust temperature By lowering the reaction temperature (e.g., from 37°C to 25°C or even 16°C) this allows for lower synthesis which enables more precise folding.
Troubleshooting Strategy: Add Chaperones/Detergents: Supplementing the reaction with purified chaperones (like DnaK/J or GroEL/ES) or mild detergents (like Brij-35) helo to keep hydrophobic patches from sticking together.
Troubleshooting Strategy: Redox Shifting If your protein requires disulfide bonds, add a mixture of reduced and oxidized glutathione (GSH/GSSG) to simulate the environment of the endoplasmic reticulum.
Troubleshooting Strategy: Check codon usage by making sure the template is optimised for the host system.
Suboptimal reaction condition
Troubleshooting Strategy: Optimise you CFPS system by checking you have a strict balance of Mg and K concentrations as well as the correct reaction pH.
Design an example of a useful synthetic minimal cell as follows:
1. Pick a function and describe it. a. What would your synthetic cell do? What is the input and what is the output?
b. Could this function be realized by cell-free Tx/Tl alone, without encapsulation?
c. Could this function be realized by genetically modified natural cell?
d. Describe the desired outcome of your synthetic cell operation.
2. Design all components that would need to be part of your synthetic cell. a. What would be the membrane made of?
b. What would you encapsulate inside? Enzymes, small molecules.
c. Which organism your Tx/Tl system will come from? Is bacterial OK, or do you need a mammalian system for some reason? (hint: for example, if you want to use small molecule modulated promotors, like Tet-ON, you need mammalian)
d. How will your synthetic cell communicate with the environment? (hint: are substrates permeable? or do you need to express the membrane channel?)
3. Experimental details
a. List all lipids and genes. (bonus: find the specific genes; for example, instead of just saying “small molecule membrane channel” pick the actual gene.)
b. How will you measure the function of your system?
Freeze-dried cell-free systems can be incorporated into all kinds of materials as biological sensors or as inducible enzymes to modify the material itself or the surrounding environment. Choose one application field — Architecture, Textiles/Fashion, or Robotics — and propose an application using cell-free systems that are functionally integrated into the material. Answer each of these key questions for your proposal pitch:
1. Write a one-sentence summary pitch sentence describing your concept.
2. How will the idea work, in more detail? Write 3-4 sentences or more.
3. What societal challenge or market need will this address?
4. How do you envision addressing the limitation of cell-free reactions (e.g., activation with water, stability, one-time use)?
Freeze-dried cell-free reactions have great potential in space, where resources are constrained. As described in my talk, the Genes in Space competition challenges students to consider how biotechnology, including cell-free reactions, can be used to solve biological problems encountered in space. While the competition is limited to only high school students, your assignment will be to develop your own mock Genes in Space proposal to practice thinking about biotech applications in space!
For this particular assignment, your proposal is required to incorporate the BioBits® cell-free protein expression system, but you may also use the other tools in the Genes in Space toolkit (the miniPCR® thermal cycler and the P51 Molecular Fluorescence Viewer). For more inspiration, check out https://www.genesinspace.org/ .
1. Provide background information that describes the space biology question or challenge you propose to address. Explain why this topic is significant for humanity, relevant for space exploration, and scientifically interesting. (Maximum 100 words)
2. Name the molecular or genetic target that you propose to study. Examples of molecular targets include individual genes and proteins, DNA and RNA sequences, or broader -omics approaches. (Maximum 30 words)
3. Describe how your molecular or genetic target relates to the space biology question or challenge your proposal addresses. (Maximum 100 words)
4. Clearly state your hypothesis or research goal and explain the reasoning behind it. (Maximum 150 words)
5. Outline your experimental plan - identify the sample(s) you will test in your experiment, including any necessary controls, the type of data or measurements that will be collected, etc. (Maximum 100 words)