Week 11 HW: Bioproduction & Cloud Labs
Part A: The 1,536 Pixel Artwork Canvas | Collective Artwork
1. My contribution
Unfortunately, I was not able to contribute a pixel to the collective artwork, as I was in the middle of midterm exams at my university during that period, which limited my availability to participate.
2. What I liked about the project
I really liked the project because of its biological foundation and particularly its connection to cell-free fluorescent protein optimization and how it was used for a global pixel artwork designed by HTGAA students :)
3. What could be improved for next year
For future versions, it could be interesting to include a live chat feature so participants can coordinate in real time and create more elaborate and intentional designs. Additionally, increasing the number of pixels beyond the 1,536 used in this edition could allow for more detailed and realistic compositions.
Part B: Cell-Free Protein Synthesis | Cell-Free Reagents
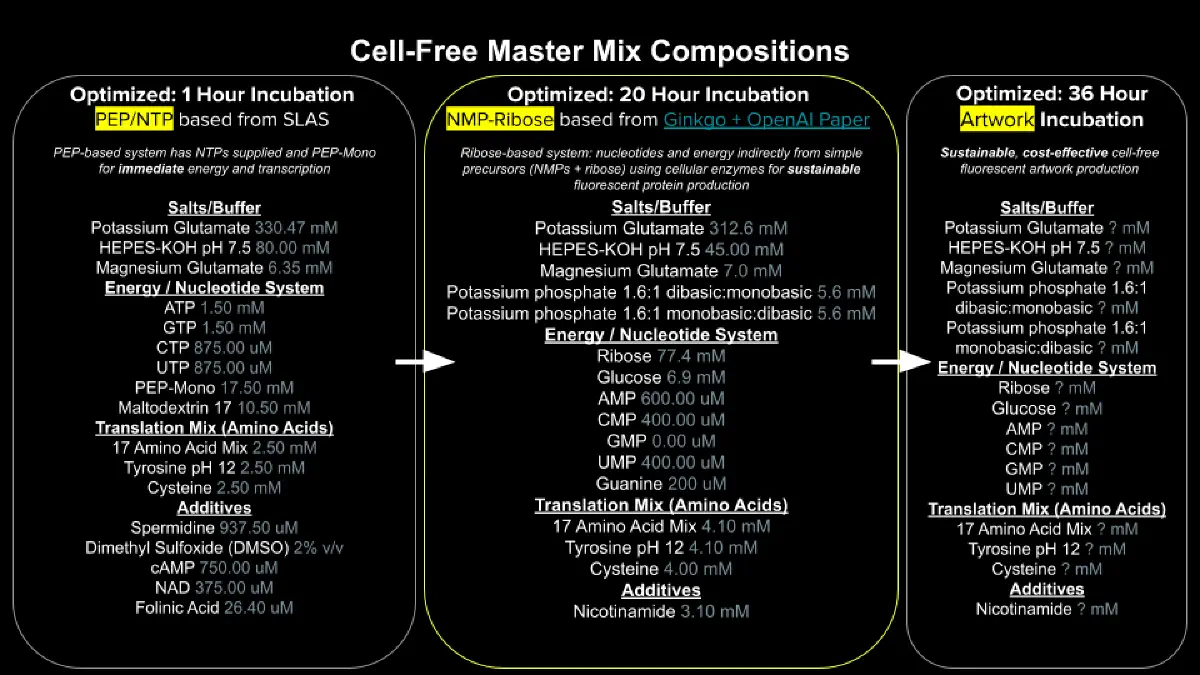
- Referencing the cell-free protein synthesis reaction composition (the middle box outlined in yellow on the image above, also listed below), provide a 1-2 sentence description of what each component’s role is in the cell-free reaction.

E. coli Lysate
BL21 (DE3) Star Lysate (includes T7 RNA Polymerase)
The lysate provides the essential molecular machinery required for transcription and translation, including ribosomes, metabolic enzymes, cofactors, and tRNAs. The presence of T7 RNA polymerase enables efficient transcription of target genes under T7 promoter control.
Salts / Buffer
- Potassium Glutamate
Maintains ionic strength and mimics intracellular conditions, thereby stabilizing macromolecular interactions and supporting enzymatic activity.
- HEPES-KOH pH 7.5¨
Serves as a buffering agent to maintain a stable pH, which is critical for optimal enzyme function during transcription and translation.
- Magnesium Glutamate
Functions as an essential cofactor for ribosomes and polymerases, playing a key role in both transcriptional and translational processes.
- Potassium Phosphate Monobasic / Dibasic
Contributes to buffering capacity and provides phosphate ions necessary for nucleotide metabolism and energy transfer reactions.
Energy / Nucleotide System
- Ribose
Acts as a precursor for nucleotide biosynthesis, supporting sustained RNA production over extended reaction times.
- Glucose
Serves as a metabolic energy source, enabling ATP regeneration through endogenous enzymatic pathways present in the lysate.
- AMP, CMP, GMP, UMP
Provide nucleotide monophosphates that can be phosphorylated into their corresponding triphosphates, which are required substrates for RNA synthesis.
- Guanine
Functions as a precursor in nucleotide salvage pathways, allowing for the biosynthesis of GMP and subsequently GTP for transcription.
Translation Mix (Amino Acids)
- 17 Amino Acid Mix
Supplies the majority of amino acids required for protein synthesis during translation.
- Tyrosine
Provided separately due to solubility and stability constraints; essential for incorporation into nascent polypeptides.
- Cysteine
Added separately due to its susceptibility to oxidation; plays a critical role in protein structure through disulfide bond formation.
Additives
- Nicotinamide
Supports redox balance and enzymatic activity by contributing to NAD⁺-dependent metabolic processes within the reaction. Backfill
- Nuclease-Free Water
Used to adjust the final reaction volume while preventing nucleic acid degradation.
- Describe the main differences between the 1-hour optimized PEP-NTP master mix and the 20-hour NMP-Ribose-Glucose master mix shown in the Google Slide above. (2-3 sentences)
The 1-hour PEP-NTP system relies on the direct addition of high-energy phosphate donors and nucleotide triphosphates, enabling rapid and high-yield protein synthesis within a short time frame. However, this approach is limited by the rapid depletion of energy substrates and accumulation of inhibitory byproducts.
In contrast, the 20-hour NMP-ribose-glucose system employs a metabolically sustained strategy in which substrates such as ribose and glucose support continuous nucleotide regeneration and ATP production. This results in prolonged reaction stability and sustained protein expression over extended periods.
- Bonus question: How can transcription occur if GMP is not included but Guanine is?
Transcription can still occur in the absence of externally supplied GMP because guanine can be converted into GMP through endogenous nucleotide salvage pathways present in the lysate. The resulting GMP can then be phosphorylated to GTP, which serves as the direct substrate for RNA polymerase during transcription.
References:
Carlson, E. D., Gan, R., Hodgman, C. E., & Jewett, M. C. (2012). Cell-free protein synthesis: Applications come of age. Biotechnology Advances, 30(5), 1185–1194. https://doi.org/10.1016/j.biotechadv.2011.09.016
Swartz, J. R. (2012). Transforming biochemical engineering with cell-free biology. AIChE Journal, 58(1), 5–13. https://doi.org/10.1002/aic.13701