Week 3 HW: Lab Automation


Week 3: Lab Automation
Lab Automation assignment
This week, we explored lab automation and its applications in synthetic biology.
1. Assignment: Python Script for Opentrons Artwork
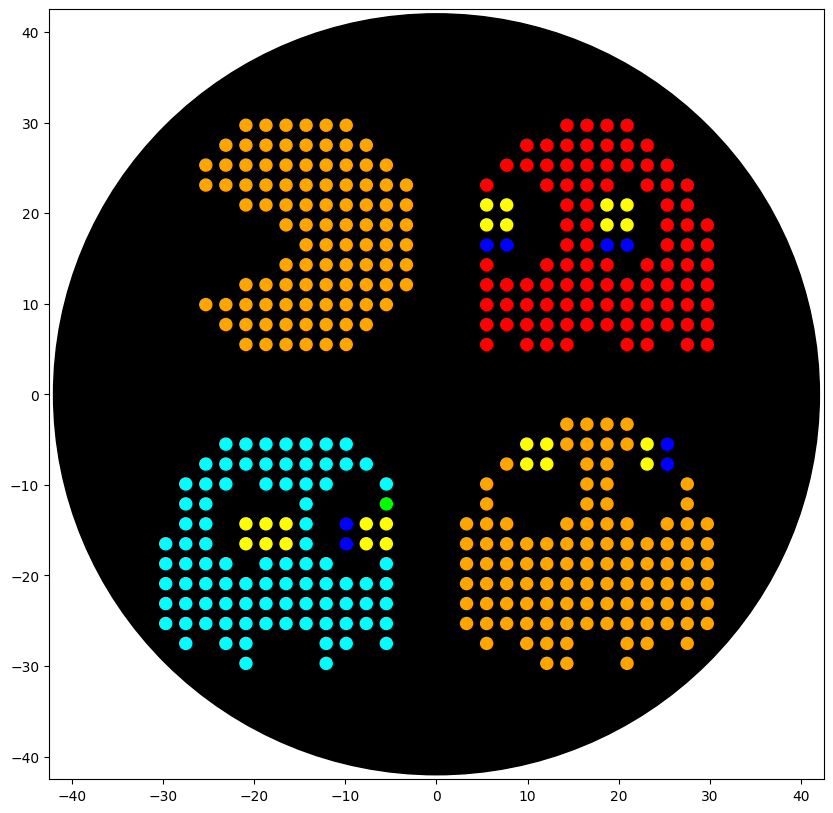
I designed a “Pac-Man” themed piece of Bio-Art for the Opentrons OT-2 robot. The design features Pac-Man and several ghosts, mapped onto a circular canvas representing the output of the lab’s liquid handling.
Google Colab Link: HTGAA Week 3 - Pac-Man Opentrons Art
Python Script Logic:
The script iterates through a matrix of coordinates, assigning specific colored liquids (Yellow for Pac-Man, Red/Blue/Cyan/Orange for ghosts) to designated wells. I used the Opentrons Python API (v2) to handle the aspirate and dispense operations with the P300 single-channel pipette.
2. Post-Lab Questions
1. Published Paper Review
Paper: Implementation of an open-source robotic platform for SARS-CoV-2 testing by real-time RT-PCR (Amen et al., Scientific Reports, 2021)
Summary: This paper details the implementation of a high-throughput diagnostic workflow for SARS-CoV-2 using Opentrons OT-2 liquid-handling robots. The authors developed a system that automates the transition from sample preparation to RT-PCR setup, allowing for up to 2,400 tests per day with a small footprint.
Novelty: The primary novelty lies in the use of open-source hardware to solve a critical capacity crisis during a global pandemic. Traditionally, such high-throughput testing required million-dollar proprietary systems. By utilizing the OT-2, the researchers created a “reproducible and flexible” diagnostic workstation that is affordable and can be rapidly deployed worldwide. This highlights how automation tools like Opentrons can democratize advanced molecular biology applications that were previously limited to elite institutions.
2. Automation for Final Project
For my final project, I am developing Next-Generation LuxR Biosensors with shifted specificities. I intend to use lab automation to bridge the gap between “Design” and “Test” in my DBTL cycle.
The Plan:
- Automated Mutant Screening: I will use the Opentrons OT-2 to setup 96-well plates containing my library of LuxR variants.
- Dynamic Range Characterization: The robot will perform automated serial dilutions of various AHL induce (Acyl-homoserine lactones) across 8 orders of magnitude (from 1 pM to 100 μM). This level of precision is difficult to achieve manually without significant error.
- Combinatorial Analysis: To study crosstalk, I will automate the mixing of multiple different AHLs and variants in a single plate to see how the sensors behave in “noisy” chemical environments.
Automation Workflow (Inspired by Example 2):
- OT-2: Transfer LuxR variant expression constructs into designated wells of a 96-well plate.
- OT-2: Perform serial dilutions of 5 different AHL analogs and dispense into the variant-containing wells.
- Plate Reader: (Manual or remote integration) Measure fluorescence over a 12-hour period to generate real-time induction curves.
- Data Analysis: Use Python to automatically calculate the EC50 (half-maximal effective concentration) for each variant-ligand pair.
Pseudocode Idea:
3. Final Project Ideas
As part of the assignment, I have proposed the following three directions for my individual final project. These ideas explore different aspects of synthetic biology, from allosteric control to molecular transport and stimuli-responsive pores.
AI-Designed Allosteric Switches for Synthetic Cell Control
Designing molecular switches that sense specific small molecules (inducers) and trigger gene expression or enzymatic activity within an artificial cell.

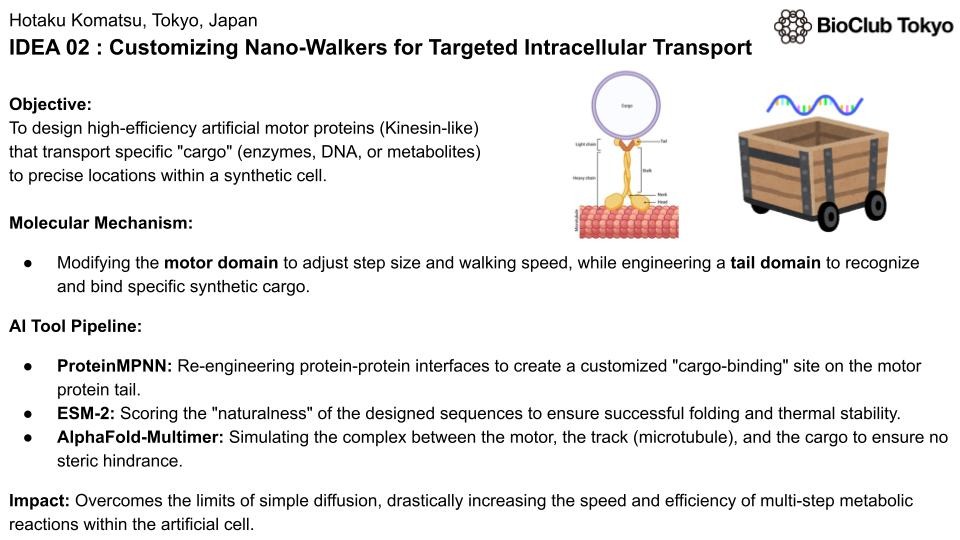
Customizing Nano-Walkers for Targeted Intracellular Transport
Designing high-efficiency artificial motor proteins (Kinesin-like) that transport specific "cargo" (enzymes, DNA, or metabolites) to precise locations within a synthetic cell.

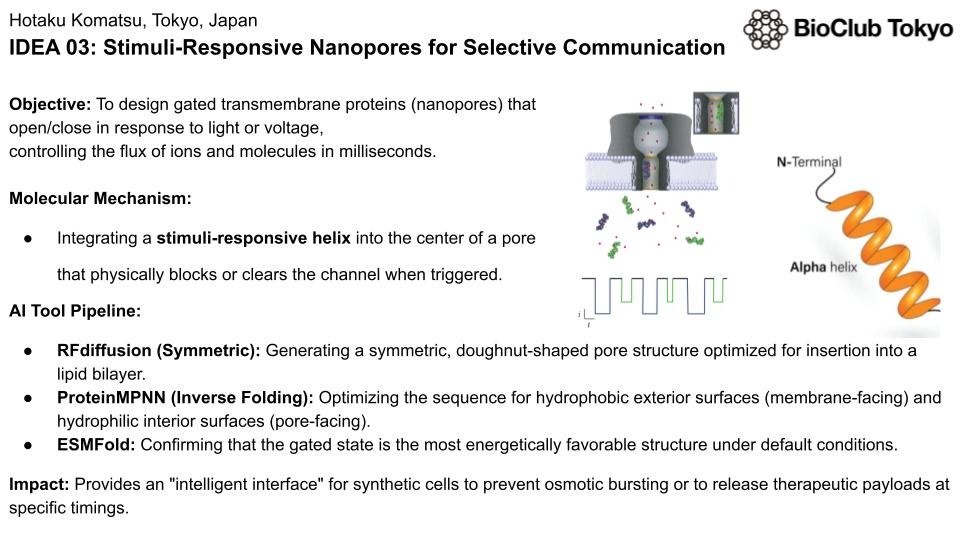
Stimuli-Responsive Nanopores for Selective Communication
Designing gated transmembrane proteins (nanopores) that open/close in response to light or voltage, controlling the flux of ions and molecules in milliseconds.