Week 2 HW: DNA Read, Write and Edit


Important
Part 1: Benchling & In-silico Gel Art
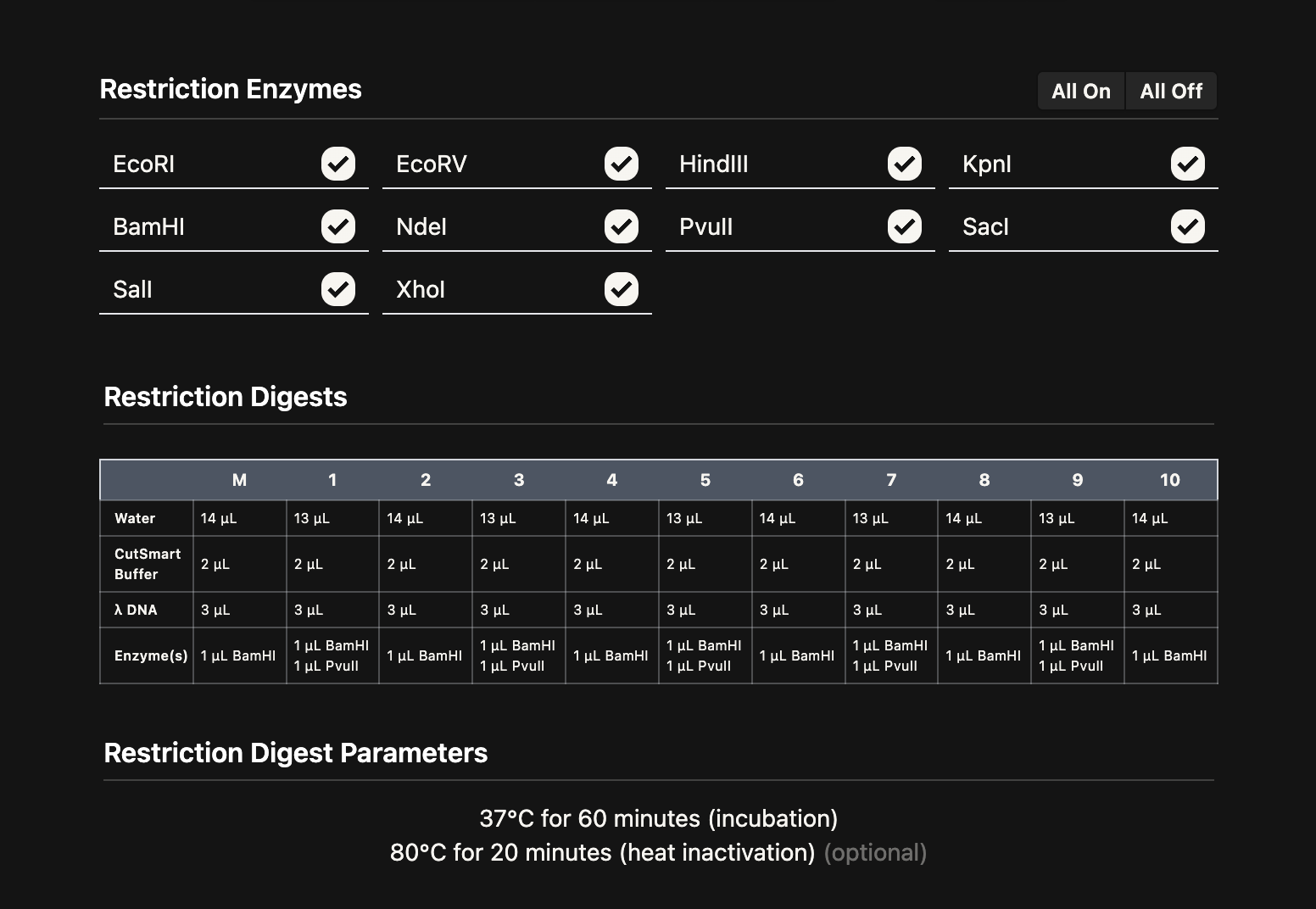
Using Benchling, I imported the Lambda DNA and began simulating the Restriction Enzyme Digestion with the following enzymes: EcoRI-HF, HindIII-HF, BamHI-HF, KpnI-HF, EcoRV-HF, SacI-HF, SalI-HF.
Here is the Virtual Digest (left) and Linear Sequence Map (right) I produced in Benchling:
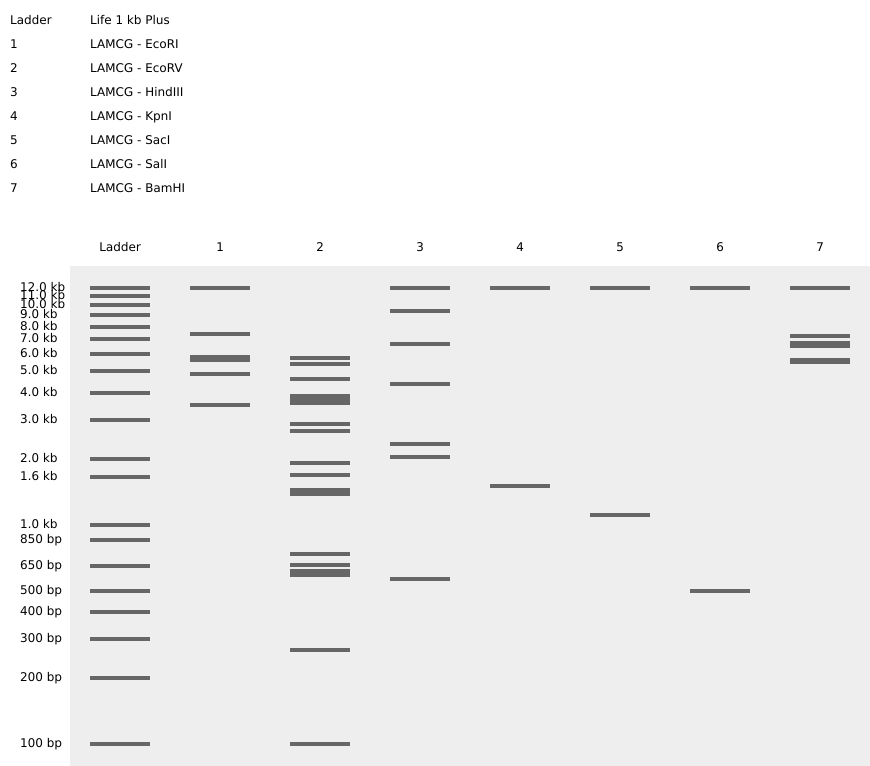

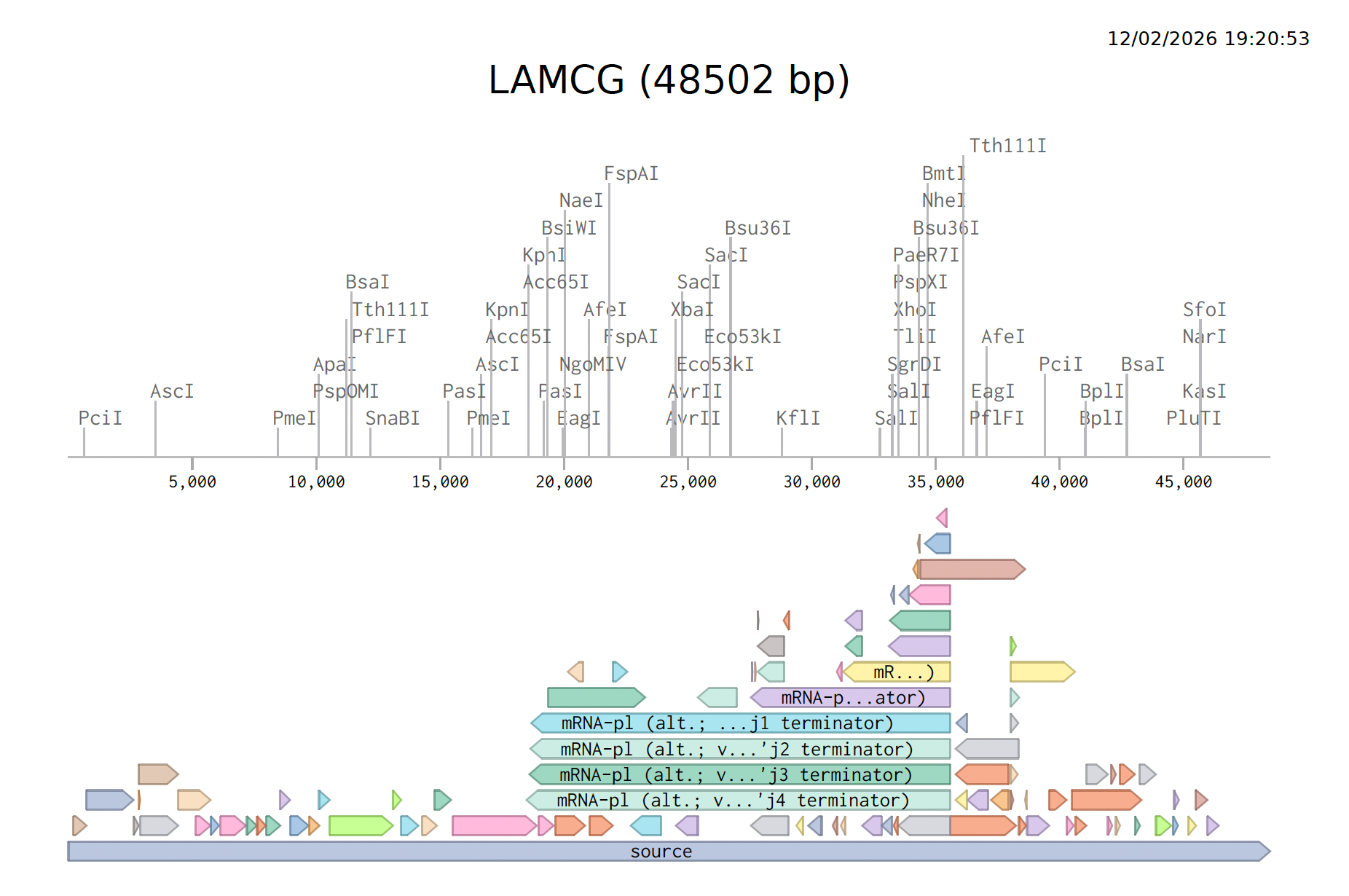
Click here to see the linear sequence map in Benchling
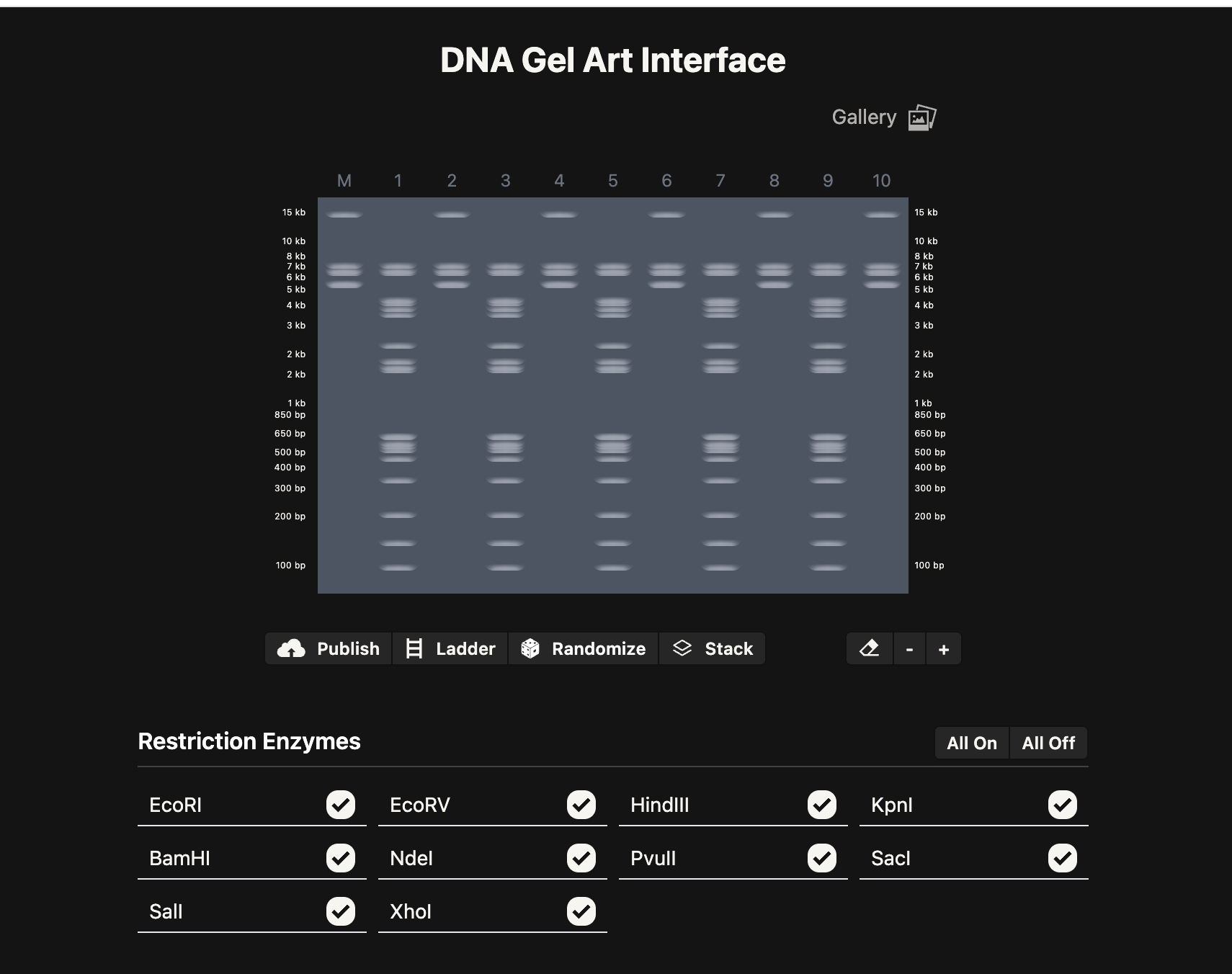
Next, I create a pattern/image in the style of Paul Vanouse’s Latent Figure Protocol artworks using this Ronan’s iterating tool. This is preparation for a Gel Electrophoresis lab, a fundamental laboratory technique used to separate and analyze DNA but with the added fun of using the process to make DNA Gel art.
I made this funky design:
Part 3: DNA Design Challenge
Choose your protein:
I have been thinking a lot about Chronobiology. I have an interest in our connection to the temporalities and rhythms of more than human species. Brainstorming for my final project, I am thinking about ways that I could use speculative bio-design to explore the patterns of presence and absence of other species and make tangible specific patterns of activity that are less visible, illusive or little known.
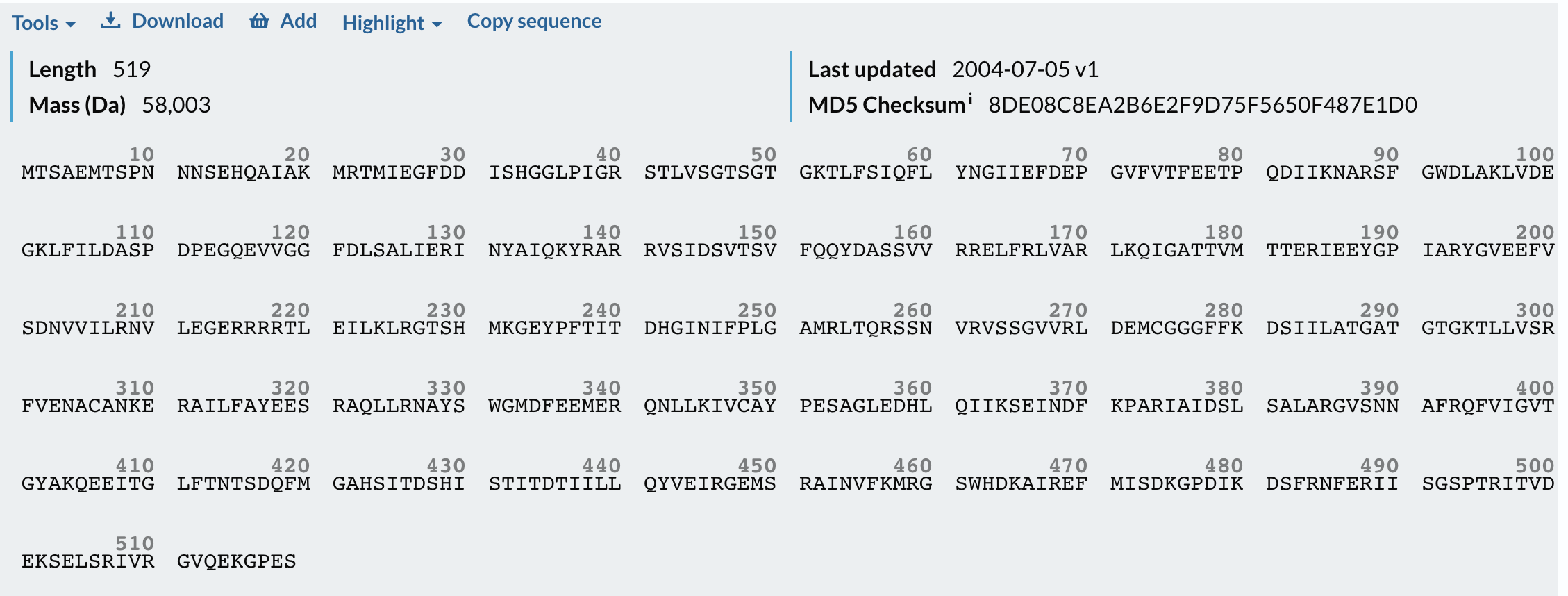
So I chose two proteins for the bio-design challenge. Below are the details, structure and DNA sequence from Uniprot
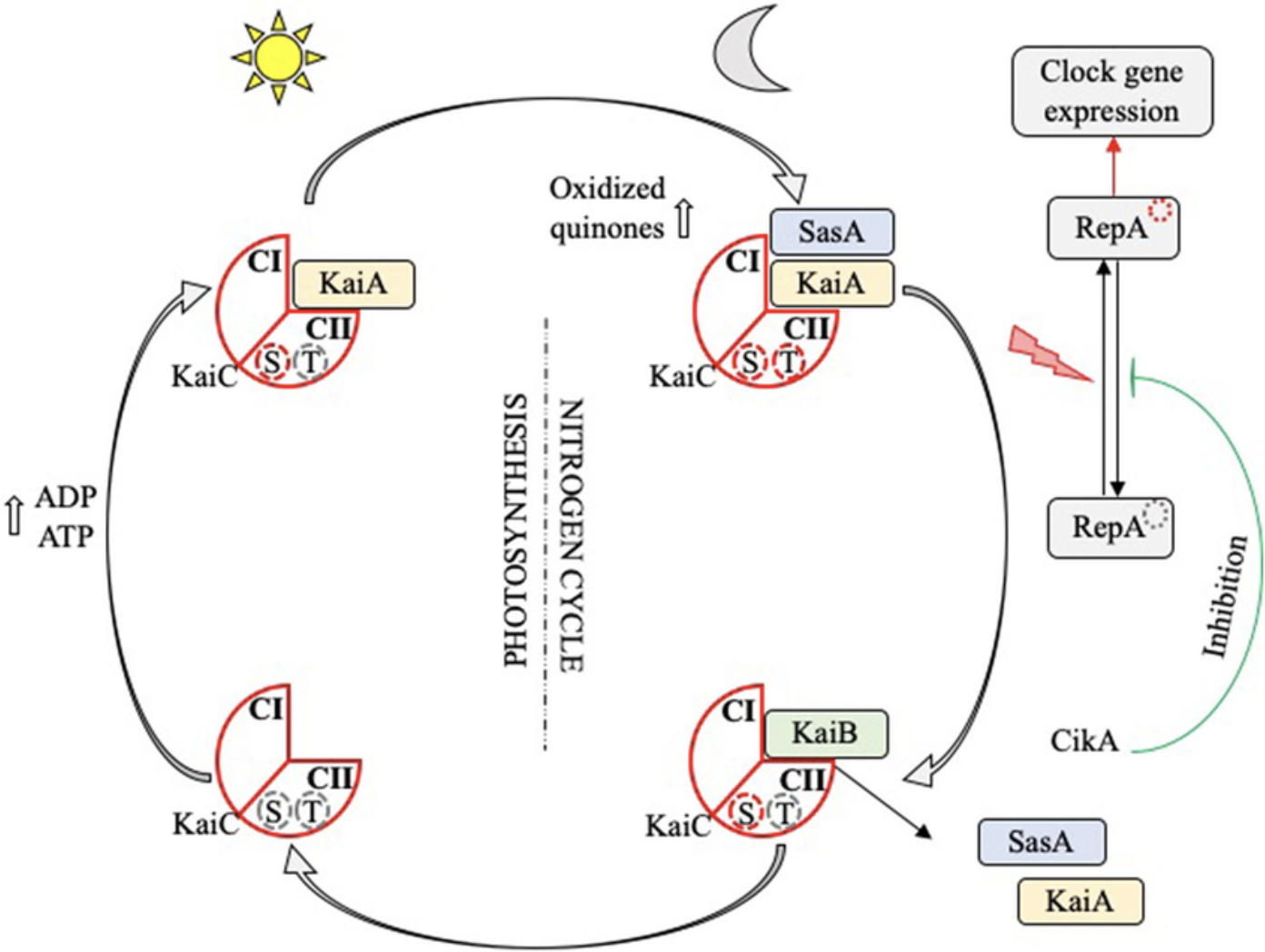
- The circadian clock oscillator protein KaiC in Synechococcus elongatus (cyanobnacteria).
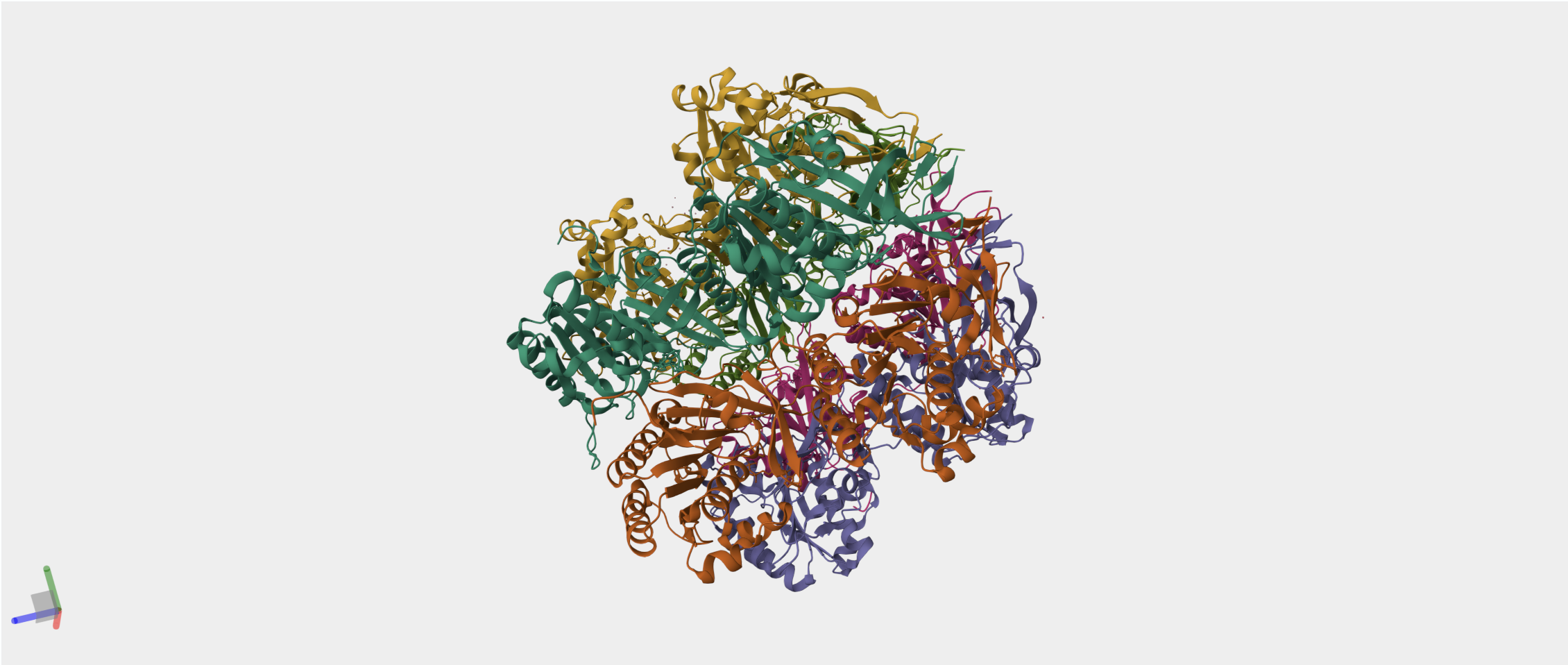

I have chosen the KaiC protein from Synechococcus elongatus PCC 7942 because it is the core component of the cyanobacterial circadian clock (The KaiABC). KaiC undergoes a precise phosphorylation and dephosphorylation cycle that repeats approximately every 24 hours, creating a reliable biological oscillator.

- Firefly Luciferase from the North American firefly Photinus pyralis that catalyzes bioluminescence.
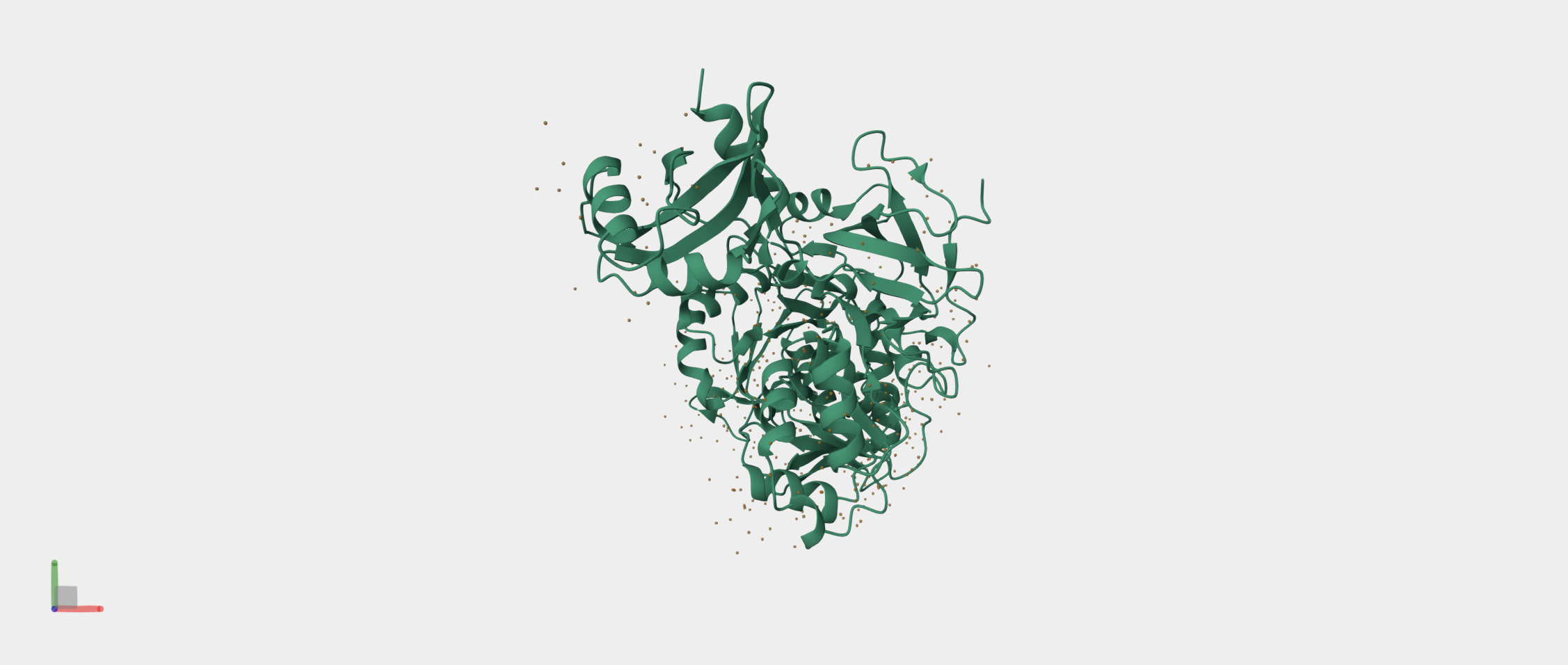

It produces the firefly’s characteristic yellow-green light through oxidation of the small molecule luciferin. I also chose to study the Firefly Luciferase sequence from Photinus pyralis because it is a well-characterized bioluminescent enzyme used extensively as a reporter in gene expression studies.
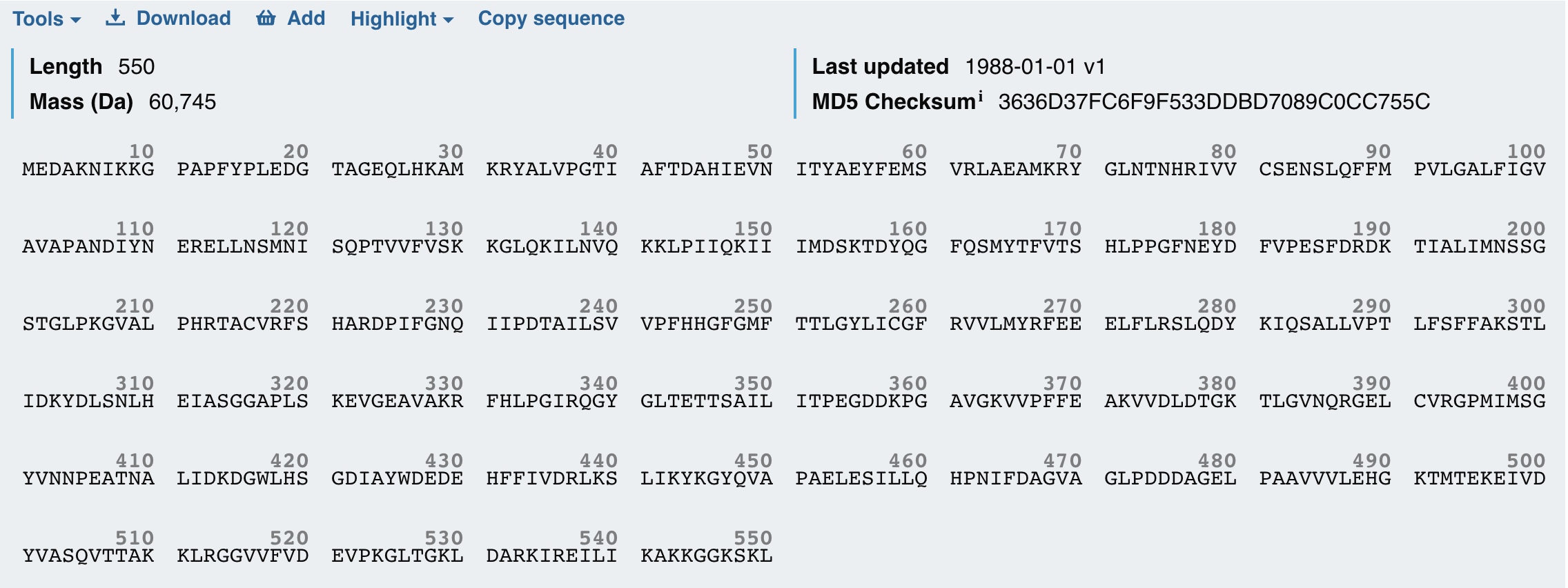

By fusing Luciferase to KaiC, I can convert these molecular oscillations into a measurable bioluminescent signal. This fusion allows real-time monitoring of the circadian rhythm, with peaks in light output corresponding to high KaiC activity and troughs corresponding to low activity. I wanted to explore both proteins today because together KaiC provides the biological timing mechanism and Luciferase provides a visible readout, making them an ideal pair for a circadian biological clock.

Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
The central dogma of molecular biology, proposed by Francis Crick, describes the flow of genetic information: DNA is transcribed into RNA, which is translated into protein (DNA>RNA>PROTEINS).
In Transcription DNA sequences are converted to RNA. This occurs in the nucleus and is mediated by RNA polymerase.
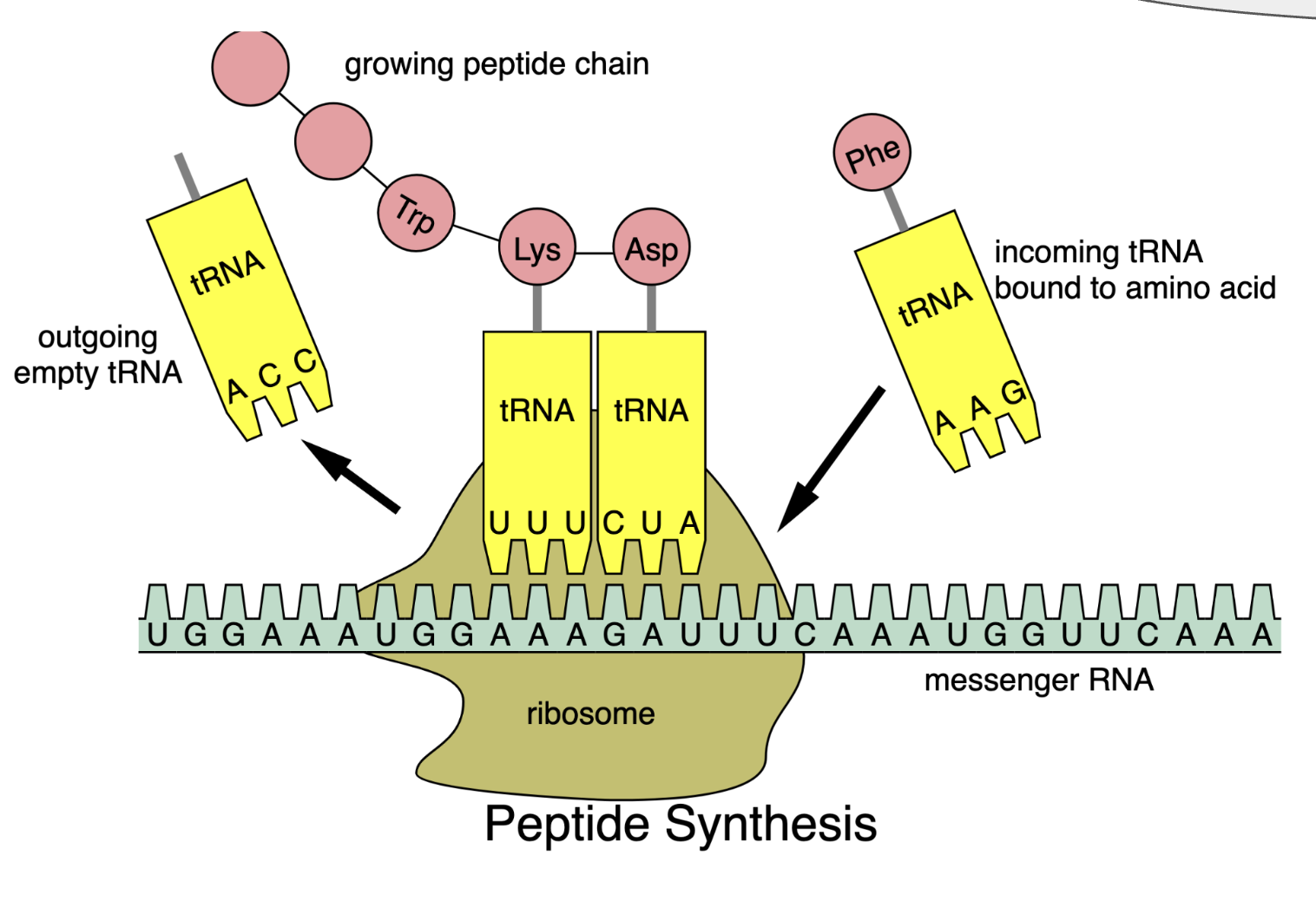
In Translation messenger RNA is decoded to synthesize a specific polypeptide chain and occurs in the cytoplasm where ribosomes read mRNA instructions to make proteins.
The Central Dogma gives us the framework to work backwards from a given protein sequence and infer the DNA sequence that the protein is derived from.
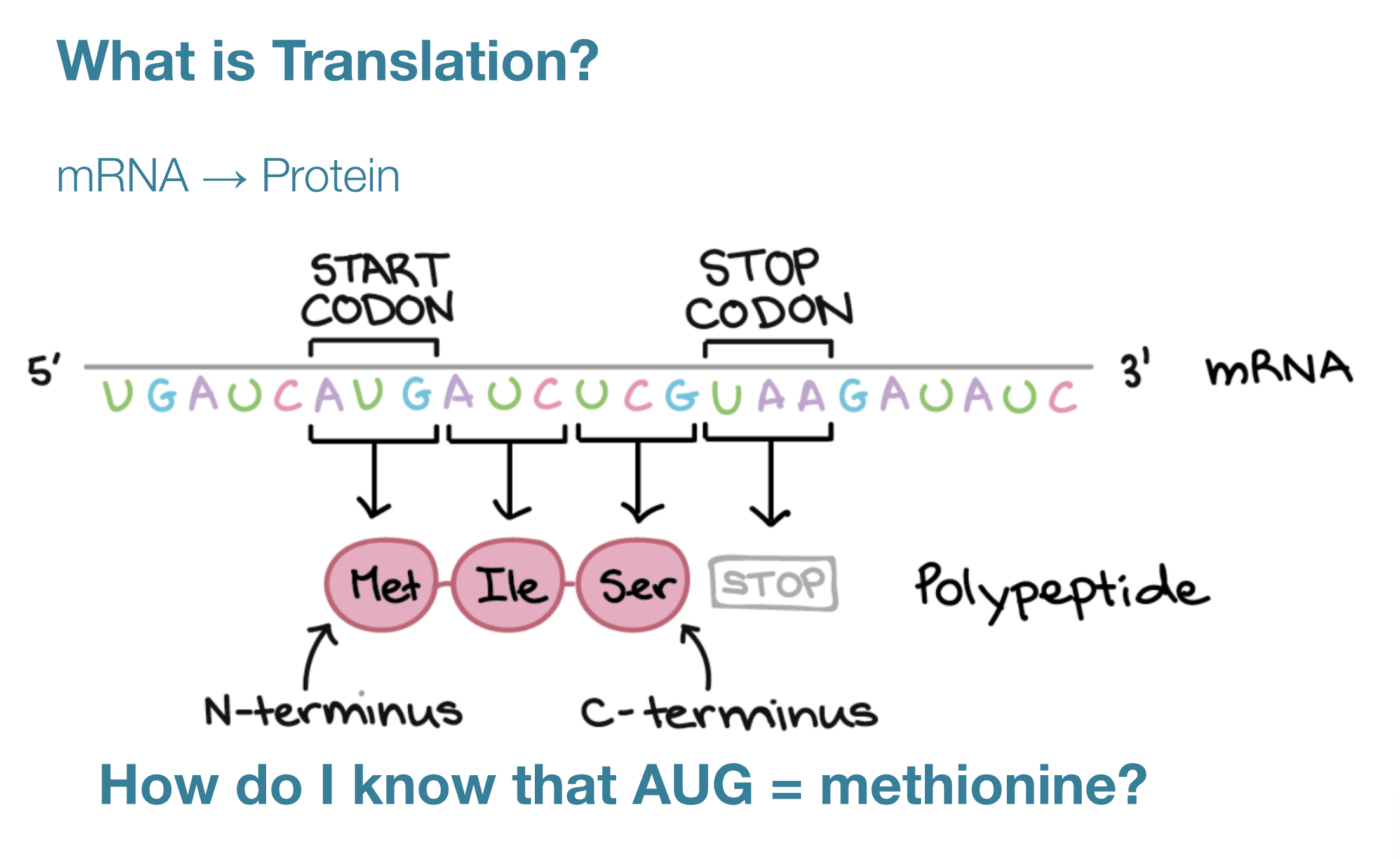
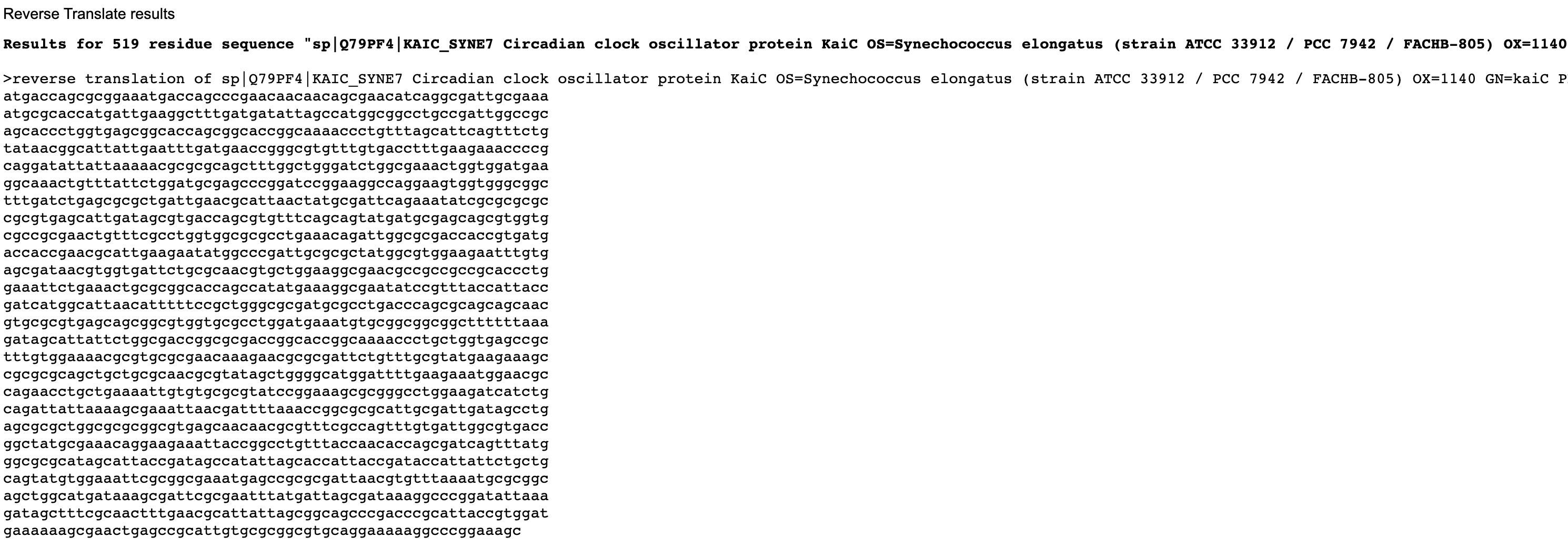
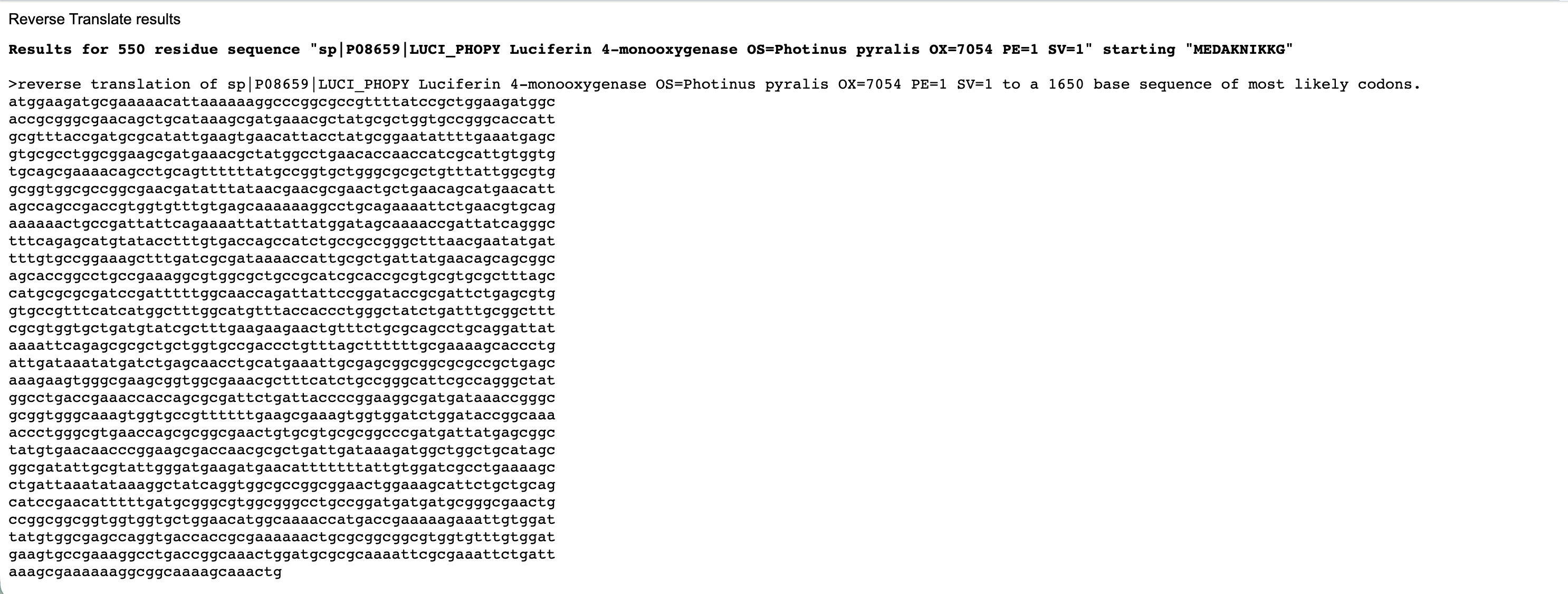
Below are the nucleotide sequences (reverse translation) for KaiC (above) and Luciferase (below). I used the online tool Sequence Manipulation Tool from Bioinformatics by entering the raw amino acid sequence for each protein.
Codon optimization: Describe why you need to optimize codon usage?
Codon optimisation is a process used to make translation more efficient and improve gene expression.
Different organisms have codon usage bias, meaning they prefer to use certain codons to specify each amino acid based on their abundance of matching tRNAs.
Codon optimisation replaces codons rare to the host organism with codons it prefers, allowing the ribosome to translate the mRNA more efficiently. This is especially important when expressing a gene from one organism in a different host, because unoptimised codons can slow translation, reduce protein yield, or cause errors in protein production.
Codon optimization: _Which organism have you chosen to optimize the codon sequence for and why?
I have chosen to optimise both KaiC and Luciferase for Escherichia coli strain K-12 by inputting the nucleotide sequence into the Integrated DNA Technologie’s Codon Optimisation Tool
This is because:
It is a standard laboratory organism that is widely available, cheap, has a well understood codon usage bias and established protocols for isolating plasmids and cloning.
Its fast growing with high protein expression efficiency.
Specifically, for a possible project where I would like to fuse Luciferase to KaiC to act as a bioluminescent reporter of circadian rhythms, codon optimisation would be important. Codon optimisation of both KaiC (from cyanobacteria) and firefly Luciferase (from eukaryotes) ensures that E. coli can efficiently translate the fusion protein using its tRNA pool, allowing high-level expression suitable for detecting and reporting circadian rhythms. The KaiC portion reliably maintains its role in the circadian oscillator, while the Luciferase portion generates strong bioluminescence that creates a good visualisation of the circadian rhythmns.


Codon optimised nucleotide sequence for KaiC.
KaiC comes from Synechococcus a prokaryotic cyanobacterium, but its codon usage bias is different from E. coli. Optimising KaiC for E. coli ensures efficient translation and more abundant protein production. Firefly Luciferase is eukaryotic meaning its codons are rare in bacteria, so without optimisation translation could be inefficient. Optimising each gene independently for E. coli ensures that translation of one protein does not impact the other. Optimising both genes for the same host ensures coordinated and efficient expression within a single engineered system, allowing it to accurately report the dynamics of the circadian clock.


Codon optimised nucleotide sequence for Firefly Luciferase.
Cell-dependent production of Luciferase in _E. coli

LIGATION:
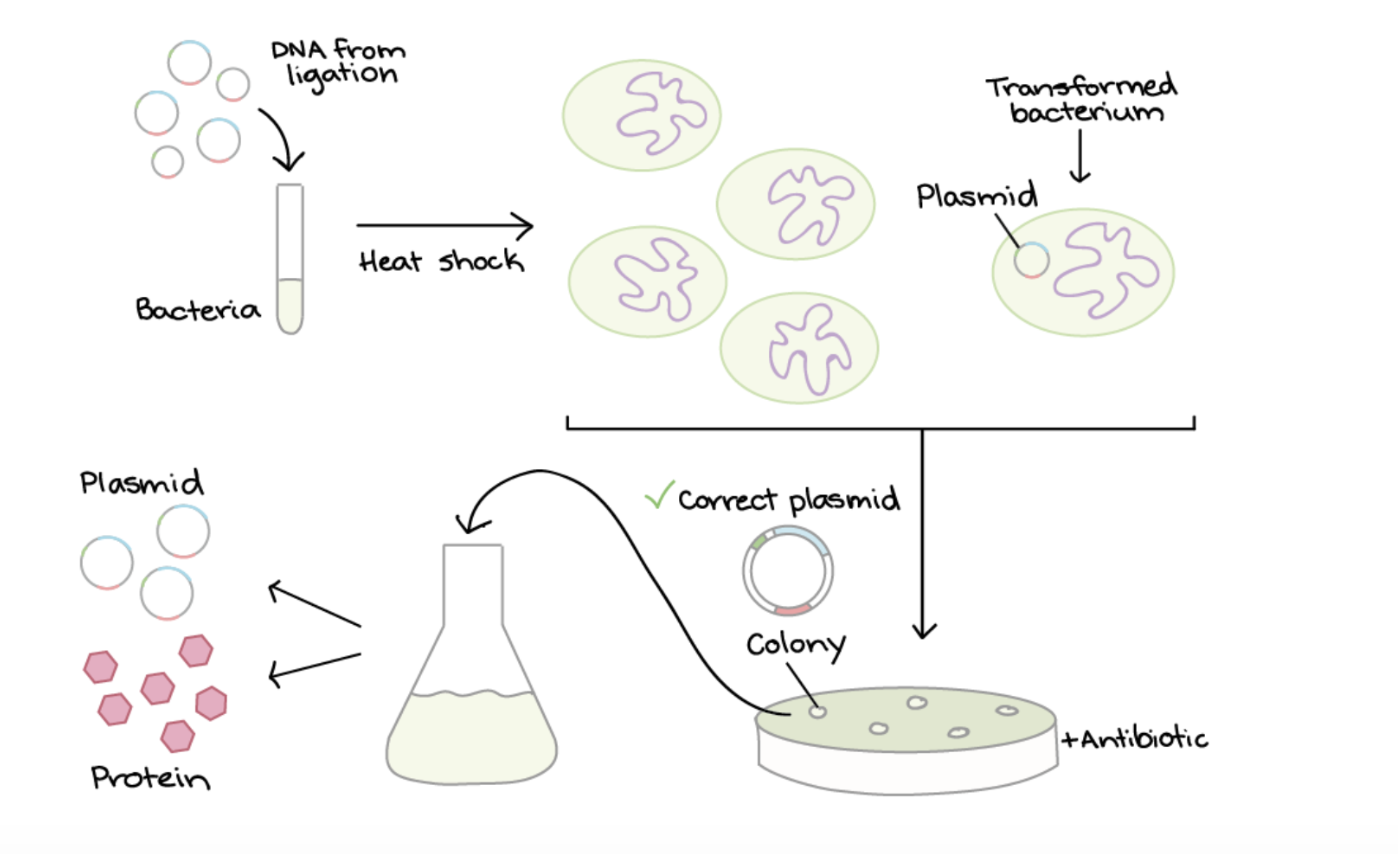
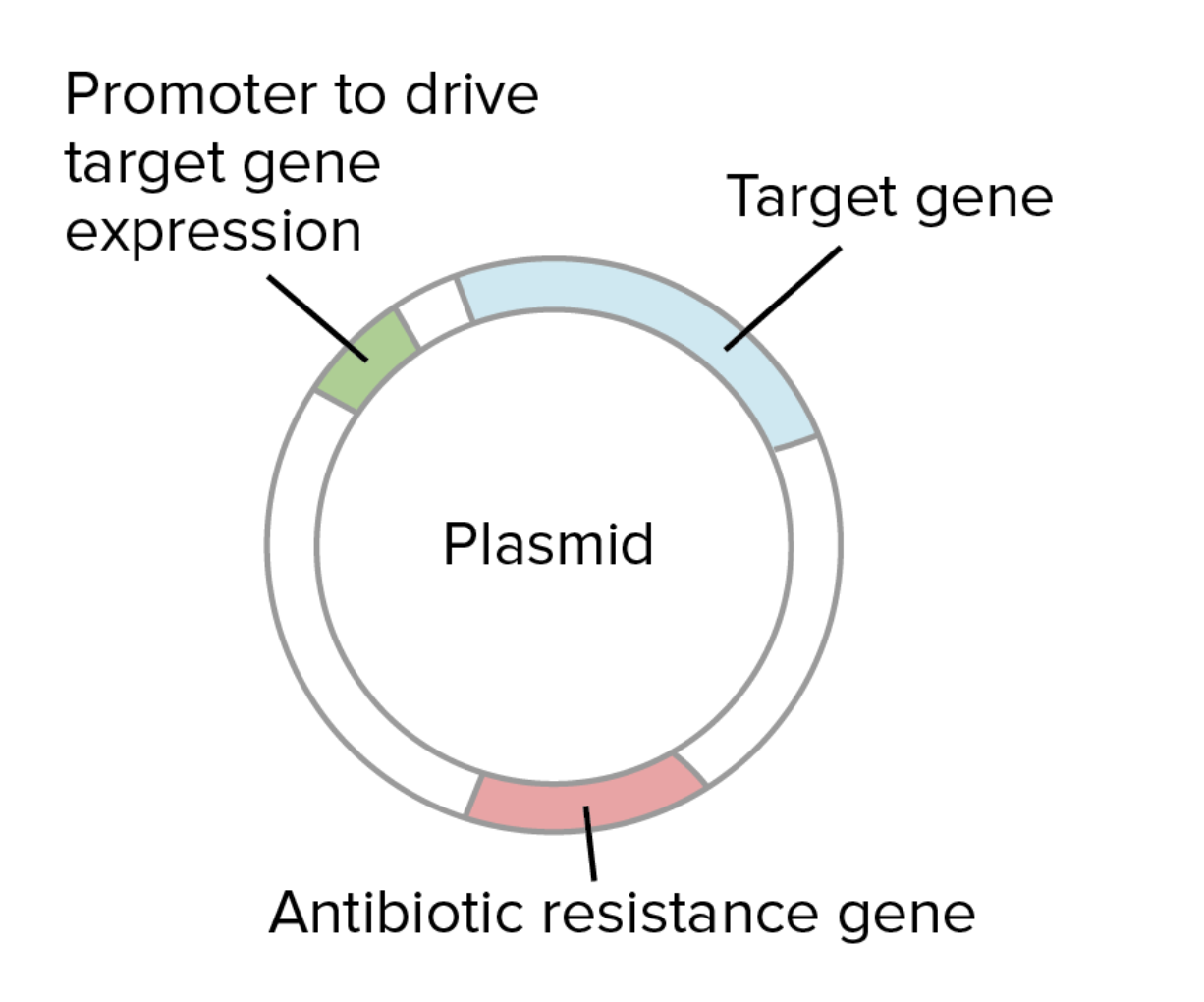
Plasmids are a small circular piece of DNA used that replicate independantly fromthe hosts chromosomal DNA. They are used to introduce foreign DNA into another cell. To produce the protein Luciferase by cell dependant production in Ecoli, first the plasmid (designed for expression in E.coli) and the Luciferase gene are cut using restriction enzymes to make compatible ends. The gene is then inserted into the the plasmid through ligation. This is where DNA ligase enzyme binds the compatible ends of plasmid and Luciferase gene. The plasmid contains a promoter to drive transcription in the E.coli and an antibiotic resistance gene for selection.
TRANSFORMATION:
This plasmid is then introduced into E. coli through transformation, where heat shock encourages some bacteria to take up the plasmid. The shock makes the bacterial membrane more permeable to the plasmids.
SELECTION:
The bacteria are grown on an antibiotic plate. The plasmids contain a antibiotic gene so only bacteria that successfully took up the plasmid survive and form bacterial cultures.
SCREENING:
Colonies are screened using PCR, restriction digest or sequencing to identify a plasmid with the Luciferase gene in the correct direction relative to the promoter and to ensure the plasmid hasn’t closed back up without taking in the gene.
PROTEIN PRODUCTION:
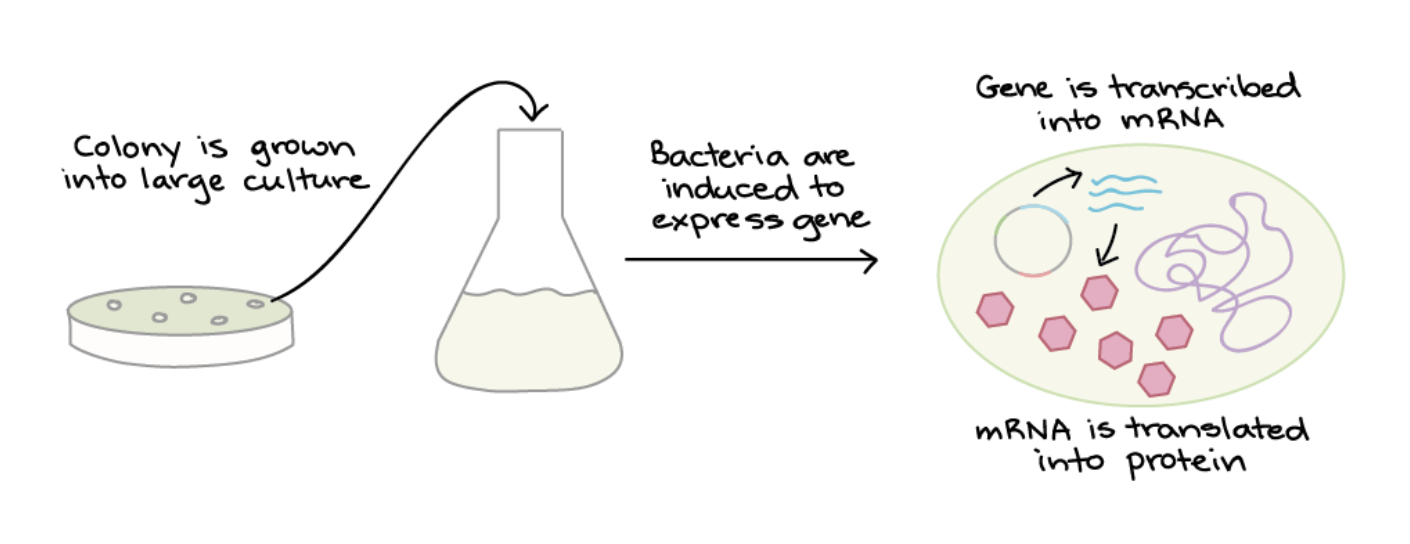

Once a colony with the correct plasmid is identified, it is grown in bulk, and protein expression is induced by adding a chemical signal. Inside the bacteria, the codon-optimised Luciferase DNA is transcribed into mRNA and then translated by bacterial ribosomes into luciferase protein. The codon optimisation ensuring that translation is efficient, even though Luciferase is originally a eukaryotic protein.The bacteria can then be lysed (split open) to release the protein.
PURIFICATION:
Finally, the Luciferase protein can either be used directly in bacterial cells or purified. Purification techniques include affinity chromatography, where antibodies specific to luciferase bind and isolate the protein.
Part 4: Prepare a Twist DNA Synthesis Order
Finally, I’ve done a test order from Twist inserting Bioscience the codon optimised sequence for Luciferase.
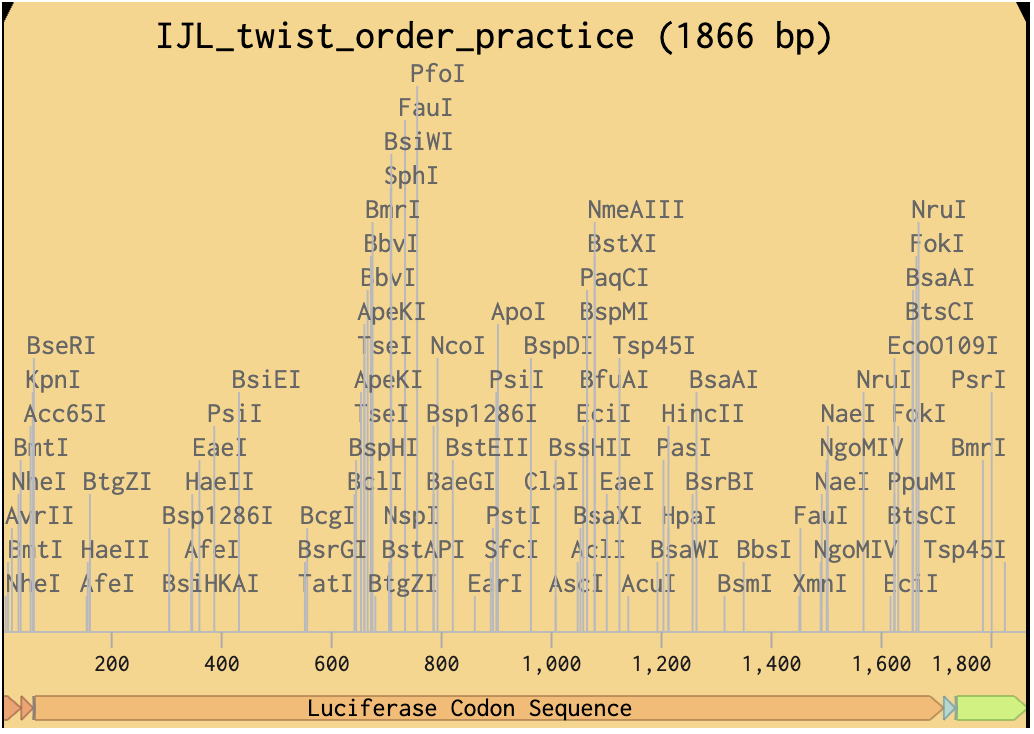
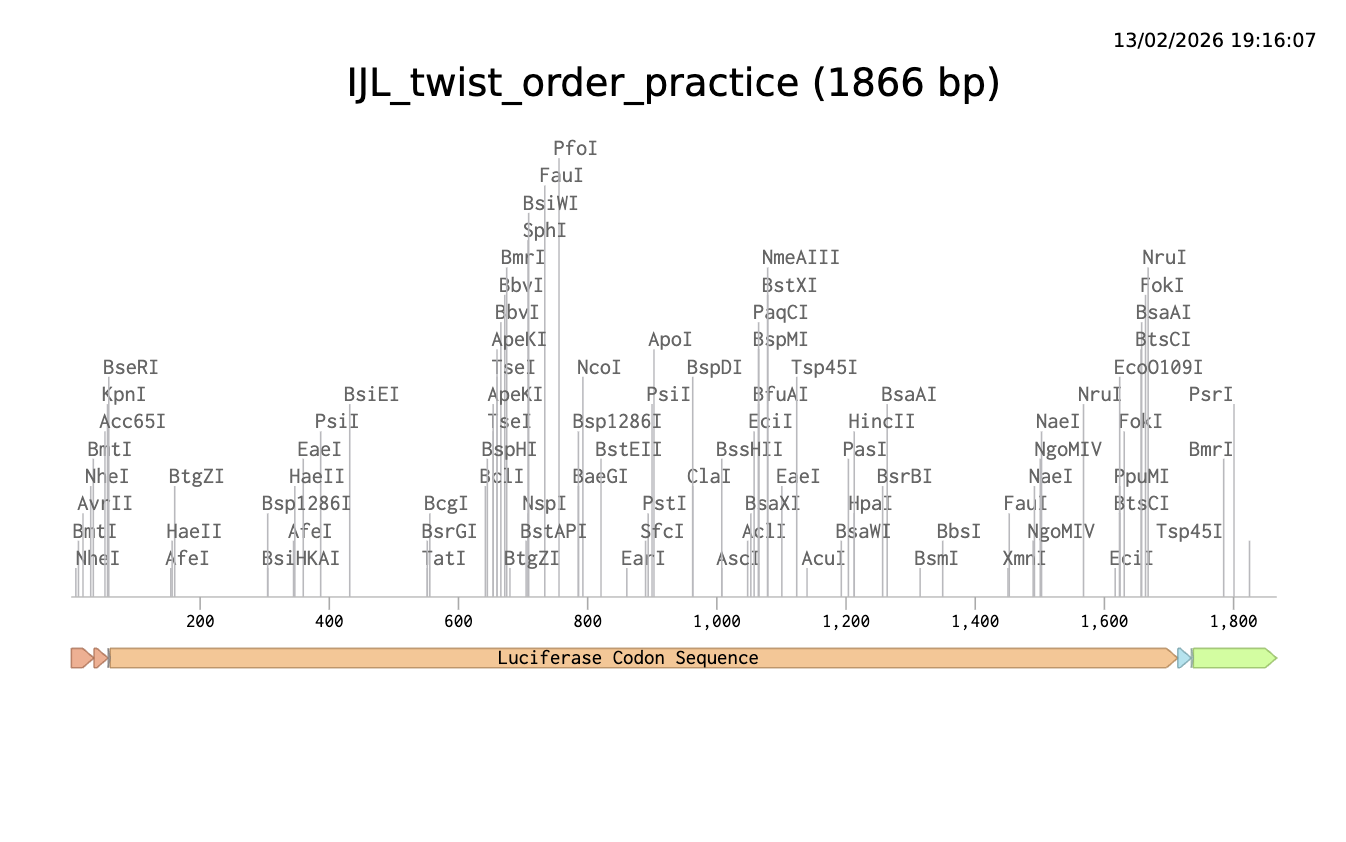
Click here to go to my linear map
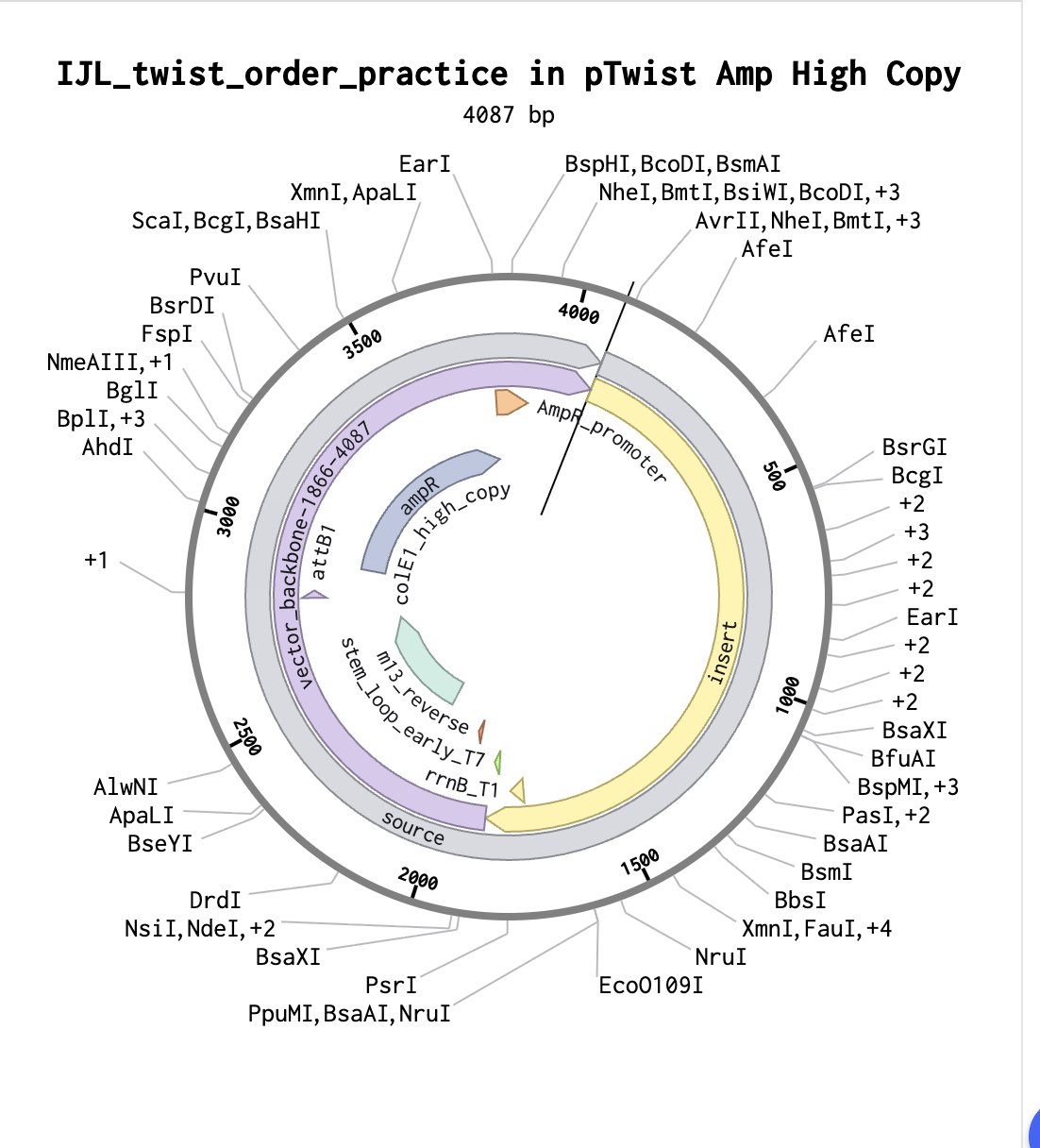
And Click here for baby’s first plasmid!
5.1 DNA Read
What DNA would you want to sequence (e.g., read) and why?
Sequencing ELF3 to track impact of climate changing on circadianflowering in plants.
I would sequence the ELF3 (EARLY FLOWERING 3) gene, which is a key player in the circadian clock of plants such as Arabidopsis thaliana’s. It’s function includes the regulation of flowering time in response to light and temperature variations, therefore mutations within the ELF3 DNA can lead to the loss of photoperiod sensitivity and shifts in seasonal flowering.
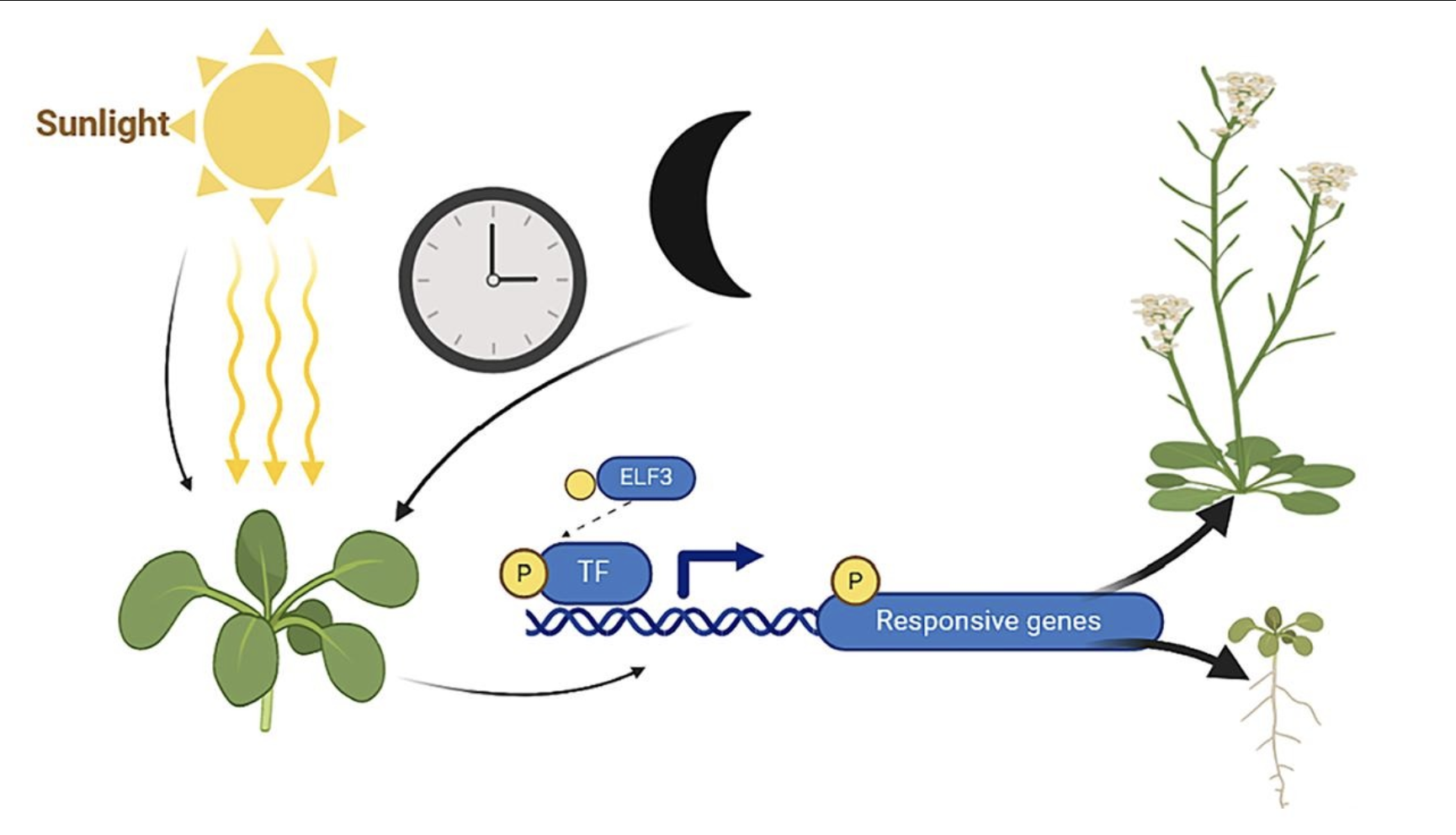

Zhu X, Wang H. Revisiting the role and mechanism of ELF3 in circadian clock modulation. Gene. 2024 Jun 30;913:148378. doi: 10.1016/j.gene.2024.148378. Epub 2024 Mar 13. PMID: 38490512.
Sequencing ELF3 allows us to identify genetic variations or mutations such as single nucleotide polymorphisms (SNPs) or insertions/deletions that could shift flowering schedules, which is particularly important for studying how climate change may disrupt plant phenology. Changes in flowering time can misalign plant/pollinator interactions, affecting ecosystems and crops so it is a crucial thing to understand.
ELF3 would be interesting to sequence because its variations make invisible molecular changes visible and can show illusive shifts in the rhythms of plant cycles. Through comparative sequencing of ELF3 across different plant populations, climates or even historical samples, we could track shifts in flowering schedules in different conditions and relate gene variations to observable plant traits that respond to climate (e.g., flowering date). This would be interesting to infer how plants are responding to environmental pressures or predicting how future climate conditions may effect flowering cycles.
What technology or technologies would you use to perform sequencing on your DNA and why?
Illumina Sequencing. Next Generation Sequencing (2nd gen).


It has a similar principle to Sanger sequencing which was historically used in combination with positional cloning to first isolate the ELF3 gene. Both methods use fluorescently-labelled nucleotides and identifies each nucleotide by its fluorescent tag. However, while Sanger sequencing handles one DNA fragment at a time, Illumina allows millions of fragments to be sequenced simultaneously in a single run. For my project, I would use Illumina sequencing to read ELF3 DNA because:
- High accuracy for detecting variants, single nucleotide polymorphisms (SNPs) and small insertions/deletions in ELF3 (approx 99.9% short reads).
- Widely used in circadian biology studies and with plant DNA samples
- High throughput which is suitable for single-gene sequencing across many plant samples.
Illumina sequencing works on the principle of sequencing by synthesis (SBS). It identifies DNA bases as they are added to a DNA strand. Each of the four DNA bases is labelled with a unique fluorescent dye, allowing the sequencing system to detect which nucleotide has been added during each cycle. The system captures images of these signals which are then used to determine the exact sequence of the DNA fragment.
Essential Steps:
DNA extraction: The input is genomic DNA isolated from plant tissue in the leaf or seeds. It is important the nucleic acids are of high quality so there will be a quality control check in preparation.
Fragmentation: The DNA is broken down into smaller pieces suitable for Illumina sequencing (typically 200–500 bp) using processes such as mechanical shearing or enzyme digestion.
Adapter ligation: Sequencing adapters are attached to both ends of each fragment to ensure they bind to the sequencing flow cell. They can also be used for barcoding multiple samples.
Library amplification: Bridge PCR is used to bend each DNA strand to form a bridge over a chip. This amplifies the bridge creating clusters at each spot and generates a strong, clear signal.
5. Sequencing by Synthesis: Fluorescently labeled nucleotides are added one by one to the growing DNA strand. Each nucleotide emits a fluorescence as it attaches, specifying the base (base calling) and allowing the sequence to be determined over cycles of synthesis.
6. Analysis: Images collected from each cycle are converted into base sequences by analysing fluorescent signals. The sequences are compared to a reference genome of ELF3 to identify variants.
By sequencing ELF3 in this way, we can generate a detailed map of how plants’ circadian clock genes vary with environmental conditions and over time, providing insights into the elusive rhythms of plants.
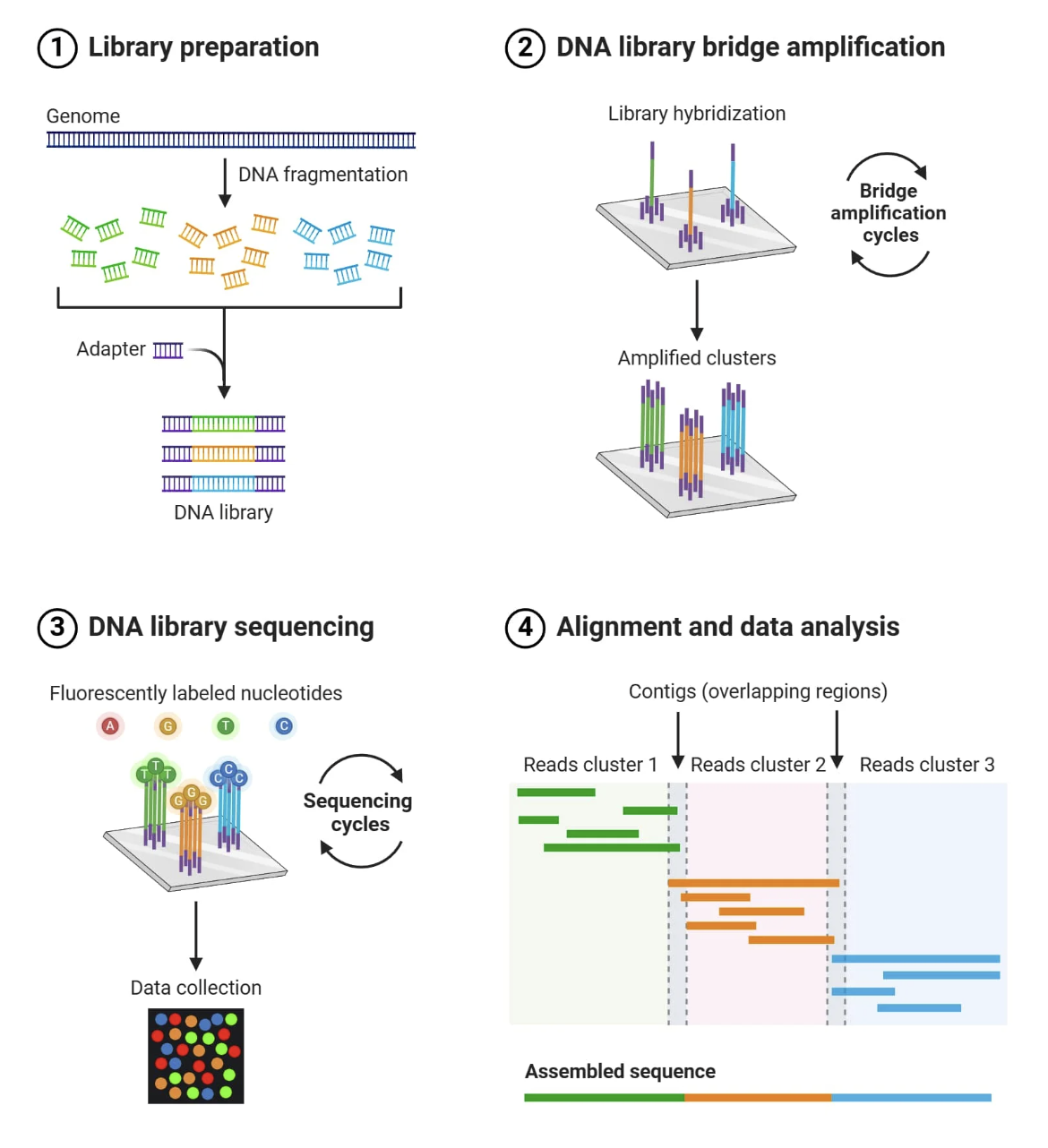
https://microbenotes.com/illumina-sequencing/
5.2 DNA Write – Fibroin for Bio-printing
What DNA would you want to synthesize (e.g., write) and why?
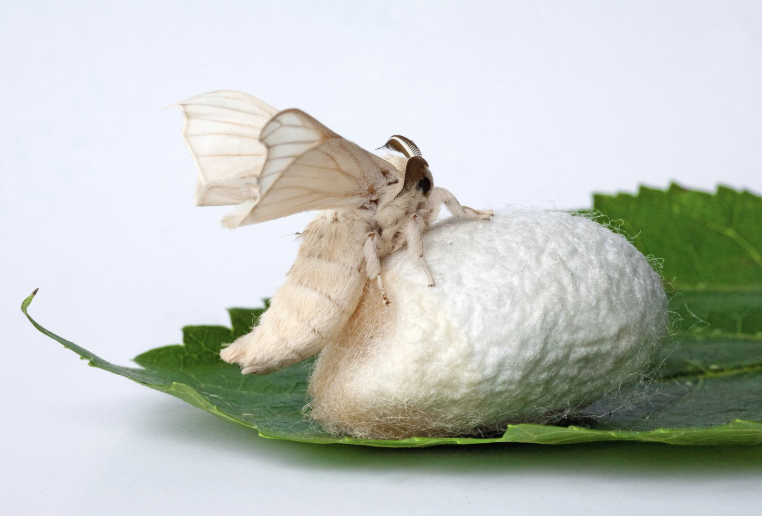
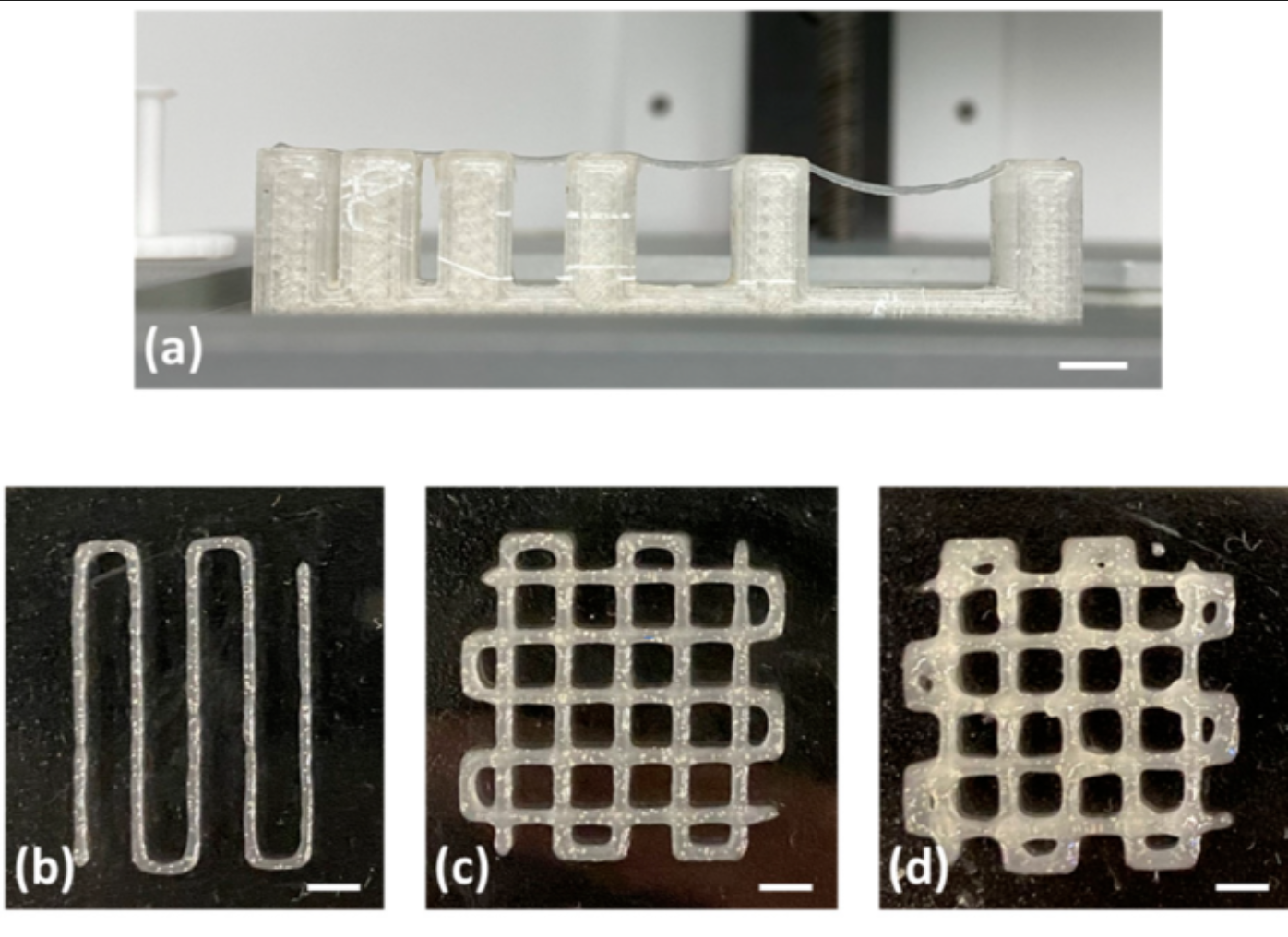


Image left: Fibrinogen-Based Bioink for Application in Skin Equivalent 3D Bioprinting. Image right: silkworm
The DNA I would like to synthesise is the fibroin gene (FibH) from silkworms (Bombyx mori) codon optimised for an Ecoli host. This gene encodes the structural protein fibroin, the main component of silk.
Fibroin has amazing mechanical properties that would be interesting for experimentation for biomaterials and bio-printing to create transparent, complex or bioactive structures. Fibroin is non-toxic and biodegradable, however, unlike its partnering gene Sericin it cannot be extracted from waste product of the silk production industry. By synthesising fibroin DNA, I could produce recombinant fibroin in an Ecoli host for sustainable and cruelty free bio-fabrication.
Some of the qualities of Fibroin that make it interesting are:
High mechanical strength: β-sheet crystalline regions give stiffness and shape retention
Transparency: can form clear films and scaffolds
Compatible with composites: can be mixed with water, polymers, hydrogels, nonparticles.
Biocompatible: widely used in tissue engineering, non toxic, biodegradable
Tuneable properties: could be engineered for responsiveness or added functionality. Recombinant production allows precise control over sequence, length etc. enabling tuneable transparency, stiffness, and degradation rates in printed structures.
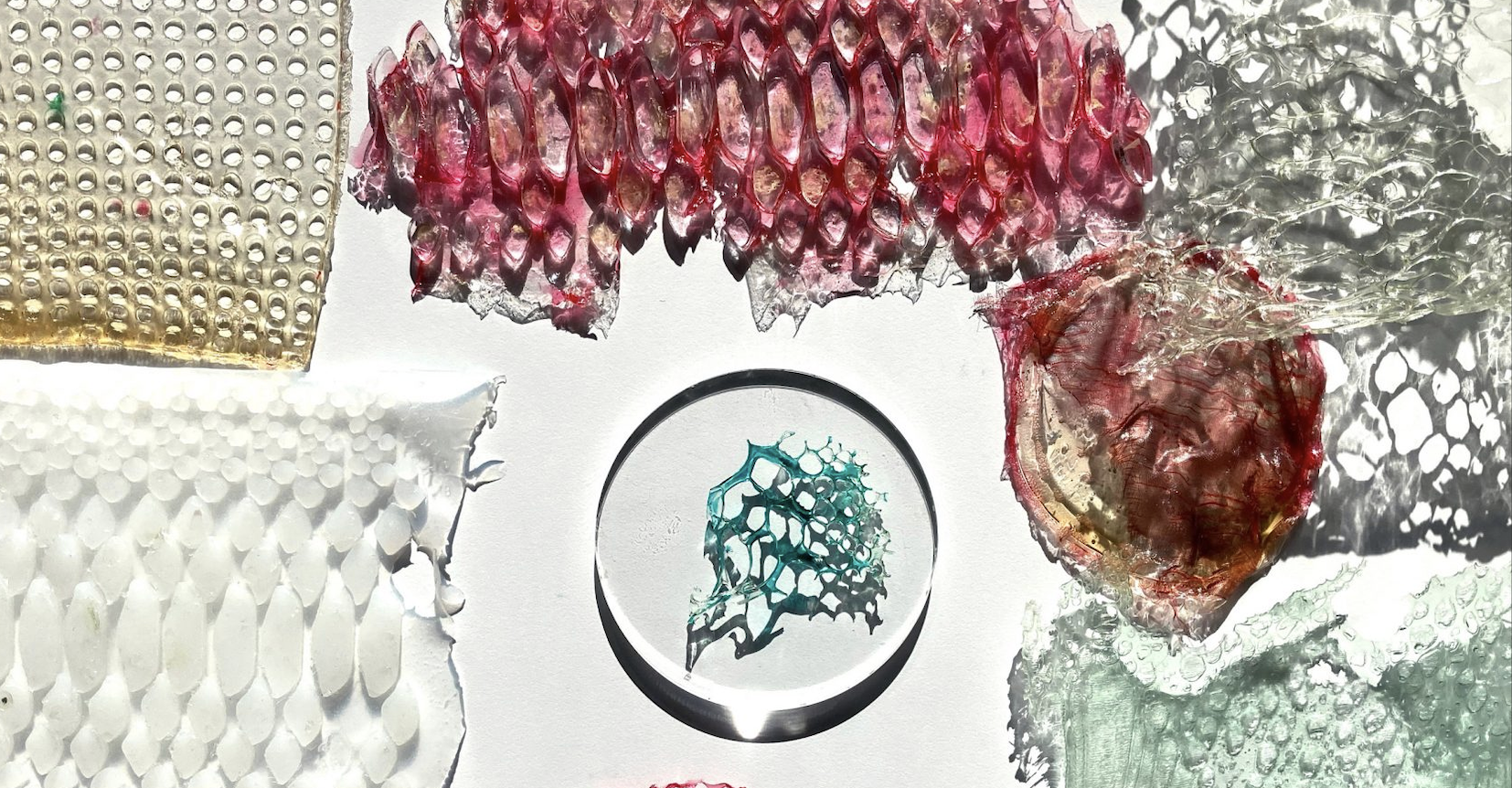


Scarlett Yang Sericin Bioplastics
What technology or technologies would you use to perform this DNA synthesis and why?
I would use oligonucleotide synthesis via Twist Bioscience. This is because unlike past methods of DNA synthesis Twist writes DNA:
Has very high throughput and extremely accurate oligo synthesis. Twist uses silicon-based DNA synthesis which allows them to make 1 million oligos per chip and up to 9,600 genes. This is perfect for assembling long or repetitive genes like FibH.
Its high fidelity reduces errors in the gene sequence to an error rate of 1:5000 bases which allows for longer sequences to be synthesised. FibH is a very long and repetitive gene which is tricky to synthesise accurately. They also handle codon optimization and cloning into plasmids for insertion into ecoli host, saving me a lot of time.
It is a process which requires them to synthesis very little reagent meaning the process is cheap and sustainable.
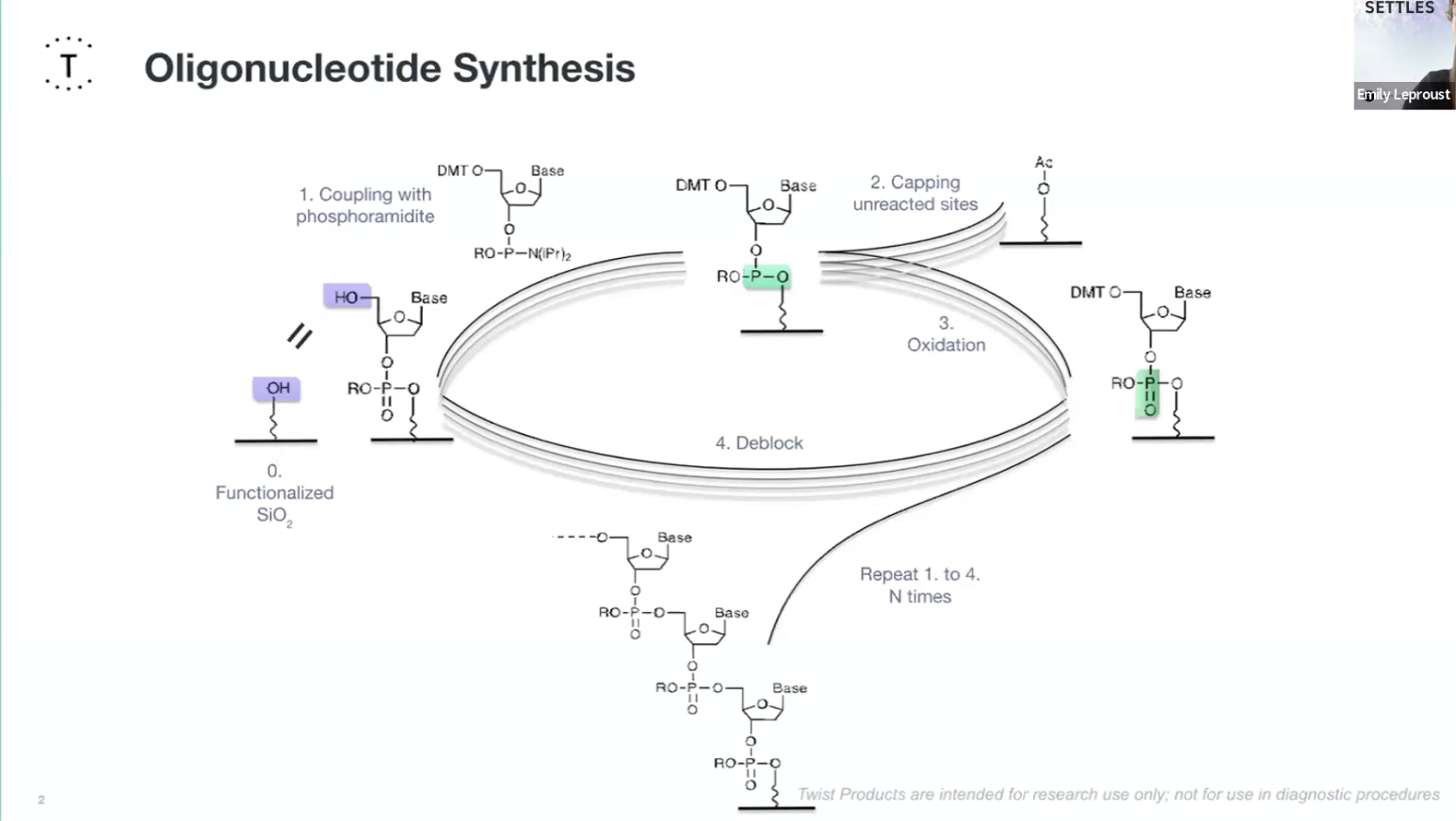

Oligionucleotide Synthesis slide Emily Le Proust
Essential Steps
Design the sequence in Benchling and codon-optimize fibroin DNA for Ecoli’s codon usage bias.
Oligonucleotide synthesis: Twist synthesises hundreds of short DNA fragments (oligos) that come together to form the entire gene sequence.
- Deprotection: The protecting group on the 5′ end is removed from the support-bound nucleoside leaving a reactive 5′-OH group.
- Coupling: This couples to a nucleotide base to form a phosphite triester bond.. This needs to be a very efficient reaction and that is extremely complete.
- Capping: Unreacted 5′-OH groups are capped to ensures that only one base can be added at a time and any unreacted sites are chemically blocked to prevent them causing errors in future cycles. The phosphide is then oxidised into a P5 phosphate creating the natural bone of DNA.
- Oxidisation: The unstable phosphite triester bond is oxidised to form a stable phosphorus phosphodiester bond creating the backbone of DNA.
- Deblocking: finally they deblock the 5" end to regenerate an O-H for the next cycle.
- Post-Synthesis Processing: Twist will undergo cleavage, where the completed oligonucleotide is detached from the solid support, deprotection, where the protecting groups on the base residues are removed and purification of the final product and quality control from mass spectrometry.
Limitations
While Twist can produce many constructs in parallel, very long or highly repetitive sequences are more challenging to synthesize and may need extra optimisation.
5.3 DNA Edit
What DNA would you want to edit and why?
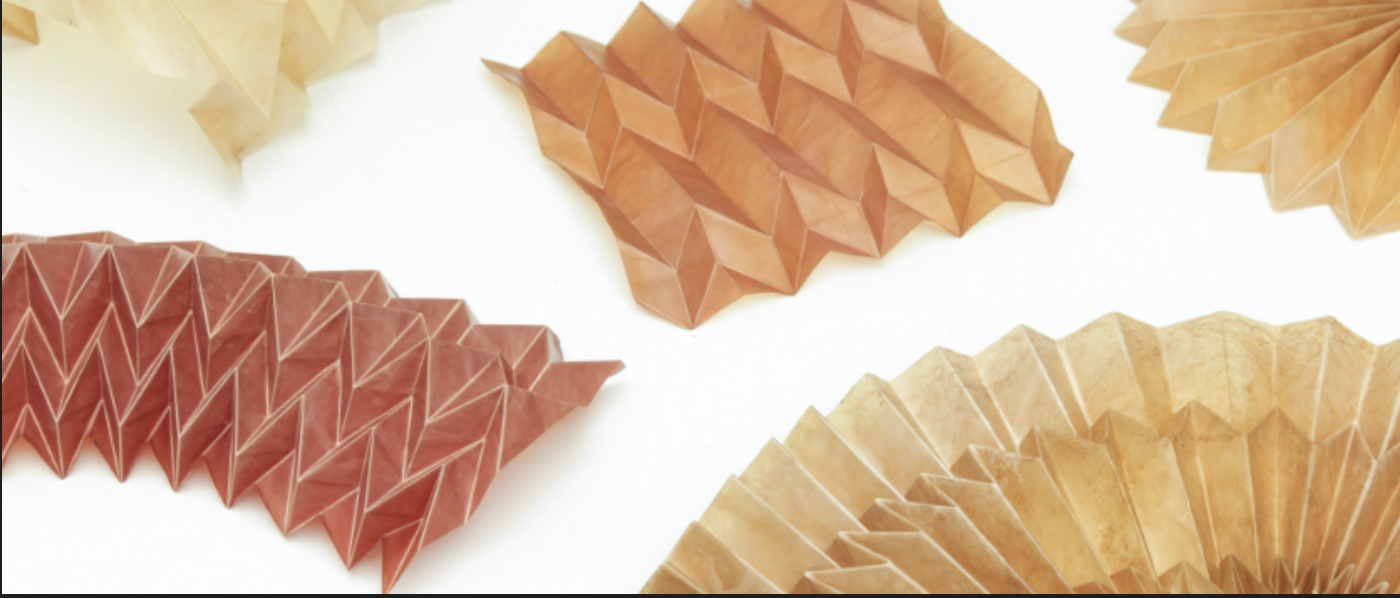


Studio Lionne van Deursen exhibits material created by microorganisms at Milan Design Week
I am very interested in bio and bio-hybrid soft robotics, bio-actuated movement and responsive structures. I am intrigued by the possible applications of genetic editing to design self-actuating and responsive materials and systems.
As an entry point to this idea I have been researching self-folding biomaterials.
So I would like to edit cellulose-producing bacteria Komagataeibacter to create self-folding, pleated bacterial cellulose by genetically programming differences in water retention.
Specifically, I would insert the BslA gene from Bacillus subtilis along with a secretion tag, so that the bacteria secrete BslA protein that binds to cellulose. This protien increases the materials hydrophobicity meaning it will retain less water and shrinks less and becomes more rigid when dry.
Unengineered bacterial cellulose is highly hydrophilic and shrinks significantly during drying and is more pliable.
By layering the two types of cellulose you can create a bilayer structure with a differential shrinkage and controlled folding. By cutting and assembling layers strategically, pleats or origami folds can be programmed. This creates a fully bio-fabricated, biodegradable actuator.
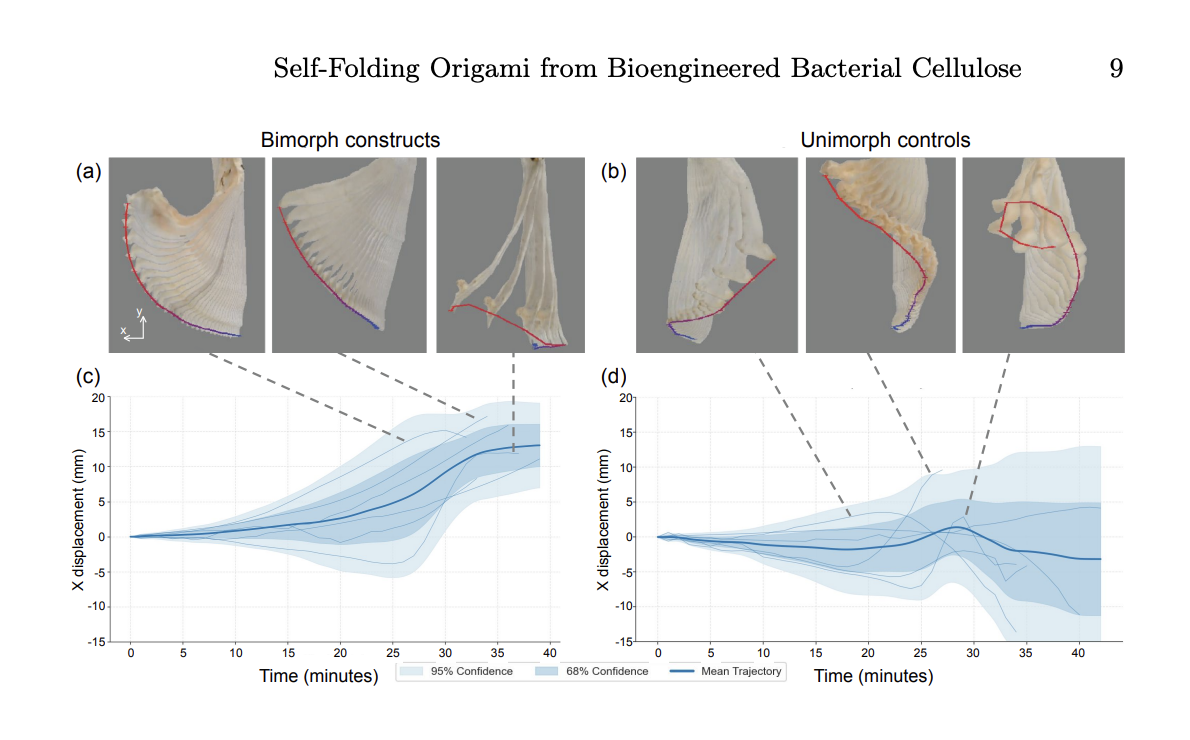

Renewable Self-Folding Origami Constructed from Bioengineered Bacterial Cellulose, Yitong Tseo, Morgan Guempel, Cathy Hogan and Ian Hunter
I think this has interesting applications for design, fashion and sculpture to create a programmable, structured material that is fully degradable. Lots of research has also been done for use in soft robotics for biomedical application. It shows the broader concept of DNA editing to change the genetic instructions of an organism to engineer functional and sustainable materials.
What technology or technologies would you use to perform these DNA edits and why?
Plasmid-based gene expression
I would use a plasmid-based system to introduce the BslA gene into Komagataeibacter. This method allows controlled expression of a non-native protein.
Plasmids are circular DNA molecules that can carry a gene of interest along with regulatory elements, such as promoters, signal peptides and terminators all required for this project. They will replicate independently inside the the bacterial cellulose, providing multiple copies of the gene for strong expression.
This method allows precise control over when and how much BslA is expressed without permanently altering the bacterial genome. It is easier to implement and is ideal for experiments because it allows rapid testing. CRISPR is probably overkill but for long-term, wide-scale production integrating the gene into the genome via CRISPR might be more stable.
The Essential Steps:
Prepare DNA sequence: In Benchling assemble the DNA sequence and design regulatory elements e.g a promotor, a signal peptide to direct secretion of BslA outside the cell and a terminator. Previous studies have used Promoter (PLux) that turns on BslA expression in response to chemical signaling, Signal peptide (N22 tag) which directs the BslA-CBM fusion protein to the CsgG gene for secretion in to the cellulose and the LuxR gene
Copy the DNA (PCR): using PCR take a small amount of the BslA gene template, add primers and run it through a thermal cycler to produce billions of copies.
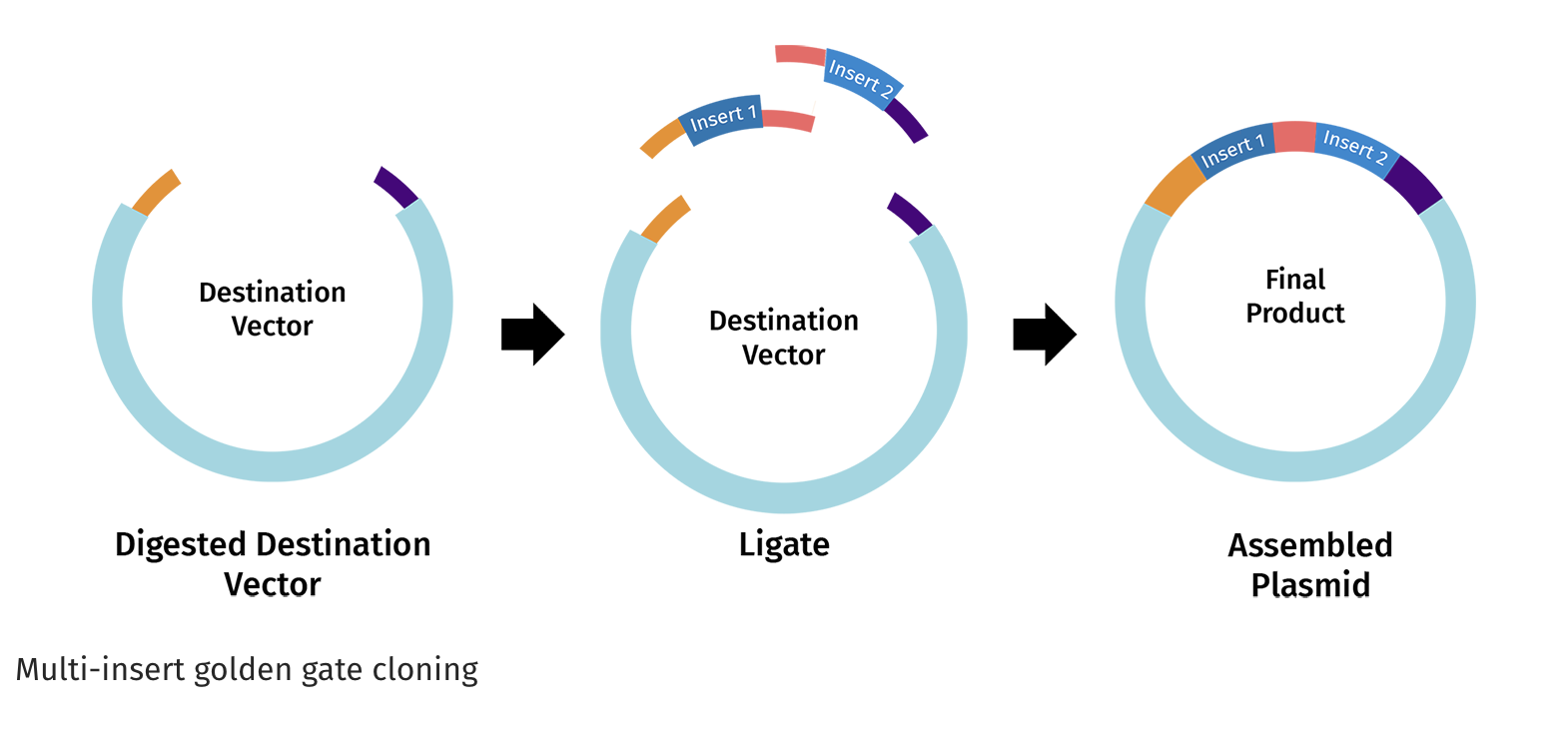

https://www.snapgene.com/guides/golden-gate-assembly
Assemble the plasmid: Use Golden Gate cloning, which employs restriction enzymes to create cuts at precise sites and create unique overhangs (sticky ends) and ligase complimetry sticky ends together to assemble the BslA gene into a plasmid backbone in one reaction with accuracy.
Transformation: Shock the cells via chemical treatment or electroporation to make them competent and deliver the plasmid. Antibiotic resistance genes on the plasmid allow selection of bacteria that have successfully taken up the plasmid.
Expression and secretion: Grow the engineered bacteria in nutrient media, as cellulose is produced, BslA is secreted and binds to the cellulose surface.
The Limitations
The limitations of this process include inconsistency of BsIA production across the bacterial cellulose population as plasmids replicate independently and the number of plasmids per cell can vary.
Stabilityis also an issue as the gene is not permenently integrated into the genome so the plasmids could be lost, for long term or industrial-scale production a genome-integrated systems (e.g. CRISPR knock-in), which maintain stable expression over generations would be better.
References
Zhu, X., & Wang, H. (2024). Revisiting the role and mechanism of ELF3 in circadian clock modulation. Gene, 913, 148378. https://doi.org/10.1016/j.gene.2024.148378
Covington, M. F., Panda, S., Liu, X. L., Strayer, C. A., Wagner, D. R., & Kay, S. A. (2001). ELF3 modulates resetting of the circadian clock in Arabidopsis. Plant Cell, 13(6), 1305–1315. https://doi.org/10.1105/tpc.13.6.1305
Zahn, T., Zhu, Z., Ritoff, N., Krapf, J., Junker, A., Altmann, T., Schmutzer, T., Tüting, C., Kastritis, P. L., Babben, S., Quint, M., Pillen, K., & Maurer, A. (2023). Novel exotic alleles of EARLY FLOWERING 3 determine plant development in barley. Journal of Experimental Botany, 74(12), 3630–3650. https://doi.org/10.1093/jxb/erad127
Richard, L. J., Giordano, V. R., Leite, V. B. P., Wigge, P. A., & Hanson, S. M. (2024). Molecular dynamics simulations illuminate the role of sequence context in the ELF3-PrD-based temperature sensing mechanism in plants. eLife, 13, RP102410. https://doi.org/10.7554/eLife.102410.1
Millar, A. J., Straume, M., Chory, J., Chua, N. H., & Kay, S. A. (1995). Circadian clock mutants in Arabidopsis identified by luciferase imaging. Science, 267, 1161–1163. https://doi.org/10.1126/science.7855595
Hicks, K. A., Albertson, T. M., & Wagner, D. R. (2001). EARLY FLOWERING3 encodes a novel protein that regulates circadian clock function and flowering in Arabidopsis. Plant Cell, 13(6), 1281–1292. https://doi.org/10.1105/tpc.13.6.1281
MicrobeNotes. (n.d.). Illumina sequencing. Retrieved from https://microbenotes.com/illumina-sequencing/
Lee, J., Park, S., Lee, S., Kweon, H. Y., Jo, Y. Y., Kim, J., Chung, J. H., & Seonwoo, H. (2023). Development of silk fibroin-based non-crosslinking thermosensitive bioinks for 3D bioprinting. Polymers (Basel), 15(17), 3567. https://doi.org/10.3390/polym15173567
Trucco, D., Sharma, A., Manferdini, C., Gabusi, E., Petretta, M., Desando, G., Ricotti, L., Chakraborty, J., Ghosh, S., & Lisignoli, G. (2021). Modeling and fabrication of silk fibroin-gelatin-based constructs using extrusion-based three-dimensional bioprinting. ACS Biomaterials Science & Engineering, 7(7), 3306–3320. https://doi.org/10.1021/acsbiomaterials.1c00410
Tseo, Y., Guempel, M., Hogan, C., & Hunter, I. (2025). Renewable self-folding origami constructed from bioengineered bacterial cellulose. bioRxiv. https://doi.org/10.1101/2025.0.0
University of the West of England. (n.d.). Research develops environmentally-friendly e-textiles. Retrieved from https://www.uwe.ac.uk/news/research-develops-environmentally-friendly-e-textiles