Week 11 HW: Bioproduction & Cloud Labs


Important
Part A: The 1,536 Pixel Artwork Canvas | Collective Artwork
Contribute at least one pixel to this global artwork experiment before the editing ends on Sunday 4/19 at 11:59 PM EST.
I contributed 10 (I’m impatient and don’t know how to code haha!)
- Make a note on your HTGAA webpages including:
What you contributed to the community bioart project (e.g., “I made part of the DNA on the bottom right plate”)
I didn’t take note of which ones but they were pretty!
What you liked about the project?
I liked the excitement and working together. It was fun to not know what the picture would look like and see it evolve over time and every time you refreshed!
What about this collaborative art experiment could be made better for next year?
It took a while for me to grasp what we were doing and the onward workflow- maybe a simpler initial explanation with more layman language/ graphics would have been helpful.
Part B: Cell-Free Protein Synthesis | Cell-Free Reagents
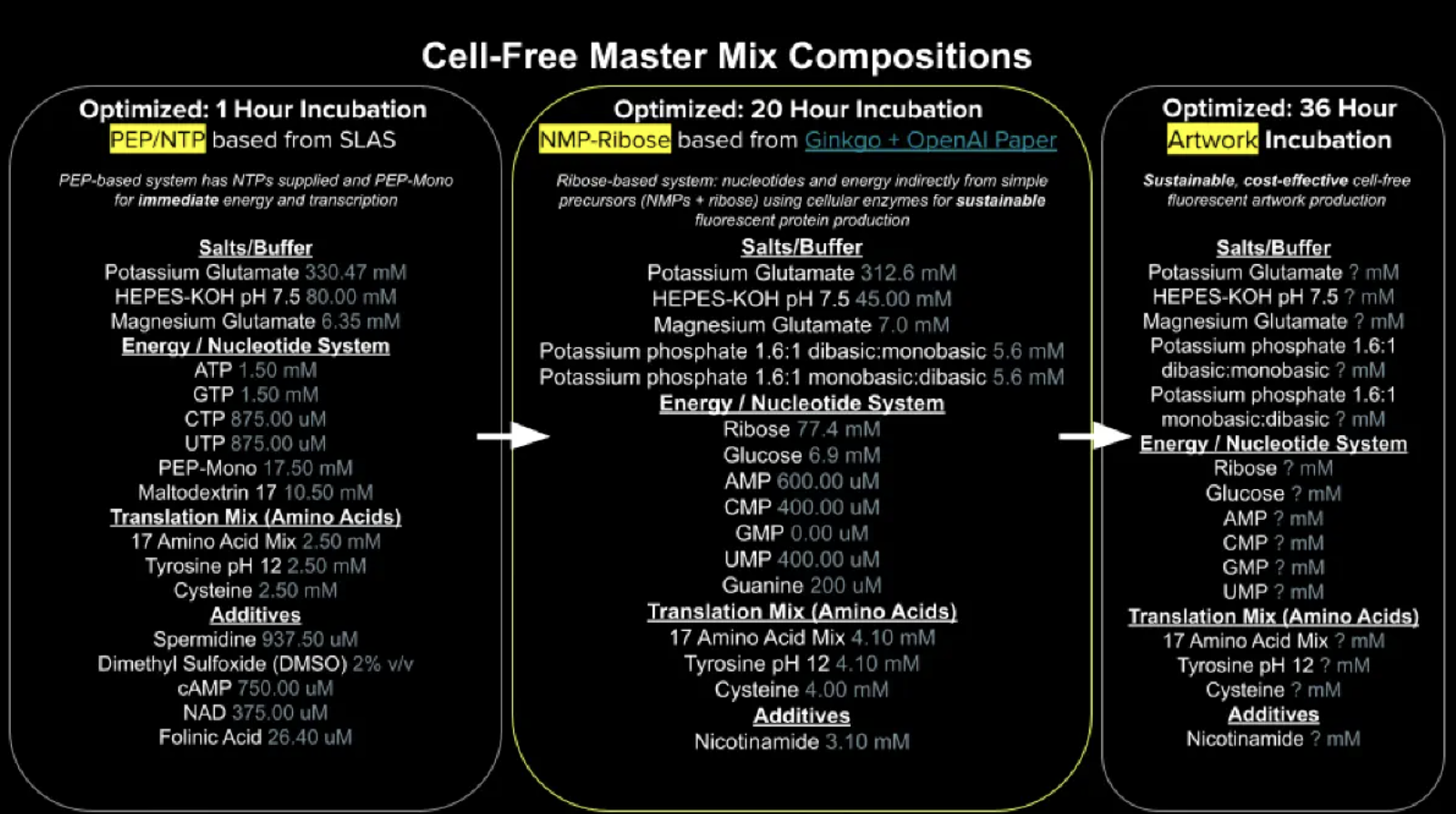

Referencing the cell-free protein synthesis reaction composition (the middle box outlined in yellow on the image above, also listed below), provide a 1-2 sentence description of what each component’s role is in the cell-free reaction.
E. coli Lysate
BL21 (DE3) Star Lysate (includes T7 RNA Polymerase)
This is a optimised, high yield E.coli cell free extract which provides all the machinery needed for transcription and translation e.g ribosomes, transcription factors, tRNAs, metabolic enzymes, chaperones. The DE3 indicates the strain carries the gene for T7 RNA polymerase. T7 RNA polymerase catalyses the formation of RNA from the DNA template, driving protein synthesis. T7RNA polymerase is one of the most efficient polymerases as it can complete a transcription cycle without requiring additional protein factors (McManus et al. 2019). The strain also carries a mutated rne gene (rne131) encoding a truncated RNase E enzyme, which reduces mRNA degradation and increases mRNA stability which makes the lysate higher yield.
Salts/Buffer
Potassium Glutamate:
A source of potassium ions for the reaction. These ions balance the charges from nucleic acid phosphate groups and other ionic compounds in the system, preservation of the ionic balance is essential for some protein nucleic acid interactions e. g ribosome stability (Jewett and Swartz, 2004). Glutamate can produce reducing equivalents such as NADH, which have a role in the generation of ATP via oxidative phosphorylation. This means that glutamate also contributes to energy regeneration in cell-free reactions (Jewett et al. 2008).
HEPES-KOH pH 7.5
Used to maintain the optimal, physiological pH of ph7.5 for the cell free system. As Smith et al. highlight in their 2026 paper highlights, pH drift, particularly acidification from metabolic byproducts is a major cause of titer collapse and the HEPES buffer had an outsized positive impact on yield (Smith et al. 2026)
Magnesium Glutamate
A source of magnesium ions for the reaction. The decrease in magnesium ions in the reaction due to the accumulation of inorganic phosphates causes protein synthesis to stop prematurely. Adding magnesium ions to CFPS has been shown to elongate the reaction time (Kim and Kim, 2009). Glutamate again contributes as an energy source in cell free reactions.
Potassium phosphate monobasic and dibasic
Together, these form a phosphate buffer that stabilises pH and supplies inorganic phosphate donors as a substrate for nucleotide regeneration pathways. This is particularly relevant in the NMP-based system (Nucleoside Monophosphate) we are using where phosphate drives NMP to NDP to NTP conversion.
Energy / Nucleotide System
Ribose
In an NMP cell-free system, ribose acts as the structural backbone needed to build and recycle NTPs. It is the source for the synthesis of phosphoribosyl pyrophosphate (PRPP) via the enzyme ribose-phosphate diphosphokinase. This activated PRPP intermediate acts as the ribose donor for phosphoribosyltransferases, which facilitate the salvage and conversion of the NMPs into NTPs required for transcription and translation (Banfalvi, 2021).
This PRPP route is most relevant to its relationship with guanine, where both Olsen et al and Smith et al found that GMP could be replaced with a combination of guanine and ribose without sacrificing protein yield. Ribose provides thet backbone that phosphoribosyltransferase enzymes use to convert free guanine into GMP, which is then phosphorylated to GTP (Smith et al. 2026)
Glucose
Is a cost effective energy source for the ATP regeneration, the system utilises glycolysis pathways present in the cell lysate to metabolise glucose and drive high-level protein synthesis.
Nucleoside Monophosphates: AMP, CMP, GMP, UMP
The source for all four NTPs required for transcription. The lysate’s endogenous kinase activities (adenylate kinase, nucleoside diphosphate kinase etc.) phosphorylate these to their di and triphosphate forms to get the NTPs. Olsen et al. established that NMPs provide a far better cost-to-titer ratio for a nucleotide regeneration strategy compared to adding pre-formed NTPs or pre-phosphorylated compounds (e.g PEP).
Guanine
is a nucleotide base added in combination with ribose as a cost-effective replacement for GMP, which is phosphorylated to GTP.
Translation Mix (Amino Acids)
17 Amino Acid Mix
Provides standard amino acid building blocks required for translation of desired protein. Glutamate, tyrosine, and cysteine are excluded from this mix because they require special handling (Whittaker, 2014).
Tyrosine
A amino acid required for the synthesis of certain proteins. It is supplied separately because of its very low aqueous solubility at neutral pH, requiring preparation at pH 12 and reduced concentration to prevent precipitation (Smith et al. 2026).
Cysteine
A amino acid required for the synthesis of certain proteins. Supplied separately because it is a highly reactive to oxidation in solution forming disulphide bonds and cystine. It must be handled carefully to maintain the cysteine pool for incorporation into proteins.
Additives
Nicotinamide
Nicotinamide is added to maintain high levels of NAD⁺/ NADH. Enzymes in the lysate convert nicotinamide to NAD⁺, replenishing the NAD⁺/NADH pool needed to sustain ATP regeneration throughout the reaction for transcription and translation (Jewett et al. 2008).
Backfill
Nuclease Free Water
Used to bring each reaction well to the defined total volume, ensuring all reactions are at equivalent concentration and volume without introducing RNase contamination that would degrade the mRNA template and collapse protein production.
References
Thermo Fisher Scientific. (n.d.). BL21 Star™ (DE3)pLysS One Shot™ chemically competent E. coli (C602003): FAQs. Retrieved April 24, 2026, from https://www.thermofisher.com/order/catalog/product/C602003/faqs
BenchChem Technical Support Team. (2025, December). The multifaceted role of potassium phosphate dibasic in molecular biology: A technical guide. BenchChem. https://pdf.benchchem.com/151/The_Multifaceted_Role_of_Potassium_Phosphate_Dibasic_in_Molecular_Biology_A_Technical_Guide.pdf
Harman, J. (2022). Development of a cell-free strategy for the directed evolution of enzymes for high-value natural products (MScRes thesis, University of Kent). https://doi.org/10.22024/UniKent/01.02.99209. Available at: https://kar.kent.ac.uk/99209/1/88Thesis_-_Joshua_Harman_complete.pdf
McManus, J. B., Emanuel, P. A., Murray, R. M., & Lux, M. W. (2019). A method for cost-effective and rapid characterization of engineered T7-based transcription factors by cell-free protein synthesis reveals insights into the regulation of T7 RNA polymerase-driven expression. Archives of Biochemistry and Biophysics, 674, 108045. https://doi.org/10.1016/j.abb.2019.07.010
Jewett, M. C., & Swartz, J. R. (2004). Mimicking the Escherichia coli cytoplasmic environment activates long-lived and efficient cell-free protein synthesis. Biotechnology and Bioengineering, 86(1), 19–26. https://doi.org/10.1002/bit.20026
Jewett, M. C., Calhoun, K. A., Voloshin, A., Wuu, J. J., & Swartz, J. R. (2008). An integrated cell‐free metabolic platform for protein production and synthetic biology. Molecular Systems Biology, 4(1), 220. https://doi.org/10.1038/msb.2008.57
Bánfalvi, G. (2021). Prebiotic pathway from ribose to RNA formation. International Journal of Molecular Sciences, 22(8), 3857. https://doi.org/10.3390/ijms22083857
Kim, H.-C., & Kim, D.-M. (2009). Methods for energizing cell-free protein synthesis. Journal of Bioscience and Bioengineering, 108(1), 1–4. https://doi.org/10.1016/j.jbiosc.2009.02.007
Whittaker, J. W. (2013). Cell-free protein synthesis: The state of the art. Biotechnology Letters, 35(2), 143–152. https://doi.org/10.1007/s10529-012-1075-4
Smith, A. A., Wong, E. L., Donovan, R. C., Chapman, B. A., Harry, R., Tirandazi, P., Kanigowska, P., Gendreau, E. A., Dahl, R. H., Jastrzebski, M., Cortez, J. E., Bremner, C. J., Morales Hemuda, J. C., Dooner, J., Graves, I., Karandikar, R., Lionetti, C., Christopher, K., Consiglio, A. L., … Shetty, R. P. (2026). Using a GPT-5-driven autonomous lab to optimize the cost and titer of cell-free protein synthesis. Preprint. https://doi.org/10.64898/2026.02.05.703998
Olsen, M. L., Copeland, C. E., Sundberg, C. A., Aw, R., Shaver, Z. M., Rao, G., Swartz, J. R., Karim, A. S., & Jewett, M. C. (2025). Design-driven optimization of low-cost reagent formulations for reproducible and high-yielding cell-free gene expression. bioRxiv. https://doi.org/10.1101/2025.08.01.668204
Describe the main differences between the 1-hour optimised PEP-NTP master mix and the 20-hour NMP-Ribose-Glucose master mix shown in the Google Slide above. (2-3 sentences)
The most fundamental difference between the two systems is their energy and nucleotide strategy. In the 1 hour system pre-formed NTPs (ATP, GTP, CTP, UTP) are used with PEP Mono as an immediate, high energy phosphate donor for rapid NTP regeneration. This enabling fast transcription and translation within 1 hour but at significantly higher reagent cost.
The 20 hour NMP/Ribose system relies on the lysate’s endogenous kinase machinery to phosphorylate cheap NMP to NTPs, with ribose, glucose and inorganic phosphate (from the potassium phosphate buffer). This is a slower but more cost-effective nucleotide regeneration strategy over 20 hours, as established by Olsen et al and Smith et al.
The 1 hour system also contains additional costly additives such as NAD⁺, folinic acid, cAMP, spermidine and DMSO, as the system relies on exogenously supplied cofactors, whereas the 20 hour system instead supplies nicotinamide as a cheap NAD⁺ precursor and relies on the lysate’s endogenous enzymes to maintain cofactors throughout a longer reaction.
Part C: Planning the Global Experiment | Cell-Free Master Mix Design
Given the 6 fluorescent proteins we used for our collaborative painting, identify and explain at least one biophysical or functional property of each protein that affects expression or readout in cell-free systems. (Hint: options include maturation time, acid sensitivity, folding, oxygen dependence, etc) (1-2 sentences each)
While sfGFP is designed to fold significantly faster than GFP variants and is much more stable, the chemical maturation of its chromophore, which involves cyclisation, dehydration and oxidation is still constrained by the need for molecular oxygen to modify the peptide backbone. This could be a dependancy that could be supplemented in the cell free system, so fluorescence is not reduced by poorly oxygenated or high-density cell-free reactions even when protein expression is strong.
mRFP1 has a relatively slow chromophore maturation kinetics and lower folding efficiency compared with newer red FPs such as mFruit and mCherry. Like all red fps it also requires an additional oxidation step to form the complete chromophore structure responsible for its red fluorescence and is prone to being dim and bleaching quickly, which often results in weak fluorescence output within the limited time window typical of TX–TL cell-free reactions. It also has low acid sensitivity.
Has a moderate acid sensitivity meaning optimal folding occurs under conditions that avoid extreme pH, so fluorescence intensity can decrease if the cell-free reaction becomes acidic during transcription–translation. Kaida and Miura, 2012 also show that mKO2 is particularly sensitive to oxygen availability compared to mAG (a monomeric Azami Green fluorescent protein), meaning that when oxygen levels drop (hypoxic conditions), mKO2 fluorescence drops off more sharply and quickly.
mTurquoise has a relatively low extinction coefficient (30,000 M⁻¹cm⁻¹) compared to other fluorescent proteins means it is less bright per molecule than alternatives, potentially limiting detection sensitivity at low expression levels.
mScarlet-I matures faster than previous RFPs but still undergoes a multi-step red chromophore maturation pathway that is slower and more oxygen-dependent than GFP proteins, often limiting early fluorescence accumulation in cell-free reactions. It also has a moderate acid sensitivity meaning pH drift during ATP consumption and transcriptional activity in cell-free systems could reduce fluorescence.
Electra2 belongs to the blue fluorescent protein class, which generally exhibits lower intrinsic brightness than green/red fluorescent proteins, and its chromophore formation proceeds through a non-fluorescent intermediate that limits its rate, delaying fluorescence development after translation. Blue fluorescent proteins often have lower photostability and are harder to detect with standard imaging systems, which can reduce signal strength in lysate-based cell-free reactions.
Part D: Hypothesis and Master-mix Experiment 1
Create a hypothesis for how adjusting one or more reagents in the cell-free mastermix could improve a specific biophysical or functional property you identified above, in order to maximize fluorescence over a 36-hour incubation. Clearly state the protein, the reagent(s), and the expected effect.
Protein:
mScarlet_I
Property:
Moderate acid-sensitivity and slower oxygen dependant maturation
Explanation:
mScarlet_I flourescence is limited by its sensitivity to pH drift which can occur as cell-free reactions metabolize glucose over 36 hours. The accumulation of organic acids (e.g. acetate, lactate) lowers the pH, which protonates the chromophore and quenches fluorescence prematurely. In addition, the maturation of its red chromophore requires an additional step of oxidative cyclisation which is energy intensive and can be limited by metabolic exhaustion.
Hypothesis and Expected Effects:
I hypothesise that the combination of 55 mM HEPES-KOH and 5.0 mM Nicotinamide will extend the window of mScarlet_I peak fluorescence compared to either reagent alone by simultaneously stabilising the reaction pH of 7.5 and sustaining the NAD+/NADH pool over 36 hours. This is because the increased HEPES-KOH will provide a greater buffering capacity to counteract ph drift and prevents acid induced quenching of the chromophore. Simultaneously, increasing the Nicotinamide sustains the metabolic energy required for the energy intensive multi-step chromophore maturation. I further hypothesise that in increasing these reagents in the mixture, I am slightly altering the ionic strength of the solution. Therefore, I am adding a slight increase of Magnesium Glutanate for ribosomal stability and therefore the efficiency of the translation. I am using a factorial experiment design that tests each reagent individually and in combination, ensuring that any observed improvements are attributable to specific reagent interactions.
The second phase of this lab will be to define the precise reagent concentrations for your cell-free experiment. You will be assigned artwork wells with specific fluorescent proteins and receive an email with instructions this week (by April 24). You can begin composing master mix compositions here.
I choose 8 wells of mScarlet_I and defined the precise concentrations of the master-mix across them to validate my hypothesis. I used a factorial experiment design with negative control in order to idetify which combination of reagent adjustments optimises the peak flourescence of the protein.


| Well | HEPES-KOH | Nicotinamide | Mg-Glutamate | Experiment |
|---|---|---|---|---|
| 1 Q3-B13 | 45.0 mM | 3.125 mM | 6.975 mM | Control |
| 2Q3-C13 | 55.0 mM | 3.125 mM | 6.975 mM | Variable A: Does +10mM HEPES increase flourescence |
| 3Q3-D13 | 45.0 mM | 5.000 mM | 6.975 mM | Variable B: Does +1.875mM Nic increase flourescence |
| 4Q3-E13 | 45.0 mM | 3.125 mM | 7.600 mM | Variable C: Does +0.625mM Mg increase flourescence |
| 5Q3-B11 | 55.0 mM | 5.000 mM | 6.975 mM | Combined Variables(A+B): Does HEPES + Nicotinamide increase flourescence |
| 6Q3-C11 | 55.0 mM | 3.125 mM | 7.600 mM | Combined Variables(A+C): Does HEPES + Mg increase flourescence |
| 7Q3-D11 | 45.0 mM | 5.000 mM | 7.600 mM | Combined Variables(B+C): Does Nicaotinamide + Mg increase flourescence |
| 8Q3-F4 | 55.0 mM | 5.000 mM | 7.600 mM | Combined Variables(A+B+C): Do Does HEPES + Nicaotinamide + Mg increase flourescence |
References
Andrews BT, Schoenfish AR, Roy M, Waldo G, Jennings PA. The rough energy landscape of superfolder GFP is linked to the chromophore. J Mol Biol. 2007 Oct 19;373(2):476-90. doi: 10.1016/j.jmb.2007.07.071. Epub 2007 Aug 15. PMID: 17822714; PMCID: PMC2695656.
Balleza, E., Kim, J. M., & Cluzel, P. (2018). Systematic characterization of maturation time of fluorescent proteins in living cells. Scientific Reports, 8, 1448. https://doi.org/10.1038/s41598-018-19355-0
Fraikin, N., Couturier, A., Mercier, R., & Lesterlin, C. (2025). A palette of bright and photostable monomeric fluorescent proteins for bacterial time-lapse imaging. Science Advances, 11(16), eads6201. https://doi.org/10.1126/sciadv.ads6201
Kaida, A., & Miura, M. (2012). Differential dependence on oxygen tension during the maturation process between monomeric Kusabira Orange 2 and monomeric Azami Green expressed in HeLa cells. Biochemical and Biophysical Research Communications, 421(4), 855–859. https://doi.org/10.1016/j.bbrc.2012.04.102
Goedhart, J., von Stetten, D., Noirclerc-Savoye, M. et al. Structure-guided evolution of cyan fluorescent proteins towards a quantum yield of 93%. Nat Commun 3, 751 (2012). https://doi.org/10.1038/ncomms1738
Papadaki, S., Wang, X., Wang, Y., Zhang, H., Jia, S., Liu, S., Yang, M., Zhang, D., Jia, J. M., Köster, R. W., Namikawa, K., & Piatkevich, K. D. (2022). Dual-expression system for blue fluorescent protein optimization. Scientific Reports, 12(1), 10190. https://doi.org/10.1038/s41598-022-13214-0