Week 10 HW: Imaging and Measurement
For properly formatted equations, read the version on GitHub.
Final Project
Please identify at least one (ideally many) aspect(s) of your project that you will measure. It could be the mass or sequence of a protein, the presence, absence, or quantity of a biomarker, etc.
I would measure both fluorescence and OD600. Fluorescence will tell me how well the antirepressors are being expressed, and OD600 will tell me how much the antirepressors affecting bacterial growth.
Please describe all of the elements you would like to measure, and furthermore describe how you will perform these measurements.
Fluorescence: measured using a plate reader.
OD600: measured using a spectrophotometer.
What are the technologies you will use (e.g., gel electrophoresis, DNA sequencing, mass spectrometry, etc.)? Describe in detail.
Plate reader and spectrophotometer.
Waters Part I — Molecular Weight
Based on the predicted amino acid sequence of eGFP (see below) and any known modifications, what is the calculated molecular weight? You can use an online calculator like the one at https://web.expasy.org/compute_pi/.
eGFP Sequence:
MVSKGEELFTG VVPILVELDG DVNGHKFSVS GEGEGDATYG KLTLKFICTT GKLPVPWPTL VTTLTYGVQC FSRYPDHMKQ HDFFKSAMPE GYVQERTIFF KDDGNYKTRA EVKFEGDTLV NRIELKGIDF KEDGNILGHK LEYNYNSHNV YIMADKQKNG IKVNFKIRHN IEDGSVQLAD HYQQNTPIGD GPVLLPDNHY LSTQSALSKD PNEKRDHMVL LEFVTAAGIT LGMDELYKLE HHHHHH
Note: This contains a His-purification tag (HHHHHH) and a linker (the LE before it).
Mw = 28006.60 Da
Calculate the molecular weight of the eGFP using the adjacent charge state approach described in the recitation. Select two charge states from the intact LC-MS data (Figure 1) and:
1. Determine $z$ for each adjacent pair of peaks $(n, n + 1)$ using:
$$z = \frac{\frac{m}{z_{n+1}}}{\frac{m}{z_n} - \frac{m}{z_{n+1}}}$$
$$z(778.2277, 757.3019) = \frac{757.3019}{778.2277 - 757.3019} = 36.1899$$
$$z(1037.4423, 1000.5021) = \frac{1000.5021}{1037.4423 - 1000.5021} = 27.0844$$
2. Determine the MW of the protein using the relationship between $\frac{m}{z_n}$, $MW$, and $z$.
$$MW = z * \frac{m}{z_n} - z*H$$
H = mass of proton (1.0073 Da)
$$MW = 36.1899 * 778.2277 - 36.1899 * 1.0073 = 28127.50 \ Da$$
$$MW = 27.0844 * 1037.4423 - 27.0844 * 1.0073 = 28071.22 \ Da$$
3. Calculate the accuracy of the measurement using the deconvoluted MW from 2.2 and the predicted weight of the protein from 2.1 using:
$$Accuracy = \frac{|MW_{experiment} - MW_{theory}|}{MW_{theory}}$$
$$Accuracy = \frac{|28127.50 - 28006.60|}{28006.60} = 0.43%$$
$$Accuracy = \frac{|28071.22 - 28006.60|}{28006.60} = 0.23%$$
Can you observe the charge state for the zoomed-in peak in the mass spectrum for the intact eGFP? If yes, what is it? If no, why not?
No, because there are many peaks closely clustered together that aren’t fully resolved, so you cannot observe the charge state of the intact eGFP.
Waters Part III — Peptide Mapping - primary structure
How many Lysines (K) and Arginines (R) are in eGFP? Please circle or highlight them in the eGFP sequence given in Waters Part I question 1 above. (Note: adding the sequence to Benchling as an amino acid file and clicking biochemical properties tab will show you a count for each amino acid).
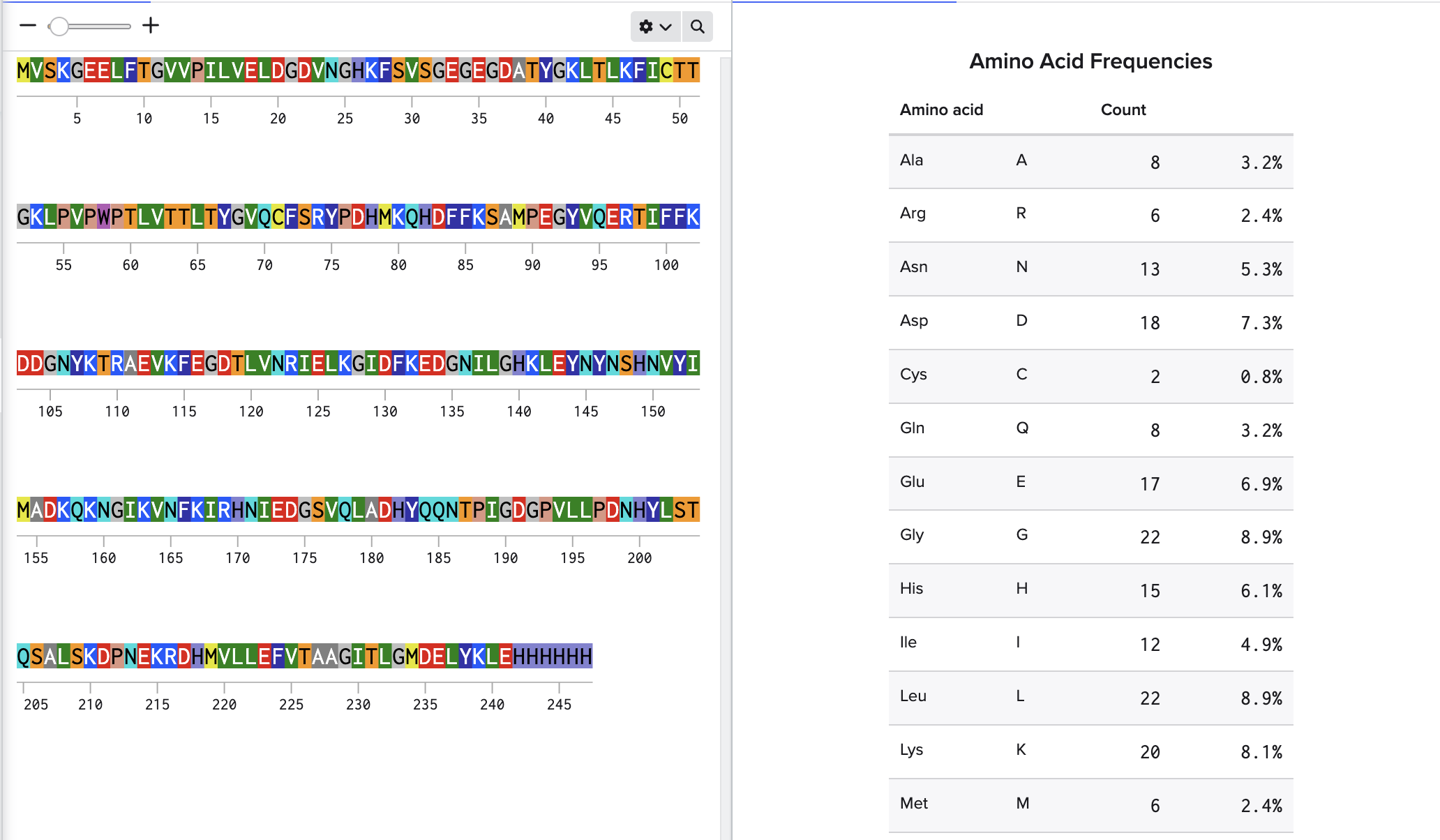

MVSKGEELFTG VVPILVELDG DVNGHKFSVS GEGEGDATYG KLTLKFICTT GKLPVPWPTL VTTLTYGVQC FSRYPDHMKQ HDFFKSAMPE GYVQERTIFF KDDGNYKTRA EVKFEGDTLV NRIELKGIDF KEDGNILGHK LEYNYNSHNV YIMADKQKNG IKVNFKIRHN IEDGSVQLAD HYQQNTPIGD GPVLLPDNHY LSTQSALSKD PNEKRDHMVL LEFVTAAGIT LGMDELYKLE HHHHHH
Lysines: 20
Arginines: 6
How many peptides will be generated from tryptic digestion of eGFP?
19 peptides
Based on the LC-MS data for the Peptide Map data generated in lab (please use Figure 5a as a reference) how many chromatographic peaks do you see in the eGFP peptide map between 0.5 and 6 minutes? You may count all peaks that are >10% relative abundance.
~18 peaks that are >10% relative abundnace.
Assuming all the peaks are peptides, does the number of peaks match the number of peptides predicted from question 2 above? Are there more peaks in the chromatogram or fewer?
It almost matches the number of predicted peptides. It is one fewer.
Identify the mass-to-charge ($\frac{m}{z}$) of the peptide shown in Figure 5b. What is the charge ($z$) of the most abundant charge state of the peptide (use the separation of the isotopes to determine the charge state). Calculate the mass of the singly charged form of the peptide ($[M + H]^+$) based on its $\frac{m}{z}$ and $z$.
$$\frac{m}{z} = 525.76712$$
$$z = 2$$
$$M = z * \frac{m}{z} - z * H$$
$$M = 2 * 525.76712 - 2 * 1.0073 = 1049.5196 \ Da$$
$$[M + H]^+ = 1049.52 + 1.0073 = 1050.5269 \ Da$$
Identify the peptide based on comparison to expected masses in the PeptideMass tool. What is mass accuracy of measurement? Please calculate the error in ppm.
Peptide: FEGDTLVNR
$$Accuracy = \frac{|1050.5269 - 1050.5214}{1050.5214} = 5.23 \ ppm$$
What is the percentage of the sequence that is confirmed by peptide mapping?
90.7%
Waters Part IV — Oligomers
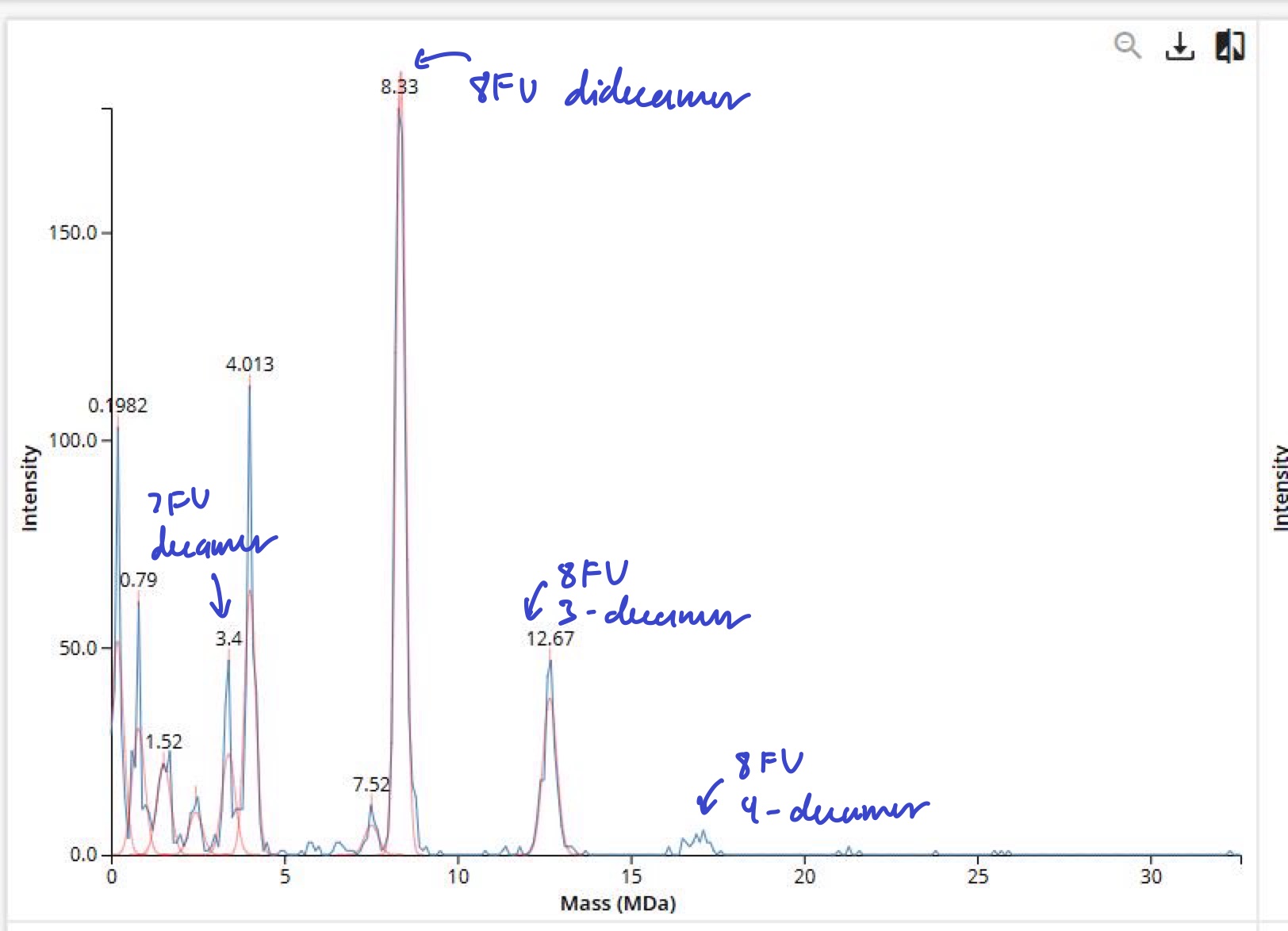

Waters Part V — Did I make GFP?
From screenshot in lab document (LEHHHHHH peptide):
| Theoretical | Observed/measured on the Intact LC-MS | PPM Mass Error | |
|---|---|---|---|
| Molecular weight (kDa) | 1.083498 | 1.083492 | 5.26 |