Week 6 HW: Genetic Circuits Part I
DNA Assembly
What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
DNA polymerase - amplifies DNA by adding nucleotides
dNTPs - bases added by DNA polymerase
Buffer - maintains pH and contains cofactors the DNA polymerase requires
What are some factors that determine primer annealing temperature during PCR?
Primer length and GC content.
There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
PCR uses primers to specify which regions of DNA are targeted through amplification. Restriction enzyme digests do not amplify DNA but use restriction enzymes to cut DNA in specific places, potentially cutting out a region of interest. PCR is more flexible in where the primers can bind, so it is easier to extract a region of interest. PCR would also be used when a lot of the region of interest is needed, since it amplifies DNA. Restriction digests are useful for linearizing a plasmid, for example, or creating sticky ends.
How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning? The ends of the fragments must be identical. This can be done by positioning PCR primers appropriately.
How does the plasmid DNA enter the E. coli cells during transformation?
Heat-shocking the E. coli will create pores in the membrane so that DNA can enter.
Describe another assembly method in detail (such as Golden Gate Assembly)
Explain the other method in 5 - 7 sentences plus diagrams (either handmade or online).
Golden Gate assembly uses a restriction enzyme that cuts outside of the enzyme binding site to create sticky ends without scarring. Primers are designed to amplify each fragment used in assembly and to add on the enzyme binding and cut site. The fragments are amplified, then a restriction digest is run to create the sticky ends before assembly. Usually, the fragments are first cloned into a plasmid for stability before being used for assembly.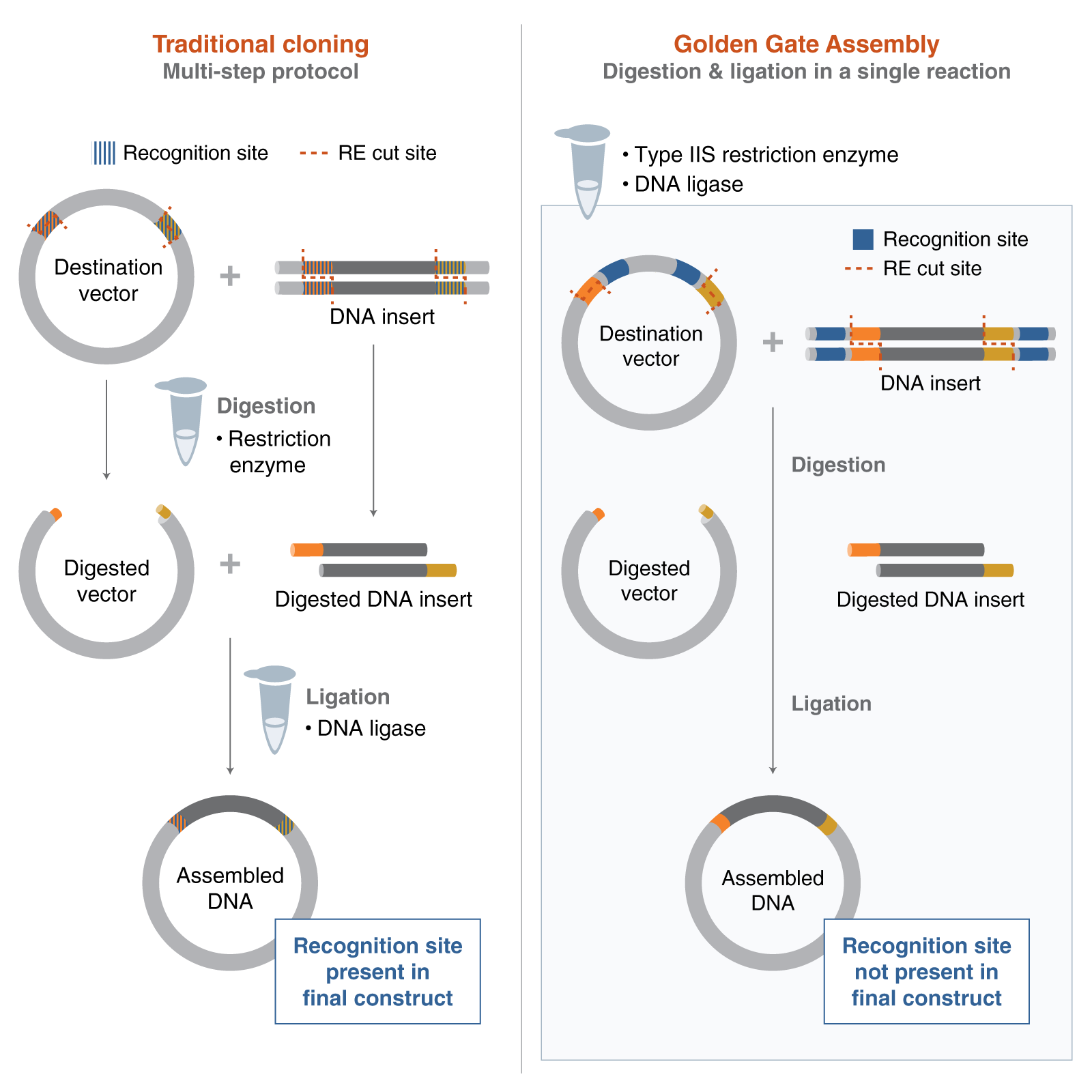

Model this assembly method with Benchling or Asimov Kernel!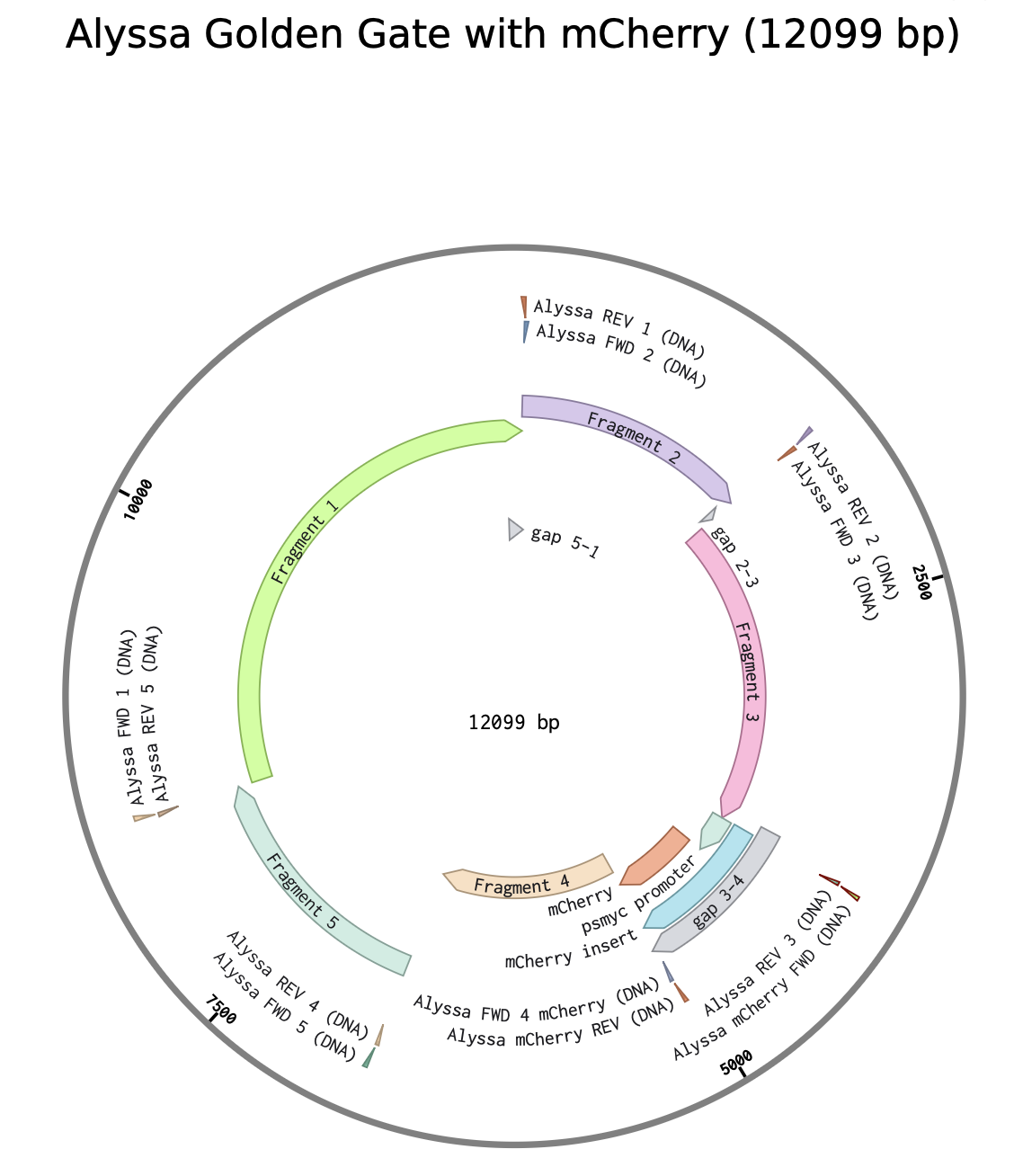

Asimov Kernel
Copied from my notebook in Asimov:
Recreated Repressilator
The graphs appear the same for the repressilator in the Bacterial Parts Demo repository and the one that I’ve created, which makes sense because the same DNA parts and simulation conditions were used.
Construct 1
This construct behaves how I expect it to because there is only one promoter and RBS, so a singular on/off switch. Since there is nothing in this system that would inhibit the expression of LacI, the expression remains constant throughout the time period, which is reflected in the two graphs on the right.
Construct 2
I would expect TetR and LitR levels to start high then decrease as they repress themselves.
The graphs somewhat match my predictions. The RNA graph does have a high concentration at the beginning then rapidly decrease, though I’m not sure why the concentration then stays constant. The protein graph shows less of a change in concentration after the initial spike and also stays constant after dropping. I also don’t know why there would be more expression of TetR than LitR. Maybe the pTetR promoter is stronger?
Construct 3
I would expect this construct to function similarly to the repressilator where expression oscillates over time.
This did not match my prediction, and I’m not sure why. Expression levels mostly rise then stay level throughout time, though some genes are highly expressed initially, then decrease to a stable level.