DNA Design
 Final Project Slide by Isobel Leonard
Final Project Slide by Isobel LeonardDNA Design Goal:
A biosensor for the simple, colour change based detection of salicylate and naphthalene in the environmental air.
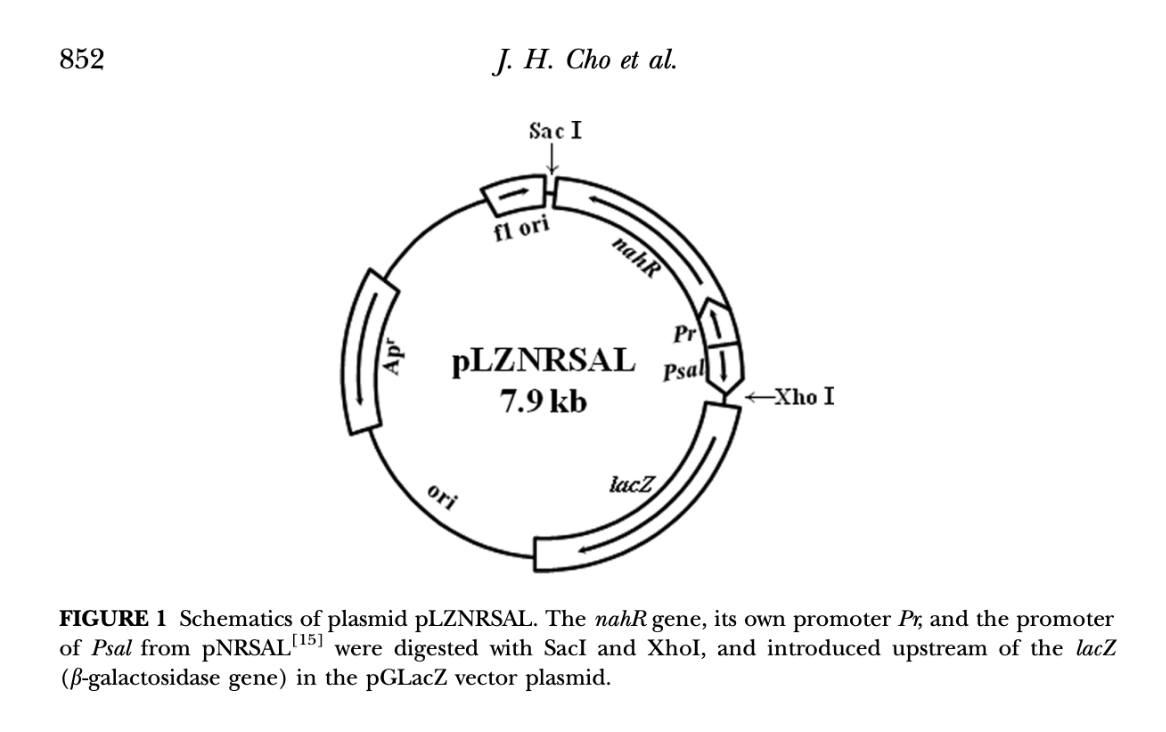
Based on the methodology of Cho et al (2014) and Park et al (2005), using the NahR/Pr/Psal lower operon from Pseudomonas putida’s Napthalene degrading plasmid NH7 and connecting it to the LacZ reporter gene that produces red-ß-D-galactopyranoside, hydrolysed in CPRG to provide a clear visible signal. The operon uses a constitutive promoter Pr and inducible promoter Psal in opposite directions.
I am trying to create a similarly functioning plasmid in Benchling for an in Silico Aim 1 using the digest and ligation assembly tool.
Reference Paper for Methodology on plasmid construction:
Reference paper for Nah/Pr/Psal Operon:
Plasmid diagram from Cho et al 2014:

In my attempts to create a similar functioning plasmid in silico I have followed the following workflow and then met some problems!
Circuit Logic:
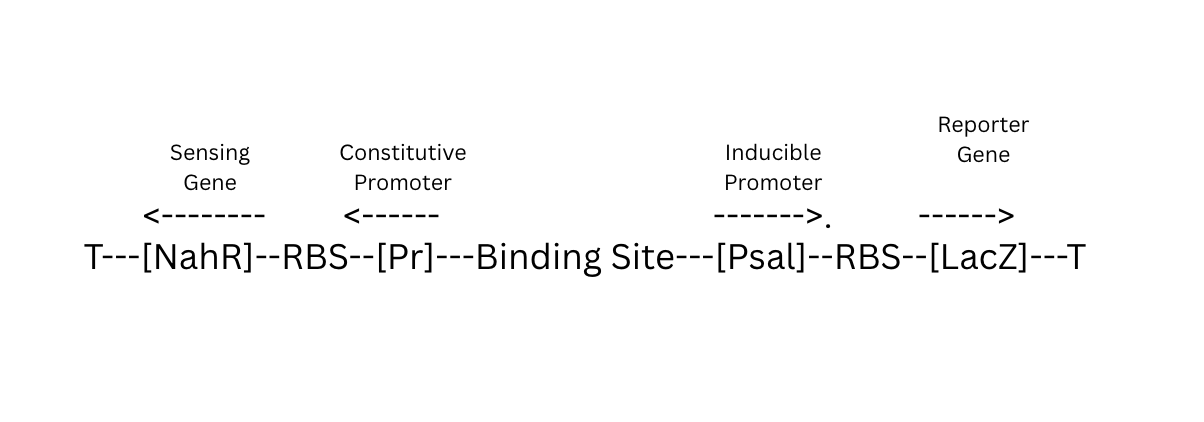


Biobits I need:
Backbone
pGL3b basic (2.9kb) digested and linearised with SaII and HindII and Luciferase reporter gene removed.
Reporter Gene:
LacZ fragment from psV-beta-Galactosidase. (3.7 kb)
Cut out with restriction enzymes SaII and HindIII and ligated into pGL3b basic to make plasmid PGLacZ.
Sensing NahR operon from Pseudomonas putida (1.3 kb)
Containing nahR/Pr/Psal and I have found variants on the sequence located in these places:
Design and add SacI and Xhol cut sites and ligated into plasmid PGLacZ.
Workflow:
Step 1:
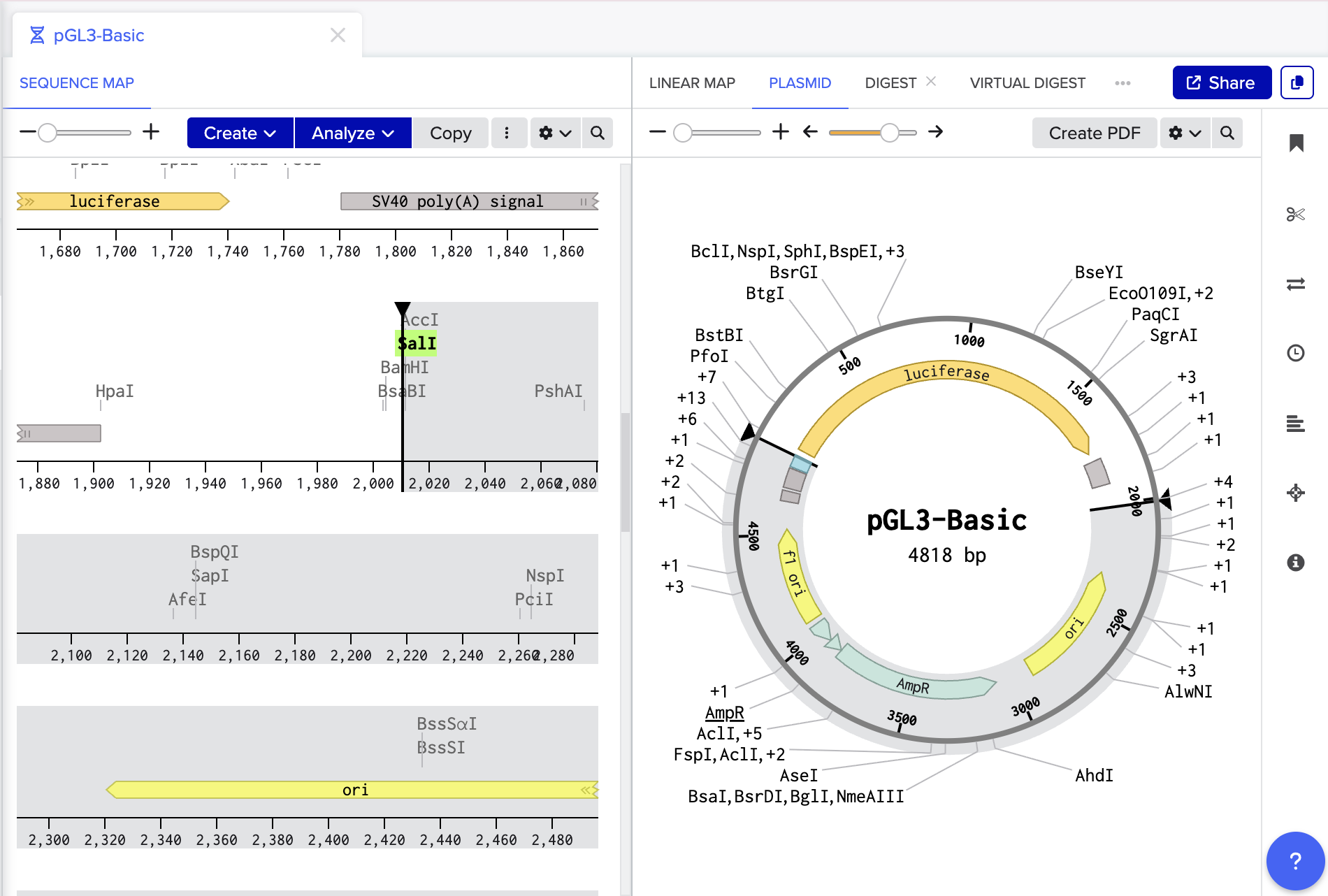
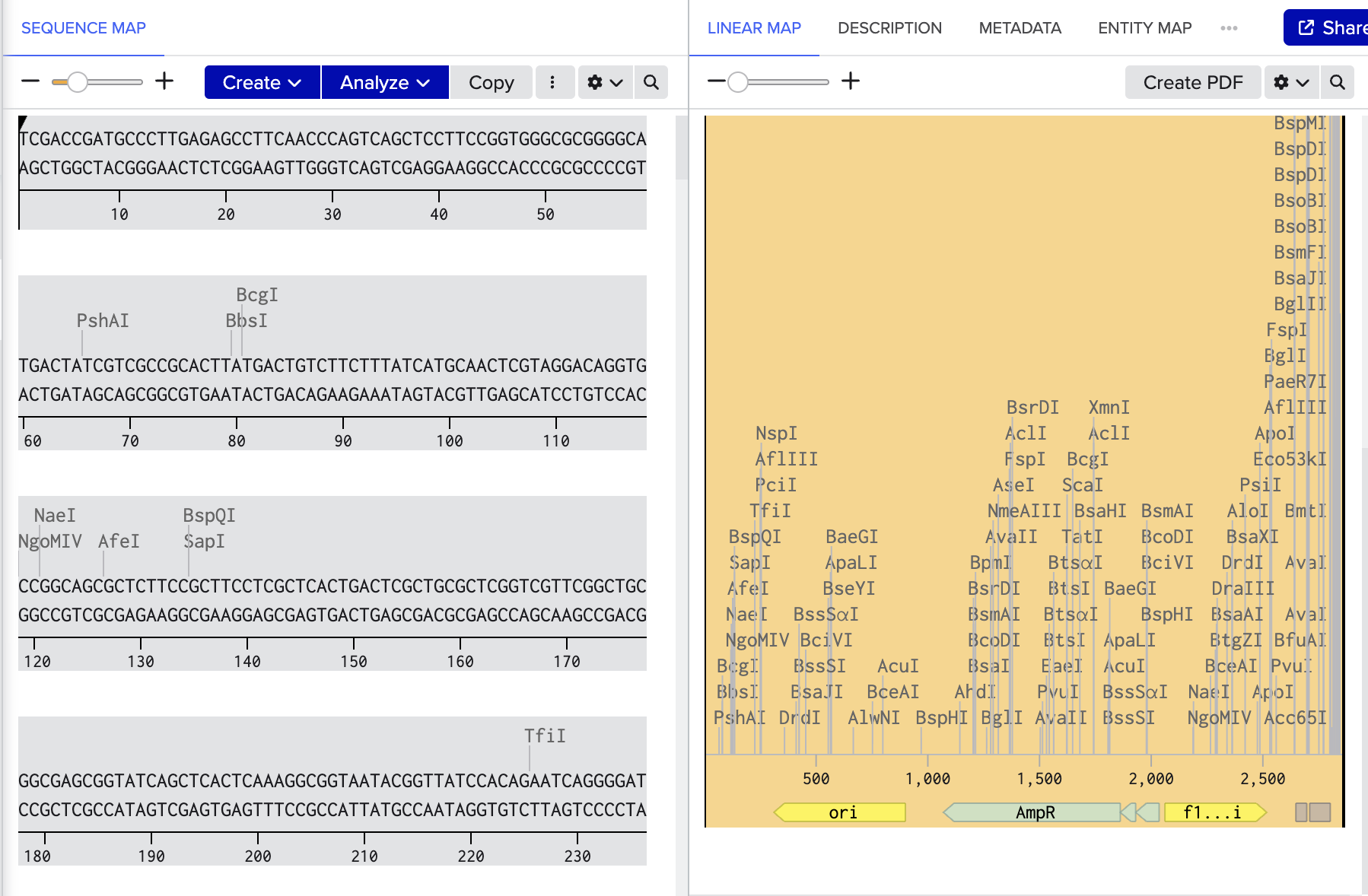
- Import pGL3- basic into benchling
- Digest with SalI and HindIII
- Remove luciferase and linearise maintaining origin of replication and antibiotic resistant gene.


Benchling link: https://benchling.com/s/seq-kf6LTrbodved1t066n13?m=slm-UxQtbtTBRIF5SaInZm2z
Step 2:
- Import psV-beta-Galactosidase
- Cut out LacZ fragment with restriction enzymes SalI and HindIII
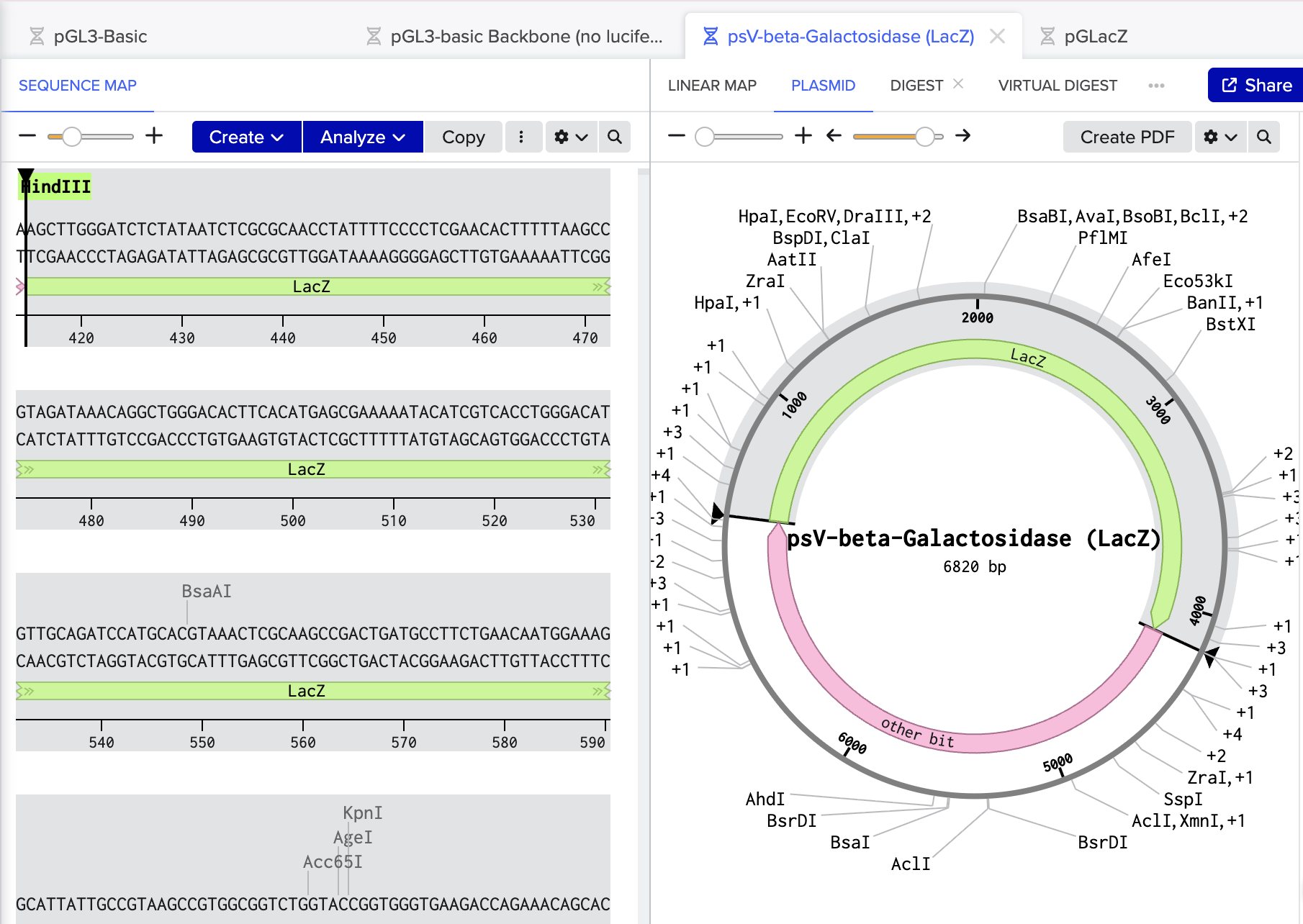

Benchling link: https://benchling.com/s/seq-R5yEkXzifvA87rugfx2z?m=slm-7vqIszJaDNRN5P8LbkGW
Step 3:
- Ligate intermediary plasmid using Assembly wizard
- select pGL3-Basic as backbone and lacZ as insert
- Ligate to form pGLacZ
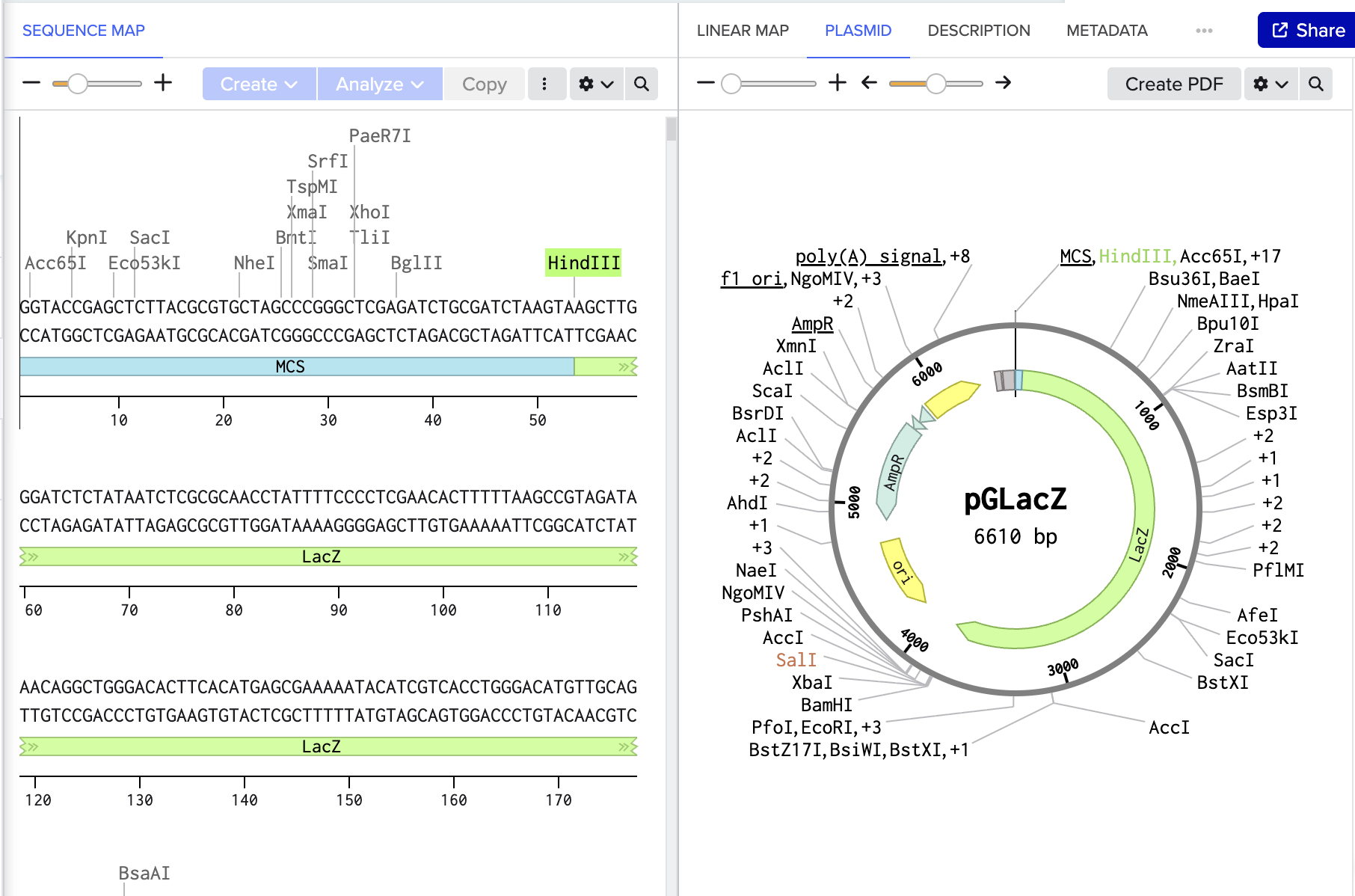

Benchling link: https://benchling.com/s/seq-VPPhB7LPfBWE65dQCNoa?m=slm-IZFORq41ceeD4uLW8Cra
Step 4:
- Locate sequence for NahR/Pr/Psal from naphthalene degrading gene cluster from P. putida and annotate.
Versions of sequence I have found:
GenBank: AY294313
- Add restriction binding sites for SacI and Xhol
- Insert upstream of LacZ
Here is where I have hit my problems
Problem 1: Unannotated sequence fragments
As there are no annotated and document sequences for each bit of the NahR/Pr/Psal operon e,g NahR gene, Pr Promoter, Psal promoter and the binding sites used, I don’t know which bit of this gene cluster from GenBank AY294313 is the bit I need.
In addition, I don’t know how to check all the bits I need for a functional sensor e.g RBS, Pr promoter, Psal Promoter are all included in the sequence (which they should be) and if they are the right way round and how to annotate them! I’ve been doing my head in a little trying to work it out!
In the Genbank info it says Gene NahR is 305-1207 and the next gene nahG starts at 1363. Meaning the promoters and binding sites must be between 1207 and 1363 but I am unsure how to identify?
I have tried my best to identify parts of the sequence in this benchling file by cross referencing between different versions of the Nah/Pr/Psal sequence across GenBank and iGEM but I don’t think I’m doing well and if this is a good approach:
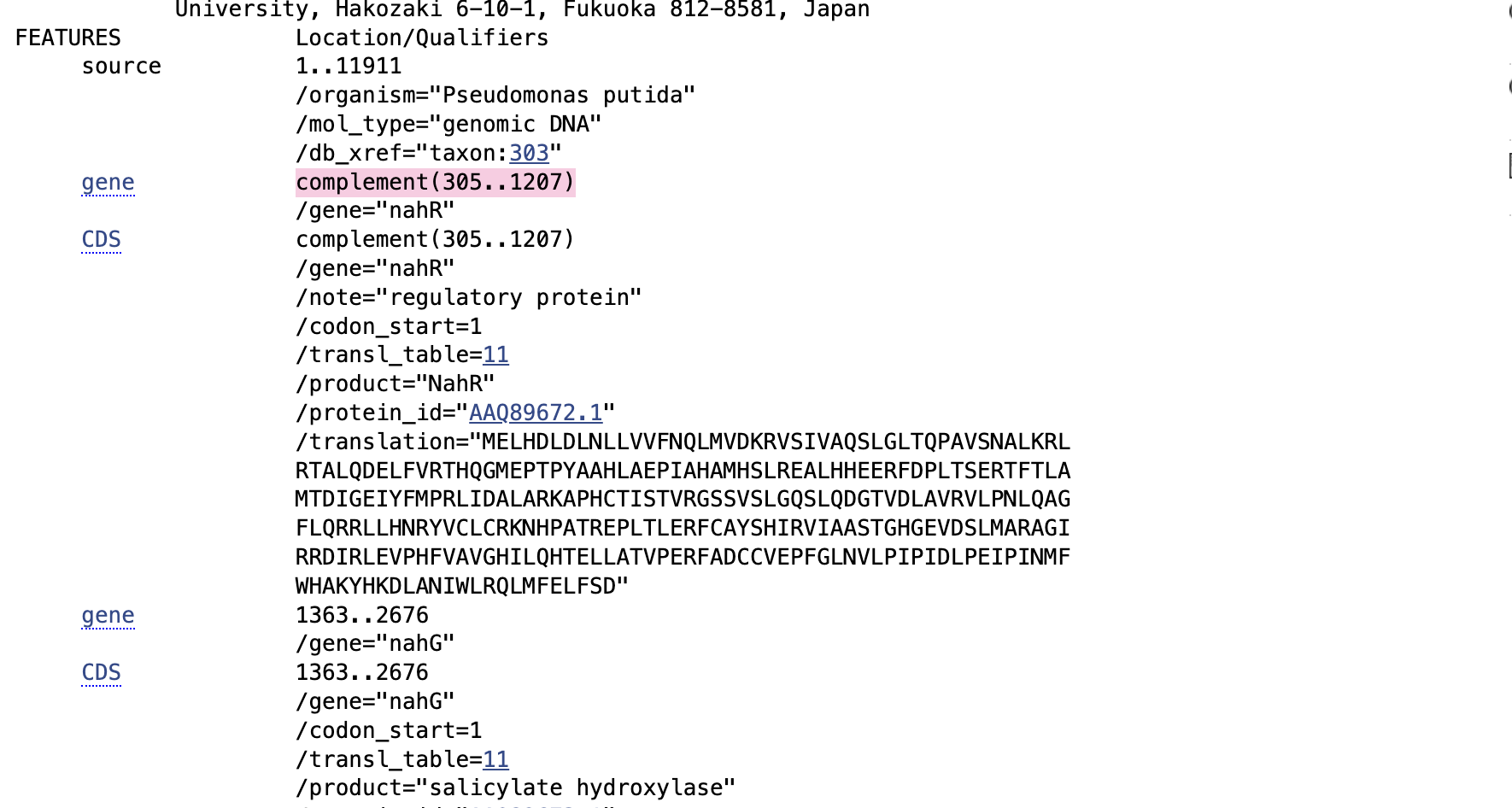

These two iGEM entries give a bit more info but are different from the GenBank sequence and I still don’t know how to tell where everything is??
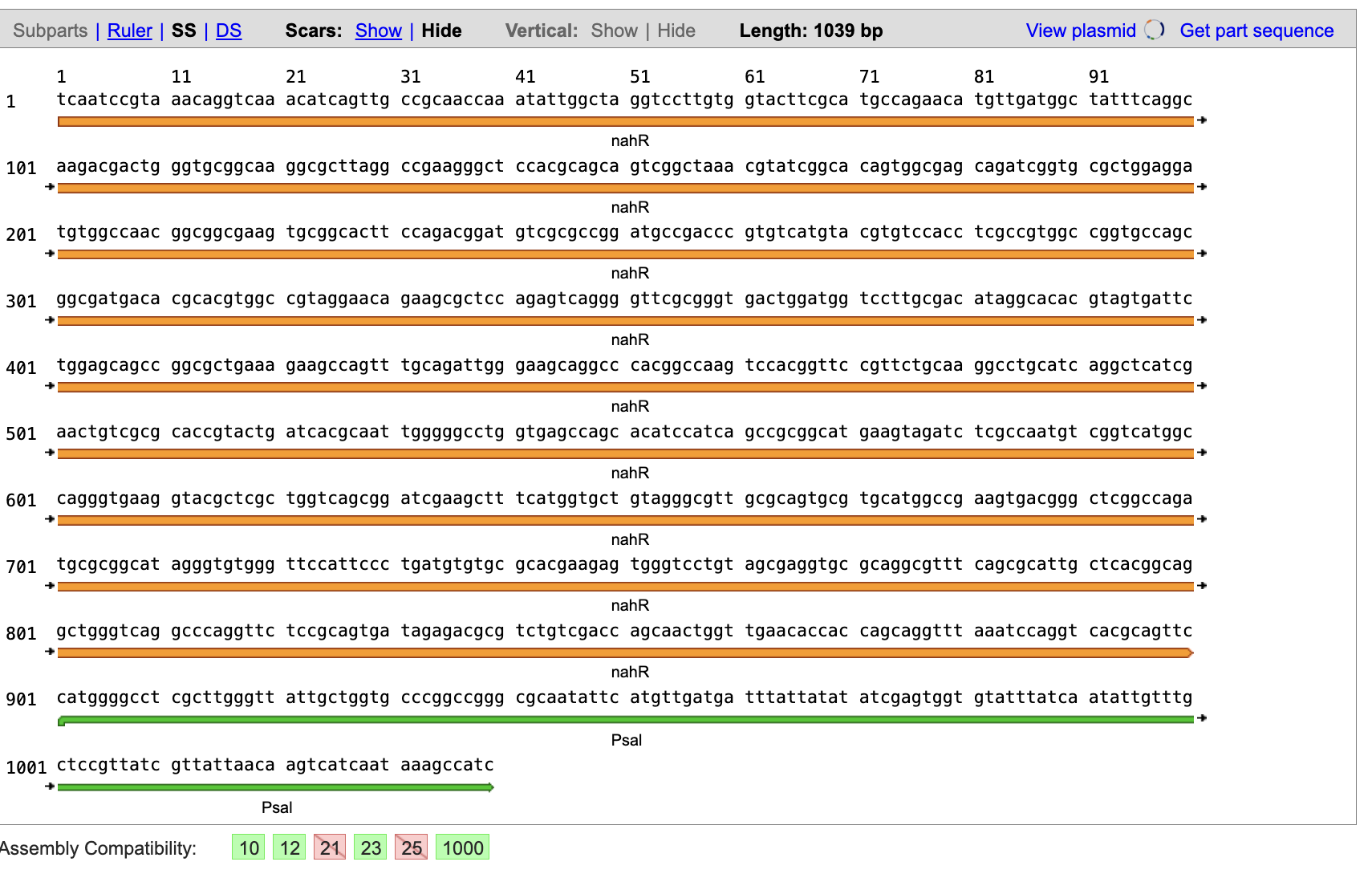



I have the same problem with the LacZ I digested from psV-beta-Galactosidase as although I can see the start and stop codon for the gene, I can’t tell if there is a RBS at the start of the sequence and which one it is?
I don’t know if the problem is that I am taking pieces from different vectors and instead I should try and build the sequence blocks myself with e.g Promoter, RBS, start codon, CDS etc. However, I still have the problem that I am unable to find the sequence for NahR, Pr or Psal promoter within the gene clusters and I wouldn’t know what RBS to use for the LacZ gene.
Problem 2: Multiple SacI cut sites
I have noticed (which is not documented in the paper I am following) that I have 2 SacI cut sites in my new plasmid PGLacZ, because the LacZ I have taken and ligated in has another SacI cut site in it. Meaning that if I were to digest at Xhol and SacI to insert the NahR/Pr/Psal region as described I would get another cut in LacZ.
However, I reason that I can just digest and ligate the pieces in a different order. First digest at Xhol and SacI while the LacZ is not in the backbone. Then digest at HindIII and SalI to remove luciferase and ligate in LacZ.
Is this okay to do??? Or is it better to use Gibson Assembly?? If so how do I design the overlaps for NahR/Pr/Psal so it goes specifically where it needs to go upstream of LacZ.
Problem 3: Cell Free System
I am looking into the possibility of innovating with the biosensor by creating a protocol to use the plasmids in a cell free system to create a safer and more stable bio-sensing public sculptures.
I am interested in merging the research I have already done with the research we learnt about in Cell free systems week from these two papers:
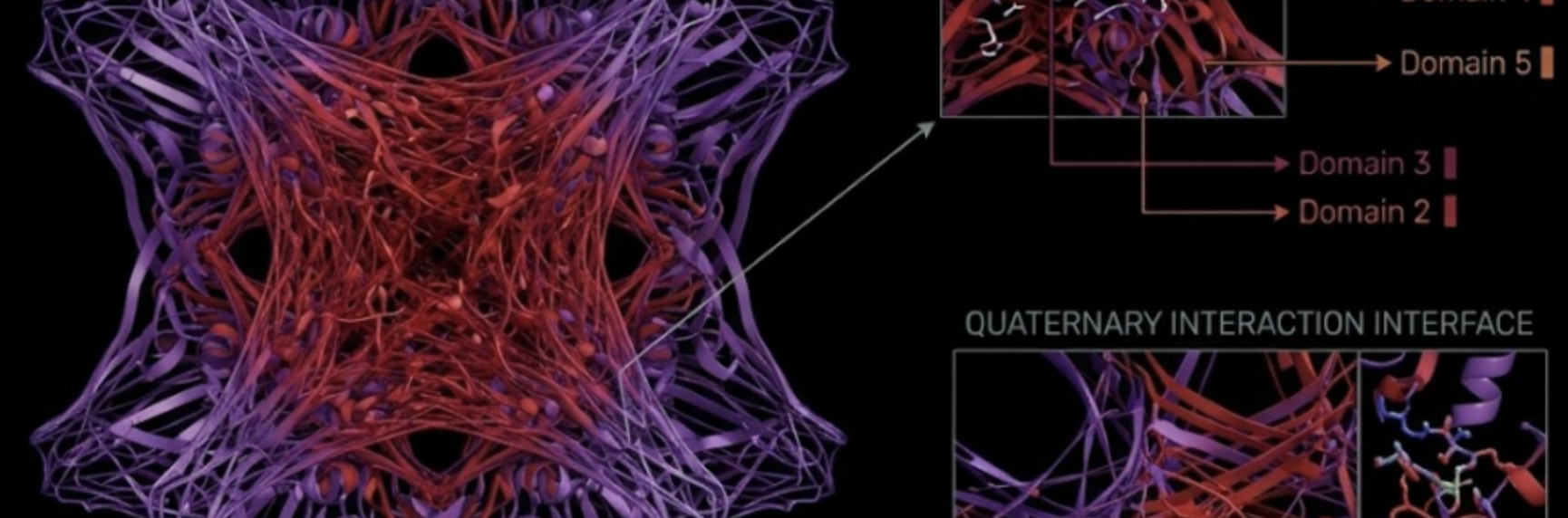
- Ho, G., Kubušová, V., Irabien, C., Li, V., Weinstein, A., Chawla, S., Yeung, D., Mershin, A., Zolotovsky, K., & Mogas-Soldevila, L. (2023). Multiscale design of cell-free biologically active architectural structures. Frontiers in Bioengineering and Biotechnology, 11, 1125156. https://doi.org/10.3389/fbioe.2023.1125156
Where the possibility of cell free biosensors being 3D printed ina biopolymer matrix for architectural structures is discussed.
- Nguyen, P. Q., Soenksen, L. R., Donghia, N. M., Angenent-Mari, N. M., de Puig, H., Huang, A., Lee, R., Slomovic, S., Galbersanini, T., Lansberry, G., Sallum, H. M., Zhao, E. M., Niemi, J. B., & Collins, J. J. (2021). Wearable materials with embedded synthetic biology sensors for biomolecule detection. Nature Biotechnology, 39, 1366–1374.https://doi.org/10.1038/s41587-021-00950-3
Where a cell free system with LacZ and CPRG was successfully designed as a wearable colourmetric bio-sensor.
However, I am a bit stumped on how to create a plasmid design for a cell free system and if the promoters I am using for the NahR operon are compatible with a cell free system as they are from Pseudomonas putida not Ecoli (would it work with Ecoli RNA polymerase or is there such thing as using Pseudomonas putida RNA polymerase in CFPS?). Is this a possible next step or is my goal not compatible?? Is there a way to optimise what I am doing so it would work??
Thank you if you read this far!! :)