Week 9 HW: Cell Free Systems
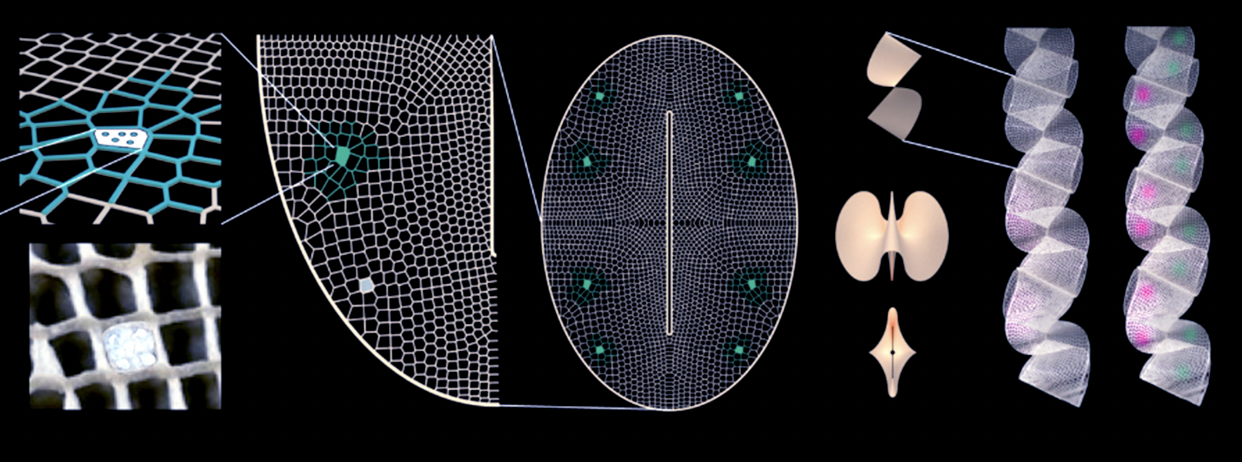


Important
General Homework Questions
Explain the main advantages of cell-free protein synthesis over traditional in vivo methods, specifically in terms of flexibility and control over experimental variables. Name at least two cases where cell-free expression is more beneficial than cell production.
In traditional in vivo methods of protein synthesis, the cell is a “black box” that prioritises its own survival (Adamala, 2026). In CFPS, there is greater flexibility and control due to these advantages:
- Direct Access
In a living cell, the cell membrane acts as a barrier. To change the internal environment, you have to rely on transport proteins or passive diffusion. In CFPS you can directly add or remove components during the reaction e.g change salt concentration, adjust the ph or add a specific enzyme, allowing for a greater tune-ability over experimental variables.
- Elimination of host cell bias
Living cells have their own metabolic priorities (like maintaining membrane integrity, energy conservation) which often conflict with the goal of synthesising a target protein. In CFPS there isn’t the restriction of the host cells bias, 100% of the energy and raw materials are dedicated to protein production.
CASE 1:
The production of proteins that are difficult to express such as toci or membrane proteins acts as a bottleneck to protein yeild in traditional methods. In vivo systems often fail when expressing proteins that interfere with the host’s essential functions such as pore forming toxins or antimicrobial peptides which are designed to puncture bacterial membranes. In a living host, the expression of these molecules leads to rapid membrane depolarisation and cell death (Katzen et al., 2005).
As CFPS operates without living cells, this allows for the synthesis of membrane active proteins (Lyukmanova et al., 2012). Additionally, researchers can aid the cell-free reaction with specific detergents or lipids to assist in the proper folding and stabilisation of complex membrane proteins, a level of environmental tuning that is impossible within the constraints of a living cell (Wuu and Swartz, 2008).
CASE 2:
In addition, the direct access to the cell-free environment allows control over the tRNA and synthetase levels, meaning the system can incorporate modified or unnatural amino acids (e.g. ncAAs such as canavanine), which would otherwise trigger the cell’s stress response or lead to non-functional products (Worst et al. 2015) (Cui et al.2020). This is used to create proteins with new chemical properties. For example, these ncAAs allow for the precise attachment of analysis tags, such as fluorophores for molecular imaging or functionalised probes for investigating protein-small molecule interactions (Parker and Pratt, 2020) (Streit et al. 2025).
- Efficiency
In cell-based methods, every change requires a new transformation and days of cell growth. In CFPS, there is the immediate reaction upon mixing components, you can produce a desired protein in hours rather than days. This high efficiency makes CFPS advantageous for protein engineering, mutagenesis studies and enzyme screening applications.
Describe the main components of a cell-free expression system and explain the role of each component.
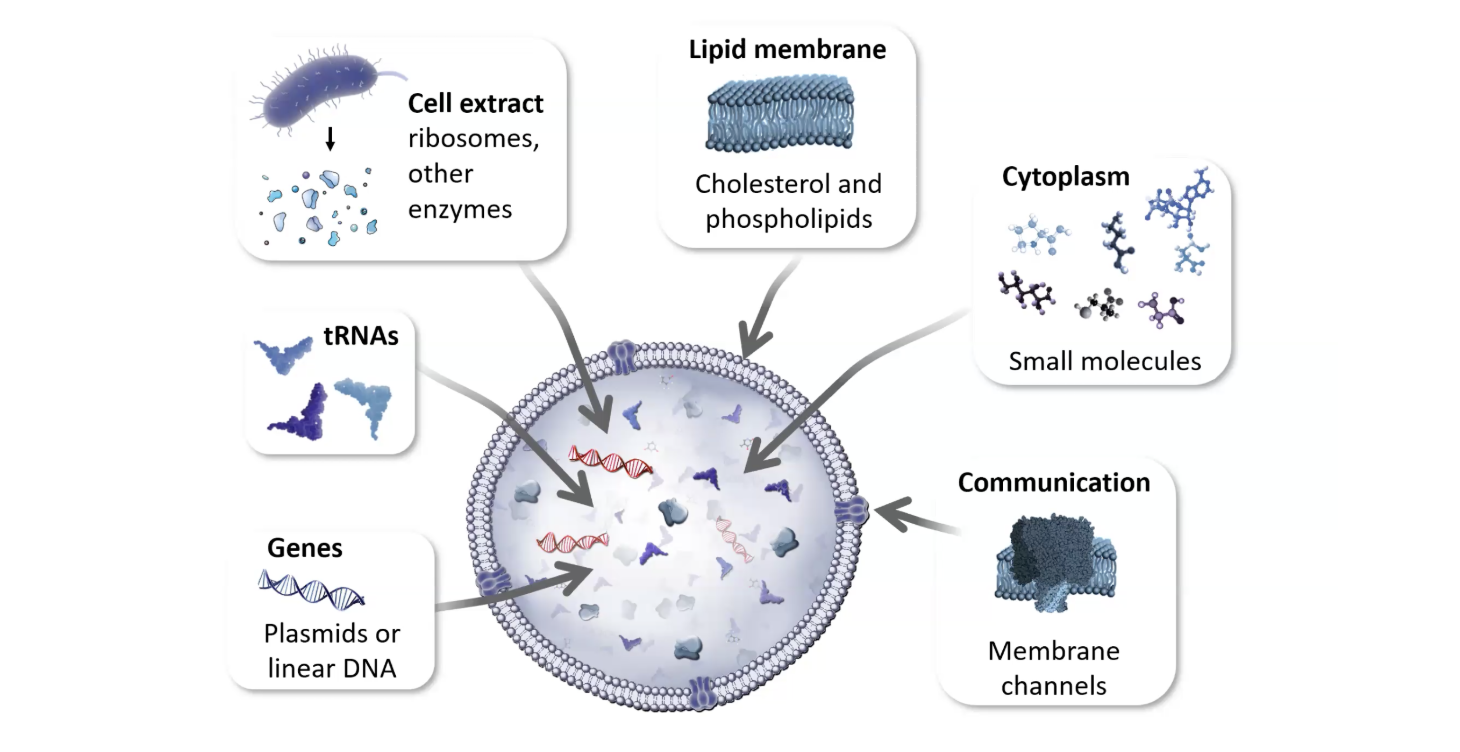

Lecture Slide by Kate Adamala
1. Plasmids or Linear DNA
The genetic template provides the encoded information for protein synthesis. It consists of a coding sequence for the target protein, regulated by a specific promoter and ribosome binding site. These sequences are recognized by the transcriptional and translational machinery to initiate protein production.
2. Cell Extract
The cell extract is a complex mixture of biomolecular machinery harvested from a host organism (e.g., E. coli) including:
RNA Polymerase: Catalyses the transcription of DNA into messenger RNA (mRNA).
Ribosomes: Coordinate the translation of mRNA into polypeptide chains.
Other Enzymes: Include aminoacyl-tRNA synthetases, chaperones for protein folding and metabolic enzymes that facilitate energy regeneration.
3. tRNAs
tRNA is charged with a specific amino acid and utilises its anticodon to recognise the corresponding codon on the mRNA strand, ensuring the accurate primary structure of the protein produced.
4. The cytoplasm: Small Molecules and Reaction Buffer
The reaction environment contains small molecules to sustain the biochemical reaction:
Amino Acids: The raw building blocks required to construct the protein chain.
Energy Source: High-energy molecules (like Phosphoenolpyruvate or Creatine Phosphate) are used to regenerate ATP and GTP, which power the transcription and translation processes.
NTPs (Nucleoside Triphosphates): The building blocks for RNA synthesis.
Salts and Buffers: Magnesium and potassium salts are essential for ribosome stability, while buffers maintain the pH required for enzymatic activity.
5. Lipid Membrane (Cholesterol and Phospholipids)
This turns a chemical reaction into a cell. By encapsulating the extract in a lipid bilayer (liposome), you create a protected micro-environment.
-Phospholipids form the main structure. -Cholesterol is often added to modulate the fluidness and stability of that membrane.
6. Communication (membrane channels)
Selective pores that allow the cell to interact with its environment by letting specific small molecules pass through the lipid barrier.
Why is energy provision regeneration critical in cell-free systems? Describe a method you could use to ensure continuous ATP supply in your cell-free experiment.
Energy regeneration is fundamental to cell free protein synthesis because translation is a very energy consuming process, requiring the hydrolysis of at least four high-energy phosphate bonds per peptide bond formed (two ATP for aminoacyl-tRNA charging; two GTP for ribosomal elongation and translocation). Without energy provision, the initial ATP pool is depleted within minutes, leading to reaction stalling. In addition, the accumulation of inorganic, metabolic byproducts exerts a negative effect, inhibiting enzymatic activity and further compromising protein synthesis efficiency (Ganesh and Maerkl, 2024 )
A approach for ATP regeneration in a cell-free protein synthesis system is to create a secondary energy substrate coupled with an enzymatic regeneration pathway e.g Phosphoenolpyruvate (PEP) as a high-energy phosphate donor and the enzyme Pyruvate Kinase (PK) as the catalyst. As ATP is consumed and produces ADP, the Pyruvate Kinase enzyme facilitates the transfer of a phosphate group from the PEP substrate directly back to the ADP. This cycle maintains a high ATP:ADP ratio, preventing the reaction from stalling due to energy depletion or the accumulation of byproducts (Kim and Swartz, 2001).
Compare prokaryotic versus eukaryotic cell-free expression systems. Choose a protein to produce in each system and explain why.
The essential dichotomy comes down to high yield prokaryotic systems (E.coli) vs the high fidelity eukaryotic systems (Wheat Germ, Rabbit Reticulocyte). Prokaryotic systems are cost-effective factory for simple, soluble protein manufacturing, allowing for high concentration yields and continuous-flow reactions. They also allow for efficient incorporation of non-natural amino acids, due to easy depletion of natural amino acids in the lysate.
However, they lacks the machinery (PDI, chaperones, microsomal membranes) for correct folding and processing of complex eukaryotic proteins, often resulting in inclusion bodies or inactive constructs. Conversely, eukaryotic systems ensure functional correctness of complex proteins through the integrated post-translational machinery in mammalian systems, including ribosomes, tRNA pools and PTM enzymes. However, their yield constraints and high lysate cost create an accessibility issue. (CD Biosynsis )
Protein Choice: GFP
For Prokaryotic systems: GFP is a small, structurally robust protein that does not require complex post-translational modifications or disulfide bonds to function and florese. E. coli lysates offer superior speed, high yield and lower cost to produce this simple reporter protein(Zemella et al. 2015).
Protein Choice: scFv Antibody Fragments e.g anti-HER2 scFv
scFv antibody fragments e.g anti-HER2 scFv fragments are better suited to eukaryotic cell-free systems such as Sf21 or rabbit reticulocyte lysates because they require correct disulphide bond formation and assisted folding to remain functional. Unlike E. coli lysates, eukaryotic lysates contain microsomal vesicles, chaperones and oxidative folding environments that support antibody maturation.
Stech et al. (2014) demonstrated that targeting scFv constructs into microsomal vesicles using a signal peptide significantly improved antigen-binding activity compared with proteins expressed in open lysate conditions. This shows that eukaryotic CFPS systems are preferable for producing structurally complex, disulphide-bonded proteins requiring post-translational processing, even though they produce lower yields than prokaryotic systems.
How would you design a cell-free experiment to optimise the expression of a membrane protein? Discuss the challenges and how you would address them in your setup.
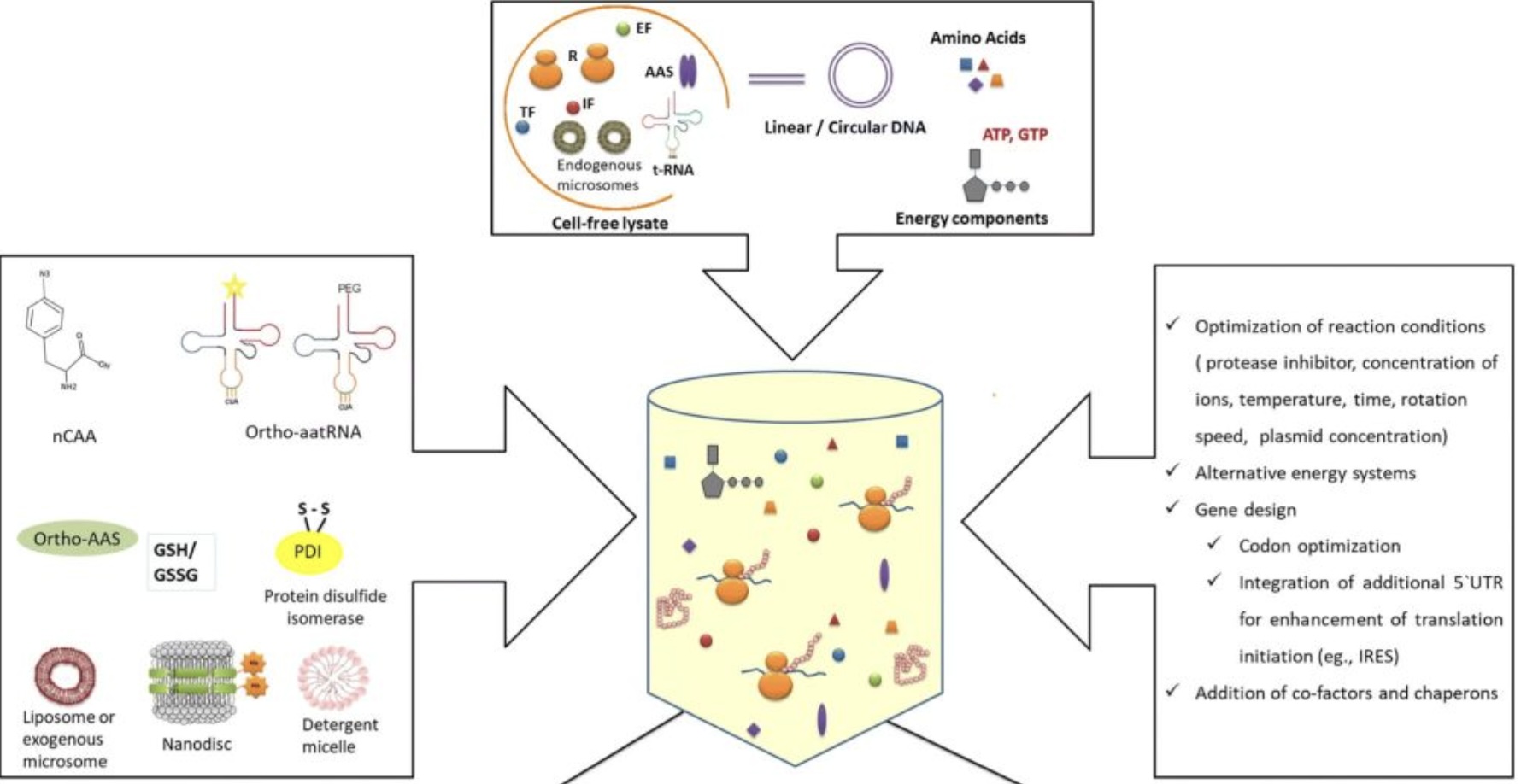

The overall process of cell-free protein synthesis. (Dondapati S K, et al. 2020)
1. System choice
I would choose a eukaryotic lysate, specifically Sf21 insect cell extract, as it naturally contains microsomes housing the Sec61 translocon, a protein channel that physically threads the membrane protein into the lipid bilayer, ensuring correct orientation and folding (Itskanov and Park, 2023).
2. Optimisation
I would optimise the DNA sequence through codon optimisation matched to the host lysate’s tRNA availability to prevent ribosomal stalling. I would also attach a short N-terminal tag to act as a ribosome handle, reducing mRNA secondary structures and accelerating translation initiation (Lyukmanova et al., 2012; Steinkühler et al., 2024). A C-terminal sfGFP tag would also be included as a folding reporter, in-gel fluorescence after SDS-PAGE immediately distinguishes folded from aggregated protein without requiring a Western blot for every condition tested (Drew et al. 2008)
3. Lipid environment
Since membrane proteins cannot fold in water, a lipid environment must be provided to prevent aggregation upon synthesis (Carpenter et al. 2008). I would use GOA-based lipid sponge droplets to provide a massive 3D hydrophobic surface area, which is particularly effective for achieving high yields with small membrane proteins (Jiang et al., 2024). Alongside this, MSP-nanodiscs would be included for detergent-free stabilisation, maintaining the protein in a native-like lipid bilayer patch, the best option for any downstream functional or structural work (Denisov and Sligar, 2017).
4. Reaction conditions
To maximise yield I would use a CECF (continuous exchange cell-free) format, where a dialysis membrane continuously replenishes NTPs and amino acids while removing inhibitory byproducts. I would incubate at 30°C with high-speed shaking, as static conditions can drop yields due to the heavy lipid components settling at the bottom of the tube. I would also include PEG 8000 as a molecular crowding agent to mimic the dense intracellular environment, promoting correct folding over aggregation (Schwarz et al. 2007)
5. Folding validation
Screening using the green fluorescent protein fusion strategy.
Challenges and how they were addressed:
Aggregation: Membrane proteins have hydrophobic regions that repel water, without a lipid environment they will clump together and precipitate out of solution. This is addressed by incorperating the lipid sponge droplets and MSP nanodiscs.
Low yield: Membrane proteins express poorly in cell free systems. This is addressed by the CECF format to ensure the supply of energy and raw materials and removal of byproduct and optimal reaction conditions maintained with the high speed shaking and PEG 8000.
Misfolding: Without the correct machinery, membrane proteins won’t fold correctly. This is addressed by the Sec61 translocon, codon optimisation and addition of ribosome handle that have been studied measures to improve membrane protein folding.
Validating folding: The expression of the protein doesn’t guarantee functionality, so the inclusion of the sfGFP reporter allows for screening of correct protein folding.
Imagine you observe a low yield of your target protein in a cell-free system. Describe three possible reasons for this and suggest a troubleshooting strategy for each.
1. Ribosomal stalling due to rare codons.
If the DNA sequence contains codons that are rarely used by the host lysate, the ribosome stalls or drops off entirely, reducing the amount of full-length protein produced. This is particularly common when expressing mammalian or human proteins in insect cell extracts.
Troubleshoot:
Lower the temperature of the reaction to 30C (or even lower). This will slow translation down, allowing time for the correct tRNA to be recruited. Re-synthesise the gene with codon optimisation matched to the host lysate’s tRNA pool and to eliminate any rare codons and replace them with more common ones (Shoba, 2025).
2. Depletion of energy substrates
In a standard batch reaction, NTPs and amino acids are consumed and inhibitory byproducts accumulate, shutting down transcription and translation long before the protein reaches useful yields.
Troubleshoot:
Switch to CECF format, where a dialysis membrane continuously replenishes substrates and removes byproducts, significantly extending the reaction and improving yields (Zemella et al. 2015).
3. Protein misfolding
If membrane proteins misfold during synthesis, it will be targeted for degradation by proteases present in the lysate, reducing the yield. Some proteins require post-translational modification, chaperones, or even disulfide bridges to fold properly.
Troubleshoot:
Introduce chaperones to assist co-translational folding. Ensure an adequate lipid environment is present via nanodiscs or lipid sponge droplets. Use the sfGFP folding reporter to quickly identify which conditions improve the ratio of folded to misfolded protein. For proteins that require post-translational modifications, use rabbit reticulocyte system (with canine microsomal membranes) or wheat germ extract would be a better choice. (Shoba, 2025)
REFERENCES:
Worst E. G., Exner M. P., De Simone A., Schenkelberger M., Noireaux V., Budisa N., et al. (2015). Cell-free expression with the toxic amino acid canavanine. Bioorg. Med. Chem. Lett. 25.
Khambhati K, Bhattacharjee G, Gohil N, Braddick D, Kulkarni V, Singh V. Exploring the Potential of Cell-Free Protein Synthesis for Extending the Abilities of Biological Systems. Front Bioeng Biotechnol. 2019 Oct 11;7:248. doi: 10.3389/fbioe.2019.00248. PMID: 31681738; PMCID: PMC6797904.
Cui, Z., Johnston, W. A., & Alexandrov, K. (2020). Cell-free approach for non-canonical amino acids incorporation into polypeptides. Frontiers in Bioengineering and Biotechnology, 8.
Parker, C. G., & Pratt, M. R. (2020). Click chemistry in proteomic investigations. Cell, 180(4), 605–632.
Streit, M., Budiarta, M., Jungblut, M., & Beliu, G. (2025). Fluorescent labeling strategies for molecular bioimaging. Biophysical Reports, 5, 100200.
Katzen, F., Chang, G., & Kudlicki, W. (2005). The past, present and future of cell-free protein synthesis. Trends in Biotechnology, 23(3), 150-156.
Lyukmanova, E. N., et al. (2012). Lipid–protein nanodiscs for cell-free production of integral membrane proteins. Methods in Enzymology, 506, 255-276.
Wuu, J. J., & Swartz, J. R. (2008). High yield cell-free production of integral membrane proteins without surfactants. Biochimica et Biophysica Acta (BBA)-Biomembranes, 1778(10), 2350-2361.
Kim, D. M., & Swartz, J. R. (2001). Regeneration of adenosine triphosphate from glycolytic intermediates for cell-free protein synthesis. Biotechnology and Bioengineering, 74(4), 309–316.
Ganesh, R. B., & Maerkl, S. J. (2024). Towards self-regeneration: Exploring the limits of protein synthesis in the protein synthesis using recombinant elements (PURE) cell-free transcription–translation system. ACS Synthetic Biology, 13(8), 2555–2566.
CD Biosynsis. (n.d.). A comparative guide: Prokaryotic vs. eukaryotic cell-free expression systems for eukaryotic proteins. Retrieved April 5, 2026, from https://www.biosynsis.com/a-comparative-guide-prokaryotic-vs-eukaryotic-cell-free-expression-systems-for-eukaryotic-proteins.html
Zemella, A., Thoring, L., Hoffmeister, C., & Kubick, S. (2015). Cell-free protein synthesis: Pros and cons of prokaryotic and eukaryotic systems. ChemBioChem, 16(17), 2420–2431. https://doi.org/10.1002/cbic.201500340
Hodot, R., et al. (2023). “Cell-Free Protein Synthesis: A Powerful Tool for the Expression of Membrane Proteins.” Frontiers in Bioengineering and Biotechnology.
Jiang, S., Çelen, G., Glatter, T., Niederholtmeyer, H., & Yuan, J. (2024). “A cell-free system for functional studies of small membrane proteins.” Journal of Biological Chemistry.
Denisov, I. G., & Sligar, S. G. (2017). “Nanodiscs in Membrane Biology and Drug Discovery.” Chemical Reviews.
Carpenter, E. P., et al. (2008). Overcoming the challenges of membrane protein crystallography. Current Opinion in Structural Biology, 18(5), 581–586.
Itskanov, S., & Park, E. (2023). Mechanism of protein translocation by the Sec61 translocon complex. Cold Spring Harbor Perspectives in Biology, 15(1), a041250. https://doi.org/10.1101/cshperspect.a041250
Lyukmanova, E. N., Shenkarev, Z. O., Khabibullina, N. F., Kulbatskiy, D. S., Shulepko, M. A., Petrovskaya, L. E., Arseniev, A. S., Dolgikh, D. A., & Kirpichnikov, M. P. (2012). N-terminal fusion tags for effective production of G-protein-coupled receptors in bacterial cell-free systems. Acta Naturae, 4(4), 58–64.
Steinkühler, J., Peruzzi, J. A., Krüger, A., Villaseñor, C. G., Jacobs, M. L., Jewett, M. C., & Kamat, N. P. (2024). Improving cell-free expression of model membrane proteins by tuning ribosome cotranslational membrane association and nascent chain aggregation. ACS Synthetic Biology, 13(1), 129–140. https://doi.org/10.1021/acssynbio.3c00357
Drew, D., Newstead, S., Sonoda, Y., Kim, H., von Heijne, G. and Iwata, S. (2008) ‘GFP-based optimization scheme for the overexpression and purification of eukaryotic membrane proteins in Saccharomyces cerevisiae’, Nature Protocols, 3(5), pp. 784–798. doi: 10.1038/nprot.2008.44.
Schwarz, D., Junge, F., Durst, F., Frölich, N., Schneider, B., Reckel, S., Sobhanifar, S., Dötsch, V., & Bernhard, F. (2007). Preparative scale expression of membrane proteins in Escherichia coli-based continuous exchange cell-free systems. Nature Protocols, 2(11), 2945–2957. https://doi.org/10.1038/nprot.2007.426
Shoba (2025) ‘Solved: Low yields in cell-free protein synthesis’, Bitesize Bio. Available at: https://bitesizebio.com/10234/solvedlow-yields-in-cell-free-protein-synthesis/ (Accessed: 5 April 2026).
Creative Biostructure. (n.d.). Cell-free membrane protein synthesis. Creative Biostructure. https://www.creative-biostructure.com/cellfree-membrane-protein-synthesis.htm
Homework question from Kate Adamala
Design an example of a useful synthetic minimal cell as follows:
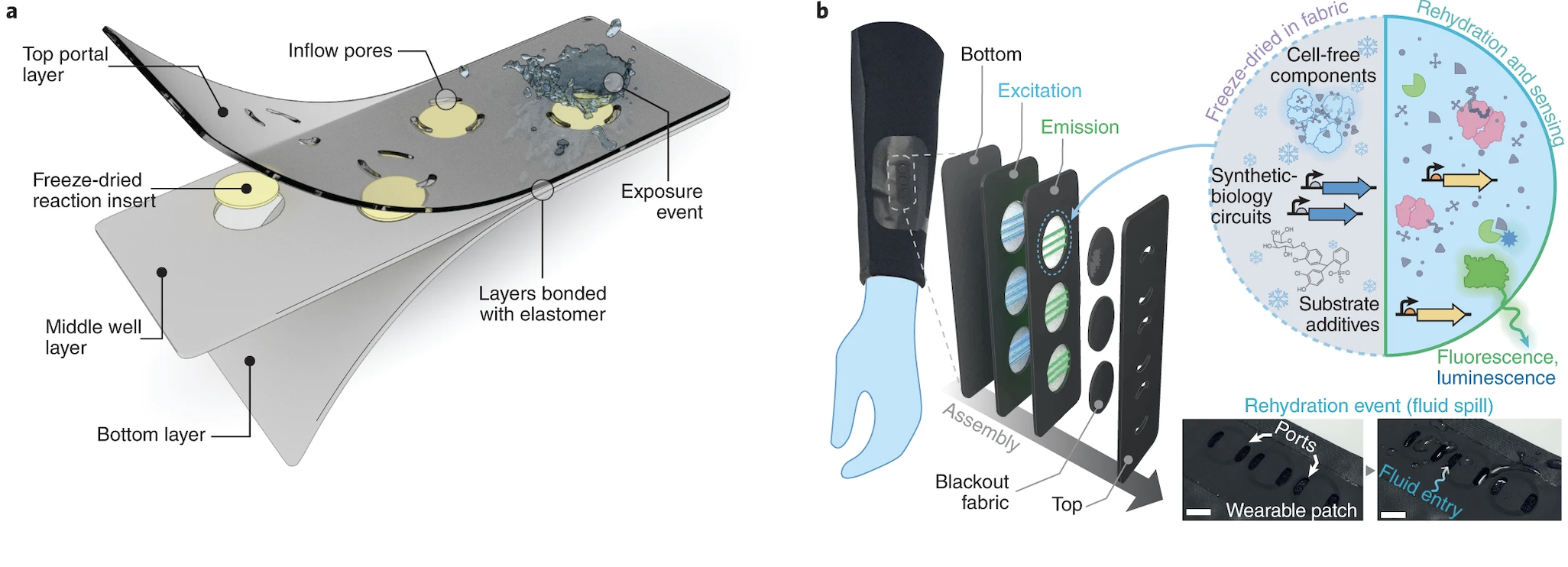

Wearable textile-embedded biosensors relying on freeze-dried cell-free synthetic biology. From: https://www.nature.com/articles/s41551-022-00869-3/figures/1
1. Pick a function and describe it.
The synthetic minimal cell functions as a gaseous biosensor designed to detect Polycyclic Aromatic Hydrocarbons (PAHs), for this case specifically naphthalene, in the atmosphere and create a visible, colourmetric output. This is linked to ideas I am exploring in my final project, related to creating a bio-hybrid public sculptures that offer a visible read out to atmospheric pollutants in order to create social engagement with air pollution inequalities.
2. What would your synthetic cell do? What is the input and what is the output?
The synthetic minimal cell acts as an environmental reporter. Its function is to indicate urban air quality by detecting specific toxic markers (PAHs) and providing a visual, colorimetric readout.
Input: Gas-phase Polycyclic Aromatic Hydrocarbons (PAHs). Output: Colorimetric change (Yellow to Red) via the hydrolysis of CPRG by the enzyme $\beta$-galactosidase.
3. Could this function be realised by cell-free Tx/Tl alone, without encapsulation?
Yes it could technically work, NahR would detect the PAH metabolite, activates transcription of lacZ and β-galactosidase hydrolyses CPRG to give a colour change. This chemistry doesn’t inherently require encapsulation to function.
But practically for my application no. For a public-facing sculpture you need the reaction to be stable over time, contained and accessible to gaseous PAH. An unprotected solutions would degrade quickly, evaporate or be contaminated. For a real application (like a public sensor), encapsulation in a solid/gel is preferable.
Could this function be realized by genetically modified natural cell?
Yes, NahR-lacZ constructs have been expressed in E. coli as whole-cell biosensors but a synthetic minimal cell is advantageous for public art installations (Cho et al. 2015). It is bio-contained and non-living, meaning it cannot replicate, mutate or pose a bio-safety risk to the public environment and it doesn’t require nutrients and waste disposal to survive.
Describe the desired outcome of your synthetic cell operation.
A stable, portable and safe minimal cell that:
- Detects environmental PAHs in real time.
- Produces a visible colour change without requiring handling of live cells.
- Is long-lasting and stable
- Can communicate with the environment .
Design all components that would need to be part of your synthetic cell.
1. What would be the membrane made of?
A liposome composed of phosphatidylcholine and cholesterol (7:3) provides a stable bilayer permeable to small, hydrophobic PAHs without requiring a protein channel (Plant et al, 1987).. Cholesterol reduces membrane leakage and improves stability across variable temperatures, important for different deployment contexts. Moreover, as noted in the Shin et al. (2005), PC membranes, when protected by a cryoprotectant like sucrose, can survive the freeze-drying process and rehydrate successfully when they encounter atmospheric moisture in the public installation.
2. What would you encapsulate inside? Enzymes, small molecules.
- NahR transcription factor and lacZ gene – incorporated in a plasmid under a PAH-inducible promoter (e.g. Psal) to detect environmental PAHs and trigger β-galactosidase expression.
- Cell-free Tx/Tl machinery (E. coli extract) – including ribosomes, RNA polymerase, translation factors (initiation, elongation, and release factors), aminoacyl-tRNA synthetases, and tRNAs. This machinery enables transcription of the plasmid and translation of β-galactosidase.
- CPRG substrate – a chromogenic substrate for β-galactosidase, which produces a visible color change when cleaved.
- NTPs, amino acids – provide the building blocks.
- Mg-glutamate and K-glutamate — ions that support ribosome activity and maintain optimal chemical environment for transcription and translation.
- 3-phosphoglyceric acid (3-PGA) - regenerates ATP to sustain transcription and translation throughout the reaction, without accumulating inhibitory phosphate byproducts (Caschera and Noireaux, 2014).
Which organism your Tx/Tl system will come from? Is bacterial OK, or do you need a mammalian system for some reason? _(hint: for example, if you want to use small molecule modulated promotors, like Tet-ON, you need mammalian)
Bacterial (E. coli) extract is sufficient here, the NahR-Psal promoter system is prokaryotic and functions natively in E. coli based cell-free systems. A mammalian system is not needed as there are no mammalian-specific promoters (e.g. Tet-ON) or PTMs required.
How will your synthetic cell communicate with the environment? _(hint: are substrates permeable? or do you need to express the membrane channel?)
PAHs are sufficiently hydrophobic to passively diffuse across the lipid bilayer membrane into the synthetic cell, where they interact with the NahR transcription factor and trigger lacZ expression. No membrane channel is required.
Experimental details
1. List all lipids and genes. _(bonus: find the specific genes; for example, instead of just saying “small molecule membrane channel” pick the actual gene.)
LIPIDS:
- POPC (1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine)- primary bilayer lipid, permeable to hydrophobic PAHs
- Cholesterol (7:3 POPC:cholesterol molar ratio)- tightens the bilayer, reduces leakage and improves stability across variable temperatures
GENES:
- NahR encodes the NahR transcription factor from Pseudomonas putida, binds PAH and activates transcription at the Psal promoter.
- lacZ encodes β-galactosidase from E. coli, placed under the Psal promoter, hydrolyses CPRG to produce the colorimetric output
- Psal promoter - PAH-inducible regulatory sequence from Pseudomonas putida that NahR binds to activate lacZ transcription
How will you measure the function of your system?
- Colorimetric assay : absorbance at 570 nm to quantify CPRG hydrolysis (yellow to pink/red) as a direct readout of PAH detection. Include control tests to known PAH concentrations and compare colour change intensity.
REFERENCES:
Cho, J. H., Lee, D. Y., Lim, W. K., & Shin, H. J. (2014). A recombinant Escherichia coli biosensor for detecting polycyclic aromatic hydrocarbons in gas and aqueous phases. Journal of Environmental Science and Health, Part A, 49(13), 1521–1527. https://doi.org/10.1080/10826068.2014.887577
Shin, H. J., Park, H. H., & Lim, W. K. (2005). Freeze-dried recombinant bacteria for on-site detection of phenolic compounds by color change. Journal of Biotechnology, 119(1), 36–43. https://doi.org/10.1016/j.jbiotec.2005.06.002
Plant, A. L., Knapp, R. D., & Smith, L. C. (1987). Mechanism and rate of permeation of cells by polycyclic aromatic hydrocarbons. Journal of Biological Chemistry, 262(6), 2514–2519.
Caschera, F., & Noireaux, V. (2014). Synthesis of 2.3 mg/ml of protein with an all Escherichia coli cell-free transcription-translation system. Biochimie, 99, 162–168. https://doi.org/10.1016/j.biochi.2013.11.025
Homework question from Peter Nguyen
Again, I will be basing this answer on my final project idea.
Write a one-sentence summary pitch sentence describing your concept.
A series of bio-sensing public sculptures that offer a naked eye, colourmetric response to air pollutants (PAHs) in order to encourage engage with air quality injustice.
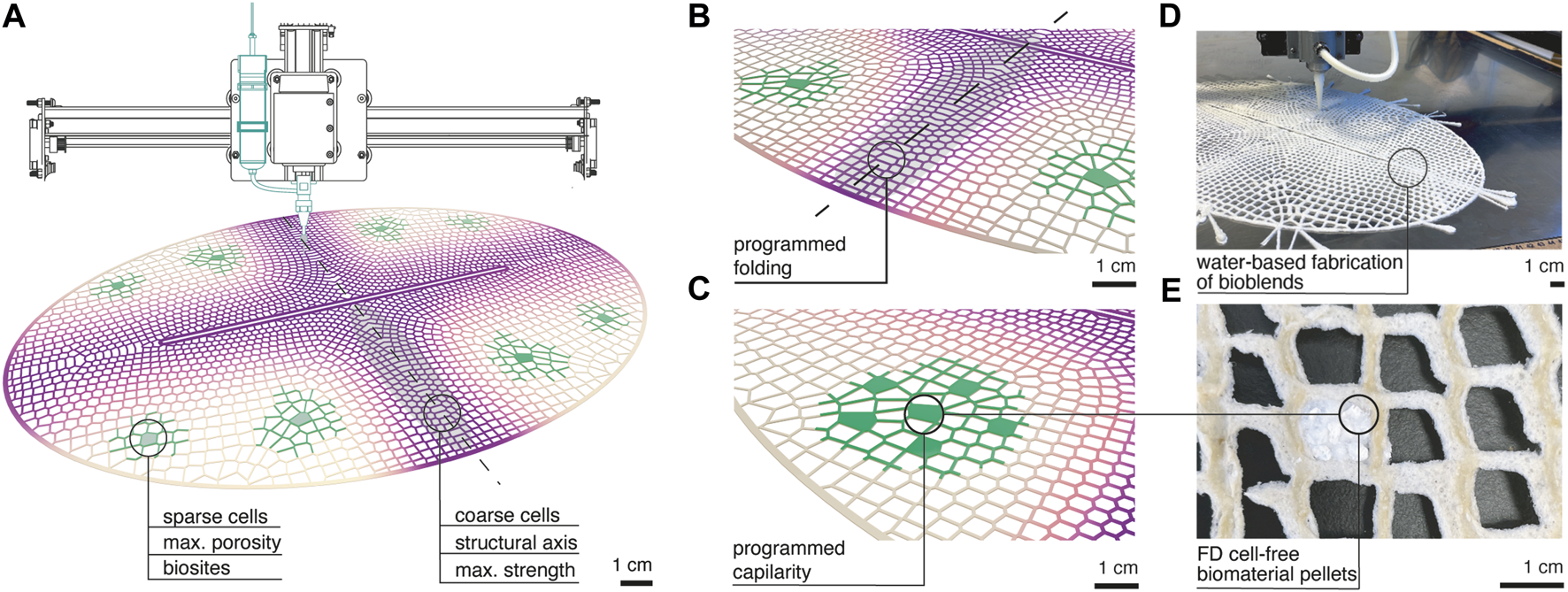

Cell-free biologically active architectural structures. From: https://www.frontiersin.org/journals/bioengineering-and-biotechnology/articles/10.3389/fbioe.2023.1125156/full/1
How will the idea work, in more detail? Write 3-4 sentences or more.
The sculptures will contain freeze-dried cell-free bio-sensing modules embedded in a 3D printed structure made of a biopolymer matrix (sodium alginate or silk fibroin) (Ho et al. 2023). The modules contain a freeze-dried cell-free NahR-lacZ genetic circuit. When PAHs diffuse passively into the module, the NahR transcription factor binds and triggers β-galactosidase expression, hydrolysing the CPRG substrate and producing a visible yellow to red colour change proportional to PAH concentration (Cho et al. 2014). The sculptures will be strategically placed in public areas across urban environments and across global cities, creating a living, real-time visualisation of invisible pollution translating an abstract environmental justice issue into a visceral, public-facing experience that requires no scientific literacy to interpret. I am keen for the sculptures to be presented together to aid visual comparison and discussion of systems that contribute to inequalities.
What societal challenge or market need will this address?
Air pollution disproportionately affects low-income and marginalised urban communities, yet the data that demonstrates this inequality is largely inaccessible to the public and communicated through abstract data, unactionable for the communities most affected. This project address this challenge by translating invisible, structural environmental injustice into a visible, intuitive experience that anyone can engage with regardless of background. There is a growing need for public-facing environmental monitoring tools that go beyond data and create emotional and social engagement with air quality as a justice issue. It would be interesting to push this further in future in to remediation also.
How do you envision addressing the limitation of cell-free reactions (e.g., activation with water, stability, one-time use)?
Stability: the cell-free bio-sensing modules will be freeze-dried with sucrose as a cryoprotectant, which protects the liposome membrane and encapsulated machinery during storage and extends shelf life significantly (Shin et al., 2005)
Activation with water: As explored by Ho et al. (2023) the biosites are embedded in a porous biopolymer lattice designed with specific porosity and capillarity properties, meaning atmospheric water and humidity travels through the lattice structure to the biosites naturally upon exposure to moisture, rehydrating and activating the cell-free reaction without any handling.
One-time use: the biosensors could be designed as replaceable cartridges modules embedded within the sculpture, allowing periodic replacement without dismantling the installation itself? However, this is a limitation I am going to need to research further.
REFERENCES
Cho, J. H., Lee, D. Y., Lim, W. K., & Shin, H. J. (2014). A recombinant Escherichia coli biosensor for detecting polycyclic aromatic hydrocarbons in gas and aqueous phases. Journal of Environmental Science and Health, Part A, 49(13), 1521–1527. https://doi.org/10.1080/10826068.2014.887577
Ho, G., Kubušová, V., Irabien, C., Li, V., Weinstein, A., Chawla, S., Yeung, D., Mershin, A., Zolotovsky, K., & Mogas-Soldevila, L. (2023). Multiscale design of cell-free biologically active architectural structures. Frontiers in Bioengineering and Biotechnology, 11, 1125156. https://doi.org/10.3389/fbioe.2023.1125156
Shin, H. J., Park, H. H., & Lim, W. K. (2005). Freeze-dried recombinant bacteria for on-site detection of phenolic compounds by color change. Journal of Biotechnology, 119(1), 36–43. https://doi.org/10.1016/j.jbiotec.2005.06.002
Homework question from Ally Huang


From: https://www.the-scientist.com/stem-cells-age-faster-in-space-73357
Provide background information that describes the space biology question or challenge you propose to address. Explain why this topic is significant for humanity, relevant for space exploration, and scientifically interesting. _(Maximum 100 words)
During long-duration spaceflight, astronauts experience skeletal muscle atrophy caused by microgravity, reducing strength, endurance and motor performance(Wang, 2013). The early molecular events driving this process remain poorly understood and current studied biomarkers focus on those detectable after muscle loss has begun. Studying human muscle precursor cells in real space conditions allows investigation of how microgravity disrupts muscle differentiation and regeneration. Muscle-specific microRNAs (myomiRs), which regulate satellite-cell activity, change before measurable tissue loss occurs and therefore provide early indicators of muscle adaptation during long-duration space missions (Di Filippo et al. 2024) . This project proposes to detect coordinated changes in myomiRs in human muscle precursor cells in real microgravity using freeze-dried BioBits cell-free systems to better understand how microgravity alters muscle gene regulation and support development of future diagnostic and preventative countermeasures to muscle atrophy for astronauts.


Japanese astronaut Satoshi Furukawa exercising on the T2 treadmill, the Cycle Ergometer with Vibration Isolation and Stabilization System (CEVIS), and the Advanced Resistive Exercise Device (ARED) in the ISS. From: https://www.nature.com/articles/s41526-021-00145-9
Name the molecular or genetic target that you propose to study. Examples of molecular targets include individual genes and proteins, DNA and RNA sequences, or broader -omics approaches. (Maximum 30 words)
A panel of muscle-specific microRNAs miR-1, miR-133a, miR-133b and miR-206, key regulators of satellite cell differentiation, myotube formation and skeletal muscle regeneration during early muscle adaptation to microgravity (Di Fillippo et al 2014).
Describe how your molecular or genetic target relates to the space biology question or challenge your proposal addresses. (Maximum 100 words)
These myomiRs regulate satellite cell activation, proliferation and differentiation, which are essential for maintaining skeletal muscle mass. Studies show their expression changes early during exposure to microgravity, before measurable muscle tissue loss occurs (Di Fillippo et al 2014). Measuring changes in miR-1, miR-133a, miR-133b and miR-206 in human muscle precursor cells cultured in real microgravity conditions will help identify how microgravity disrupts muscle regeneration pathways. Understanding these early regulatory changes provides insight into the molecular events driving muscle atrophy and supports development of targeted countermeasures for long-duration missions.
Clearly state your hypothesis or research goal and explain the reasoning behind it. (Maximum 150 words)
I hypothesise that exposure of human muscle precursor cells to real microgravity will produce downregulation in muscle-specific microRNAs miR-1, miR-133a, miR-133b and miR-206, reflecting early disruption of gene regulatory pathways controlling muscle atrophy. Detecting these changes using freeze dried BioBits cell-free protein expression reactions combined with miniPCR amplification will demonstrate a compact and space compatible strategy for studying regulators of muscle differentiation and regeneration whose dysregulation contributes to atrophy in space. This experiment aims to improve understanding of the molecular mechanisms underlying microgravity induced muscle atrophy by identifying early regulatory responses in muscle precursor cells. I aim to establishing a workflow for detecting myomiRs in real microgravity, using portable cell-free systems. This will support future studies investigating muscle adaptation in space and help guide development of effective countermeasures to maintain astronaut musculoskeletal health during long-duration missions.
Outline your experimental plan - identify the sample(s) you will test in your experiment, including any necessary controls, the type of data or measurements that will be collected, etc. (Maximum 100 words)
Human muscle precursor cells isolated from skeletal muscle biopsies on Earth will be expanded and cryopreserved before launch in space. Cells will be differentiated under microgravity conditions alongside parellel Earth gravity controls. RNA will be collected at defined timepoints and target myomiRs (miR-1, miR-133a, miR-133b, miR-206) reverse transcribed and amplified using miniPCR. Amplified products will activate a panel of freeze-dried BioBits cell-free toehold switch reporter reactions, each specific to one myomiR. Fluorescence output measured with the P51 Molecular Fluorescence Viewer will be compared between microgravity and control samples to determine relative expression changes. Negative controls will confirm assay specificity.
REFEERENCES
Wang XH. MicroRNA in myogenesis and muscle atrophy. Curr Opin Clin Nutr Metab Care. 2013 May;16(3):258–66. doi: 10.1097/MCO.0b013e32835f81b9. PMID: 23449000; PMCID: PMC3967234.
Di Filippo, E. S., Chiappalupi, S., Falone, S., Dolo, V., Amicarelli, F., Marchianò, S., Carino, A., Mascetti, G., Valentini, G., Piccirillo, S., Balsamo, M., Vukich, M., Fiorucci, S., Sorci, G., & Fulle, S. (2024). The MyoGravity project to study real microgravity effects on human muscle precursor cells and tissue. npj Microgravity, 10, 92. https://doi.org/10.1038/s41526-024-00392-1
Vitry, G., Finch, R., Mcstay, G., Behesti, A., Déjean, S., Larose, T., Wotring, V., & da Silveira, W. A. (2022). Muscle atrophy phenotype gene expression during spaceflight is linked to a metabolic crosstalk in both the liver and the muscle in mice. Communications Biology, 5, 1056. https://doi.org/10.1038/s42003-022-04006-1
Teodori, L., Costa, A., Campanella, L., & Albertini, M. C. (2019). Skeletal muscle atrophy in simulated microgravity might be triggered by immune-related microRNAs. Frontiers in Physiology, 9, 1926. https://doi.org/10.3389/fphys.2018.01926