Week 3 HW: Lab Automation


Important
FINAL PROJECT IDEAS: 3 Initial Proposals.
Final Project Ideas by Isobel LeonardHomework Part 1: Python Script for Opentrons Artwork
This week we are creating a Python file to run on an Opentrons OT-2 liquid handling robot to create flourescent designs. This is achieved by depositing E.coli genetically engineered to express different fluorescent proteins onto black agar plates. The flouresence will be visible under UV light and so will our designs!
I started first quite ambitious and used The Automation Art Interface to upload a image of Hello Kitty.
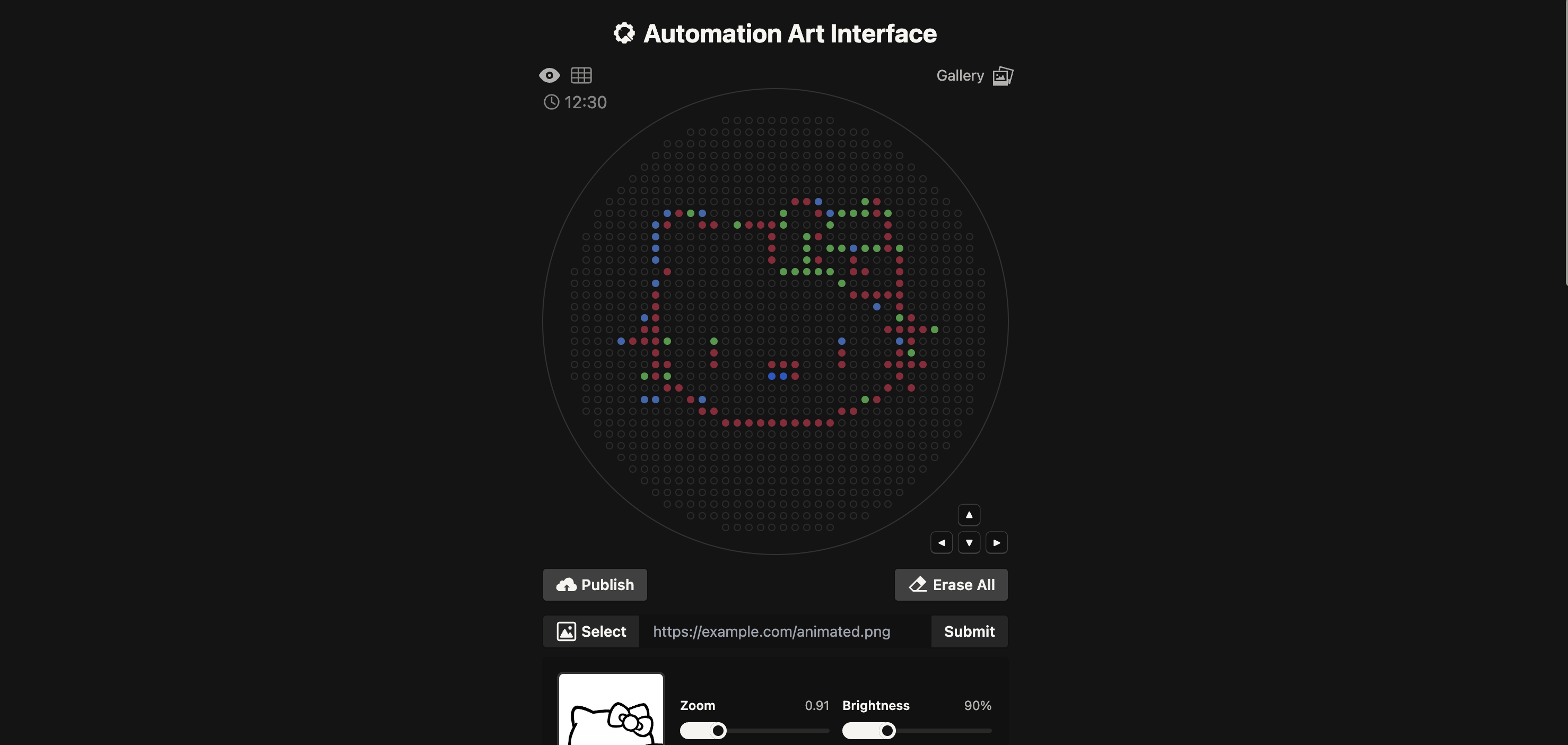

I have no experience programming or coding so this was quite a daunting task and I ended up needing to use Google Gemini to assist. I gave it some of the example code e.g the mathematical heart example and DNA helix as well as the “important notes” highlighted in the Opentrons Collab. Then I gave it the co-ordinates generated in the Automation Art Interface and asked it to assigned each group of co-ordinate to Red, green and orange. I then gave it “Your code” template and asked it to help complete it with my co-ordinates.
CLICK HERE TO EXPAND CODE
from opentrons import types
metadata = { # see https://docs.opentrons.com/v2/tutorial.html#tutorial-metadata
'author': '',
'protocolName': '',
'description': '',
'source': 'HTGAA 2026 Opentrons Lab',
'apiLevel': '2.20'
}
##############################################################################
### Robot deck setup constants - don't change these
##############################################################################
TIP_RACK_DECK_SLOT = 9
COLORS_DECK_SLOT = 6
AGAR_DECK_SLOT = 5
PIPETTE_STARTING_TIP_WELL = 'A1'
well_colors = {
'A1' : 'Red',
'B1' : 'Green',
'C1' : 'Orange'
}
def run(protocol):
##############################################################################
### Load labware, modules and pipettes
##############################################################################
# Tips
tips_20ul = protocol.load_labware('opentrons_96_tiprack_20ul', TIP_RACK_DECK_SLOT, 'Opentrons 20uL Tips')
# Pipettes
pipette_20ul = protocol.load_instrument("p20_single_gen2", "right", [tips_20ul])
# Modules
temperature_module = protocol.load_module('temperature module gen2', COLORS_DECK_SLOT)
# Temperature Module Plate
temperature_plate = temperature_module.load_labware('opentrons_96_aluminumblock_generic_pcr_strip_200ul',
'Cold Plate')
# Choose where to take the colors from
color_plate = temperature_plate
# Agar Plate
agar_plate = protocol.load_labware('htgaa_agar_plate', AGAR_DECK_SLOT, 'Agar Plate') ## TA MUST CALIBRATE EACH PLATE!
# Get the top-center of the plate, make sure the plate was calibrated before running this
center_location = agar_plate['A1'].top()
pipette_20ul.starting_tip = tips_20ul.well(PIPETTE_STARTING_TIP_WELL)
##############################################################################
### Patterning
##############################################################################
###
### Helper functions for this lab
###
# pass this e.g. 'Red' and get back a Location which can be passed to aspirate()
def location_of_color(color_string):
for well,color in well_colors.items():
if color.lower() == color_string.lower():
return color_plate[well]
raise ValueError(f"No well found with color {color_string}")
# For this lab, instead of calling pipette.dispense(1, loc) use this: dispense_and_detach(pipette, 1, loc)
def dispense_and_detach(pipette, volume, location):
"""
Move laterally 5mm above the plate (to avoid smearing a drop); then drop down to the plate,
dispense, move back up 5mm to detach drop, and stay high to be ready for next lateral move.
5mm because a 4uL drop is 2mm diameter; and a 2deg tilt in the agar pour is >3mm difference across a plate.
"""
assert(isinstance(volume, (int, float)))
above_location = location.move(types.Point(z=location.point.z + 5)) # 5mm above
pipette.move_to(above_location) # Go to 5mm above the dispensing location
pipette.dispense(volume, location) # Go straight downwards and dispense
pipette.move_to(above_location) # Go straight up to detach drop and stay high
###
### YOUR CODE HERE to create your design
###
# Coordinate sets for Hello Kitty outline
point_sets = [
('Red', [(3.3, 23.1),(5.5, 23.1),(18.7, 23.1),(-18.7, 20.9),(7.7, 20.9),(18.7, 20.9),(-20.9, 18.7),(-14.3, 18.7),(-12.1, 18.7),(-5.5, 18.7),(-3.3, 18.7),(-1.1, 18.7),(20.9, 18.7),(-1.1, 16.5),(7.7, 16.5),(20.9, 16.5),(-1.1, 14.3),(20.9, 14.3),(-1.1, 12.1),(7.7, 12.1),(14.3, 12.1),(23.1, 12.1),(-20.9, 9.9),(14.3, 9.9),(16.5, 9.9),(23.1, 9.9),(23.1, 7.7),(-23.1, 5.5),(14.3, 5.5),(16.5, 5.5),(18.7, 5.5),(20.9, 5.5),(23.1, 5.5),(-23.1, 3.3),(23.1, 3.3),(-23.1, 1.1),(25.3, 1.1),(-25.3, -1.1),(-23.1, -1.1),(20.9, -1.1),(23.1, -1.1),(25.3, -1.1),(27.5, -1.1),(-27.5, -3.3),(-25.3, -3.3),(-23.1, -3.3),(25.3, -3.3),(-23.1, -5.5),(-12.1, -5.5),(12.1, -5.5),(23.1, -5.5),(-23.1, -7.7),(-20.9, -7.7),(-12.1, -7.7),(-1.1, -7.7),(1.1, -7.7),(3.3, -7.7),(12.1, -7.7),(20.9, -7.7),(23.1, -7.7),(25.3, -7.7),(27.5, -7.7),(-23.1, -9.9),(3.3, -9.9),(23.1, -9.9),(-20.9, -12.1),(-18.7, -12.1),(20.9, -12.1),(25.3, -12.1),(-16.5, -14.3),(18.7, -14.3),(-14.3, -16.5),(-12.1, -16.5),(12.1, -16.5),(14.3, -16.5),(-9.9, -18.7),(-7.7, -18.7),(-5.5, -18.7),(-3.3, -18.7),(-1.1, -18.7),(1.1, -18.7),(3.3, -18.7),(5.5, -18.7),(7.7, -18.7),(9.9, -18.7)]),
('Green', [(7.7, 23.1),(-20.9, 20.9),(-14.3, 20.9),(9.9, 20.9),(-23.1, 18.7),(-23.1, 16.5),(-23.1, 14.3),(14.3, 14.3),(-23.1, 12.1),(-23.1, 7.7),(18.7, 3.3),(-25.3, 1.1),(-29.7, -3.3),(12.1, -3.3),(23.1, -3.3),(-25.3, -14.3),(-23.1, -14.3),(-14.3, -14.3)]),
('Orange', [(16.5, 23.1),(-16.5, 20.9),(1.1, 20.9),(12.1, 20.9),(14.3, 20.9),(16.5, 20.9),(20.9, 20.9),(-7.7, 18.7),(1.1, 18.7),(9.9, 18.7),(5.5, 16.5),(5.5, 14.3),(9.9, 14.3),(12.1, 14.3),(16.5, 14.3),(18.7, 14.3),(23.1, 14.3),(5.5, 12.1),(1.1, 9.9),(3.3, 9.9),(5.5, 9.9),(7.7, 9.9),(9.9, 9.9),(12.1, 7.7),(23.1, 1.1),(29.7, -1.1),(-20.9, -3.3),(-12.1, -3.3),(25.3, -5.5),(-25.3, -9.9),(-20.9, -9.9),(16.5, -14.3)]),
('Red', [(-1.1, -9.9),(1.1, -9.9)])
]
import numpy as np
for color, points in point_sets:
pts = np.array(points)
pts[:,0] -= np.mean(pts[:,0])
pts[:,1] -= np.mean(pts[:,1])
radii = np.sqrt(pts[:,0]**2 + pts[:,1]**2)
scale = 40 / np.max(radii)
pts *= scale
pipette_20ul.pick_up_tip()
cell_well = location_of_color(color)
for i, (x, y) in enumerate(pts):
if i % 20 == 0:
pipette_20ul.aspirate(min(20, len(pts)-i), cell_well)
adjusted_location = center_location.move(types.Point(x, y))
dispense_and_detach(pipette_20ul, 1, adjusted_location)
pipette_20ul.drop_tip()
# Don't forget to end with a drop_tip()
RESULTS:
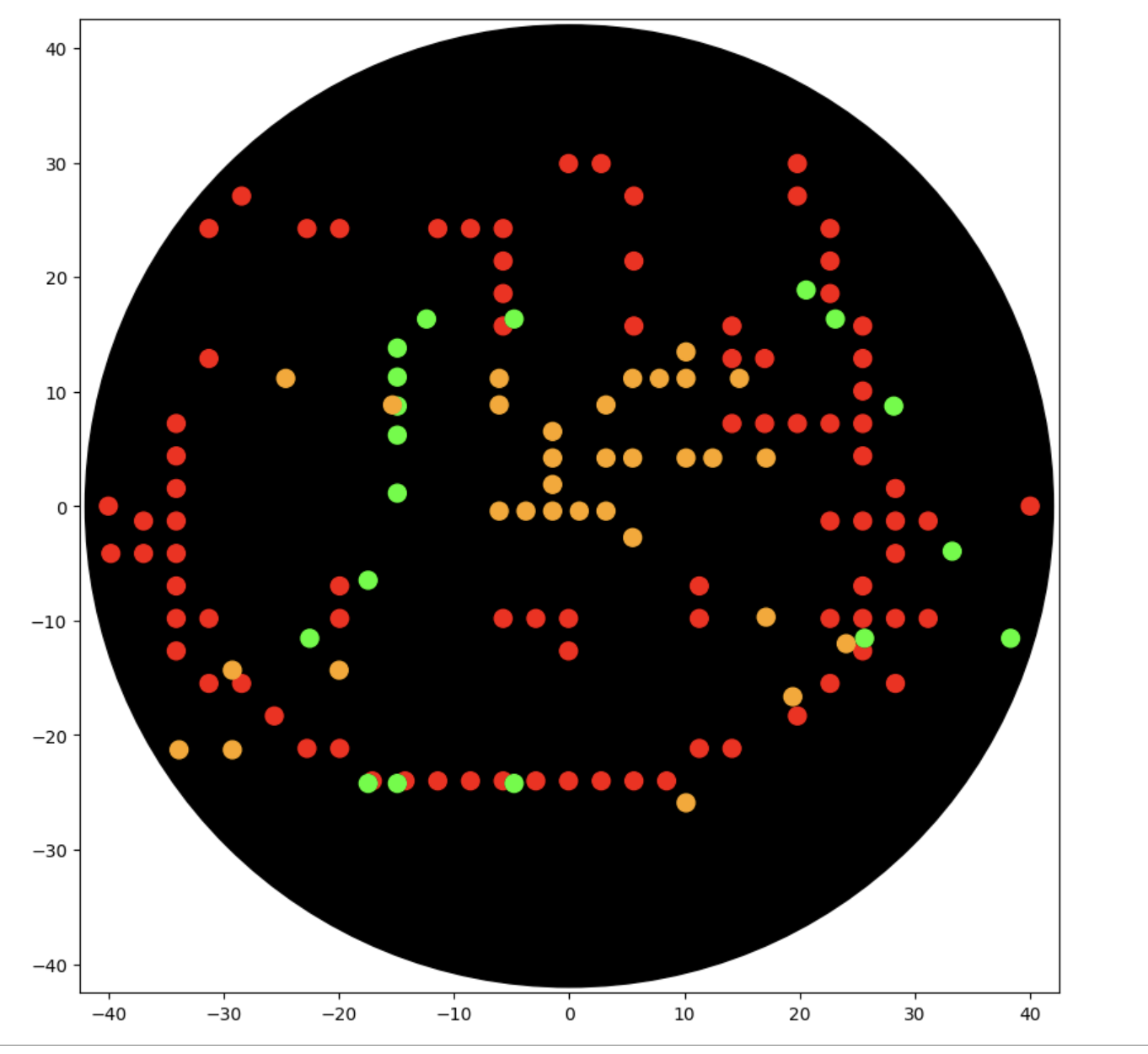
Kitty went quite wonky and I battled for quite some time with Gemini, but I think it is something wrong with the scaling that I didn't understand. I kept getting issues with my indentation, so I decided to pivot and try a simpler design.
PIVOT!
Instead I tried a little lady face: Published Here.
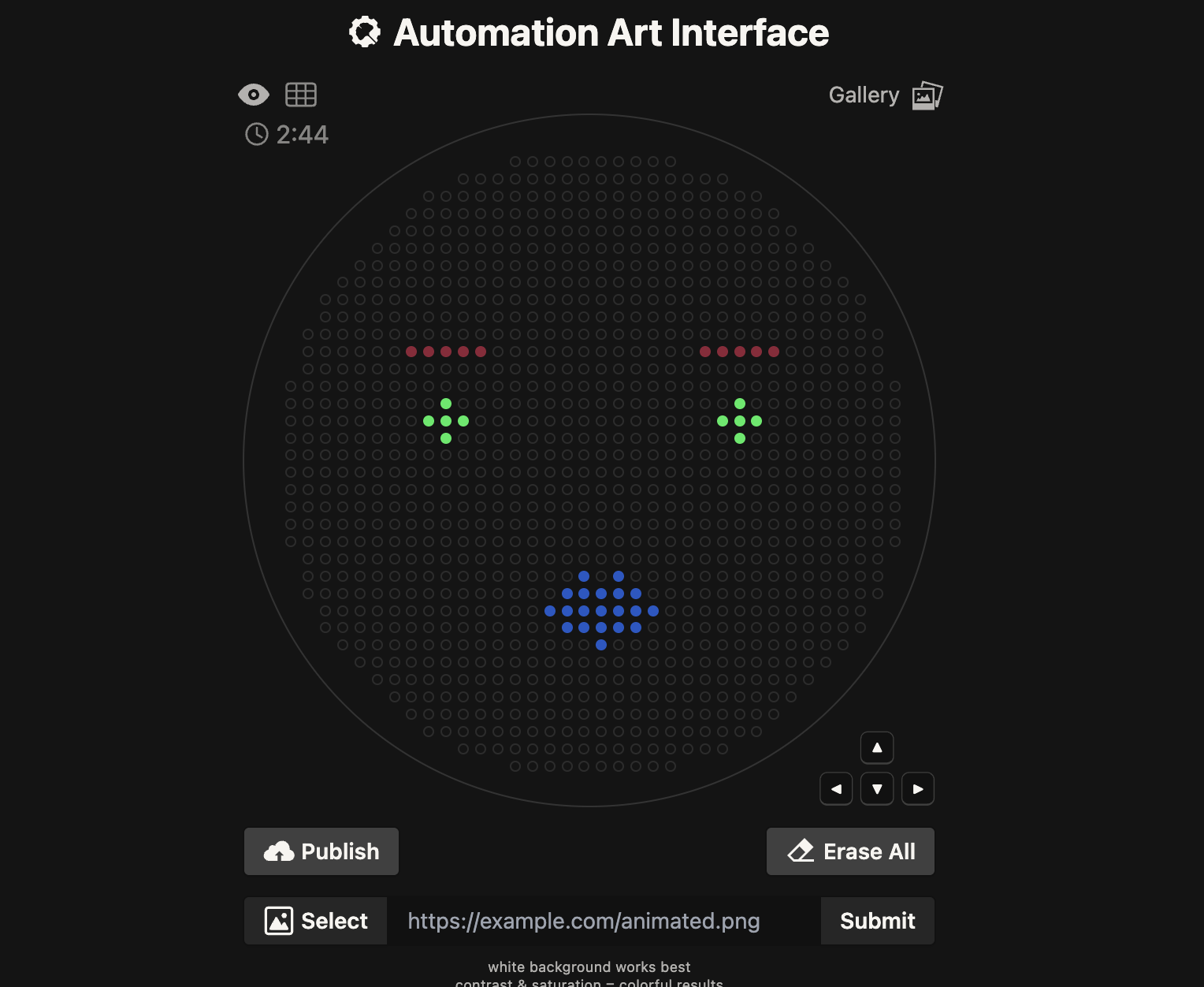
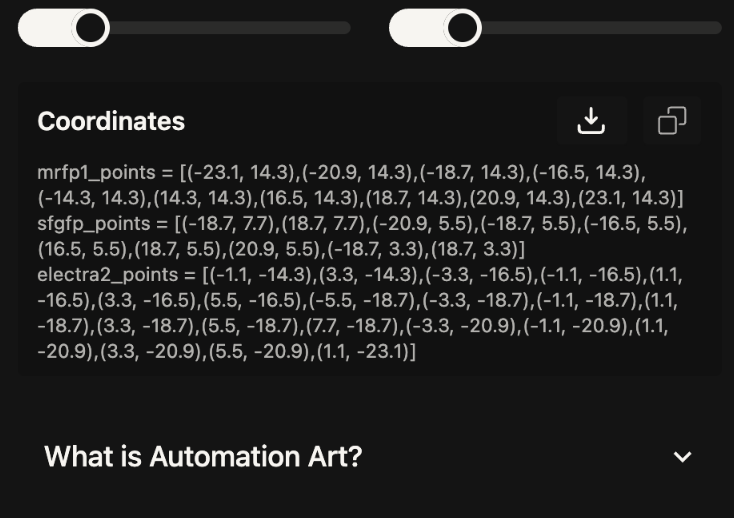
Again, I used Google Gemini in the same way feeding it the example codes and then the new co-ordinates catagorised into each colour- red, green and orange. I initially had an issue that the Opentrons library wasn’t available so I added the top line on the code from troubleshooting with Gemini and it worked.
CLICK HERE TO EXPAND CODE
# 1. Install the library (Required for Google Colab)
!pip install opentrons
from opentrons import types
metadata = {
'author': 'Your Name',
'protocolName': 'Microbial Art Assignment',
'description': 'Simple design with Red, Green, and Orange dots',
'source': 'HTGAA 2026 Opentrons Lab',
'apiLevel': '2.20'
}
##############################################################################
### Robot deck setup constants
##############################################################################
TIP_RACK_DECK_SLOT = 9
COLORS_DECK_SLOT = 6
AGAR_DECK_SLOT = 5
PIPETTE_STARTING_TIP_WELL = 'A1'
well_colors = {
'A1' : 'Red',
'B1' : 'Green',
'C1' : 'Orange'
}
def run(protocol):
# Load labware
tips_20ul = protocol.load_labware('opentrons_96_tiprack_20ul', TIP_RACK_DECK_SLOT, 'Opentrons 20uL Tips')
pipette_20ul = protocol.load_instrument("p20_single_gen2", "right", [tips_20ul])
temperature_module = protocol.load_module('temperature module gen2', COLORS_DECK_SLOT)
temperature_plate = temperature_module.load_labware('opentrons_96_aluminumblock_generic_pcr_strip_200ul', 'Cold Plate')
color_plate = temperature_plate
agar_plate = protocol.load_labware('htgaa_agar_plate', AGAR_DECK_SLOT, 'Agar Plate')
center_location = agar_plate['A1'].top()
pipette_20ul.starting_tip = tips_20ul.well(PIPETTE_STARTING_TIP_WELL)
# Helper functions
def location_of_color(color_string):
for well, color in well_colors.items():
if color.lower() == color_string.lower():
return color_plate[well]
raise ValueError(f"No well found with color {color_string}")
def dispense_and_detach(pipette, volume, location):
above_location = location.move(types.Point(z=5))
pipette.move_to(above_location)
pipette.dispense(volume, location)
pipette.move_to(above_location)
### YOUR DESIGN DATA ###
red_points = [(-23.1, 14.3),(-20.9, 14.3),(-18.7, 14.3),(-16.5, 14.3),(-14.3, 14.3),(14.3, 14.3),(16.5, 14.3),(18.7, 14.3),(20.9, 14.3),(23.1, 14.3)]
green_points = [(-18.7, 7.7),(18.7, 7.7),(-20.9, 5.5),(-18.7, 5.5),(-16.5, 5.5),(16.5, 5.5),(18.7, 5.5),(20.9, 5.5),(-18.7, 3.3),(18.7, 3.3)]
orange_points = [(-1.1, -14.3),(3.3, -14.3),(-3.3, -16.5),(-1.1, -16.5),(1.1, -16.5),(3.3, -16.5),(5.5, -16.5),(-5.5, -18.7),(-3.3, -18.7),(-1.1, -18.7),(1.1, -18.7),(3.3, -18.7),(5.5, -18.7),(7.7, -18.7),(-3.3, -20.9),(-1.1, -20.9),(1.1, -20.9),(3.3, -20.9),(5.5, -20.9),(1.1, -23.1)]
# Drawing Red
if red_points:
pipette_20ul.pick_up_tip()
for i in range(0, len(red_points), 20):
batch = red_points[i:i+20]
pipette_20ul.aspirate(len(batch), location_of_color('Red'))
for x, y in batch:
target = center_location.move(types.Point(x=x, y=y))
dispense_and_detach(pipette_20ul, 1, target)
pipette_20ul.drop_tip()
# Drawing Green
if green_points:
pipette_20ul.pick_up_tip()
for i in range(0, len(green_points), 20):
batch = green_points[i:i+20]
pipette_20ul.aspirate(len(batch), location_of_color('Green'))
for x, y in batch:
target = center_location.move(types.Point(x=x, y=y))
dispense_and_detach(pipette_20ul, 1, target)
pipette_20ul.drop_tip()
# Drawing Orange
if orange_points:
pipette_20ul.pick_up_tip()
for i in range(0, len(orange_points), 20):
batch = orange_points[i:i+20]
pipette_20ul.aspirate(len(batch), location_of_color('Orange'))
for x, y in batch:
target = center_location.move(types.Point(x=x, y=y))
dispense_and_detach(pipette_20ul, 1, target)
pipette_20ul.drop_tip()
Results:
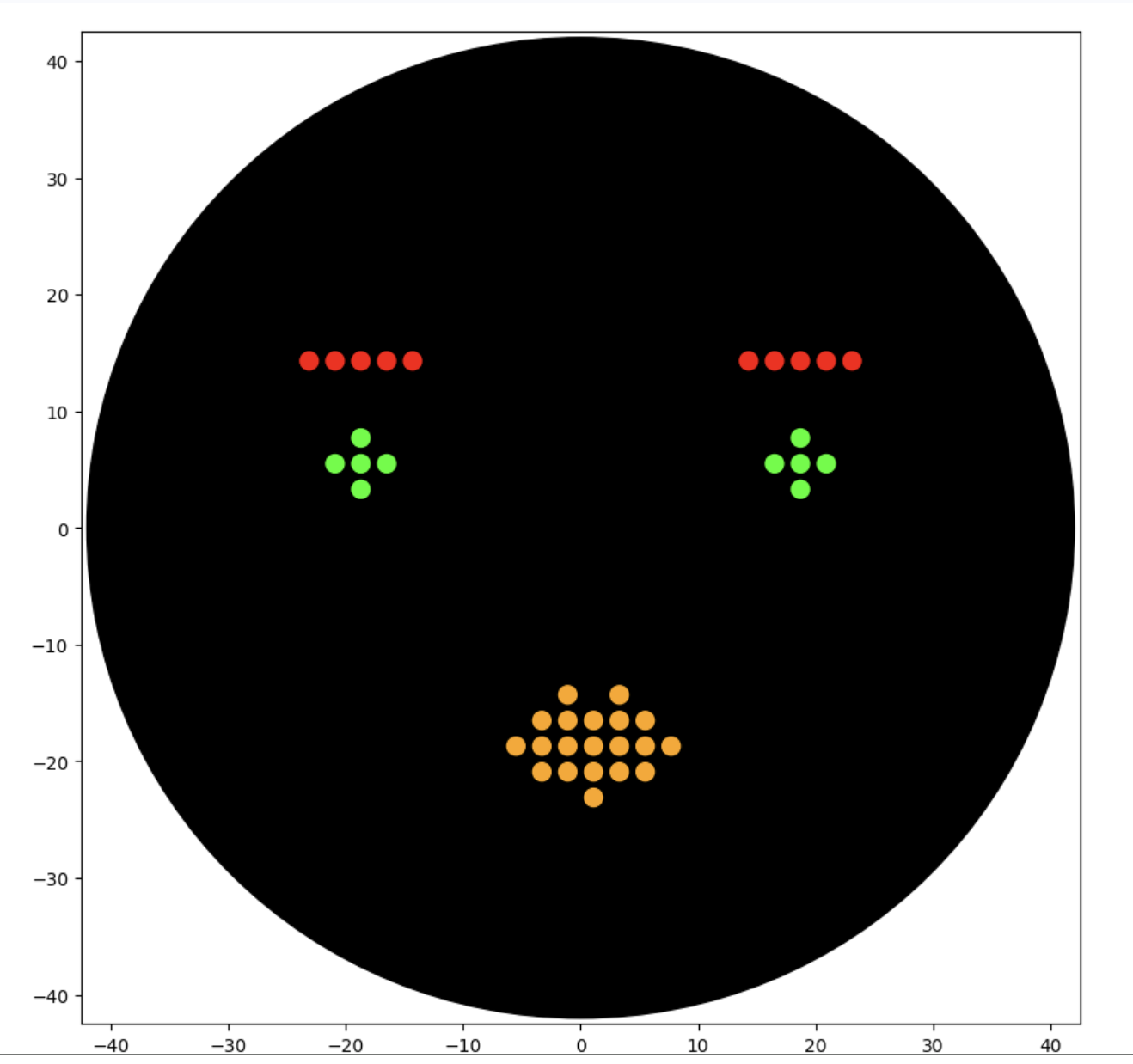
This worked well and although not as cute as the kitty, I am happy with the design.
Preparing for Lifelabs Opentron
Finally, I needed to update my Opentron design to work with the colours we had available at Lifelabs London Node. This was A1: pink, B1 : blue and C1: purple.
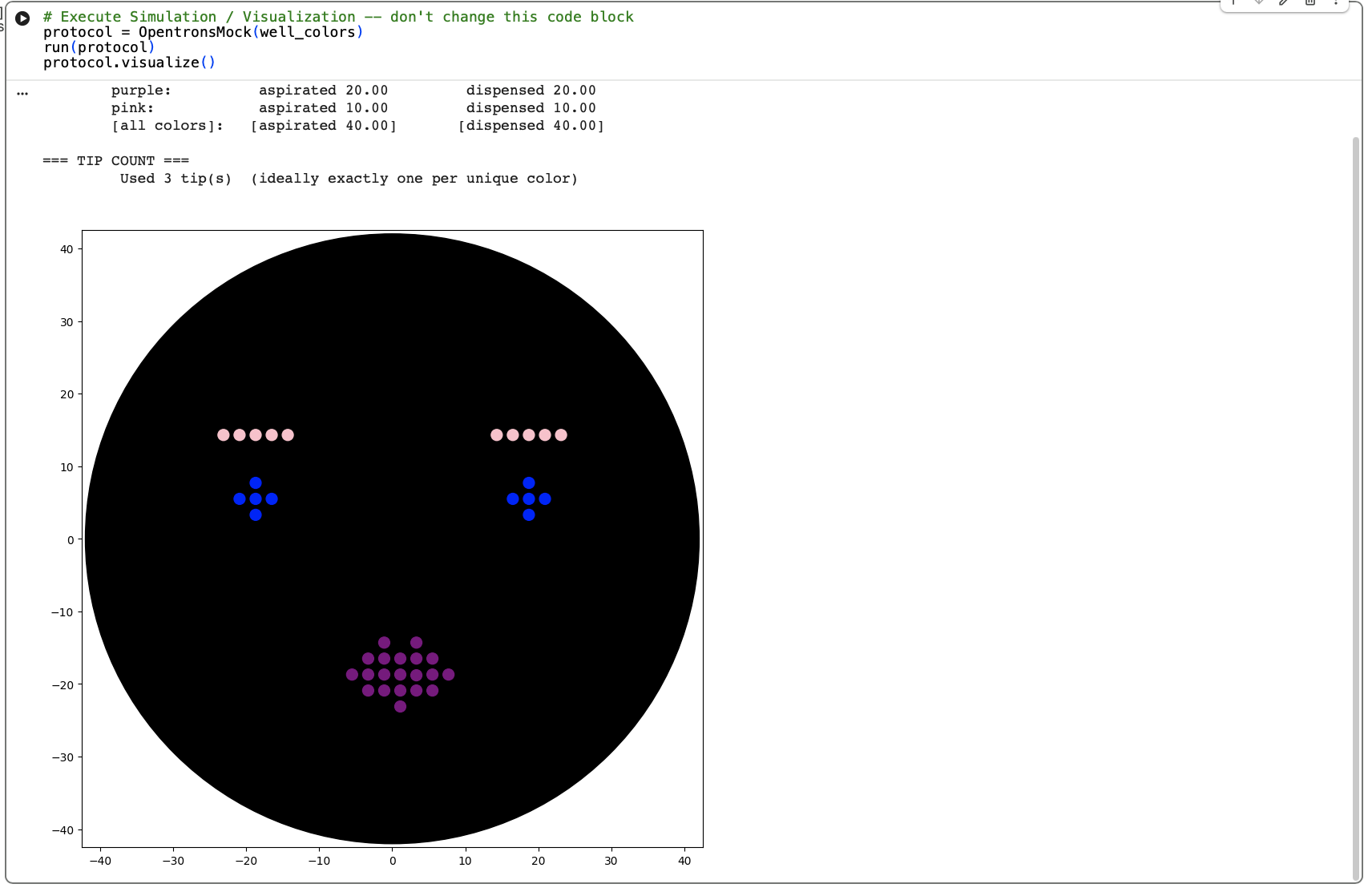

CLICK HERE TO EXPAND CODE
from opentrons import types
metadata = { # see https://docs.opentrons.com/v2/tutorial.html#tutorial-metadata
'author': 'isobel jo leonard',
'protocolName': 'ladyface',
'description': 'opentron protocol',
'source': 'HTGAA 2026 Opentrons Lab',
'apiLevel': '2.20'
}
##############################################################################
### Robot deck setup constants - don't change these
##############################################################################
TIP_RACK_DECK_SLOT = 9
COLORS_DECK_SLOT = 6
AGAR_DECK_SLOT = 5
PIPETTE_STARTING_TIP_WELL = 'A1'
well_colors = {
'A1' : 'pink',
'B1' : 'blue',
'C1' : 'purple'
}
def run(protocol):
##############################################################################
### Load labware, modules and pipettes
##############################################################################
# Tips
tips_20ul = protocol.load_labware('opentrons_96_tiprack_20ul', TIP_RACK_DECK_SLOT, 'Opentrons 20uL Tips')
# Pipettes
pipette_20ul = protocol.load_instrument("p20_single_gen2", "right", [tips_20ul])
# Modules
temperature_module = protocol.load_module('temperature module gen2', COLORS_DECK_SLOT)
# Temperature Module Plate
temperature_plate = temperature_module.load_labware('opentrons_96_aluminumblock_generic_pcr_strip_200ul',
'Cold Plate')
# Choose where to take the colors from
color_plate = temperature_plate
# Agar Plate
agar_plate = protocol.load_labware('htgaa_agar_plate', AGAR_DECK_SLOT, 'Agar Plate') ## TA MUST CALIBRATE EACH PLATE!
# Get the top-center of the plate, make sure the plate was calibrated before running this
center_location = agar_plate['A1'].top()
pipette_20ul.starting_tip = tips_20ul.well(PIPETTE_STARTING_TIP_WELL)
##############################################################################
### Patterning
##############################################################################
###
### Helper functions for this lab
###
# pass this e.g. 'Red' and get back a Location which can be passed to aspirate()
def location_of_color(color_string):
for well,color in well_colors.items():
if color.lower() == color_string.lower():
return color_plate[well]
raise ValueError(f"No well found with color {color_string}")
# For this lab, instead of calling pipette.dispense(1, loc) use this: dispense_and_detach(pipette, 1, loc)
def dispense_and_detach(pipette, volume, location):
"""
Move laterally 5mm above the plate (to avoid smearing a drop); then drop down to the plate,
dispense, move back up 5mm to detach drop, and stay high to be ready for next lateral move.
5mm because a 4uL drop is 2mm diameter; and a 2deg tilt in the agar pour is >3mm difference across a plate.
"""
assert(isinstance(volume, (int, float)))
above_location = location.move(types.Point(z=location.point.z + 5)) # 5mm above
pipette.move_to(above_location) # Go to 5mm above the dispensing location
pipette.dispense(volume, location) # Go straight downwards and dispense
pipette.move_to(above_location) # Go straight up to detach drop and stay high
###
### YOUR CODE HERE to create your design
###
# Define your design points
sfgfp_points = [(-23.1, 14.3),(-20.9, 14.3),(-18.7, 14.3),(-16.5, 14.3),(-14.3, 14.3),(14.3, 14.3),(16.5, 14.3),(18.7, 14.3),(20.9, 14.3),(23.1, 14.3)]
mrfp1_points = [(-18.7, 7.7),(18.7, 7.7),(-20.9, 5.5),(-18.7, 5.5),(-16.5, 5.5),(16.5, 5.5),(18.7, 5.5),(20.9, 5.5),(-18.7, 3.3),(18.7, 3.3)]
electra2_points = [(-1.1, -14.3),(3.3, -14.3),(-3.3, -16.5),(-1.1, -16.5),(1.1, -16.5),(3.3, -16.5),(5.5, -16.5),(-5.5, -18.7),(-3.3, -18.7),(-1.1, -18.7),(1.1, -18.7),(3.3, -18.8),(5.5, -18.7),(7.7, -18.7),(-3.3, -20.9),(-1.1, -20.9),(1.1, -20.9),(3.3, -20.9),(5.5, -20.9),(1.1, -23.1)]
# --- Design for sfgfp_points (Pink) ---
pipette_20ul.pick_up_tip()
color_source_pink = location_of_color('pink')
# Aspirate enough volume for all points (1uL per point)
pipette_20ul.aspirate(len(sfgfp_points), color_source_pink)
for x, y in sfgfp_points:
adjusted_location = center_location.move(types.Point(x, y))
dispense_and_detach(pipette_20ul, 1, adjusted_location)
pipette_20ul.drop_tip()
# --- Design for mrfp1_points (Blue) ---
pipette_20ul.pick_up_tip()
color_source_blue = location_of_color('blue')
pipette_20ul.aspirate(len(mrfp1_points), color_source_blue)
for x, y in mrfp1_points:
adjusted_location = center_location.move(types.Point(x, y))
dispense_and_detach(pipette_20ul, 1, adjusted_location)
pipette_20ul.drop_tip()
# --- Design for electra2_points (Purple) ---
pipette_20ul.pick_up_tip()
color_source_purple = location_of_color('purple')
pipette_20ul.aspirate(len(electra2_points), color_source_purple)
for x, y in electra2_points:
adjusted_location = center_location.move(types.Point(x, y))
dispense_and_detach(pipette_20ul, 1, adjusted_location)
pipette_20ul.drop_tip()
You can access the Google Collab also.
Part 2: Postlab Questions.
Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
They actually use the Opentrons in the research project I discussed in Week 1: Self-dyeing textiles grown from cellulose-producing bacteria with engineered tyrosinase expression by Walker et al (2023).
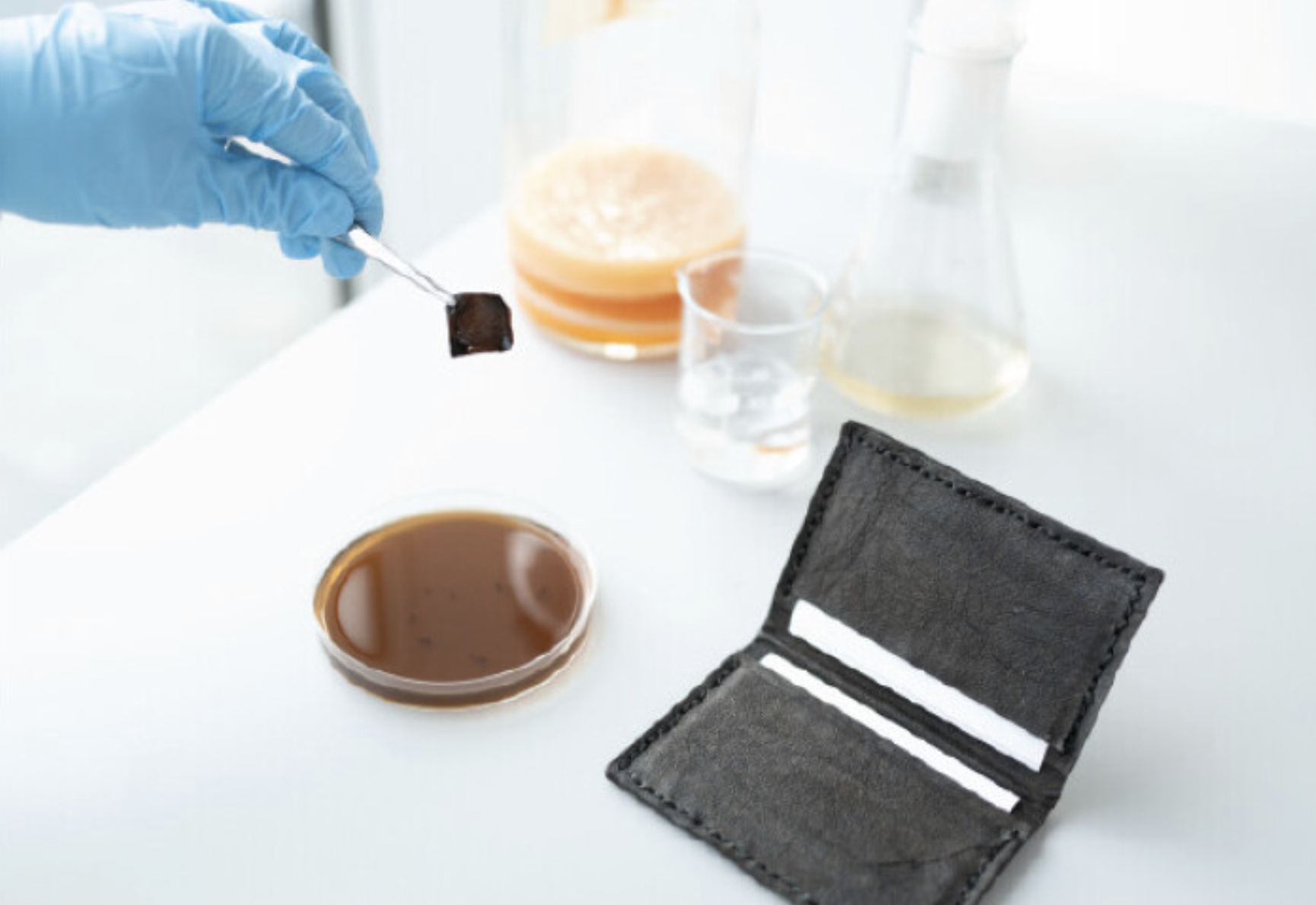

Self-dyeing textiles grown from cellulose-producing bacteria with engineered tyrosinase expression Kenneth T. Walker, Jennifer Keane, Vivianne J. Goosens, Wenzhe Song, Koon-Yang Lee, View ORCID ProfileTom Ellis doi: https://doi.org/10.1101/2023.02.28.530172
In this study, they engineered bacteria Komagataeibacter rhaeticus to produce melanin (a black pigment) in bacterial cellulose, creating a sustainable, self-dyeing textile for the fashion industry.
To measure and compare how much melanin the bacteria make under controlled conditions, they conducted a eumelanin production assay and used an Opentrons OT-2 liquid handling robot to prepare 384-well reaction plates
The robot handled transferring precise amounts of development buffer into the reaction plates using an 8-channel 300 μL OT-2 Gen2 pipette. The reaction plates were kept cold at 4°C using the OT-2 Thermo-module to slow eumelanin production during preparation.
Bacterial cells were mixed in one round of aspiration using the OT-2 Gen2 pipette, then a defined volume was transferred into each well of the 384-well plate.
After centrifugation, the Opentrons Absorbance Plate Reader Module was heated to 45°C to accelerate eumelanin production and prevent potential cell growth from affecting optical density readings. Optical density measurements were then taken over time to assess melanin production.
This setup allowed them to run hundreds of controlled reactions simultaneously and automatically measure pigment formation over time. This is novel because they are using automation to perform an experiment at this scale, which would be very difficult manually. This allows them to generate a large, consistent dataset for analysis.
Write a description about what you intend to do with automation tools for your final project. You may include example pseudocode, Python scripts, 3D printed holders, a plan for how to use Ginkgo Nebula, and more. You may reference this week’s recitation slide deck for lab automation details.
Light-Controlled textile patterning with bacterial pigment production.
One possible direction for my final project is to use light to control where pigment-producing bacteria grow on textiles and other objects, essentially creating a form of “bacterial exposure printing.” I want to explore whether projecting patterns of colored light onto a surface can direct the expression of genes that help bacteria stick and form biofilms in specific areas.
This idea is inspired by research showing that Escherichia coli can be patterned onto materials by controlling curli fiber production with light, which anchors the cells in place. If successful, this system could be further engineered so that the bacteria produce pigment only where they are induced to grow, offering a controlled, reproducible, and sustainable method for creating patterns on fabric.
Automation could be useful in this research project as a way of ensuring bacterial preparation and placement is consistent and reproducible.
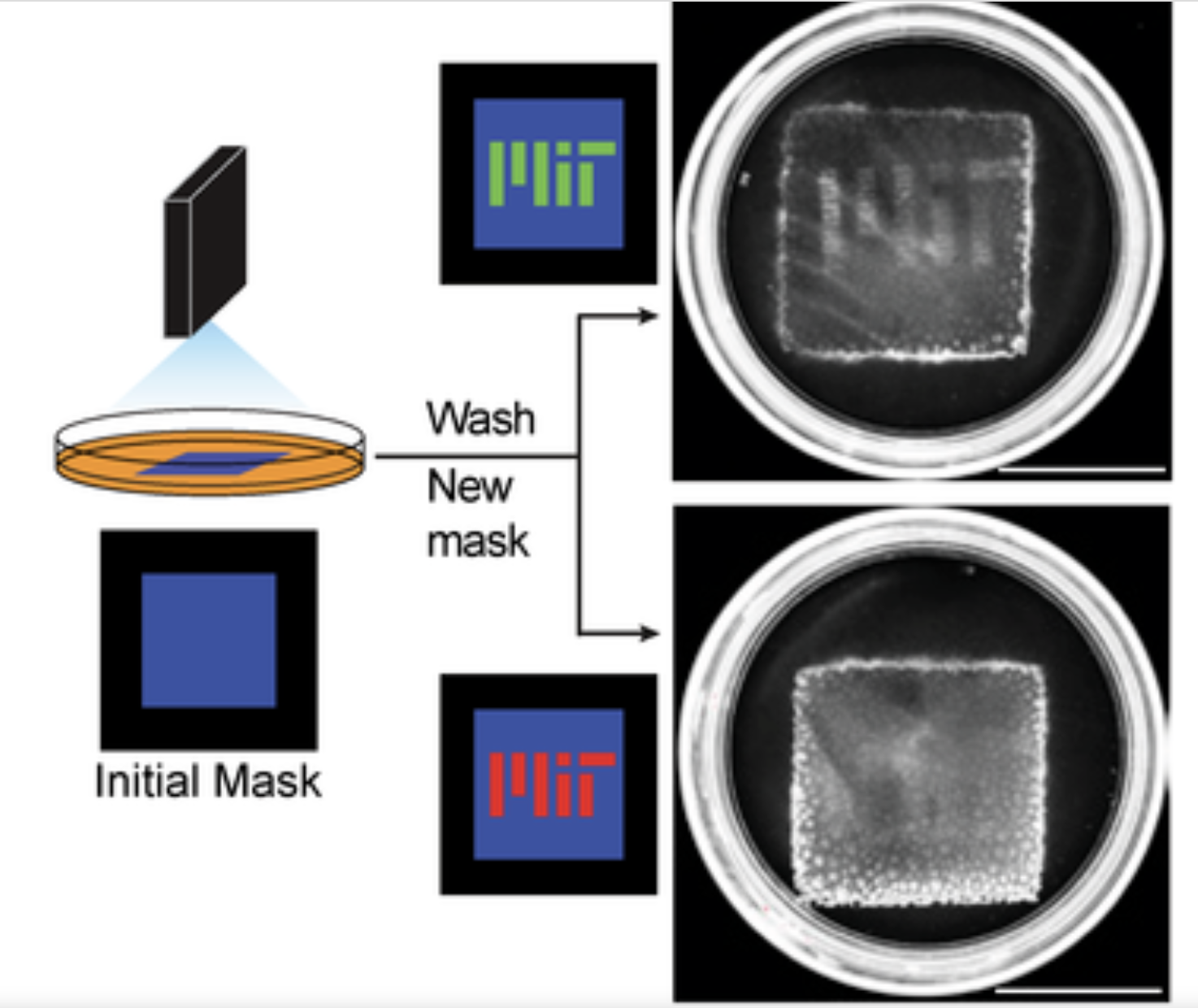
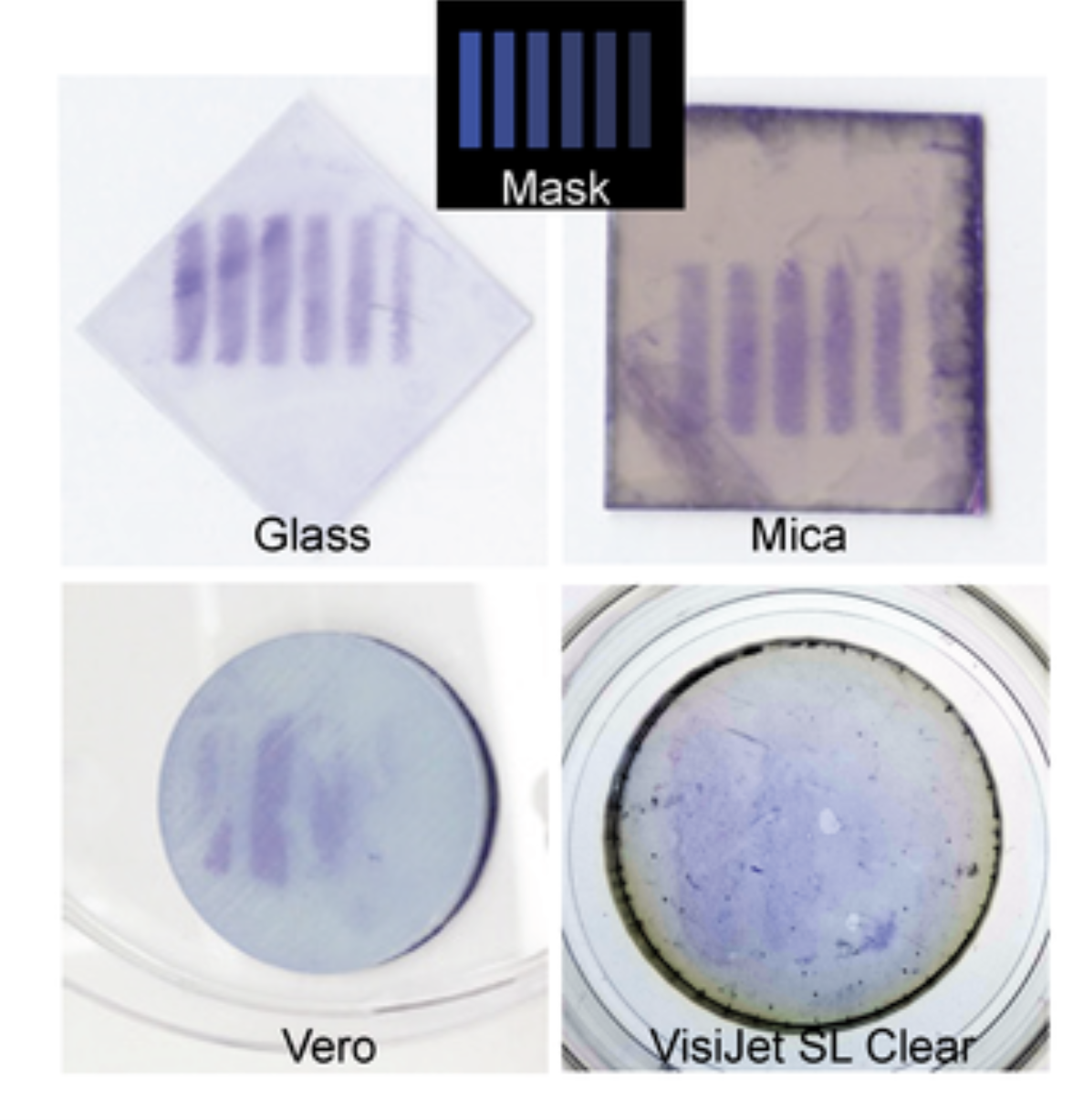
Moser, Felix, Tham, Eléonore, González, Lina M., Lu, Timothy K. and Voigt, Christopher A. 2019. "Light Controlled, High Resolution Patterning of Living Engineered Bacteria Onto Textiles, Ceramics, and Plastic." Advanced Functional Materials, 29 (30).
Automated Preparation
First, the Opentrons OT-2 could be used for automating the culture preparation before pigment patterning begins. The robot could dispense sterile growth media into wells, inoculate bacteria from starter cultures and measure identical volumes to ensure consistency.
It can also perform precise dilutions so each sample has the same concentration of bacteria. This automation reduces variables between pigment producing bacteria, so that it is possible to determine if pigment production differences are due to light patterning rather than inconsistent culture preparation. This being automated also saves time preparing lots of bacteria.
3D Printed Holders:
Textile is not a standard lab plates, so it would be necessary to design custom 3D-printed holders that fit onto the Opentron deck slots and clamp the fabric preventing it from any movement or folding that would effect the placement of the bacteria. This holder would allow the robot to treat non-standard materials like regular labware and create automated precision over an organic process.
Similarly, a custom holder for the light source could ensure that the projected patterns are always at a fixed distance and angle from the surface, providing uniform exposure for precise control over where the bacteria adheres. The exposure, duration or intensity of the light could also be programmed or automated by arduino script and sequenced with the Opentrons protocol.
Precisely Depositing Bacterial
Using the Opentrons robot, I could dispense precise amounts of pigment producing bacterial culture onto specific co-ordinates on the textile. This would allows for a controlled a base pattern. This means that there would be consistency across trials to see whether the bacteria responds to growth in the light induced areas so we could reliably compare samples.
Automated Washing
The Opentron can be used for a standardised washing cycle to gently remove excess bacteria from the textile. This would ensure an identical timescales of bacterial growth and light exposure then the same washing force to be able to reliably compare results and reproduce patterns.
Documentation
The Opentron can also be used to capture standardised images under the controlled lighting and see results of pigment production. It can also be used to analyse the images and compare pigment intensity.
References:
Moser, F., Tham, E., González, L. M., Lu, T. K., & Voigt, C. A. (2019). Light-controlled, high-resolution patterning of living engineered bacteria onto textiles, ceramics, and plastic. Advanced Functional Materials, 29(27), 201901788. https://doi.org/10.1002/adfm.201901788
Walker, K. T., Keane, J., Goosens, V. J., Song, W., Lee, K.-Y., & Ellis, T. (2023). Self‑dyeing textiles grown from cellulose‑producing bacteria with engineered tyrosinase expression [Preprint]. bioRxiv. https://doi.org/10.1101/2023.02.28.530172