Week 3 HW: Lab Automation
Python Script for Opentrons Artwork
After designing in http://opentrons-art.rcdonovan.com/ (a pig with a heart in the centre), I chose to run the Collab notebook for simplicity. I used Claude Code to help generate my coordinates into proper code, before running all.

After my first simulation I noticed “WASTING BIO-INK : more aspirated than dispensed” warnings, so I edited my code. The next simulation still seemed to be wasting ink, but Claude informed me it’s an issue with floating point rounding.
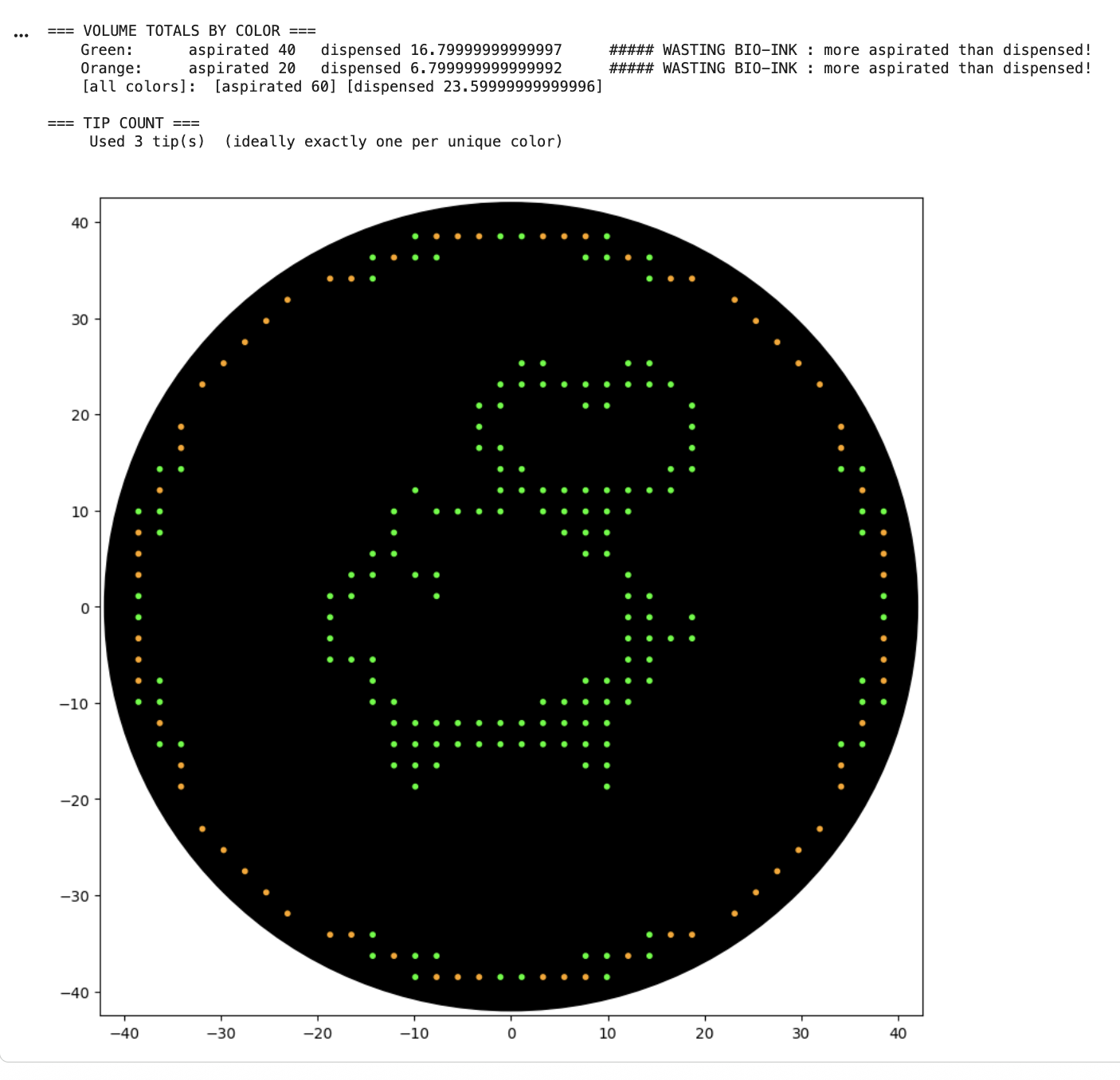

Python script link: https://colab.research.google.com/drive/1ONInhseZmTcpt755AN2gSf4Gsnn31Nd3#scrollTo=pczDLwsq64mk&line=1&uniqifier=1
Post-Lab Questions
Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
Since my project requires extensive wet-lab trial and error, I found “AssemblyTron: flexible automation of DNA assembly with Opentrons OT-2 lab robots” the most applicable paper. DNA assembly is mostly still done by hand, which makes such a meticulous process error-prone. Furthermore, it poses a high barrier to entry for those without lab or educational access. AssemblyTron is a free, open-source Python package you download for the Opentrons OT-2 that is optimized for DNA construct design protocols including: PCR setups, Golden Gate Assembly, Homology-dependent assembly, and more. The Opentron handles a wide range of tedious and time-consuming processes, allowing for better time-management and faster research progress. Lastly, as someone with a diagnosed hand tremor, I struggle constantly with contamination and resource waste (ex. repeatedly grazing tips) in the lab. This would be immensely helpful for individuals who struggle with fine-motor skills, especially for delicate procedures such as cloning methods.
Write a description about what you intend to do with automation tools for your final project. You may include example pseudocode, Python scripts, 3D printed holders, a plan for how to use Ginkgo Nebula, and more.
My plan requires a four-gene bioluminescence pathway through Golden Gate assembly. These are novel genes in plants, and will inevitably require trouble shooting via different combinations of promoters, luciferase variants, targeting signals, etc. Instead of assembling one construct at a time, (a real time-related bottleneck in this project), I could set up dozens of constructs in parallel. An example pseudocode could be as follows:
Work Cited
Bryant, J. A., Kellinger, M., Longmire, C., Miller, R., & Wright, R. C. (2023). AssemblyTron: flexible automation of DNA assembly with Opentrons OT-2 lab robots. Synthetic Biology, 8(1), ysac032. https://doi.org/10.1093/synbio/ysac032
Ai Citations (Claude 4.6)
Generate [coordinates] into this code [your code section in Collab notebook] for my opentrons project.
Edit the code to not waste bio-ink [attached screenshot]
Is it still wasting ink? [attached screenshot]
What are the cutting edge Opentrons use cases in synthetic bio right now?