Subsections of Projects
Individual Final Project
HTGAA Spring 2026
The Exoskin Project: A Spider Silk-Snow Fungus Bioactive Biomaterial for Wound Healing and Beyond

Abstract
Exoskin asks a central research question: What new material(s) and relational possibilities become available when synthetic biology is used to design co-relational constructs across biological kingdoms? How far can that co-relational construct extend its structural, functional, and environmental range? We start this research with the main players, MaSp1 spider silk and the Tremella fuciformis (snow fungus) polysaccharide, through a CBM27 carbohydrate-binding domain, creating a designed molecular anchor rather than a physical blend. The result is a platform with thus far two construct variants.
Construct 1 — BioMechSkin (CBM27_RGD_MaSp1_4x): Targets diabetic foot ulcers, which affect 15-25% of all people with diabetes and are the leading cause of non-traumatic lower limb amputation worldwide. Current dressings do not simultaneously address moisture retention, mechanical resistance, and tissue repair. This construct does: Tremella provides moisture retention and macrophage modulation, MaSp1 provides tensile strength and friction resistance, and an RGD motif recruits fibroblasts and keratinocytes directly to the wound surface. AlphaFold3 confirms CBM27 pLDDT above 90, RGD solvent exposed and distal, MaSp1 disordered as expected.
Construct 2 — Photomechanical Actuator (VVD_GGS_CBM27_MaSp1_4x): Adds a VVD LOV domain from Neurospora crassa to the same chassis. A blue light pulse triggers VVD dimerisation, contracting the silk hydrogel network without any electronic actuator. This opens the platform to soft robotics for Mars and space environments, directly addressing a NASA-identified gap in electronics-free actuation under extreme conditions. AlphaFold3 and MolProbity validation at the 92nd percentile confirm structural integrity of the construct.
Both constructs share the same CBM27-Tremella-MaSp1 backbone, establishing it as a generalizable platform for encoding function at the protein sequence level.
Long term, this project unfolds in two phases:
Phase 1: Computational and Speculative explores how many material combinations and outputs become possible by extending the co-relational design philosophy across different kingdom pairs and molecular architectures. This phase runs entirely in silico, using AlphaFold3, molecular dynamics, and design iteration to map the conceptual space of what cross-kingdom molecular design can achieve.
Phase 2: Wet Lab Validation (mid-year onwards) will test the most promising constructs through cell-free protein synthesis and composite assembly, either through Node partnership or collaboration with the University of Johannesburg where I am artist in residence. This two-step approach allows the philosophical and practical dimensions to develop in parallel.
Introduction
Exoskin asks a broad single question: can cross-kingdom molecular design produce a material whose functional range exceeds what either organism achieves alone, and what emerges when that architecture is pushed to its limits? Two constructs are proposed as answers, both built on the same CBM27-Tremella-MaSp1 backbone. The first, CBM27_RGD_MaSp1_4x, targets wound healing. The second, VVD_GGS_CBM27_MaSp1_4x, adds a light-responsive LOV domain for photomechanical actuation in extreme environments. Together they test whether a single molecular platform can span from clinical wound care to space robotics. Ethically and conceptually, the project explores boundaries between human, animal, and fungal biology and raises questions about posthuman embodiment and hybridity.
The wound care construct is optimised for diabetic foot ulcers through two functional additions to the MaSp1 spider silk core:
Broadly Exoskin has applications in tissue engineering, drug delivery and regenerative medicine. Ethically and conceptually, the project explores traditional boundaries between human, animal, and fungal biology and raise questions about posthuman embodiment, hybridity, and science.
Here it is optimised for wound healing through two functional additions to the MaSp1 spider silk core:
- CBM27 domain from Thermotoga maritima Man5 mannanase to physically anchor the silk network onto Tremella fuciformis polysaccharide matrix
- RGD motif specifically selected to recruit skin fibroblasts and keratinocytes to the wound surface
Unlike passive biomaterials, this construct actively participates in tissue repair through three simultaneous mechanisms:
- Structural support from silk nanofibrils
- Moisture retention and macrophage modulation from Tremella polysaccharide
- Integrin-mediated skin cell recruitment from the RGD motif
Section 1: Project Aims
Aim 1 - Experimental: Design and computationally validate the CBM27_RGD_MaSp1_4x fusion construct using Benchling, Twist Biosciences, and AlphaFold3. see update below from Synbio 2026 confererence regarding Sidewinder - Robinson, N.E., Zhang, W., Ghosh, R., Gerber, B., Zhang, H., Sanfiorenzo, C., Wang, S., Di Carlo, D. and Wang, K. (2026) ‘Construction of complex and diverse DNA sequences using DNA three-way junctions’, Nature, 651(8105), pp. 491-500. https://doi.org/10.1038/s41586-025-10006-0
Aim 2a - Expression: Validate through cell-free protein synthesis or bacterial expression, confirmed by SDS-PAGE and polysaccharide-binding assay. (Hook Note - The tremella polysaccharide structure consists of a main chain of β-(1→3)-linked mannopyranoside, substituted with various side chains, which your CBM27 domain is designed to recognize. Ref: nih)
Aim 2b - Composite Assembly (contingent on Aim 2a): Combine the purified protein with Tremella fuciformis polysaccharide extract to assemble the composite hydrogel and characterise by rheology and fluorescence microscopy. - Ginkgo Bioworks / Node — target partners for cell-free expression (Aim 2a); plasmid submitted to Node for synthesis
Aim 3 - Visionary: Application and prototype wound dressing. Longer term, the MaSp1 silk fibre bundle contraction properties open a pathway to bio-inspired soft robotics actuators and exoskins driven by hydrogel mechanics. A light-responsive actuation variant is under exploration. Bio kinetic sculptures for moisture harvesting in desert areas such as Karoo. Very long term Engineer Tremella fuciformis to produce the silk fusion protein endogenously, creating a self-assembling living composite.
Other Applications
| Application | Description |
|---|---|
| Tissue Engineering Scaffold | Skin, cartilage, soft tissue repair |
| Drug Delivery Matrix | Sustained localised therapeutic release |
| Soft Robotics Actuator | Bio-inspired hydrogel-driven movement |
| Wearable Biosensor Substrate | Flexible, skin-conforming sensor base |
| Self-Healing Biomaterial | Reversible CBM27-mannan re-annealing |
Leverage Sidewinder DNA Assembly Technology for Robust Synthesis of CBM27_RGD_MaSp1_4x Fusion Protein Construct
The CBM27_RGD_MaSp1_4x fusion protein design presents a DNA synthesis challenge inherent to spider silk engineering: its four tandem repeats of the MaSp1 domain create a highly repetitive sequence that conventional assembly methods cannot reliably handle. Traditional DNA assembly techniques (PCA, Gibson assembly, Golden Gate) achieve misconnection rates of 1 in 10 to 1 in 30 and fail consistently beyond 5-10 fragment assemblies. This bottleneck is particularly acute for repetitive biomaterials like spider silk.
However, Sidewinder, a DNA assembly technology recently published in Nature (January 2026) by Caltech researchers Kaihang Wang and colleagues, addresses this limitation directly. Sidewinder uses DNA three-way junctions to separate assembly instructions from the final sequence, achieving a misconnection rate of approximately 1 in 1,000,000 and enabling flawless assembly of highly repetitive constructs. The technology was prominently featured at SynBioBeta 2026 (May 6, 2026) by Genyro co-founders Adrian Woolfson and Professor Wang, who presented Sidewinder as infrastructure for translating AI-designed biology into physical systems at scale. As a future aim or complementary approach, Sidewinder could be employed to synthesize the full pET28a plasmid containing the CBM27_RGD_MaSp1_4x insert with guaranteed sequence fidelity, ensuring reliable bacterial expression (Aim 2a), accurate characterization of hydrogel mechanics (Aim 2b), and enabling rapid design-build-test-learn iterations for optimizing the Tremella-silk interaction (Aim 3).
Application or Look and Feel of the Material
After expression and purification, the protein is mixed with Tremella polysaccharide extract. The CBM27 domain automatically anchors the silk into the fungal matrix and MaSp1 self-assembles into fibres, forming a hydrogel sheet with no chemical crosslinking needed. That sheet is cut to size and placed directly on the wound. On a diabetic foot ulcer it does three things at once: Tremella keeps the wound moist and calms chronic inflammation, the silk holds the structure together under the mechanical stress of walking, and the RGD motif pulls the patient’s own skin cells into the wound to drive healing. No cells are delivered, no drugs, just a protein-polysaccharide scaffold that creates the right conditions for the body to repair itself.
Section 2: The Constructs
The fusion protein is built from four functional units arranged N to C terminus:
| Domain | Position | Function |
|---|---|---|
| CBM27 (buffered) | aa 1-176 | Anchors silk to Tremella mannan |
| GGGGS x3 linker | aa 177-188 | Domain flexibility |
| RGD motif | aa 189-193 | Skin cell recruitment |
| MaSp1 x4 repeats | aa 194-333 | Mechanical structure |
| 6xHis tag | aa 334-340 | Ni-NTA purification |
340 amino acids · ~37 kDa · cloned into pET28a at NdeI/XhoI · 6,297 bp total plasmid


CBM27_RGD_MaSp1_4x cloned into pET28a(+) at NdeI/XhoI · 6,297 bp
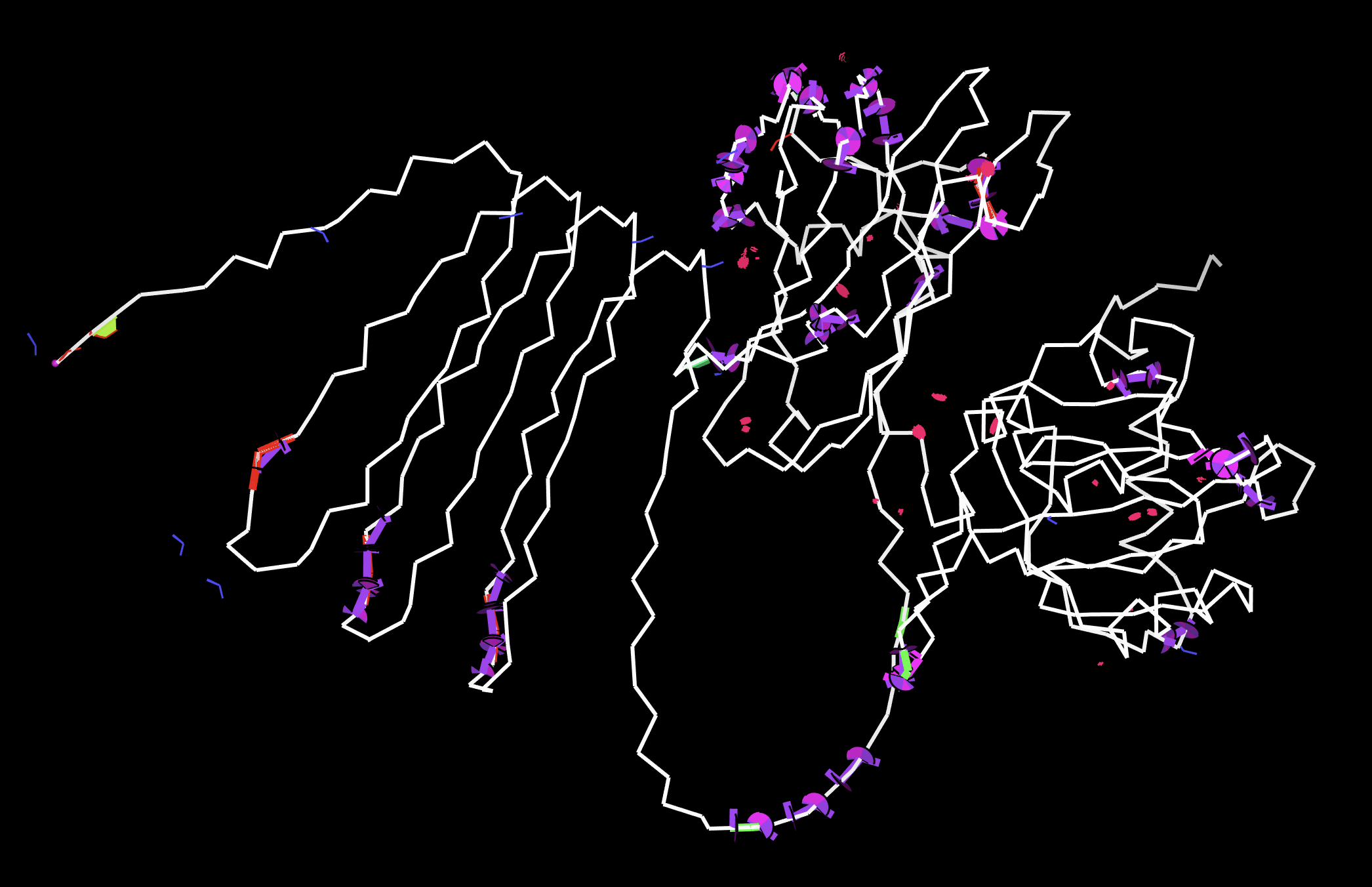
AlphaFold3 Structure Prediction of CBM27_RGD_MaSp1_4x
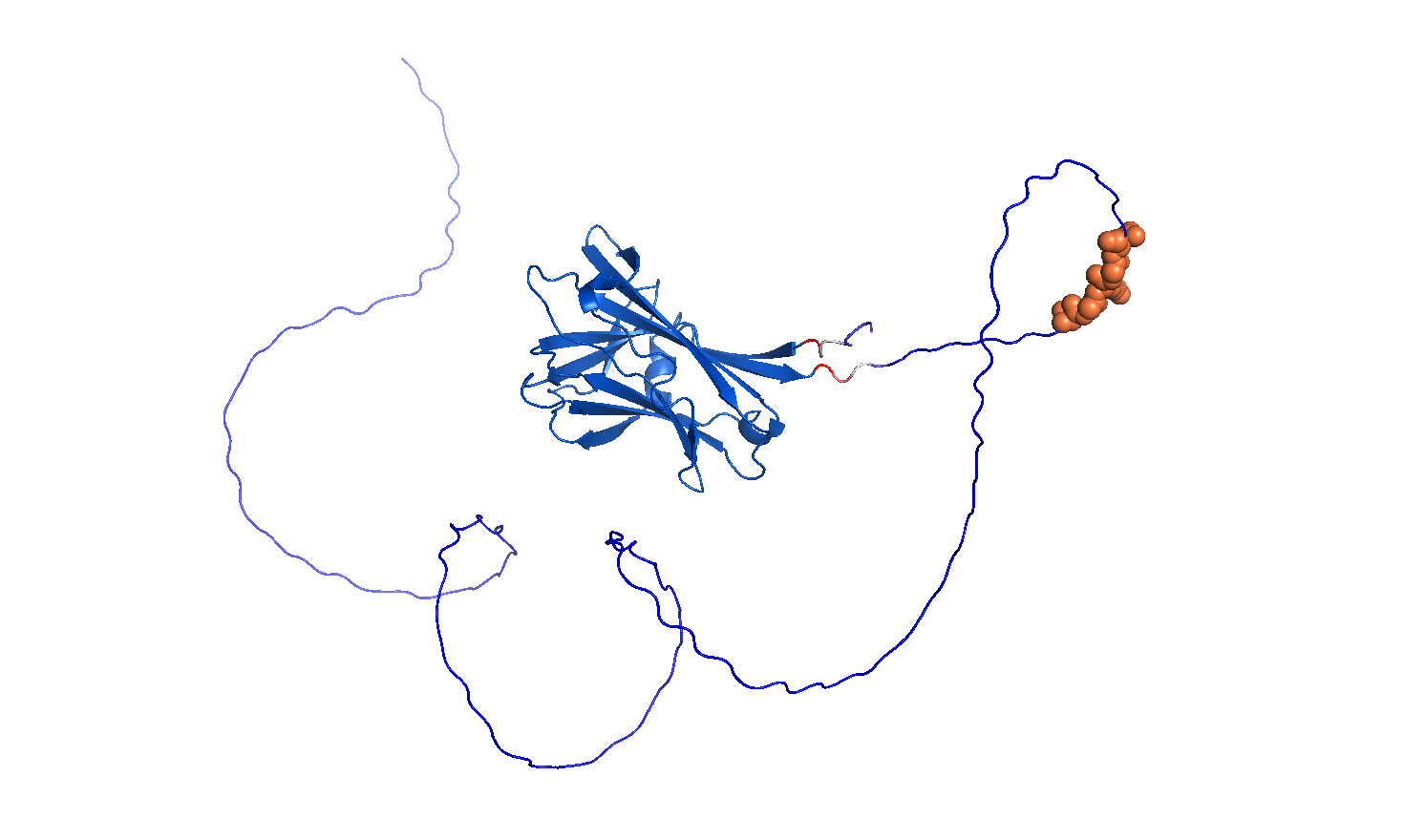
Model_0, Coloured by pLDDT confidence using standard AlphaFold convention. Rendered in PyMOL.
The AlphaFold3 prediction of my full fusion construct CBM27_RGD_MaSp1_4x reveals a structurally coherent architecture consistent with my design intent. The central dark blue beta-sandwich is the CBM27 domain (residues 1-176), predicting with very high confidence (pLDDT >90) and adopting the characteristic jellyroll fold seen in the experimentally validated TmCBM27 crystal structure (PDB: 1OF4, Boraston et al. 2003). This confirms that the CBM27 domain folds correctly within the fusion context and is not destabilised by the adjacent linker or silk repeat regions.
Extending from the CBM27 domain, the white and red transitional region marks the GGS flexible linker and the boundary into lower confidence territory. Immediately following this, the orange spheres identify the RGD motif (residues 189-193), which sits fully solvent exposed and spatially distal from the CBM27 body. This exposed geometry is the ideal configuration for integrin binding in wound care, biological accumulation in the sculpture context, and surface functionalisation in cryogenic assay conditions. The large blue disordered loops extending around the structure are the four MaSp1 silk repeat units (residues 194-334). These predict as intrinsically disordered, which is the expected and correct behaviour for recombinant spider silk repeat proteins prior to fibre assembly or hydrogel casting. The confidence gradient across the silk loops, darker navy closest to CBM27 fading to lighter periwinkle at the distal ends, reflects the decreasing positional constraint as the chain moves further from the structured anchor domain.
Taken together, this prediction computationally validates three key design features:
- CBM27 folds correctly in the fusion context
- RGD is spatially accessible
- MaSp1 remains dynamically disordered as required for silk network formation
A note on the MaSp1 prediction: It is important to note that AlphaFold3 has known limitations in predicting intrinsically disordered repeat proteins, particularly those with highly repetitive glycine and alanine rich sequences such as spider silk. The MaSp1 backbone trace shown here should not be interpreted as a meaningful 3D conformation. The disordered loops represent AlphaFold3 acknowledging that it cannot assign confident positional coordinates to these residues, not that the silk adopts this specific extended loop geometry in solution. The true behaviour of MaSp1 repeats is well established in the literature: they remain disordered in aqueous solution and assemble into beta-sheet rich nanofibrils upon shear stress, pH change, or drying (Rising et al. 2011). The computational validation of the silk region will be addressed separately through molecular dynamics simulation of the repeat unit assembly behaviour rather than single chain folding prediction, which is not the appropriate tool for this class of protein.

AlphaFold3 confidence metrics:
The overall pTM score for the full fusion construct is 0.54, which reflects the presence of the large intrinsically disordered MaSp1 repeat region rather than indicating poor prediction quality for the structured domains. pTM scores are sensitive to disordered regions and will always be suppressed in constructs containing long flexible or repeat sequences. When interpreted domain by domain, the CBM27 region (residues 1-176) shows pLDDT values consistently above 90, placing it in the very high confidence category and confirming that the folded domain is predicted reliably. The RGD motif and GGS linker score below 50 as expected for short flexible functional motifs. The MaSp1 repeats score variably across the disordered range, consistent with their known intrinsically disordered character in solution.
For comparison, the experimentally validated crystal structure of isolated TmCBM27 (PDB: 1OF4, Boraston et al. 2003, 1.6 Angstrom resolution) confirms the expected beta-sandwich jellyroll fold for this domain. The high confidence CBM27 prediction in our fusion construct (pLDDT >90, residues 1-176) is structurally consistent with this experimentally solved reference, confirming that domain-level folding is maintained even within the multi-domain fusion context. https://www.rcsb.org/structure/1OF4
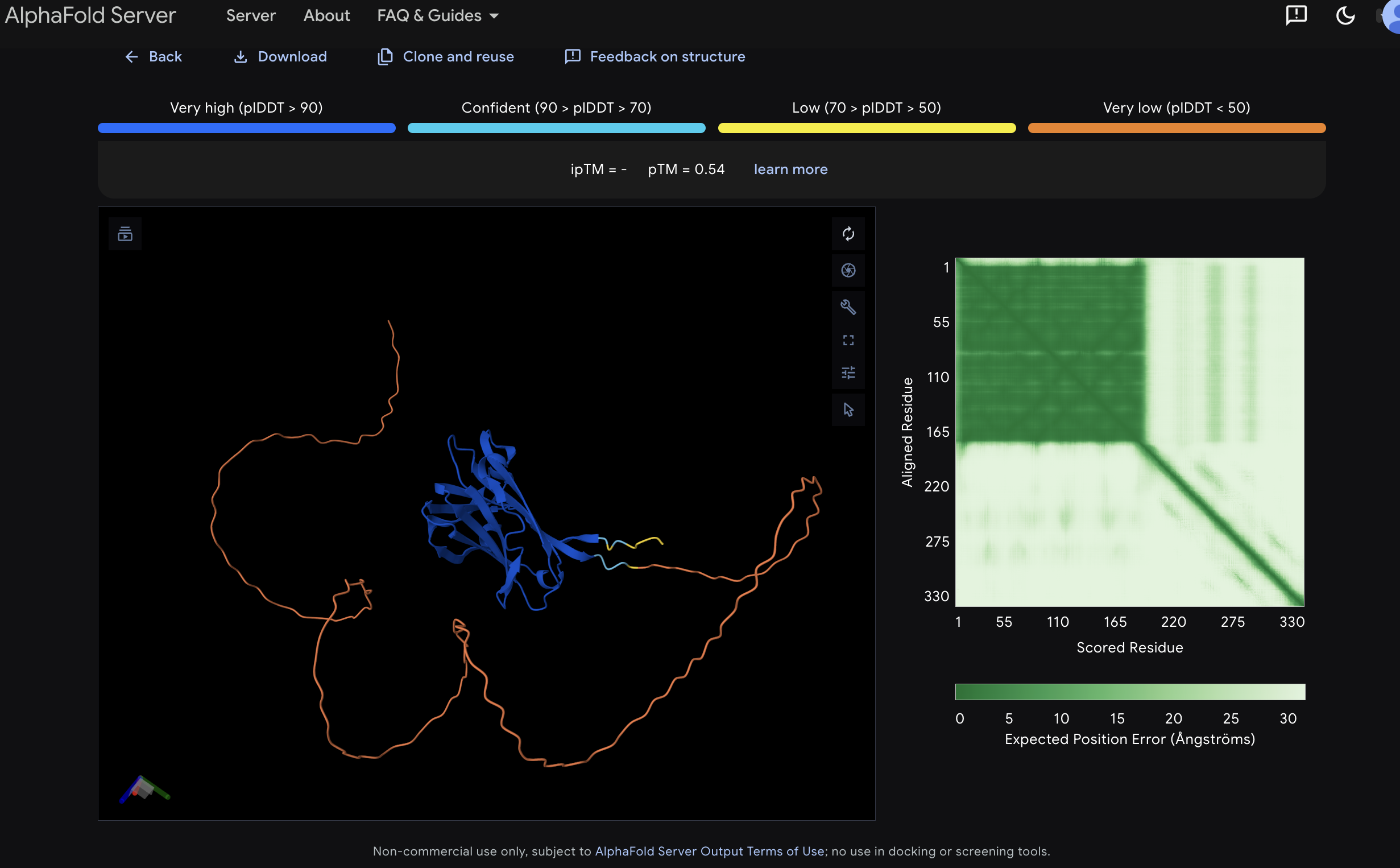
Construct 2: VVD_GGS_CBM27_MaSp1_4x
The second construct adds a light-responsive LOV domain to the same chassis, targeting photomechanical soft robotics for Mars and space environments.
| Domain | Position | Function |
|---|---|---|
| VVD (LOV domain, Neurospora crassa) | N-terminus | Light-sensitive blue light receptor; dimerises under ~450 nm, contracting the silk hydrogel network |
| GGS linker | — | Flexible linker between VVD and CBM27 |
| CBM27 | — | Anchors silk to Tremella mannan polysaccharide |
| MaSp1 x4 repeats | C-terminus | Mechanical silk scaffold |
AlphaFold3: pTM 0.45. VVD and CBM27 fold independently with high confidence. MaSp1 disordered as expected.
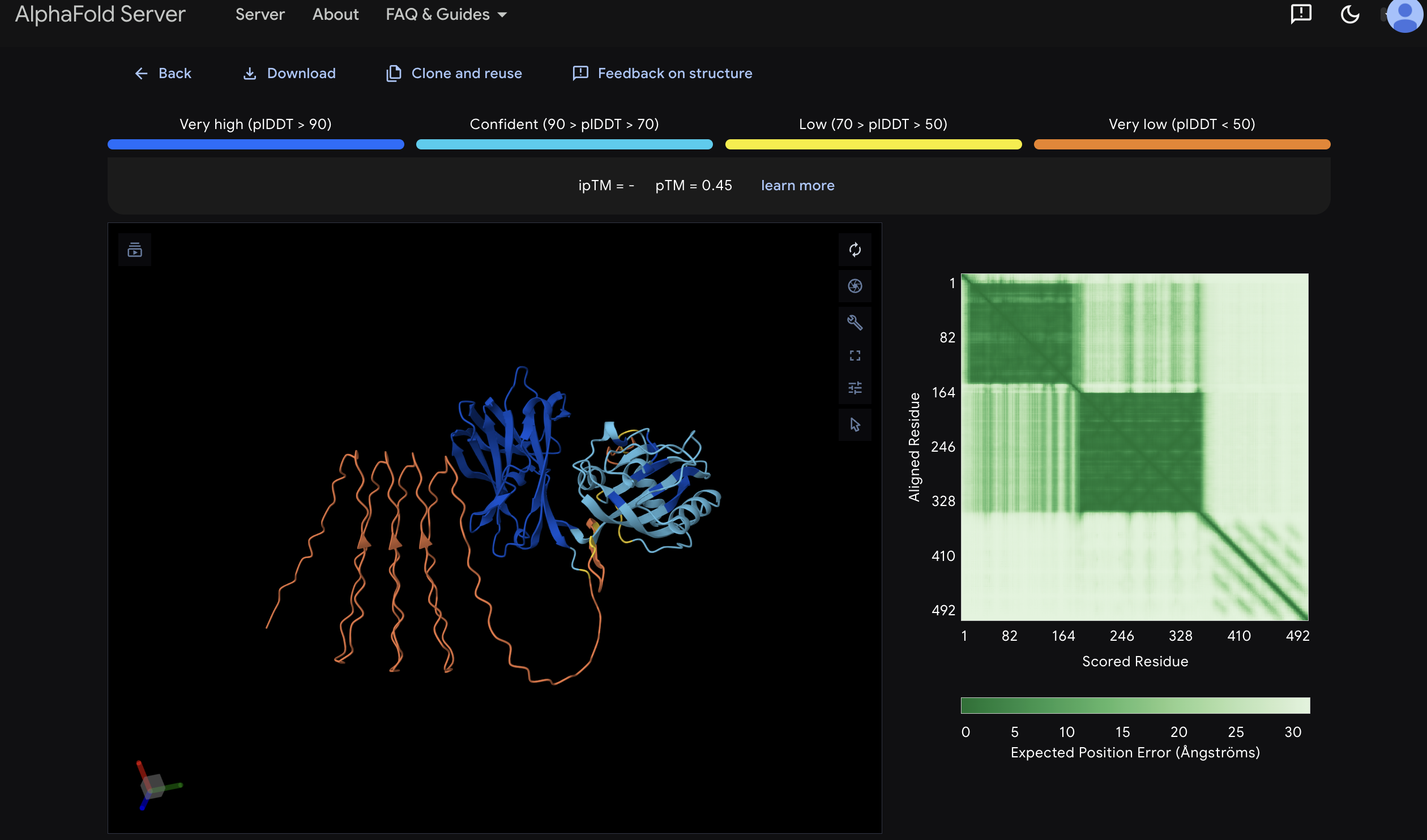
VVD_GGS_CBM27_MaSp1_4x HERO IMAGE!
AlphaFold3 Structure Prediction of VVD_GGS_CBM27_MaSp1_4x
Model_0, pTM 0.45. Coloured by pLDDT confidence using standard AlphaFold convention. Rendered in PyMOL. The AlphaFold3 prediction of the second fusion construct VVD_GGS_CBM27_MaSp1_4x reveals two independently folding structured domains consistent with the design intent. The larger domain predicting in green and cyan is the VVD LOV domain (residues 1-156), folding with confident to high pLDDT scores and adopting the characteristic PAS domain beta-sandwich fold consistent with the experimentally validated VVD crystal structures PDB 3RH8 and 3D72 (Zoltowski and Crane 2008). The smaller domain in dark blue is the CBM27 domain (residues 163-337), predicting with very high confidence pLDDT >90, consistent with the first construct prediction and the TmCBM27 crystal structure PDB 1OF4 (Boraston et al. 2003). The large disordered loops extending below are the four MaSp1 silk repeats (residues 352-493), predicting as intrinsically disordered as expected for recombinant spider silk repeat proteins prior to fibre assembly. The overall pTM of 0.45 reflects the presence of the large intrinsically disordered MaSp1 region and should not be interpreted as indicating poor prediction quality for the structured domains. The two distinct high confidence structured regions visible in the PAE matrix, one covering residues 1-82 corresponding to VVD and one covering residues 164-328 corresponding to CBM27, confirm that both functional domains fold independently and correctly within the fusion context. Critically the VVD and CBM27 domains appear in close spatial proximity in the predicted structure, consistent with the GGS linker allowing the LOV domain conformational change to propagate mechanical strain into the CBM27-anchored silk network upon blue light activation.
MolProbity (run on VVD construct only): Clashscore 4.46 at 95th percentile. MolProbity score 1.62 at 92nd percentile. Favoured rotamers 99.70%. Zero poor rotamers. Zero bad bonds. Benchmarked against 1784 experimentally solved PDB crystal structures.
MolProbity validation of the AlphaFold3 predicted structure of VVD_GGS_CBM27_MaSp1_4x scores at the 95th percentile for clashscore when benchmarked against 1784 experimentally solved crystal structures in the Protein Data Bank, confirming that the computational prediction meets the geometric quality standards of experimental structural biology. Wet lab confirmation of actual protein folding and domain function awaits cell free expression results from Ginkgo Bioworks other wetlab assay.
MolProbity confirms near-zero atomic clashes across the entire 493 residue fusion construct, with a clashscore placing the predicted structure in the 95th percentile of all experimentally solved crystal structures in the Protein Data Bank.
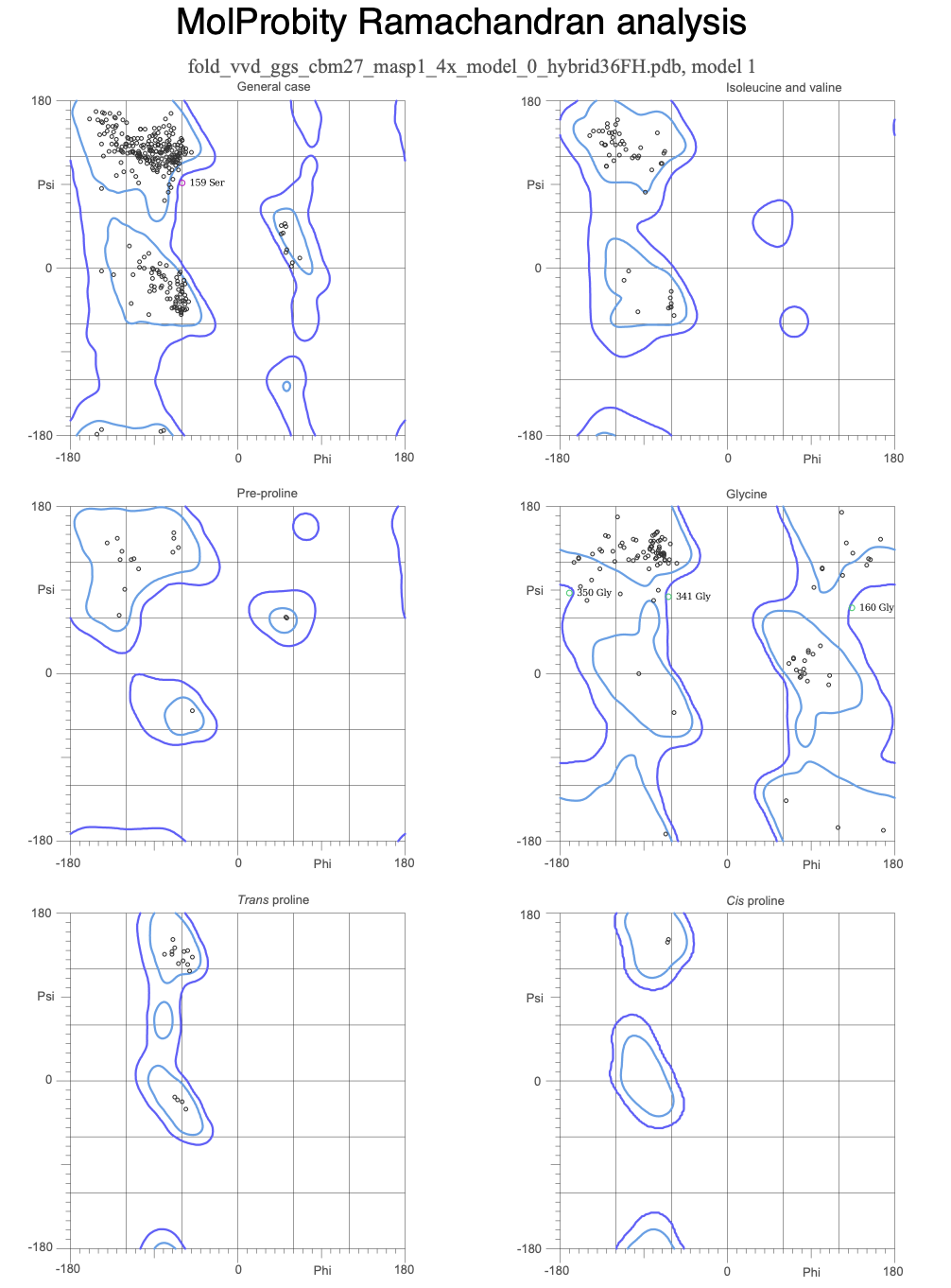
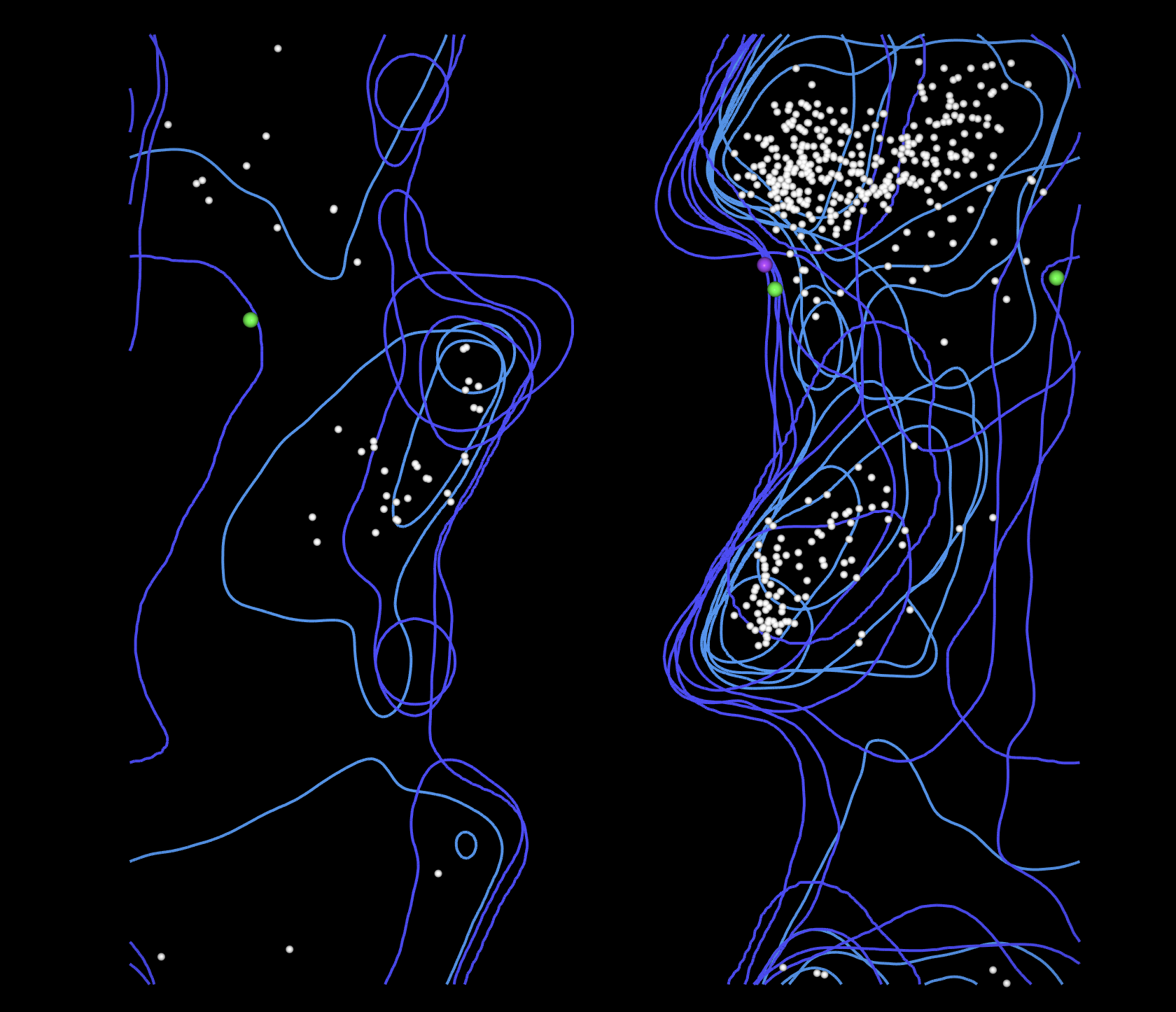
Clashscore 4.46, 95th percentile compared against N=1784 PDB structures at all resolutions. MolProbity score 1.62, 92nd percentile compared against N=27,675 PDB structures at 0-99 Angstrom resolution.
Section 3: Background
Literature Context
Recombinant MaSp1 spider silk expressed in E. coli self-assembles into fibres with tensile properties comparable to native dragline silk (Rising et al., 2011), but lacks biological signalling without post-synthesis functionalisation. Tremella fuciformis polysaccharide promotes fibroblast proliferation and moisture retention exceeding hyaluronic acid at equivalent concentrations (Huang et al., 2025). Boraston et al. (2003) confirmed that CBM27 domains can be grafted onto heterologous proteins while retaining binding specificity. No study has combined all three components into a single genetically encoded construct.
Novelty
This construct is novel in three ways: the CBM27-MaSp1 fusion has not been reported; the addition of RGD converts a passive scaffold into an active wound-healing participant; and the approach uses a designed molecular interaction rather than physical blending, a principle generalisable to other polysaccharide-CBM pairs. Detail follows below.
CBM27-MaSp1 Fusion Protein Hydrogel Scaffold
Key context for CBM27_RGD_MaSp1_4x construct, Aim 1 and Aim 2
Novelty assessment: High. The specific combination of CBM27 with MaSp1 for structural hydrogel or actuator scaffold applications is not present in current literature. The addition of an RGD motif for active cell recruitment and the use of Tremella fuciformis polysaccharide as the CBM27 binding target further distinguishes this construct from all prior art.
Relevance to construct: This note covers two distinct novelty claims within the same construct. First, the CBM27-MaSp1 fusion itself as a structural hydrogel scaffold has no direct precedent. Second, the Tremella fuciformis polysaccharide as the specific binding target for CBM27 is entirely novel in this context. Prior CBM-silk work uses cellulosic matrices from Clostridium thermocellum cellulosome systems, not fungal mannan polysaccharides. The β-(1→3)-linked mannopyranoside backbone of Tremella is chemically distinct from cellulose and represents a new class of CBM-polysaccharide pairing for silk composite materials.
RGD novelty note: The addition of the RGD integrin-binding motif to this construct converts a passive structural scaffold into an actively cell-recruiting bioaterial. No prior CBM-silk fusion work incorporates a cell adhesion motif. This three-way combination of CBM27 anchor, RGD recruitment, and MaSp1 structural backbone in a single genetically encoded construct is without precedent in the literature. Tremella novelty note: The use of Tremella fuciformis polysaccharide as the matrix phase of the composite is novel in two respects. It introduces a fungal-origin scaffold with documented cryoprotective and moisture-retention properties that cellulosic matrices do not possess, and it establishes a silk-fungal composite identity that is distinct from all prior silk-CBM work which uses bacterial or plant cellulose systems.
Closest prior art: Gomes et al. (2011), Biomaterials. MaSp1 fused with antimicrobial peptides, CBMs from Clostridium thermocellum used with antimicrobial peptides for cellulosic surfaces, not CBM27 with spider silk for hydrogel or actuator applications. No RGD, no fungal matrix. Mohammadi et al. (2019), Science Advances. ADF3 spidroin fused with CBMs from Clostridium thermocellum cellulosome, hydrogel noted as potential application but not developed. No CBM27, no MaSp1, no RGD, no fungal polysaccharide matrix. Key gaps your construct addresses: No prior work combines CBM27 specifically with MaSp1, no prior CBM-silk work targets a fungal mannan matrix, no prior CBM-silk fusion incorporates an active cell recruitment motif, and no prior work develops these composites explicitly as structural hydrogel or soft actuator scaffolds with a defined bioart or extreme environment application context.
References: Gomes et al. 2011, Biomaterials. CBM-silk antimicrobial fusion proteins. Mohammadi et al. 2019, Science Advances. Cellulose-spider silk CBM composites. https://www.science.org/doi/10.1126/sciadv.aaw2541
Photomechanical Soft Robotics for Space Environments Key context for VVD_GGS_CBM27_MaSp1_4x construct (VVD: LOV domain from Neurospora crassa), Aim 3
Novelty assessment: High. The integration of a genetically encoded LOV domain as a photomechanical actuator in a silk-fungal composite material for space environment applications is not present in current literature. This is the first proposed use of a LOV domain specifically for photomechanical actuation in harsh extraterrestrial conditions, leveraging available light flux as an electronics-free autonomous trigger.
Relevance to construct: The VVD LOV domain activates under blue light at approximately 450nm through cysteinyl-flavin adduct formation, driving homodimerisation and propagating conformational strain through the adjacent MaSp1 silk network. Martian solar spectrum at surface level retains sufficient blue light flux to activate LOV domain photochemistry, raising the possibility of fully autonomous actuation without onboard electronic light sources. This directly addresses the central gap in the space soft robotics literature, which has not considered genetically encoded photoreceptors as actuation mechanisms.
Four-layer novelty claim: This construct sits at the intersection of four properties that no prior work addresses simultaneously. Genetically encoded actuation mechanism. LOV domain photochemistry specifically. Silk-fungal composite structural material. Space environment application context.
Martian UV flux note: LOV domain activation occurs at approximately 450nm blue light. The Martian surface solar spectrum, while UV-shifted relative to Earth, retains blue light flux sufficient for LOV photochemistry. This opens the possibility of light-autonomous soft material actuation on the Martian surface without electronic control systems, a capability gap explicitly identified in the NASA ACR24 architecture technology gaps document for high-performance actuators and grippers in extreme environments.
Closest prior art: Lopez-Lopez et al. (2025), ICRA. Inflatable soft robotic arms for space debris capture using pneumatic actuation, motor-free but not photomechanical and not LOV-based. Zeng et al. (2018), Advanced Materials. Light-responsive liquid-crystal elastomers for soft microrobots, photomechanical but synthetic polymer systems, not genetically encoded, not space-environment specific. Nohooji and Voos (2025), Advanced Intelligent Systems. Compliant robotics for space, material selection focused, discusses harsh environment conditions but not photomechanical or LOV-based actuation.
Key gaps your construct addresses: No prior work uses a genetically encoded LOV domain for space soft robotics. No prior photomechanical soft robotics work targets extraterrestrial light as an autonomous trigger. No prior space soft robotics work uses a silk-fungal composite as the structural actuator material. The direct exploitation of Martian solar flux for autonomous LOV-driven silk network actuation is entirely unaddressed in the literature.
NASA technology gap alignment: NASA ACR24 Architecture Technology Gaps document identifies high-performance actuators, sensors, and interfaces for robotic joints and grippers operating in extreme cold and vacuum as an unresolved need. This construct addresses that gap through a passive, electronics-free, genetically encoded photomechanical mechanism that does not rely on lubricants, heating systems, or electronic controllers.
References: Lopez-Lopez et al. 2025: https://softroboticsforspace.eu/images/paper-submissions/Lopez-ICRA2025-Soft-Robotics-for-Space-Applications.pdf Zeng et al. 2018: https://advanced.onlinelibrary.wiley.com/doi/full/10.1002/adma.201703554 Nohooji and Voos 2025: https://www.researchgate.net/publication/390462454_Compliant_Robotics_in_Space_A_Prospective_Review_of_Soft_and_Deformable_Systems_for_Space_Missions NASA ACR24: https://www.nasa.gov/wp-content/uploads/2024/12/acr24-architecture-technology-gaps.pdf
Why It Matters
Chronic wounds cost the US healthcare system over $25 billion per year.(Add a ref/additional data ) If validated, this construct demonstrates that carbohydrate-binding modules can serve as molecular anchors between protein-based materials and naturally occurring polysaccharide scaffolds, a principle with broad applications across tissue engineering, soft robotics, and beyond.
Ethical Implications
All components are biosafe at this stage: CBM27 is from a non-pathogenic thermophile, Tremella is an edible mushroom, and the protein is produced in contained E. coli. Aim 3 (GMO fungus) would require regulatory review before any cultivation outside a controlled lab. Clinical risks including allergenic response and off-target integrin binding are noted and would require GMP-standard safety testing before patient use.
Section 4: Experimental Design
Techniques Used
- Protein design (Benchling)
- DNA construct design and codon optimisation
- Synthesis feasibility testing (IDT, Twist Biosciences)
- Twist order design (Gene Fragment and Clonal Gene)
- Sequence databases (UniProt, NCBI GenBank)
- Bioethical considerations
Industry Partners
- Twist Biosciences — synthesis feasibility and draft gene orders
- Ginkgo Bioworks — target partner for cell-free expression (Aim 2a)
- Benchling — construct assembly and sequence management
Section 5: Results
What Was Validated
The full DNA construct design and synthesis feasibility pipeline, from domain sequence acquisition through to a confirmed-orderable 4x construct on Twist.
Key Findings
The 8x MaSp1 repeat construct failed at both IDT (complexity score 53, threshold 24) and Twist (Not Accepted), due to unavoidable alanine/glycine codon poverty across 8 identical repeats. The redesigned 4x construct was confirmed orderable: Complex, $145.45, 6,297 bp. AlphaFold3 structure prediction and HADDOCK binding simulation are pending.
Data
Note: Move documentation images here
VVD_GGS_CBM27_MaSp1_4x – AlphaFold3 & MolProbity (Construct 2)
AlphaFold3: pTM 0.45. VVD and CBM27 both fold independently with high confidence. MaSp1 disordered as expected.
MolProbity (run on VVD construct only): Clashscore 4.46 at 95th percentile. MolProbity score 1.62 at 92nd percentile. Favoured rotamers 99.70%. Zero poor rotamers. Zero bad bonds. Benchmarked against 1784 experimentally solved PDB crystal structures.
Challenges
The 8x MaSp1 repeat construct failed synthesis due to alanine/glycine codon poverty across identical repeats. Redesigned to 4x, which was confirmed orderable on Twist at $145.45, 6,297 bp. Sidewinder (Wang et al., Nature 2026) directly addresses this repetitive sequence assembly problem and is flagged as a future synthesis route.
Next Steps
Validate CBM27_RGD_MaSp1_4x and VVD_GGS_CBM27_MaSp1_4x constructs through cell-free expression (Ginkgo CFPS or NEB PURExpress), SDS-PAGE, and polysaccharide binding assay. If protein expresses: mix with Tremella fuciformis polysaccharide (CAS 9075-53-0, 1% w/v in PBS pH 7.4).
References
Core Construct
[1] Xu et al. 2025, Advanced Healthcare Materials. [2] Huang et al. 2025, ACS Synthetic Biology. [3] Shahin G, 2025, MSc Thesis, LUT University. [4] Boraston et al. 2003, Structure of a Carbohydrate-Binding Module. [5] Rising et al. 2011, Cellular and Molecular Life Sciences.
Spider Silk Synthesis
[6] Robinson, N.E., Zhang, W., Ghosh, R., Gerber, B., Zhang, H., Sanfiorenzo, C., Wang, S., Di Carlo, D. and Wang, K. (2026) ‘Construction of complex and diverse DNA sequences using DNA three-way junctions’, Nature, 651(8105), pp. 491–500. https://doi.org/10.1038/s41586-025-10006-0
Closest Prior Art — CBM-Silk Composites
[7] Gomes et al. 2011, Biomaterials. CBM-silk antimicrobial fusion proteins. [8] Mohammadi et al. 2019, Science Advances. Cellulose-spider silk CBM composites. https://www.science.org/doi/10.1126/sciadv.aaw2541
Closest Prior Art — Space Soft Robotics
[9] Lopez-Lopez et al. 2025, ICRA. Inflatable soft robotic arms for space debris capture. https://softroboticsforspace.eu/images/paper-submissions/Lopez-ICRA2025-Soft-Robotics-for-Space-Applications.pdf [10] Zeng et al. 2018, Advanced Materials. Light-responsive liquid-crystal elastomers for soft microrobots. https://advanced.onlinelibrary.wiley.com/doi/full/10.1002/adma.201703554 [11] Nohooji and Voos 2025, Advanced Intelligent Systems. Compliant robotics in space. https://www.researchgate.net/publication/390462454_Compliant_Robotics_in_Space_A_Prospective_Review_of_Soft_and_Deformable_Systems_for_Space_Missions
NASA Technology Gap
[12] NASA ACR24 Architecture Technology Gaps, 2024. https://www.nasa.gov/wp-content/uploads/2024/12/acr24-architecture-technology-gaps.pdf
Initial FP Aims

 ---
---Subsections of Individual Final Project
Aim 1.Construct Design: CBM27_RGD_MaSp1 Fusion Protein
Spider Silk-Snow Fungus Bioactive Biomaterial Optimised for Wound Care — CBM27_RGD_MaSp1_4x Fusion Protein · HTGAA Spring 2026

Documented (Update)
In Benchling:
- Full 8x construct (480 aa) fully annotated for in silico validation and Ginkgo pitch
- 4x construct (340 aa) fully annotated as synthesis-ready design
In Twist:
Saved draft order for CBM27_RGD_MaSp1_4x_Fusion as a Gene Fragment at $71.61 (~£56), 1,023 bp, complexity: Complex, orderable
Saved draft order for CBM27_RGD_MaSp1_4x_pET28a as a Clonal Gene in pET28a(+) at NdeI/XhoI insertion point at $145.45 (~£115), 1,005 bp insert, 6,297 bp total plasmid, complexity: Complex, orderable
Downloaded FASTA, GenBank and text files of the optimized gene fragment sequence
Downloaded GenBank file of the complete pET28a construct
Circular plasmid map generated and documented showing all functional elements
In Benchling: (Older)
Full 8x construct (480 aa) fully annotated 4x construct (340 aa) fully annotated as synthesis-ready design
In Twist:
Saved draft order for CBM27_RGD_MaSp1_4x_Fusion at $71.61 Downloaded FASTA, GenBank and text files of the optimized sequence
In IDT:
Documented complexity failure of 8x construct (score 57.3) Documented complexity failure of identical MaSp1 repeats (score 53)
The Four-Component Logic The fusion protein is built from four functional units arranged in a deliberate order from N-terminus to C-terminus:
The CBM27 domain, sourced from the Thermotoga maritima beta-mannanase Man5 (gene TM1227), physically anchors the silk network to the Tremella polysaccharide matrix by binding its mannan backbone. Without this anchor, the silk and polysaccharide components would phase-separate in wound exudate.
The flexible GGGGS x3 linker gives the CBM27 domain rotational freedom to engage the Tremella polysaccharide chains independently from the rest of the protein. This design follows validated precedent from CBM27 fusion protein literature. The RGD motif (GRGDS), derived from the minimal integrin-binding sequence of human fibronectin, actively recruits fibroblasts and keratinocytes to the material surface to accelerate tissue repair. This converts the dressing from a passive scaffold into an active wound-healing participant.
Eight MaSp1 consensus repeat units provide the mechanical backbone of the construct, self-assembling into beta-sheet nanofibrils that give the hydrogel its toughness and structural integrity. Construct Architecture The full fusion protein is 480 amino acids with the following domain map:
DomainPositionsFunctionCBM27 (buffered)1-176Tremella polysaccharide anchorGGGGS x3 Linker177-188Domain flexibilityRGD motif189-193Cell recruitmentMaSp1 x8 repeats194-474Mechanical structure6xHis tag475-480Ni-NTA purification Molecular weight: 44,409 Da. Isoelectric point: 7.29. Sequence Source and Verification

The CBM27 domain boundaries were verified against the UniProt entry for Thermotoga maritima Man5, which annotates the Carbohydrate Binding Module 27 at residues 495-664 of the full 669 aa protein. A five-residue buffer was added to each boundary to avoid clipping structurally important residues at the domain edge, giving a final CBM27 input of 177 amino acids.
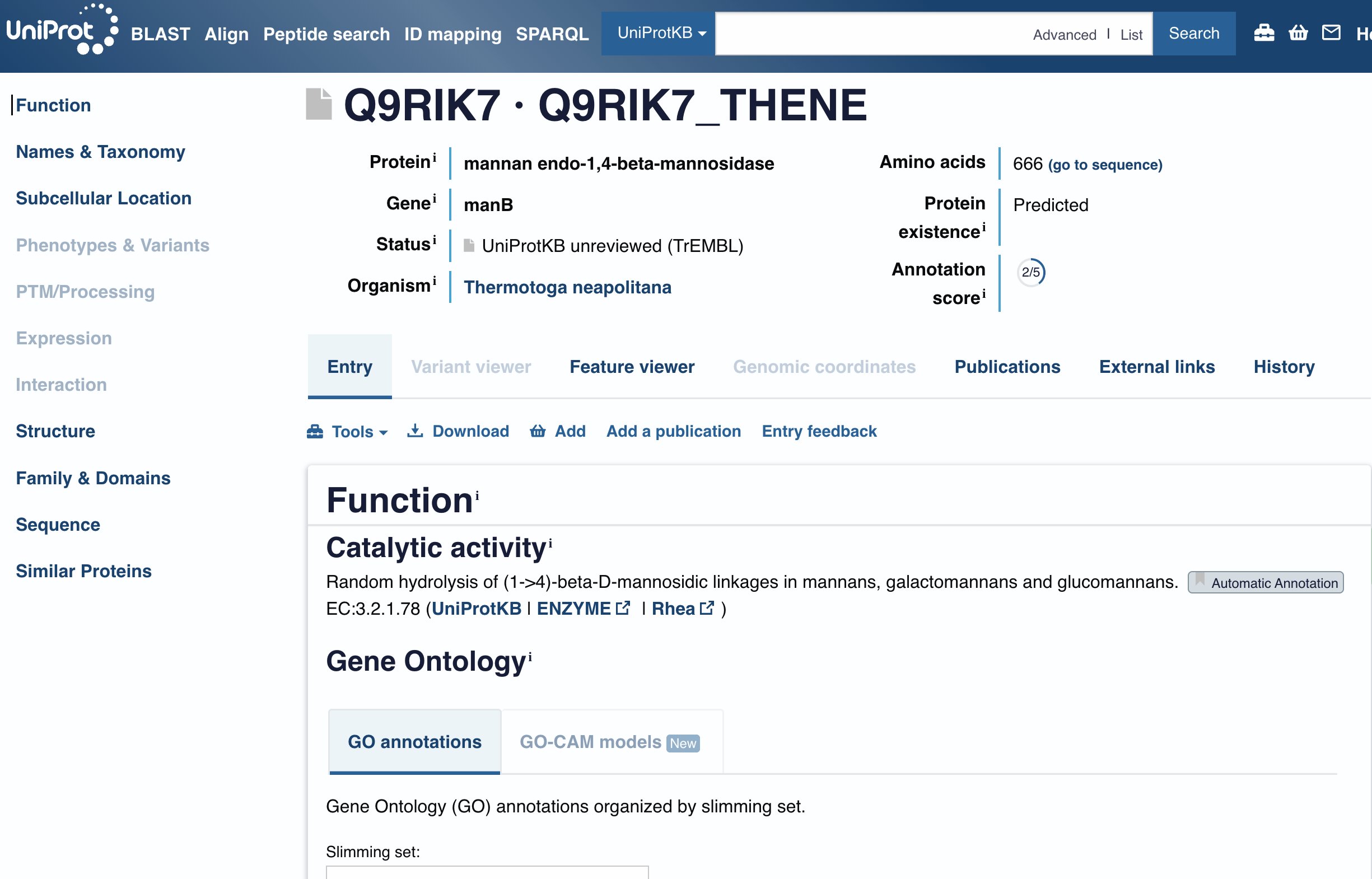
The complete construct was assembled and annotated in Benchling as an AA sequence file named CBM27_RGD_MaSp1_Fusion within the Spider Silk - Tremella project.
Synthesis Complexity and the Codon Shuffling Requirement Initial synthesis feasibility analysis via the IDT Codon Optimization Tool confirmed that direct translation of 8 identical MaSp1 repeat units generates prohibitive sequence complexity, returning a total complexity score of 57.3 against a synthesis threshold of 24. The specific failure modes identified were:
Repeated DNA sequences exceeding 13 bases appearing at multiple locations across the insert 73.5% of the overall sequence composed of repeats longer than 8 bases A 100-base window at position 1165 with 85% GC content Hairpin structures forming between identical repeat regions

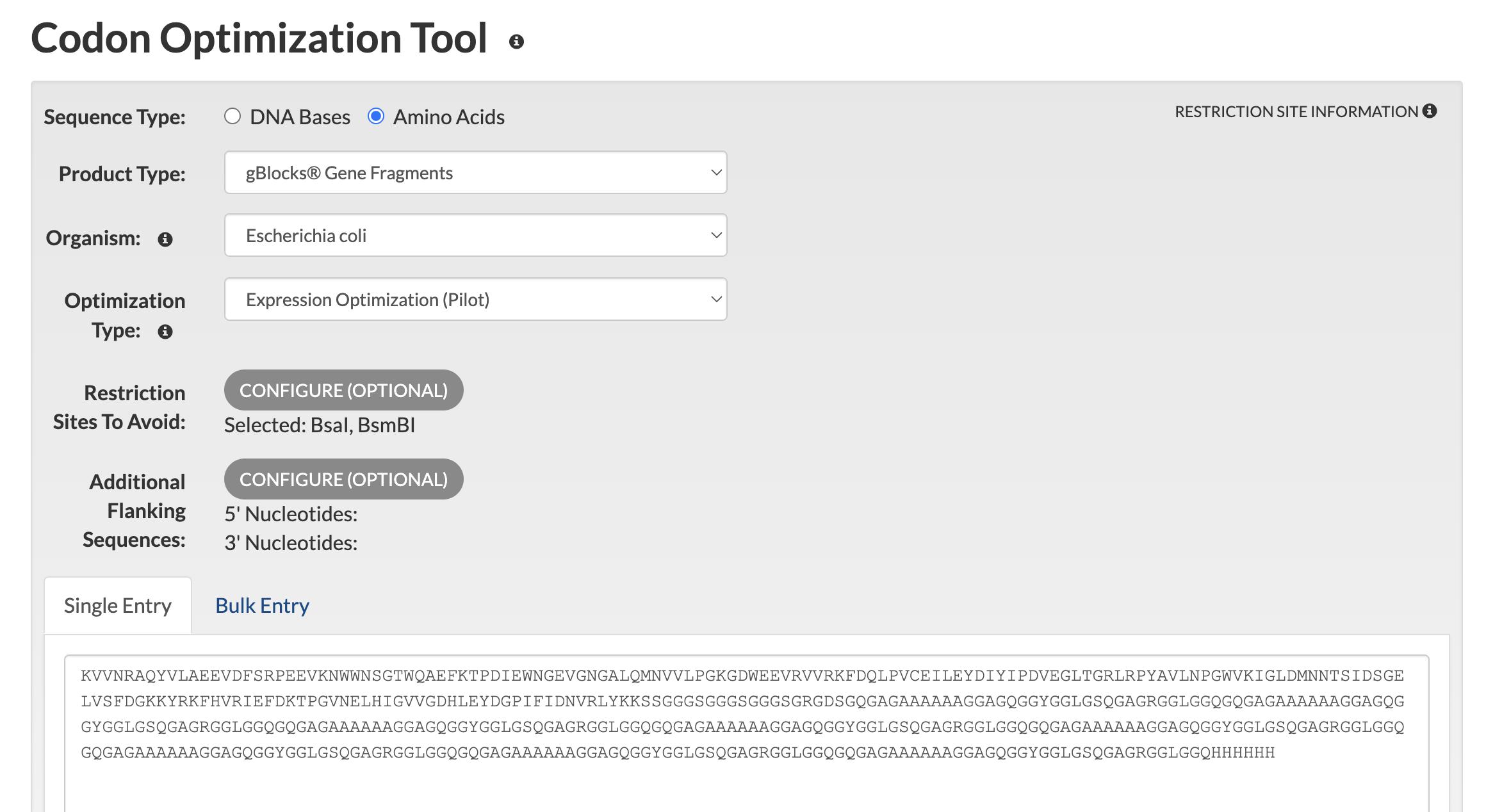


This result validates the codon shuffling strategy as a necessary design requirement rather than an optional refinement. The next step is to manually assign synonymous codons to each of the 8 MaSp1 repeats so that every repeat encodes an identical amino acid sequence but presents a distinct DNA sequence to the synthesis machinery.
1. Sequence Acquisition: CBM27 Domain
The carbohydrate binding module 27 domain was sourced from the Thermotoga maritima beta-mannanase Man5 (gene TM1227), a 669 amino acid protein. The CBM27 domain boundaries were verified against the UniProt entry, which annotates the domain at residues 495-664. A five-residue buffer was added to each boundary to avoid clipping structurally important residues at the domain edge, giving a final CBM27 input sequence of 177 amino acids spanning residues 490-669.
The extracted sequence was:
2. Construct Assembly in Benchling
The full fusion protein was assembled in Benchling as an AA sequence file named CBM27_RGD_MaSp1_Fusion within the Spider Silk - Tremella project folder. Each domain was added sequentially and annotated with a distinct colour:
| Domain | Positions | Colour | Function |
|---|---|---|---|
| CBM27 (buffered) | 1-176 | Blue | Tremella polysaccharide anchor |
| GGGGS x3 Linker | 177-188 | Grey | Domain flexibility |
| RGD motif | 189-193 | Pink | Cell recruitment |
| MaSp1 R1-R8 | 194-474 | Green | Mechanical structure |
| 6xHis tag | 475-480 | Yellow | Ni-NTA purification |
Total length: 480 amino acids. Molecular weight: 44,409 Da. Isoelectric point: 7.29.
The annotated sequence map confirmed all domains were correctly positioned and accounted for. The biochemical properties panel confirmed the sequence was 480 amino acids running from KVVN at the N-terminus to HHHH at the C-terminus.
3. Codon Optimization: CBM27 Domain
The CBM27 domain alone (177 aa) was submitted to the IDT Codon Optimization Tool with the following settings:
- Sequence type: Amino Acids
- Product type: gBlocks Gene Fragments
- Organism: Escherichia coli
- Restriction sites to avoid: BsaI (GGTCTC), BsmBI (CGTCTC)
The CBM27 domain passed initial screening with no complexity issues. The codon-optimized DNA output was 531 bp. No BsaI or BsmBI sites were introduced. Several other restriction sites were noted (PstI, SmaI, XmaI) but these are irrelevant to the Golden Gate assembly strategy and were not flagged as problems.
4. Synthesis Feasibility Testing: Full 8x Construct
IDT Test 1: Identical MaSp1 Repeats
The complete 480 amino acid sequence was submitted to IDT Codon Optimization with E. coli settings. IDT returned a complexity score of 57.3, well above the synthesis threshold of 24, with the following specific failure modes:
- A repeat sequence of 42 bases appearing at multiple locations
- 73.5% of the overall sequence composed of repeats longer than 8 bases
- A 100-base window at position 1165 with 85% GC content
- Hairpin structures forming between identical repeat regions
Result: Denied. Not synthesizable.
This confirmed the known problem with identical tandem silk repeats: even after codon optimization, the DNA homology between repeated units causes synthesis machinery slippage.
IDT Test 2: Codon-Shuffled MaSp1 Repeats
Eight synonymous codon-shuffled variants of the MaSp1 repeat were computationally generated, each encoding the identical amino acid sequence GQGAGAAAAAAGGAGQGGYGGLGSQGAGRGGLGGQ but using distinct codon assignments drawn from the E. coli K12 codon table. No two adjacent repeats shared more than 3 consecutive identical base pairs.
The full insert including CBM27, linker, RGD, all 8 shuffled repeats, 6xHis tag and stop codon was assembled as a 1,443 bp sequence and submitted to IDT gBlocks entry for complexity testing.
Result: Complexity score 53. Still Denied.
The codon shuffling reduced the score from 57.3 to 53 but was insufficient to bring it below the 24 threshold. The dominant remaining problem was a 42-base shared subsequence between two repeats in the alanine-rich region, where the limited synonymous codon options for alanine and glycine prevented sufficient DNA diversification across 8 repeats.
Twist Bioscience Test: Full 8x Construct
The same 480 amino acid sequence was submitted to Twist Bioscience’s gene ordering portal using their built-in amino acid import and codon optimization workflow with E. coli codon table, BsaI and BsmBI avoidance.
Result: Not Accepted.
Twist’s algorithm, which is more sophisticated than IDT’s and better handles repetitive sequences, also rejected the 8x construct. This was definitive confirmation that 8 identical MaSp1 repeats cannot be synthesized as a single gene fragment by any current commercial synthesis vendor.
5. Design Decision: 4x Repeat Construct
Based on the synthesis feasibility data, a second construct was designed using 4 MaSp1 repeat units instead of 8. This was created in Benchling by duplicating the original file and deleting repeats R5 through R8 (positions 334-473), producing a 340 amino acid construct named CBM27_RGD_MaSp1_4x_Fusion.
| Domain | Positions | Function |
|---|---|---|
| CBM27 (buffered) | 1-176 | Tremella polysaccharide anchor |
| GGGGS x3 Linker | 177-188 | Domain flexibility |
| RGD motif | 189-193 | Cell recruitment |
| MaSp1 R1-R4 | 194-333 | Mechanical structure |
| 6xHis tag | 334-340 | Ni-NTA purification |
6. Synthesis Feasibility Testing: 4x Construct on Twist
The 340 amino acid 4x construct was submitted to Twist Bioscience using the same amino acid import workflow. Twist performed codon optimization for E. coli with BsaI and BsmBI avoidance and returned the following result:
- Complexity: Complex
- Length: 1,023 bp
- Price: $71.61 (~£56)
- Status: Orderable
Complex in Twist terminology means the sequence is manufacturable but requires more careful synthesis handling than a standard sequence. It is not a rejection. The remaining complexity flags were minor repeat density warnings in the MaSp1 region, all categorised as warnings rather than errors.
Twist’s codon optimization was applied and the optimized sequence was downloaded in FASTA, GenBank and text formats. The order draft was saved to the Twist account.
7. Summary of Constructs and Status
| Construct | Repeats | Length | IDT Result | Twist Result | Purpose |
|---|---|---|---|---|---|
| CBM27_RGD_MaSp1_Fusion | 8x | 480 aa / 1,443 bp | Score 53, Denied | Not Accepted | In silico validation, Ginkgo pitch |
| CBM27_RGD_MaSp1_4x_Fusion | 4x | 340 aa / 1,023 bp | Not tested | Complex, $71.61, Orderable | Wet lab synthesis |
8. Backbone Vector Documentation

The insert was designed for expression in pET-28a(+) (Novagen, 5,365 bp), cloned at the NdeI/XhoI insertion points. This places the fusion protein under T7 promoter control with a C-terminal 6xHis tag provided by the vector for Ni-NTA purification. The vector carries kanamycin resistance for bacterial selection and a colE1 high copy origin of replication. The complete circular plasmid was designed directly in Twist Bioscience’s Clonal Gene ordering portal, which performs synthesis, cloning, transformation, colony picking and Sanger sequencing verification, delivering a ready-to-use sequence-verified plasmid. The full construct is 6,297 bp and was downloaded as a GenBank file and imported into Benchling as a circular DNA sequence for documentation. The plasmid is compatible with NEB PURExpress E6800 and Ginkgo Bioworks CFPS cell-free expression systems, both of which are T7 promoter driven E. coli based systems requiring no further cloning before expression.
9. Next Steps
In silico track: (see In silico page)
Submit the full 480 aa CBM27_RGD_MaSp1_Fusion sequence to AlphaFold3 to predict the 3D structure and confirm that the CBM27 domain and RGD motif are surface-exposed and not buried within the MaSp1 beta-sheet core.
Wet lab track:
The protein construct ready to be ordered from Twist. Preferred Ginkgo Bioworks (for remote lab experience) or LifeFabs assay as second option. Protocol draft started.
Protocol for Assay
Draft v 1
Experimental Protocol: CBM27_RGD_MaSp1_4x Fusion Expression and Tremella Composite Formation
Construct: CBM27_RGD_MaSp1_4x_Fusion Vector: pET28a Expression system: NEB PURExpress E6800 Protein MW: ~32 kDa
Step 1: Cell-Free Expression
Materials needed:
- PURExpress Solution A
- PURExpress Solution B
- Murine RNase Inhibitor
- Template DNA (linear PCR product or circular plasmid, 250 ng)
- Nuclease-free water
- Ice
Protocol:
- Thaw Solutions A and B on ice. Do not vortex.
- Assemble the following reaction on ice in this exact order in a 1.5 ml microcentrifuge tube:
- 10 µl Solution A
- 7.5 µl Solution B
- 0.5 µl RNase Inhibitor (20 units)
- 2 µl template DNA (250 ng)
- 5 µl nuclease-free water
- Total volume: 25 µl
- Mix gently by pipetting up and down 5 times. Do not vortex.
- Incubate at 37°C for 2 hours.
- Place on ice immediately after incubation.
Run alongside: one negative control reaction with no template DNA.
Step 2: SDS-PAGE Confirmation
Materials needed:
- 10-20% Tris-glycine precast gel
- SDS loading buffer
- Protein ladder (10-250 kDa range)
- Running buffer
- Coomassie Blue stain
Protocol:
- Take 2.5 µl of the expression reaction and add 2.5 µl SDS loading buffer.
- Heat at 95°C for 5 minutes to denature proteins.
- Load onto gel alongside protein ladder and negative control.
- Run at 200V for 35 minutes.
- Stain with Coomassie Blue for 1 hour, destain with water overnight.
- Look for a band at approximately 32 kDa.
Success criterion: A visible band at 32 kDa in the expression lane that is absent in the negative control lane.
Step 3: Tremella Composite Formation (only if Step 2 successful)
Materials needed:
- Tremella fuciformis dried polysaccharide (commercially available, e.g. Sigma or specialist supplier)
- PBS buffer pH 7.4
- Your expressed fusion protein from Step 1 (remaining ~22.5 µl)
- Microcentrifuge tubes
- Rotary mixer or gentle agitation
Protocol:
- Prepare a 1% w/v Tremella polysaccharide solution by dissolving 10 mg dried Tremella polysaccharide in 1 ml PBS pH 7.4. Stir gently at room temperature for 2 hours until fully dissolved. This produces a viscous gel solution.
- Take the remaining expressed fusion protein from Step 1 (~22.5 µl).
- Add 22.5 µl of the 1% Tremella polysaccharide solution to the protein.
- Mix gently by pipetting 10 times. Do not vortex.
- Incubate at room temperature for 30 minutes on a rotary mixer at low speed to allow CBM27 domain binding to the Tremella mannan backbone.
- Observe for gelation. The mixture should become more viscous than either component alone if the CBM27 is anchoring the silk to the Tremella matrix.
Success criterion: Visible increase in viscosity or gel formation compared to a control mixture of the same volume of PURExpress reaction without the fusion protein mixed with Tremella.
What Each Result Means
| Result | Interpretation |
|---|---|
| Band at 32 kDa on gel | Protein expressed successfully |
| No band on gel | Expression failed, check DNA template and reaction setup |
| Gel formation in Step 3 | CBM27 is anchoring silk to Tremella, composite hydrogel confirmed |
| No gel formation in Step 3 | CBM27 may not be folding correctly in cell-free system, or Tremella preparation needs optimisation |
Group Final Project


L-Protein Mutants
Problem: How to improve the stability and auto-folding of the lysis protein of an MS2-phage? This mechanism is key to understanding how phages may help address antibiotic resistance.
After going through the readings, including the group final project document a Plan A would be: (This stays within scope, MurJ and multi-target approaches seem intersting though…)
Use computational tools like AlphaFold2 or ProteinMPNN to identify mutations that improve intrinsic stability and auto-folding of the lysis protein
Target mutations that strengthen the hydrophobic core, eliminate aggregation-prone regions, or introduce stabilising interactions like salt bridges
Engineer the lysis protein to fold correctly without requiring DnaJ or any other bacterial chaperone
Design mutations that also accelerate oligomerisation or enhance membrane pore-forming activity for faster lysis
Synthesise the mutant gene via Twist, clone into plasmid using Gibson Assembly, validate structural integrity with Nuclera, then test in E. coli.