Week 2 HW: DNA Read, Write, and Edit
Part 1
Benchling & In-silico Gel Art

Part 2
No wet lab access
Part 3
DNA Design Challenge
Choose Protein
I chose the amino acid sequence of VioC - Chromobacterium violaceum for Violacein pigment.
I will reverse translate and codon optimize to amplify pigment production and thus its antimicrobial, UV-resistant properties.
sp|Q9S3U9|VIOC_CHRVO Violacein synthase OS=Chromobacterium violaceum (strain ATCC 12472 / DSM 30191 / JCM 1249 / CCUG 213 / NBRC 12614 / NCIMB 9131 / NCTC 9757 / MK) OX=243365 GN=vioC PE=1 SV=2 MKRAIIVGGGLAGGLTAIYLAKRGYEVHVVEKRGDPLRDLSSYVDVVSSRAIGVSMTVRG IKSVLAAGIPRAELDACGEPIVAMAFSVGGQYRMRELKPLEDFRPLSLNRAAFQKLLNKY ANLAGVRYYFEHKCLDVDLDGKSVLIQGKDGQPQRLQGDMIIGADGAHSAVRQAMQSGLR RFEFQQTFFRHGYKTLVLPDAQALGYRKDTLYFFGMDSGGLFAGRAATIPDGSVSIAVCL PYSGSPSLTTTDEPTMRAFFDRYFGGLPRDARDEMLRQFLAKPSNDLINVRSSTFHYKGN VLLLGDAAHATAPFLGQGMNMALEDARTFVELLDRHQGDQDKAFPEFTELRKVQADAMQD MARANYDVLSCSNPIFFMRARYTRYMHSKFPGLYPPDMAEKLYFTSEPYDRLQQIQRKQN VWYKIGRVN
Reverse translate
sp|Q9S3U9|VIOC_CHRVO Violacein synthase OS=Chromobacterium violaceum (strain ATCC 12472 / DSM 30191 / JCM 1249 / CCUG 213 / NBRC 12614 / NCIMB 9131 / NCTC 9757 / MK) OX=243365 GN=vioC PE=1 SV=2 ATGAAGCGAGCGATTATTGTCGGGGGGGGTTTAGCTGGAGGTCTAACTGCGATATACTTGGCTAAACGTGGATACGAGGT ACATGTGGTCGAGAAACGGGGCGACCCACTCAGGGACCTGTCTAGCTATGTTGATGTGGTTTCATCACGCGCAATCGGGG TCAGCATGACTGTAAGAGGCATCAAGTCAGTTTTAGCGGCCGGTATCCCCCGAGCTGAATTAGATGCCTGTGGTGAGCCA ATAGTTGCCATGGCGTTTTCCGTCGGGGGACAATATCGCATGCGGGAACTTAAACCACTCGAAGACTTCCGACCGCTTTC GCTTAACCGAGCAGCCTTCCAGAAGCTTTTGAACAAGTACGCAAACCTTGCCGGCGTACGGTACTATTTCGAACATAAAT GCCTGGATGTAGACCTGGATGGGAAATCCGTACTGATCCAAGGGAAGGACGGACAGCCGCAGCGACTTCAAGGAGATATG ATTATCGGCGCAGATGGGGCACACAGTGCAGTTCGCCAAGCGATGCAGTCAGGATTGCGGCGCTTTGAGTTTCAACAAAC GTTCTTTAGGCACGGGTATAAAACGCTGGTCCTACCCGACGCCCAAGCACTCGGGTATCGAAAGGACACGTTATATTTTT TTGGAATGGACAGCGGAGGGTTGTTCGCAGGCCGAGCCGCAACAATACCCGATGGTAGCGTGTCCATAGCTGTGTGTCTG CCCTACTCCGGCTCCCCCAGTTTGACAACCACAGATGAACCGACTATGCGTGCATTTTTCGACAGGTACTTTGGAGGTCT TCCACGGGATGCGAGGGACGAGATGCTTAGACAATTTTTAGCCAAGCCGTCTAATGATCTAATAAATGTGCGATCTTCAA CTTTTCATTACAAAGGTAACGTTCTGCTTTTAGGCGACGCCGCACATGCTACCGCGCCATTTTTAGGACAAGGCATGAAT ATGGCGTTAGAGGATGCGCGAACATTCGTAGAATTACTTGATCGCCACCAAGGCGATCAGGATAAAGCGTTTCCAGAGTT CACGGAGCTTAGAAAGGTGCAAGCGGACGCGATGCAAGATATGGCCCGGGCGAATTACGATGTTCTATCTTGCTCCAACC CGATTTTTTTTATGAGGGCGCGGTATACCCGCTACATGCACAGCAAGTTTCCGGGACTGTACCCGCCGGATATGGCCGAG AAACTGTATTTCACGTCAGAGCCGTACGATCGATTACAACAAATACAGCGCAAGCAAAACGTATGGTACAAGATAGGCAG AGTTAAT
Codon Optimize

https://en.vectorbuilder.com/tool/codon-optimization/b93b7790-7536-4d9b-a72e-02d62c3944e8.html
Next Next steps would be to embed into a seaweed matrix.
Part 4
Prepare a Twist DNA Synthesis Order
After reading more on living materials, bacterial pigments, and connecting it to my interest in light and circadian rhythms, I wanted to explore how to make a simple biological system that expresses anti-microbial or other elements only when needed, rather than all the time. So building a ’temporal’ antimicrobial system that produces a bacteria-killing peptide Magainin on a 24-hour schedule controlled by a circadian promoter RpaA. I started with just learning how to design the Magainin peptide and annotate properly.
Benchling
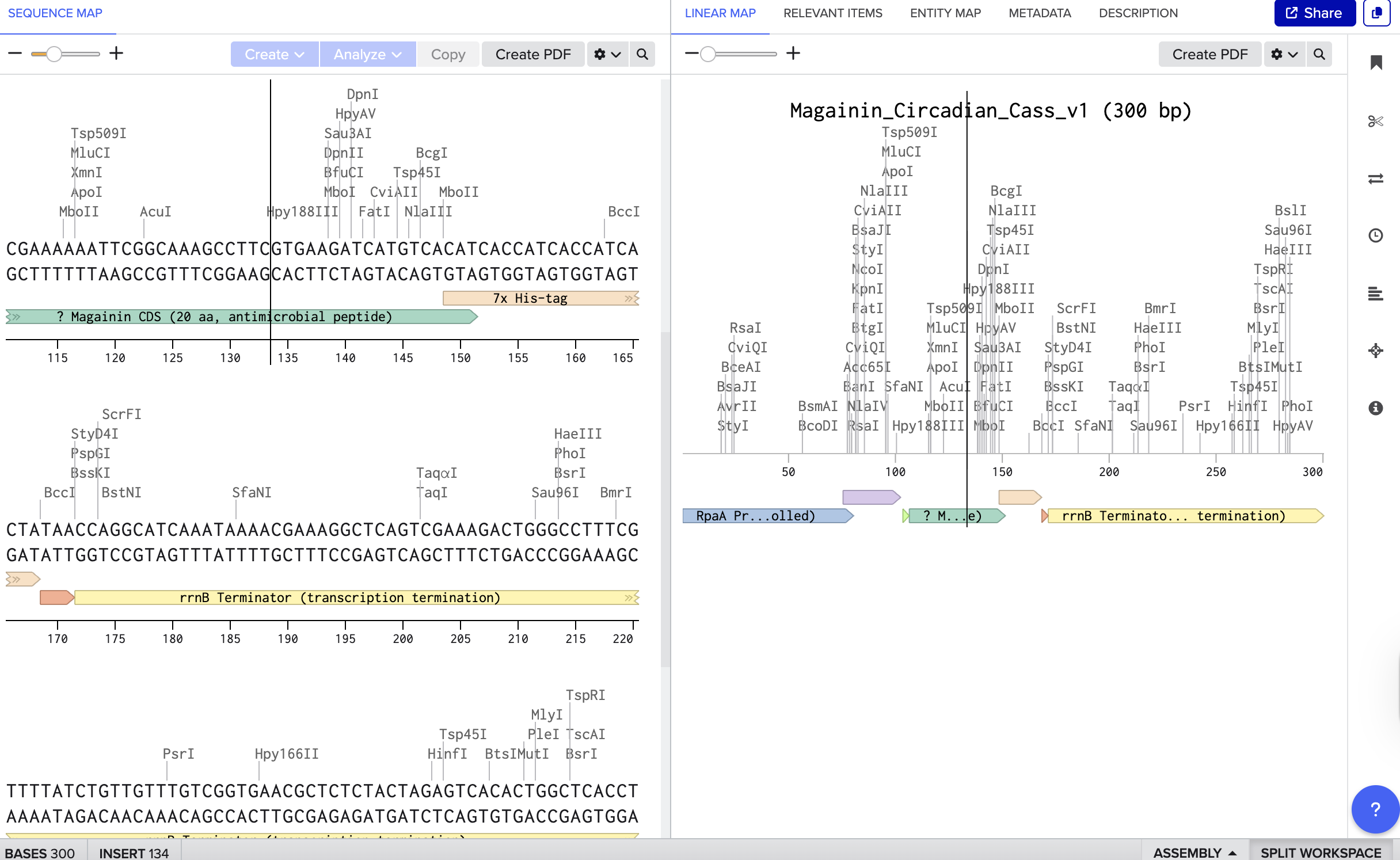
Twist
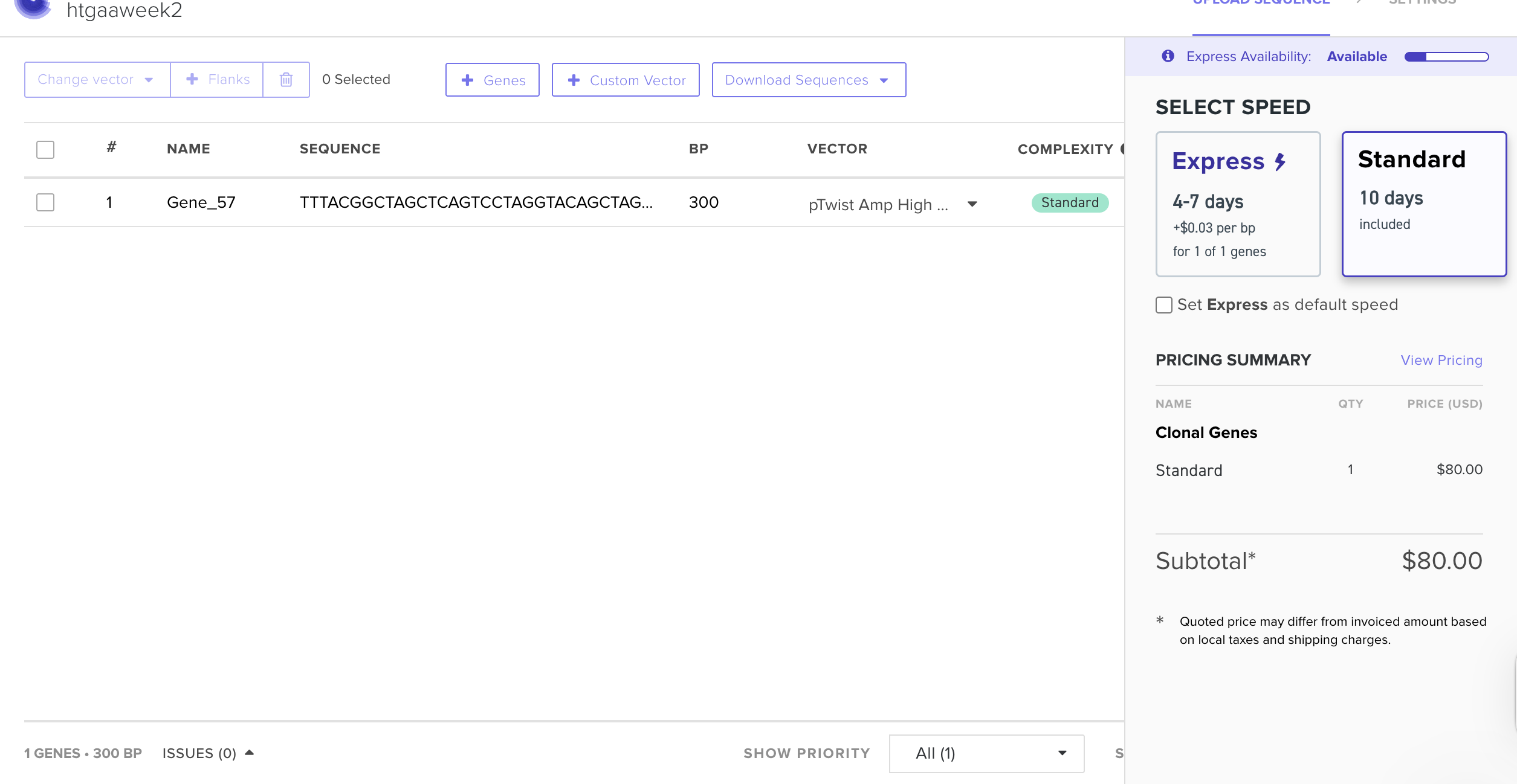
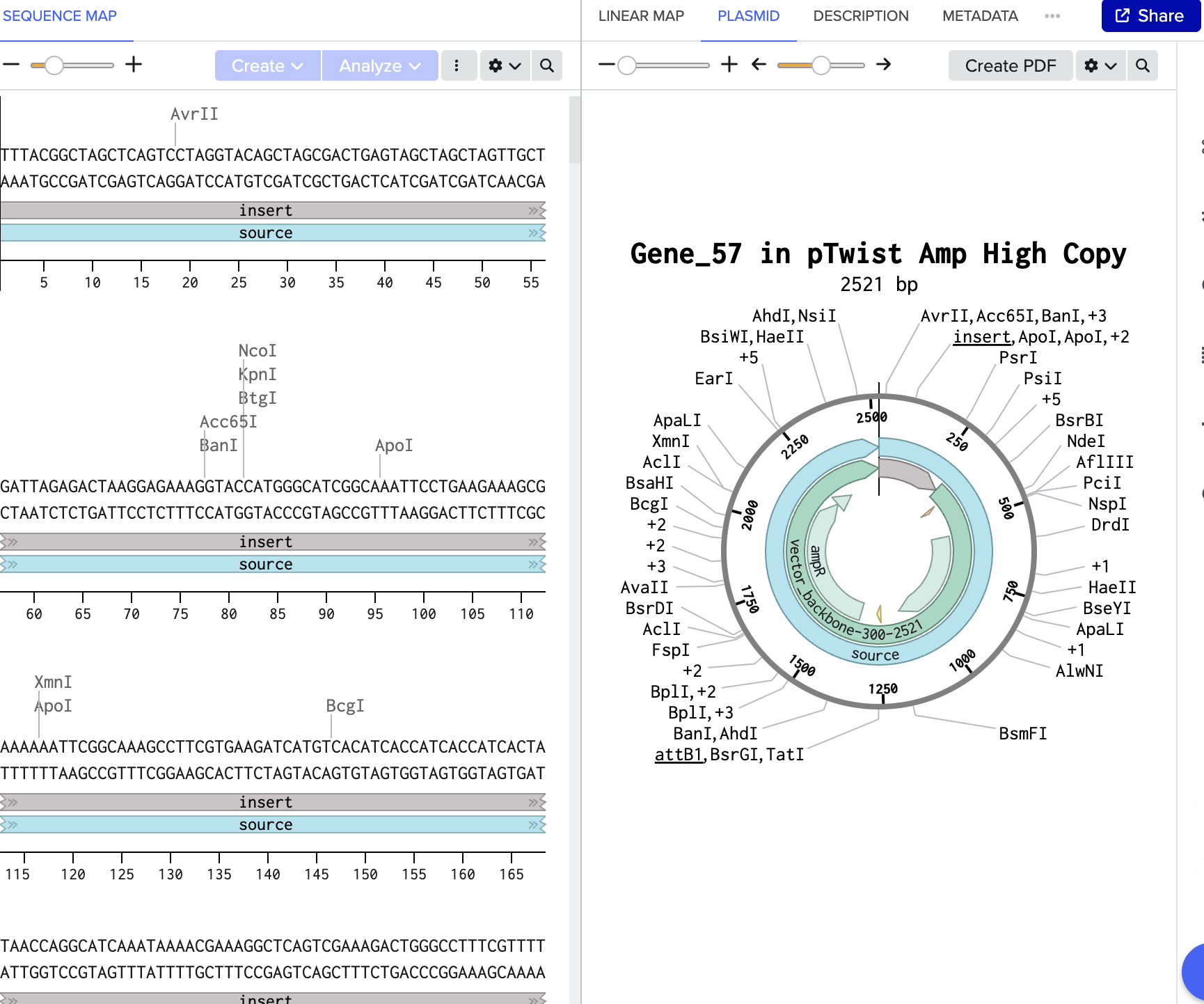
REF:
- Fang et al. (2025) - “Mechanism and reconstitution of circadian transcription in cyanobacteria”
- Salis et al. (2009) - “Automated Design of Synthetic Ribosome Binding Sites”
- Westerhoff et al. (2008) - “Structure, Membrane Orientation, Mechanism, and Function of Pexiganan (Magainin derivative)”
Part 5
DNA Read/Write/Edit
DNA Read (Sequencing)
What DNA would you want to sequence and why?
I would sequence my pLight-Circadian-Color plasmid (which contains the RpaA gene from Synechococcus elongatus, an anthocyanin color gene, and a light sensor) to check that it was made correctly before testing if bacteria with this plasmid change color on a 24-hour schedule when exposed to light.
What sequencing technology would you use?
I would use Sanger sequencing because it’s most accurate.
DNA Write (Synthesis)
What DNA would you synthesize and why?
I would synthesize my yet-to-be-completed pLight-Circadian-Color plasmid containing three genes (RpaA from Synechococcus elongatus for timing, anthocyanin for color, light sensor for activation) to test if bacteria can change color on a 24-hour schedule in response to light.
DNA Edit
What DNA would you edit and why?
After I verify the plasmid works, I would edit the RpaA promoter to make it stronger so the color changes are brighter and more noticeable on a 24-hour schedule.
What editing technology would you use?
I would use site-directed mutagenesis to make small changes to the RpaA promoter because it’s precise.
References & Resources
Lecture Materials
- Week 2 Lecture - DNA Read, Write, & Edit, George Church, Joe Jacobson, Emily Leproust
- Week 2 Lab - DNA Gel Art, February 12-13, 2026
Required Readings
- Fang et al. (2025). “Mechanism and reconstitution of circadian transcription in cyanobacteria.” Journal of Biological Chemistry
- Salis et al. (2009). “Automated Design of Synthetic Ribosome Binding Sites to Control Protein Expression.” Nature Biotechnology, 27, 946-950
- Westerhoff et al. (2008). “Structure, Membrane Orientation, Mechanism, and Function of Pexiganan (Magainin derivative).” Biochemistry
Software & Tools Used
- Benchling - DNA sequence design, annotation, and in-silico gel electrophoresis
- Twist Bioscience - DNA synthesis order preparation and optimization
- VectorBuilder Codon Optimization Tool - Reverse translation and codon optimization for violacein synthase
- UniProt - Protein sequence database (VioC entry: sp|Q9S3U9|VIOC_CHRVO)
- Imgur - Image hosting for documentation
Sequences Worked With
- VioC (Violacein synthase) from Chromobacterium violaceum strain ATCC 12472
- RpaA circadian promoter from Synechococcus elongatus
- Magainin antimicrobial peptide sequence
AI Assistance
- Claude (Anthropic) - DNA design and sequencing strategy
- Model: Claude Sonnet 4.5
- Date(s) used: February, 2026
- Tasks: Assisted with reverse translation strategy for VioC, guidance on codon optimization principles, clarified Sanger sequencing vs synthesis tradeoffs
Project Development
- Circadian-controlled antimicrobial system design (RpaA + Magainin)
- Violacein pigment amplification through codon optimization
- pLight-Circadian-Color plasmid conceptual design
Additional Resources
- Twist Bioscience synthesis guidelines and specifications
- Benchling annotation standards
- Circadian rhythm gene expression literature
Acknowledgments
- Course instructors
- TAs for Benchling platform guidance