Homework
Weekly homework submissions:
- Question 1 In the 1980’s, Keith Wood became the first to make a tobacco plant glow using firefly luciferase. However, he faced a critical limitation: they could not synthesize their own luciferin, requiring an external luciferin spray to emit light. 1 As Wood pivoted to fungal pathways, the firefly route has been largely abandoned despite its superior light efficiency. However, recent breakthrough have reopened this door, including the discovery of spontaneous benzoquinone + cysteine L-luciferin formation, and the identification of ACOT1 in D-luciferin transformation. My project aims to put these breakthroughs together, engineering plants capable of bioluminscence through firefly pathways.
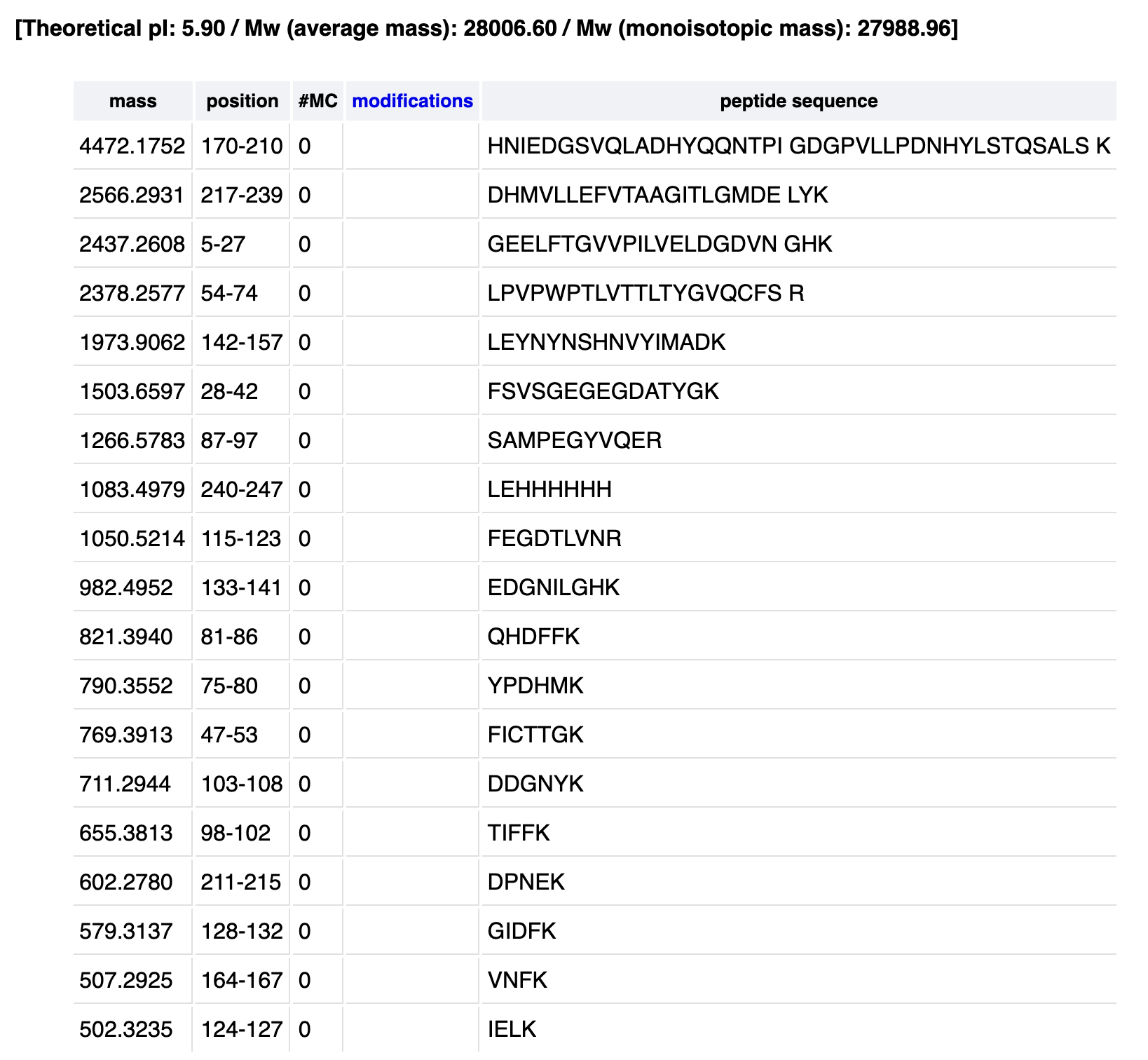
- Homework: Waters Part I — Molecular Weight eGFP Sequence: MVSKGEELFTG VVPILVELDG DVNGHKFSVS GEGEGDATYG KLTLKFICTT GKLPVPWPTL VTTLTYGVQC FSRYPDHMKQ HDFFKSAMPE GYVQERTIFF KDDGNYKTRA EVKFEGDTLV NRIELKGIDF KEDGNILGHK LEYNYNSHNV YIMADKQKNG IKVNFKIRHN IEDGSVQLAD HYQQNTPIGD GPVLLPDNHY LSTQSALSKD PNEKRDHMVL LEFVTAAGIT LGMDELYKLE HHHHHH Using the online calculator, the theoretical mass is 28006.60 Da. After subtracting the chromophore maturation (-20 Da), my predicted molecular weight comes out to 27,986.60 Da.
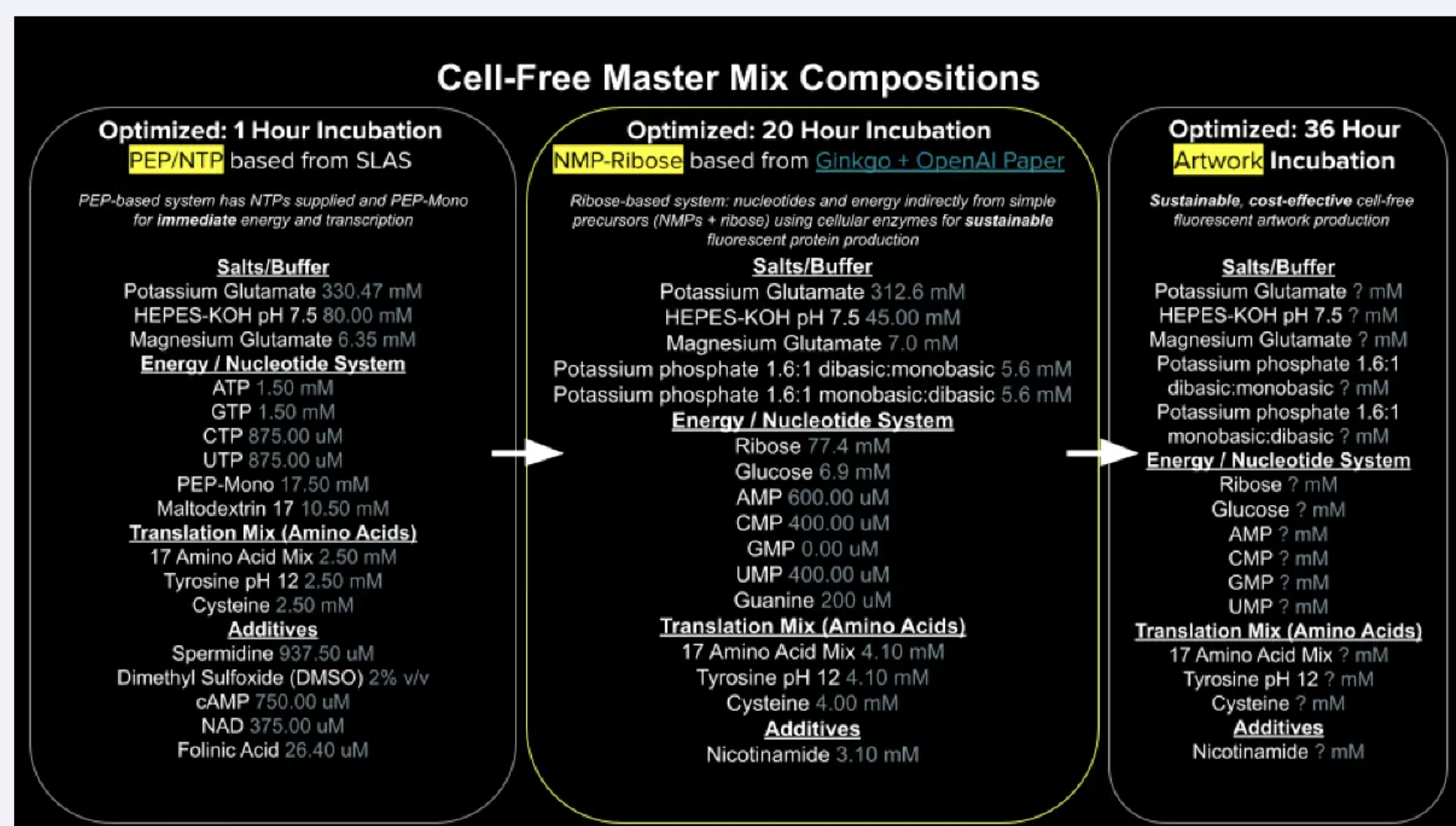
- Part A: The 1,536 Pixel Artwork Canvas | Collective Artwork I’m so sorry, I didn’t meet the deadline to submit but I will definitely be applying to be a TA this fall. :) Part B: Cell-Free Protein Synthesis | Cell-Free Reagents Referencing the cell-free protein synthesis reaction composition (the middle box outlined in yellow on the image above, also listed below), provide a 1-2 sentence description of what each component’s role is in the cell-free reaction.
- Part 1: Benchling & In-silico Gel Art If ignoring ladder and lane 1, there is the design! I tried to recreate No Face from Spirited Away. link to benchling: https://benchling.com/s/seq-8pB9vY3uTYXRrqsIKJS6?m=slm-FP5NlW0BAaqfbSmaoUg0 DNA Design Challenge For my homework, I decided to pick the enzyme that is the strongest candidate for facilitating luciferin synthesis. I picked my enzyme from my independent research. First I downloaded Fallon’s paper data from Fallon et al. 2018, “Firefly genomes illuminate parallel origins of bioluminescence in beetles” in eLife (DOI: 10.7554/eLife.36495. This experiment compared gene expression between the fat body (a firefly’s liver) versus the lantern (the organ that makes light) to find which genes are highly expressed in the lantern. I ran a filter in the file PPYR_OGS1.1_fatbody-vs…_test.txt texts, keeping only the statistically significant genes of TPM ≥ 50 and sleuth b ≥ 3. The TPM measures how actively a gene is being expressed in the lantern tissue, a higher TMP signaling a higher likelihood of luciferin expression. The sleuth is the statistical software Fallon used, that estimates log2 fold change. The higher the b sleuth, the more expression a gene has specifically in the lantern than fat body. I also ran qval ≤ 1e-10 to adjust for random noise.
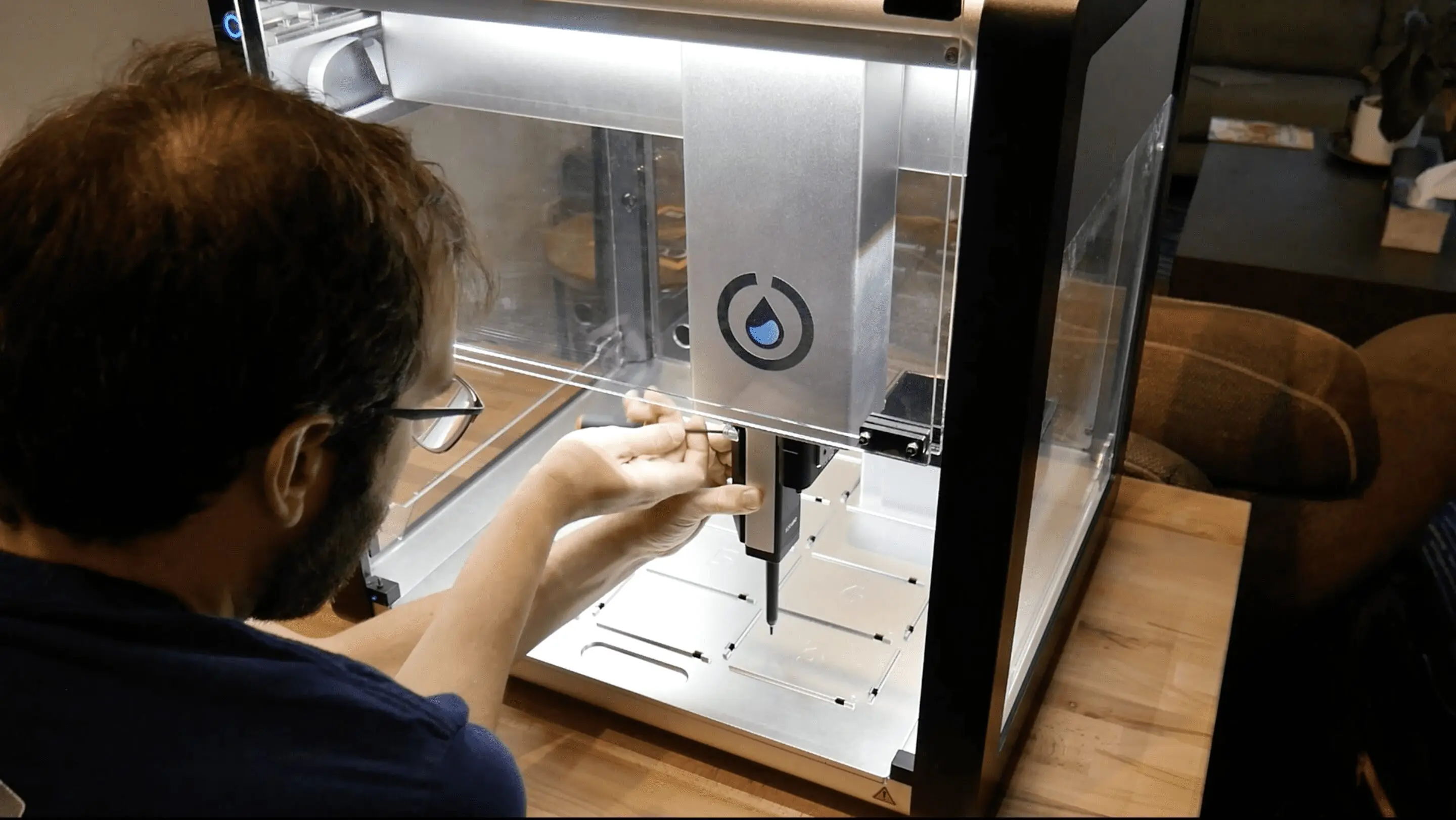
- Python Script for Opentrons Artwork After designing in http://opentrons-art.rcdonovan.com/ (a pig with a heart in the centre), I chose to run the Collab notebook for simplicity. I used Claude Code to help generate my coordinates into proper code, before running all. After my first simulation I noticed “WASTING BIO-INK : more aspirated than dispensed” warnings, so I edited my code. The next simulation still seemed to be wasting ink, but Claude informed me it’s an issue with floating point rounding.

- Part A. Conceptual Questions 1. How many molecules of amino acids do you take with a piece of 500 grams of meat? (on average an amino acid is ~100 Daltons) A dalton is a unit of mass defined as 1/12th the mass of an unbound neutral Carbon-12 atom at rest, equivalent to 1.66X10-24 grams. It is commonly used in molecular biology to calculate the mass of large molecules into grams. Meat protein is roughly 25% protein by mass, therefore 500g x 0.25 = 125g of protein since almost all amino acids are concentrated in the protein of meat.
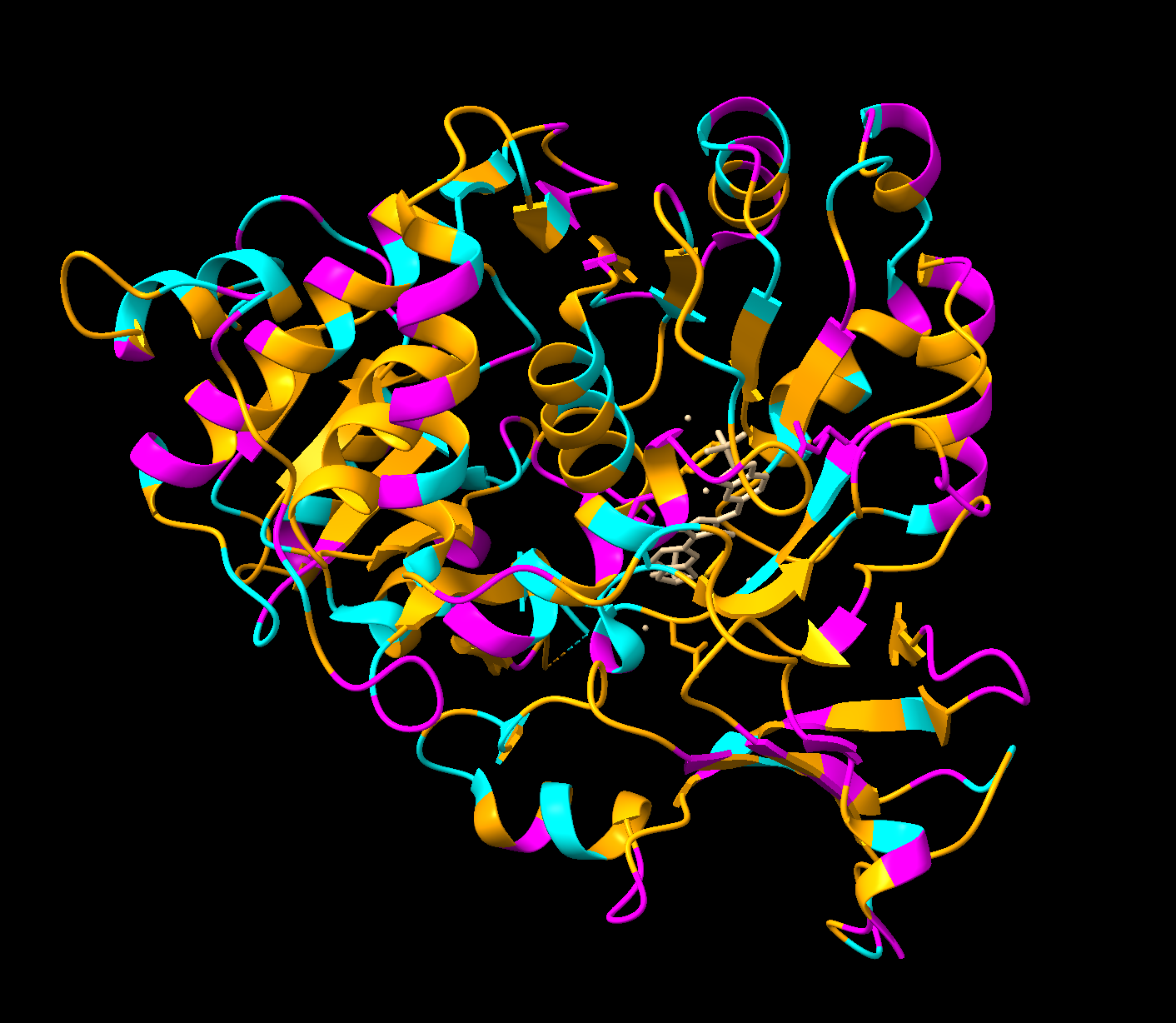
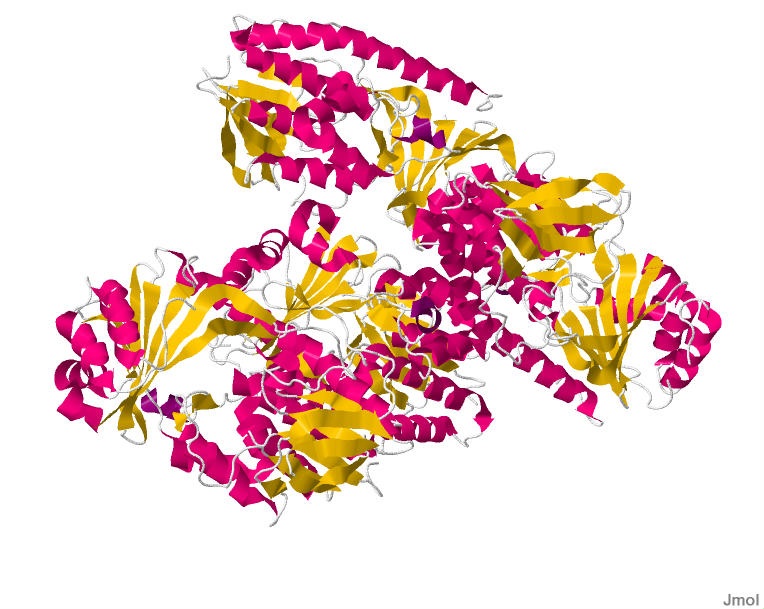
- Part A: SOD1 Binder Peptide Design (From Pranam) Part 1: Generate Binders with PepMLM sp|P00441|SODC_HUMAN Superoxide dismutase [Cu-Zn] OS=Homo sapiens OX=9606 GN=SOD1 PE=1 SV=2 MATKAVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTS AGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVV HEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ SOD1 A4V mutation: MATKVVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTS AGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVV HEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ Notestranslation begins with Methionine in eukaryotes, therefore the position of translation is technically 5 (A -> V). Protein language models train on amino acid sequences. Masked language modelling trains sequences by randomly masking some positions, and training the model to fill in the masked positions based on context. PepMLM does the same thing but to peptide binder design, where it masks all the peptides in the proteins you give, iteratively fills in those masked positions from most to least confident. Low perplexity = more confident, high perplexity = model uncertain.
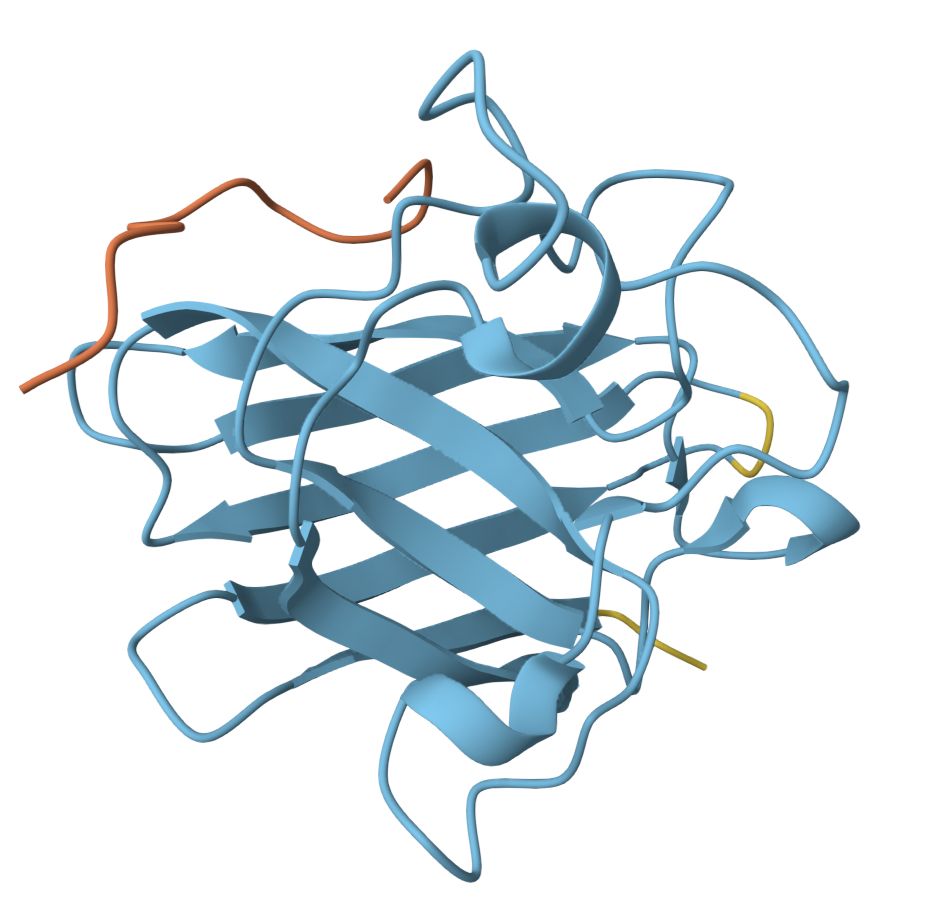
- Assignment: DNA Assembly What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose? Components included are Phusion DNA Polymerase, deoxynucleotides (dNTPs), and HF Reaction Buffer containing MgCl₂. Physion DNA polymerase is a Pyrococcus-like enzyme. The polymerase domain reads the strand and synthesizes it by adding dNTPs one at a time (through phosphodeister bond formation). This requires a primer to already be present in the template. The enhancing domain fused onto the polymerase keeps the enzyme attached to the DNA template, speeding up the process. The exonuclease domain reads this newly synthesized stand and corrects any mistakes. Deoxynucleotides (dNTPs) are the building blocks of DNA at optimized concentrations. The HF reaction buffer helps control the pH for optimal activity, while the MgCl₂ positions the dNTPs correctly and stabilizes the transition state during bond formation.
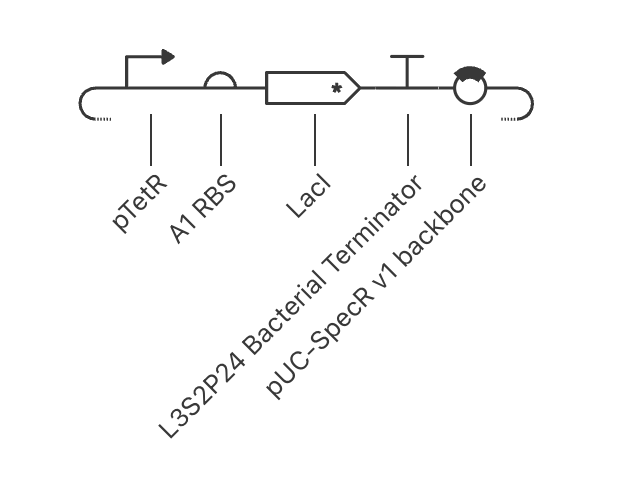
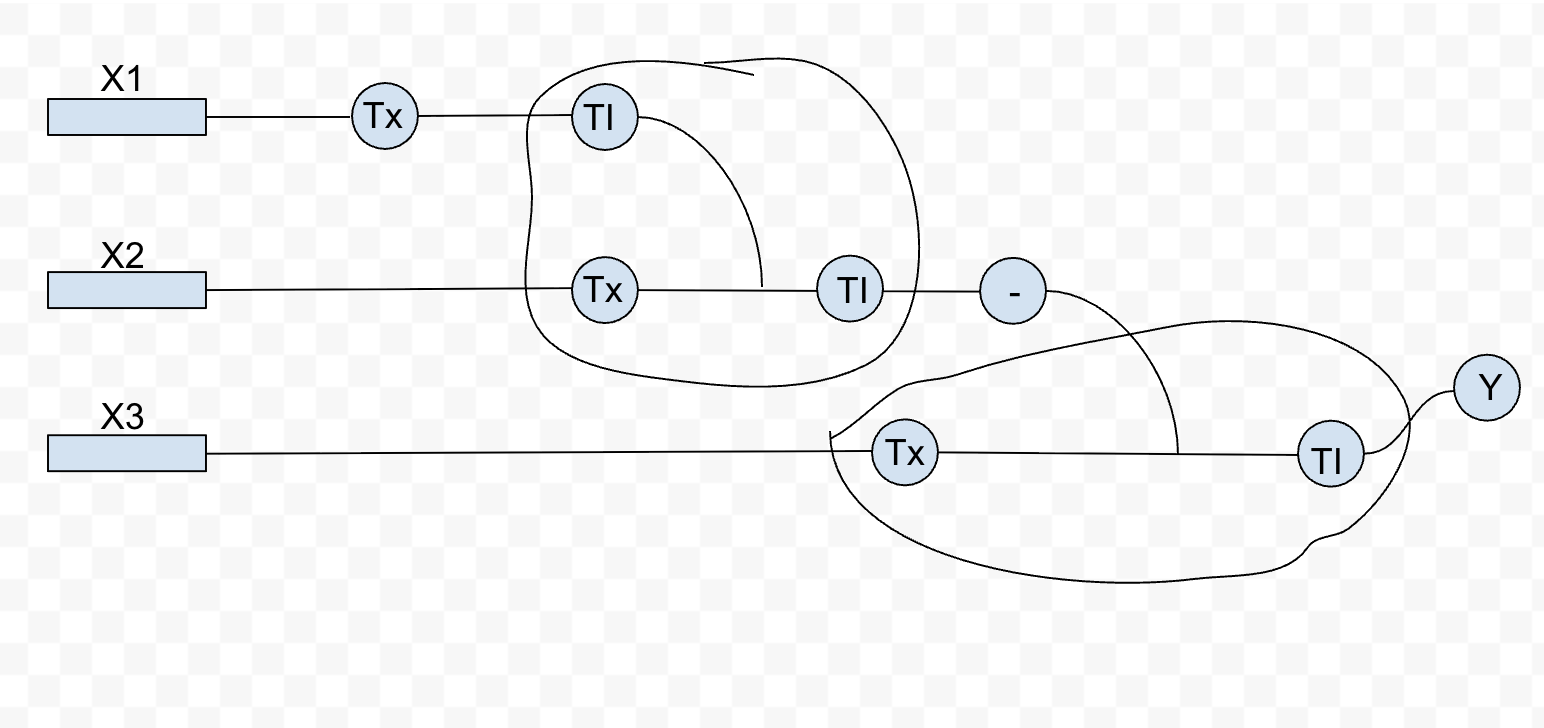
- Assignment Part 1: Intracellular Artificial Neural Networks (IANNs) What advantages do IANNs have over traditional genetic circuits, whose input/output behaviors are Boolean functions? IANNs (Intracellular Artificial Neural Networks) have several advantages over traditional genetic circuits. Booleans inherently work in a binary fashion: there is either a statement ment that makes it true/false, and therefore, either on/off. IANNs have the ability to process continuous signals, allowing for inputs to be weighted against each other before an output. This makes the system more sensitive to differences, not just whether something is true or false. This is great for cells as molecular concentrations exist on a spectrum. Another reason is its reconfigurability: you can reprogram the IANN circuit without needling to completely rebuild it. IANNs also make multi-input easier to configure, as an arbitrary number of inputs can be collapsed into one step, instead of building onto each other in booleans. Lastly, boolean circuits are static, with inherently fixed behaviour after it is built. IANNs are defined by weights, allowing for greater flexibility if you were to modify the weight thresholds.
- Homework Part A: General and Lecturer-Specific Questions Explain the main advantages of cell-free protein synthesis over traditional in vivo methods, specifically in terms of flexibility and control over experimental variables. Name at least two cases where cell-free expression is more beneficial than cell production. Cell-free protein synthesis provides multiple advantages over traditional methods due to its lack of membrane. You don’t have to cross the membrane, and the cell’s homeostasis controls don’t resist any experimentations. Furthermore, since everything is now a controllable variable, you have better control over composition (such as concentrations) of amino acids, substrates, etc. You also maintain better folding control and template flexibility (since you don’t have to clone into plasmids). Cases where this is beneficial include space travel (since cell-free synthesis allows for freeze-drying, which is more shelf-stable in storage), and anywhere else resources may be limited to run a fermenter (war zones, rural clinics, etc.) Another case is the expression of “unhostable” proteins, which would in other cases, kill its living cell. The cell-free environment means you bypass the actual cell host.