Week 2 HW: DNA Read, Write, and Edit
DNA Design Challenge
Protein chosen: tape measure protein of Alyssa1, a Mycolicibacterium phage satellite called an “Extracellular Prophage-Inducing Particle” (EPIP). Tape measure proteins are rare in phage satellites, and TEM imaging has shown that EPIPs have uniquely long tails compared to the helper phage, the phage they parasitize, suggesting the tape measure protein may contribute to a unique mechanism of parasitism.
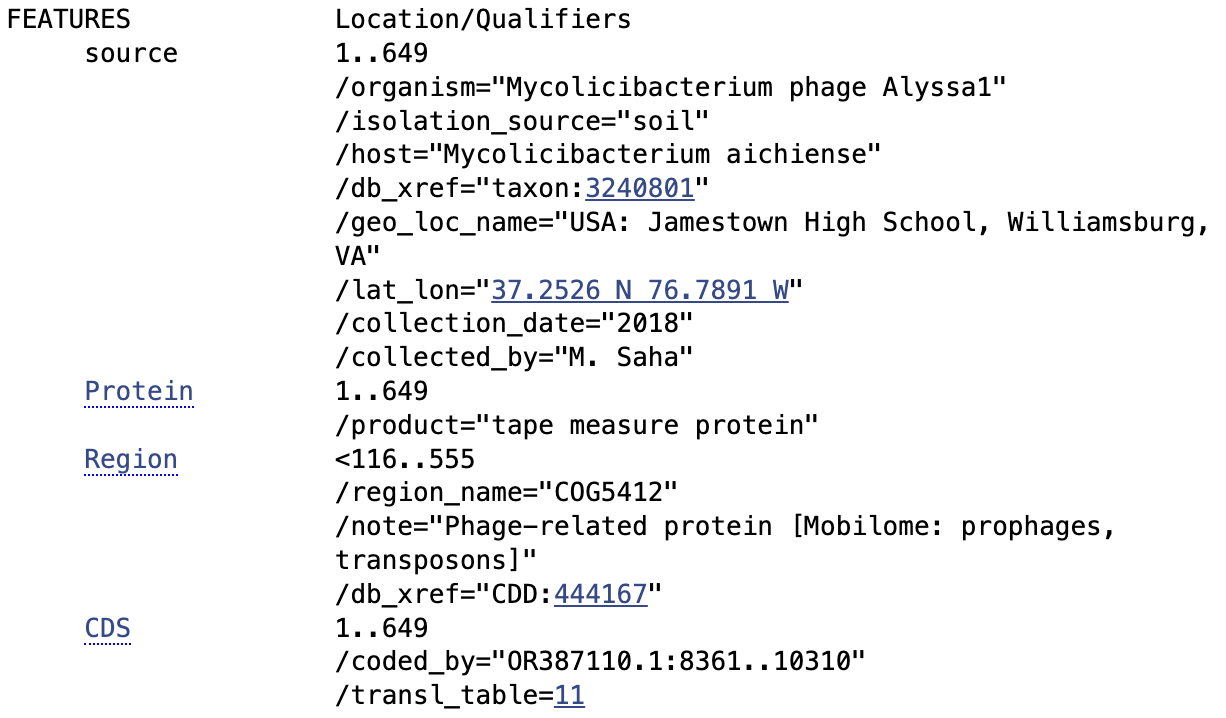

Amino acid sequence: MARKAGMSTGVEVARISVKVGPDTKHFRRELKKDLDRIEESMRAKIDVEPDMKGFRQEVQSKTKGMRTSVKVDADVDRKGFLGRIADSLSQIQPPSFGSGINPTGYAAIAAGIAALTPLIAGTLGAATTALMALPGLVAAVATPIAALTLGIDGLKAAATRLQGPFEDLKATMSSAVESQFGPVFDQLRSLFPVLKGALPSVTAGLADMAKSIADVIASPEGLAKIDTTIRNIGMALTTAAPGVGKFTDGLMGLVQSFTGKPLQGVAEWFSKTGDSFSAWVEKMTRPSWFTGKSPLEVAFGNLGDTLKTIADTLGDVGQKALDFFSDPEKIKSFRDELTLVSDAVKGIATGINGIATAYSSLPLSGEGLKGLMPIQAQLGVKMFDGLKESAAKAFAEVATMAGGFVSNIGSTFMSIGDTLSGIWGGITNAAATAFNSLVSAAQSAVSGVVSAVATIPGQVASALAGLAEAGASAGRNLVQGLVNGISGMIGSAVAKARELASSVASAVTGFLGIHSPSKLFTEIGEYVGQGFDNGLQSQLATLGQTAKAMAETVTEEFNGGLKFGADGFSTDSDNPLMQAGAGLANAPVDFAKATGKQFLSDLGISGNGVLSRALTEGIQYIFQISSVDEALSIKDRETSKNALSIVGR
Reverse translated nucleotide sequence:
atggcgcgcaaagcgggcatgagcaccggcgtggaagtggcgcgcattagcgtgaaagtgggcccggataccaaacattttcgccgcgaactgaaaaaagatctggatcgcattgaagaaagcatgcgcgcgaaaattgatgtggaaccggatatgaaaggctttcgccaggaagtgcagagcaaaaccaaaggcatgcgcaccagcgtgaaagtggatgcggatgtggatcgcaaaggctttctgggccgcattgcggatagcctgagccagattcagccgccgagctttggcagcggcattaacccgaccggctatgcggcgattgcggcgggcattgcggcgctgaccccgctgattgcgggcaccctgggcgcggcgaccaccgcgctgatggcgctgccgggcctggtggcggcggtggcgaccccgattgcggcgctgaccctgggcattgatggcctgaaagcggcggcgacccgcctgcagggcccgtttgaagatctgaaagcgaccatgagcagcgcggtggaaagccagtttggcccggtgtttgatcagctgcgcagcctgtttccggtgctgaaaggcgcgctgccgagcgtgaccgcgggcctggcggatatggcgaaaagcattgcggatgtgattgcgagcccggaaggcctggcgaaaattgataccaccattcgcaacattggcatggcgctgaccaccgcggcgccgggcgtgggcaaatttaccgatggcctgatgggcctggtgcagagctttaccggcaaaccgctgcagggcgtggcggaatggtttagcaaaaccggcgatagctttagcgcgtgggtggaaaaaatgacccgcccgagctggtttaccggcaaaagcccgctggaagtggcgtttggcaacctgggcgataccctgaaaaccattgcggataccctgggcgatgtgggccagaaagcgctggatttttttagcgatccggaaaaaattaaaagctttcgcgatgaactgaccctggtgagcgatgcggtgaaaggcattgcgaccggcattaacggcattgcgaccgcgtatagcagcctgccgctgagcggcgaaggcctgaaaggcctgatgccgattcaggcgcagctgggcgtgaaaatgtttgatggcctgaaagaaagcgcggcgaaagcgtttgcggaagtggcgaccatggcgggcggctttgtgagcaacattggcagcacctttatgagcattggcgataccctgagcggcatttggggcggcattaccaacgcggcggcgaccgcgtttaacagcctggtgagcgcggcgcagagcgcggtgagcggcgtggtgagcgcggtggcgaccattccgggccaggtggcgagcgcgctggcgggcctggcggaagcgggcgcgagcgcgggccgcaacctggtgcagggcctggtgaacggcattagcggcatgattggcagcgcggtggcgaaagcgcgcgaactggcgagcagcgtggcgagcgcggtgaccggctttctgggcattcatagcccgagcaaactgtttaccgaaattggcgaatatgtgggccagggctttgataacggcctgcagagccagctggcgaccctgggccagaccgcgaaagcgatggcggaaaccgtgaccgaagaatttaacggcggcctgaaatttggcgcggatggctttagcaccgatagcgataacccgctgatgcaggcgggcgcgggcctggcgaacgcgccggtggattttgcgaaagcgaccggcaaacagtttctgagcgatctgggcattagcggcaacggcgtgctgagccgcgcgctgaccgaaggcattcagtatatttttcagattagcagcgtggatgaagcgctgagcattaaagatcgcgaaaccagcaaaaacgcgctgagcattgtgggccgc
Codon optimized nucleotide sequence:
ATGGCCCGCAAAGCCGGCATGAGCACCGGCGTGGAAGTCGCGCGTATTAGCGTGAAAGTGGGCCCGGATACCAAACATTTTCGTCGCGAACTGAAAAAAGATCTGGATCGCATTGAAGAAAGCATGCGTGCCAAAATTGACGTAGAACCGGATATGAAAGGCTTTCGTCAGGAAGTGCAGAGCAAAACCAAAGGCATGCGCACCAGCGTGAAAGTGGATGCGGATGTGGATCGCAAAGGCTTCCTGGGCCGTATTGCCGATAGCCTGAGTCAGATTCAGCCGCCTAGCTTTGGCAGCGGCATTAACCCGACCGGTTACGCCGCGATTGCGGCGGGCATTGCCGCGCTGACCCCGCTGATTGCGGGCACCCTGGGCGCGGCGACCACCGCGCTGATGGCCCTGCCGGGCCTGGTGGCGGCGGTGGCGACCCCGATTGCGGCGCTGACTCTGGGCATTGATGGCCTGAAAGCCGCCGCGACCCGCCTGCAGGGTCCGTTTGAAGATCTGAAAGCCACCATGTCGAGCGCCGTGGAAAGCCAATTTGGCCCGGTGTTTGATCAGCTGCGTAGCCTGTTTCCGGTGCTGAAAGGCGCGCTGCCGAGCGTGACCGCGGGTCTTGCGGATATGGCGAAGAGCATTGCCGATGTGATTGCCAGCCCGGAAGGCCTGGCAAAAATTGATACCACCATCCGTAACATTGGCATGGCCCTGACCACCGCGGCCCCGGGCGTTGGCAAATTTACCGATGGTCTGATGGGCCTGGTACAGAGCTTTACCGGCAAACCGCTGCAGGGCGTGGCGGAATGGTTTAGTAAAACCGGTGATTCATTTAGCGCCTGGGTGGAAAAAATGACCCGCCCGAGCTGGTTTACCGGCAAAAGCCCGCTGGAAGTCGCATTCGGCAACCTGGGCGATACCCTGAAAACCATTGCCGATACGCTGGGCGATGTCGGCCAGAAAGCCCTGGATTTTTTTAGCGATCCGGAAAAAATTAAATCGTTTCGCGATGAACTGACCCTGGTGAGCGATGCCGTAAAAGGCATTGCCACCGGTATCAACGGCATTGCCACCGCCTACTCATCCCTGCCGCTGAGCGGCGAAGGCCTGAAAGGCCTGATGCCGATTCAGGCCCAGCTGGGCGTTAAAATGTTTGATGGTCTGAAAGAAAGCGCCGCGAAAGCCTTCGCGGAAGTTGCGACCATGGCAGGCGGCTTTGTGTCGAACATTGGCTCCACCTTTATGAGCATTGGCGATACCCTGAGCGGTATTTGGGGTGGCATTACCAATGCGGCGGCGACCGCGTTTAACAGCCTGGTTAGCGCGGCGCAGAGCGCGGTGAGCGGCGTGGTGAGCGCGGTGGCCACCATTCCGGGTCAGGTGGCGAGCGCCCTGGCGGGCCTGGCCGAAGCAGGCGCCAGCGCCGGCCGCAATCTGGTGCAGGGTCTGGTCAACGGCATTAGCGGCATGATCGGCAGCGCCGTAGCGAAAGCGCGCGAACTGGCGAGCAGCGTGGCGTCAGCGGTTACCGGCTTCCTGGGCATTCACAGCCCGAGCAAATTATTTACCGAAATTGGTGAATATGTGGGCCAGGGCTTCGATAACGGCCTGCAGAGCCAGCTGGCGACTCTGGGTCAGACCGCGAAAGCGATGGCGGAAACCGTGACCGAAGAATTTAACGGCGGCCTGAAATTTGGTGCGGACGGCTTTAGCACCGACAGCGATAATCCGTTAATGCAGGCGGGCGCCGGCCTGGCCAACGCGCCGGTGGATTTTGCGAAAGCCACCGGTAAACAGTTTCTGTCTGATCTGGGTATCAGCGGCAACGGCGTTCTGAGCCGTGCACTGACCGAAGGCATCCAGTATATTTTTCAGATTAGCAGCGTTGATGAAGCGCTGAGCATTAAAGATCGCGAAACCAGCAAAAACGCCCTGAGCATTGTGGGCCGT
This sequence has been optimized for expression in E. coli. In this instance, it would be helpful to do this as these phage satellites were discovered in Mycolicibacterium aichiense, a non-model organism, so expressing this protein in E. coli would make it easier to work with. In general, nucleotide sequences are optimized so that the protein can be more efficiently expressed in a given organism, so that working with it is easier (such as by removing restriction enzyme cut sites), or so that synthesis is easier (such as by lowering the GC content).
Prepare a Twist DNA Synthesis Order
Alyssa1 Tape Measure Protein Expression Cassette in Benchling
Sequence in Benchling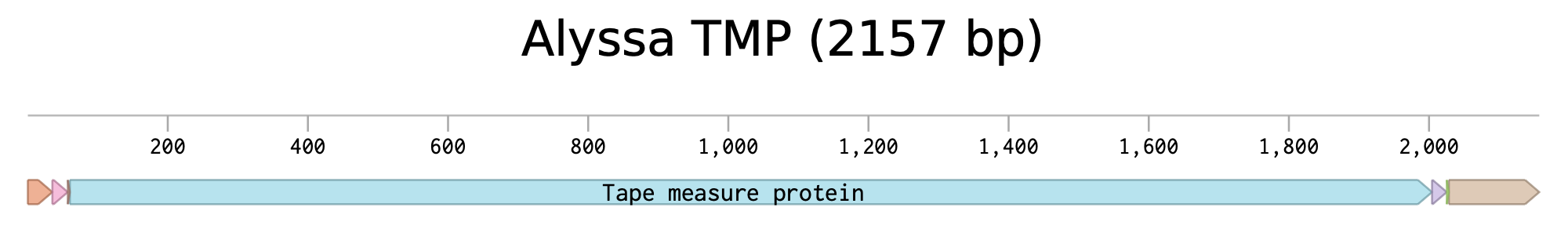

Choosing a Vector in Twist: I chose pTwist Amp High Copy because the Saha Lab works with ampicillin and if I want to express this protein in E. coli, I assume I’d want a lot of protein and for every E. coli descendent to have the plasmid, so a high copy plasmid would work best.
Alyssa1 Tape Measure Protein in Plasmid
Sequence from Twist imported into Benchling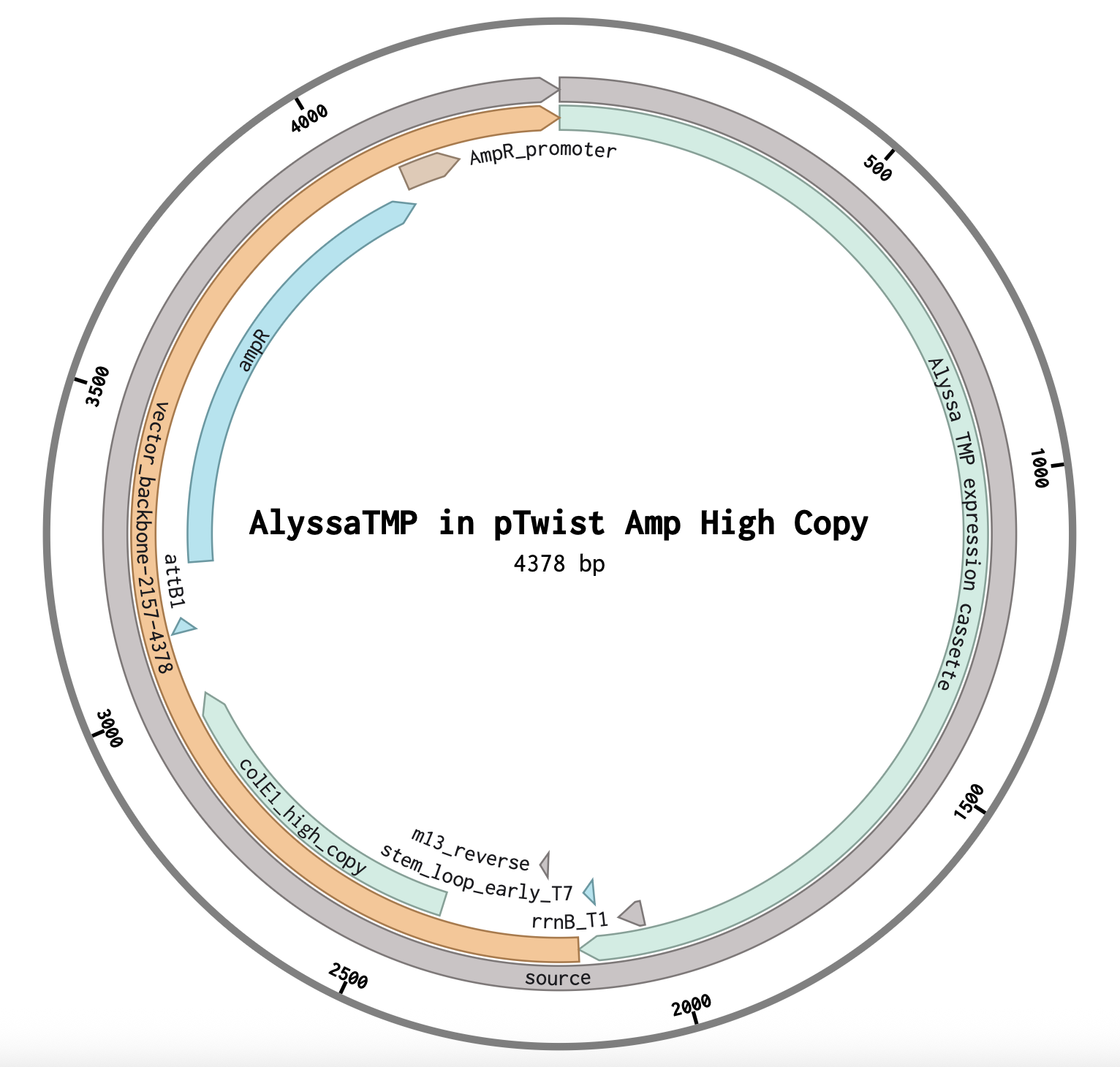

DNA Read/Write/Edit
Read
What DNA would you want to sequence (e.g., read) and why?
I would like to sequence my gut microbiome. I’ve read about how the microbiome, specifically the gut microbiome can influence a person’s behaviors and send signals to the brain. This raises some philosophical questions of whether humans are truly governed by their own thoughts or if their thoughts are being subtly influenced by bacteria in their gut. I think it would be interesting to see what species dominate my microbiome.
In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
Nanopore sequencing it can generate long reads, so it is easier to identify unique features of each species. This makes it easier to assemble genomes.
Write
What DNA would you want to synthesize (e.g., write) and why?
I have always wondered if it would one day be possible to create an organism with “no evolutionary history.” Many of the synthetic biology products/parts now are from preexisting organisms, but is it possible to create an entirely new sequence of DNA that produces proteins with no orthologs? Theoretically, this may be possible, similar to the “monkey at a type-writer” idea, where by chance a random sequence of nucleotides could not only produce a novel protein, but a novel organism. I would like to do this if only to see if it is possible.
What technology or technologies would you use to perform this DNA synthesis and why?
To design this sequence, I would need some kind of machine learning model because biology is extremely complex and cannot be modeled with traditional methods. To synthesize this DNA, I could use the most common method of DNA synthesis, phosphoramidite chemistry, to synthesize pieces of this DNA, then use an assembly technique like Gibson assembly or Golden Gate assembly to construct the genome. These methods are most accessible and are commonly used.
Essential steps: deprotection, add next nucleotide, cap to stop strands that didn’t get the new nucleotide added from growing, and oxidation.
Limitations: this method can’t make fragments longer than ~200 bp due to errors that accumulate.
Edit
What DNA would you want to edit and why? I would like to edit my dog’s DNA to make him age more slowly. My dog makes me very happy but I don’t get to see him much because I’m at college and he lives with my parents. I would love it if he could stick around long enough that I could take care of him once I am out of school.
What technology or technologies would you use to perform these DNA edits and why?
I would use CRISPR to knock in or out genes that I’d need to change. Editing an organism to change its life span would require many edits to the genome, and those edits would likely have to be precise. Because of that, CRISPR would be the best option.
Essential steps: guide RNA recognizes DNA sequence, and the Cas 9 enzyme makes a double-stranded break in the DNA.
Preparation and input: prepare the guide RNA, so I’d need to know the sequence. Inputs include cells, some sort of vector to get the CRISPR-Cas9 system into the cells, and the CRISPR-Cas9 system (enzyme and guide RNA)
Limitations: cuts in off target places, requirement of a PAM sequence, delivery mechanism (finding a vector)