Individual Final Project
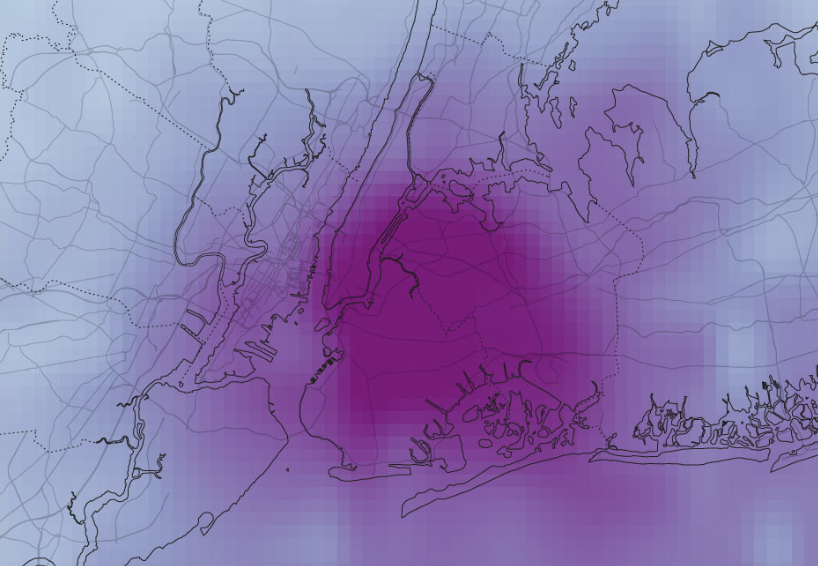
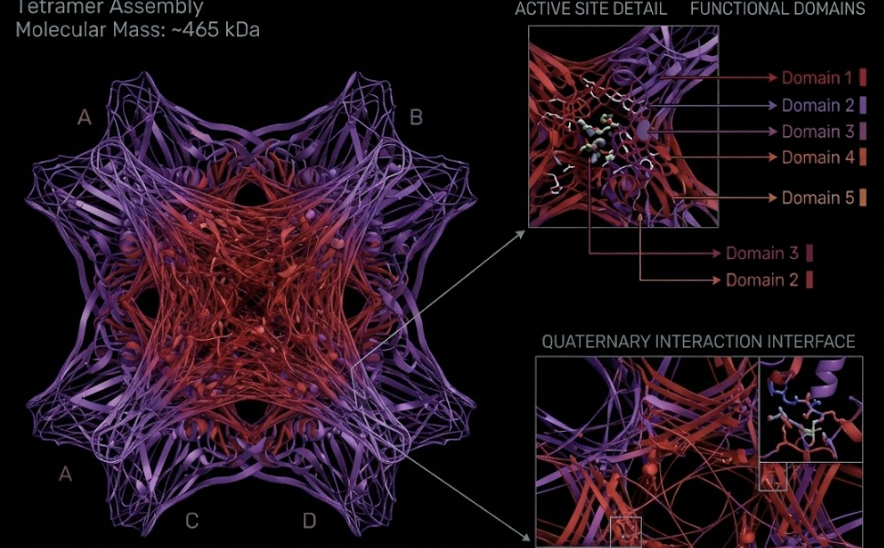
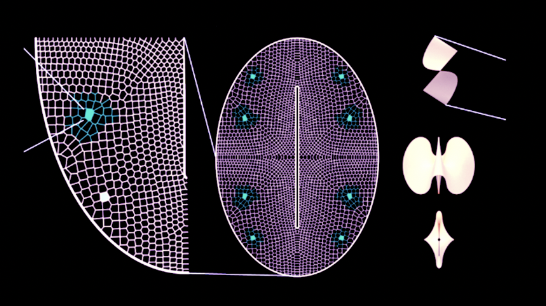
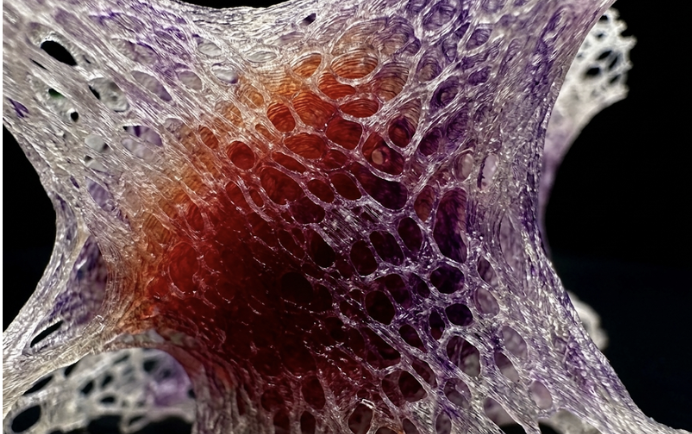

Air pollution is a pervasive oxidative stressor that disproportionately impacts marginalised urban populations, creating a silent crisis of environmental inequality. Current sensing infrastructure often lacks public visibility, leaving the physical toll of poor air quality abstract and unactionable for the communities most affected.
This conceptual project, ALVEOLI envisions a public art installation designed to create awareness and social engagement, addressing the short falls of how environmental data is communicated and experienced.
I aim to develop 3 bio-hybrid sculptures installed in 3 cities globally, translating invisible atmospheric pollutants into a high-contrast visual readout using engineered microbial biosensors.
Aim 1: Optimise and validate an E. coli extract-based cell-free protein synthesis (CFPS) system compatible with σ⁷⁰ bacterial promoters.
Optimise and validate an E. coli extract-based cell-free protein synthesis (CFPS) system using a constitutive σ⁷⁰ promoter–GFP reporter construct to confirm reliable transcription–translation performance and compatibility with downstream biosensor circuitry in a lyophilised CFPS format. Characterise expression kinetics, signal stability and reproducibility across extract batches to establish baseline system performance for zinc-responsive biosensor development.
Aim 2: Design and validate a zinc particulate responsive CFPS biosensor with colorimetric output
Design a zinc particulate-responsive biosensor based on the ZntR/PzntA regulatory system coupled to a colorimetric reporter compatible with E. coli extract-based CFPS.
Construct DNA sequences in Benchling and prepare constructs for synthesis via Twist Bioscience to confirm manufacturability and robustness of the genetic design.
Develop a lyophilisation protocol and test biosensor response under controlled laboratory conditions, characterising detection limits, signal stability, and colorimetric intensity.
Develop and evaluate a particulate capture, ion-release, and sensor activation workflow enabling conversion of PM2.5-associated zinc into detectable Zn²⁺ prior to CFPS activation, with consideration of the aesthetic and practical constraints of integration into a public artwork.
Aim 3: Integrate the zinc biosensor into a deployable sculptural sensing platform and evaluate performance under environmental conditions
Integrate the CFPS zinc biosensor into a 3D-printed sculptural sensing platform incorporating particulate capture, ion release and lyophilised reaction modules.
Evaluate biosensor performance under environmental conditions, including detection limits, response robustness and signal stability relative to environmental air sample processed in lab conditions.
Use experimental data to guide iterative optimisation strategies such as transcriptional signal amplification and the design of specific riboswitches to improve sensitivity to environmentally bioavailable concentrations of zinc particulate matter.
Finalise a comprehensive public installation proposal and prototype for bio-sensing sculptures in different cities, detailing sculpture placement, maintenance and bio-safety protocols and demonstrating the feasibility of integrating scientific bio-sensing with public art to make air pollution inequality visible and socially engaging.
Apply to funding bodies to pursue realisation of a global project.
Evaluate candidate pollutants based on:
Output: justification of final selection
Determine:
Output: selected sensing strategy compatible with CFPS transcription machinery
Establish:
Design validation experiment using a simple reporter construct to confirm:
Output: validate reliable CFPS system
Select based on:
Output: select sensing mechanism
Evaluate based on:
Output: selected reporting cassette
Determine
and draw out circuit logic.
Validate circuit design in silico prior to Twist order:
In Benchling:
Submit sequence to Twist Bioscience
Output: synthesis-ready circuit design
Test under controlled conditions:
Measure:
Evaluate compatibility with freeze-drying workflow.
Output: functional biosensor prototype
Design strategy for converting particulate-associated pollutants into detectable molecular form.
Evaluate:
Output: validated sample preparation workflow
Design integration strategy considering:
Output: deployable sensing sculpture prototype
Test system under real-world conditions:
Measure:
Output: field validation dataset
Use results to refine:
Output: second spiral biosensor design strategy
 Final Project Slide by Isobel Leonard
Final Project Slide by Isobel LeonardA biosensor for the simple, colour change based detection of salicylate and naphthalene in the environmental air.
Based on the methodology of Cho et al (2014) and Park et al (2005), using the NahR/Pr/Psal lower operon from Pseudomonas putida’s Napthalene degrading plasmid NH7 and connecting it to the LacZ reporter gene that produces red-ß-D-galactopyranoside, hydrolysed in CPRG to provide a clear visible signal. The operon uses a constitutive promoter Pr and inducible promoter Psal in opposite directions.
I am trying to create a similarly functioning plasmid in Benchling for an in Silico Aim 1 using the digest and ligation assembly tool.
Reference Paper for Methodology on plasmid construction:
Reference paper for Nah/Pr/Psal Operon:
Plasmid diagram from Cho et al 2014:
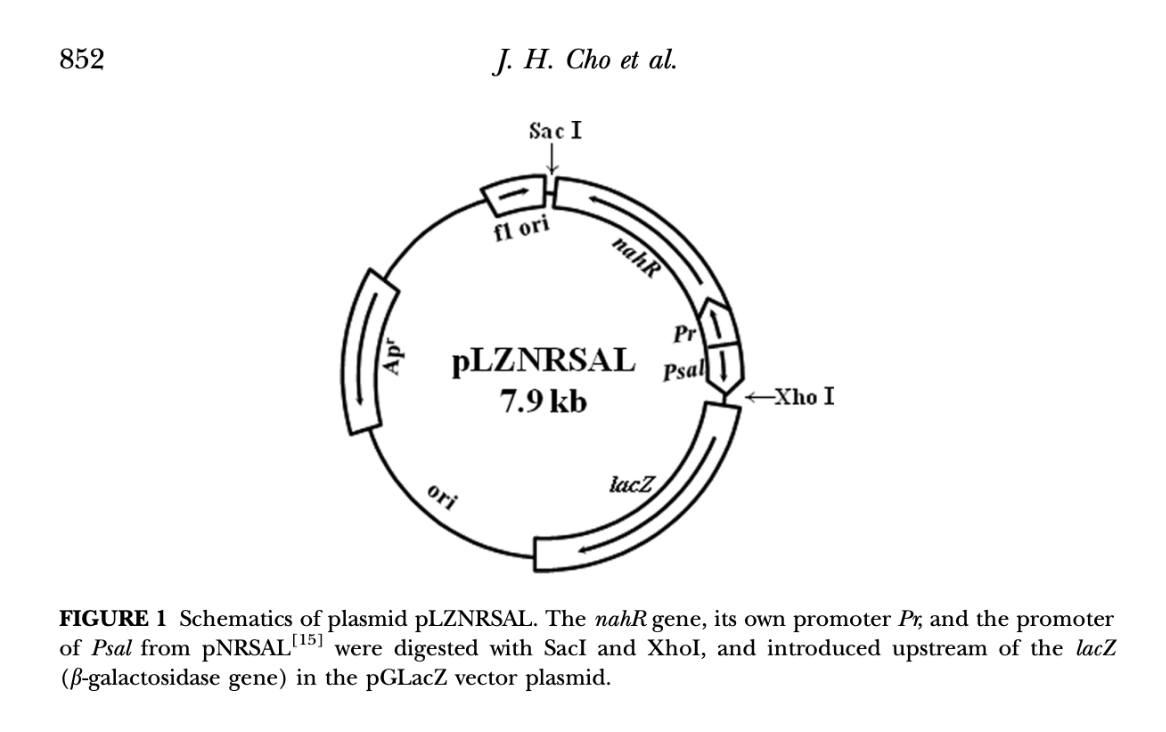

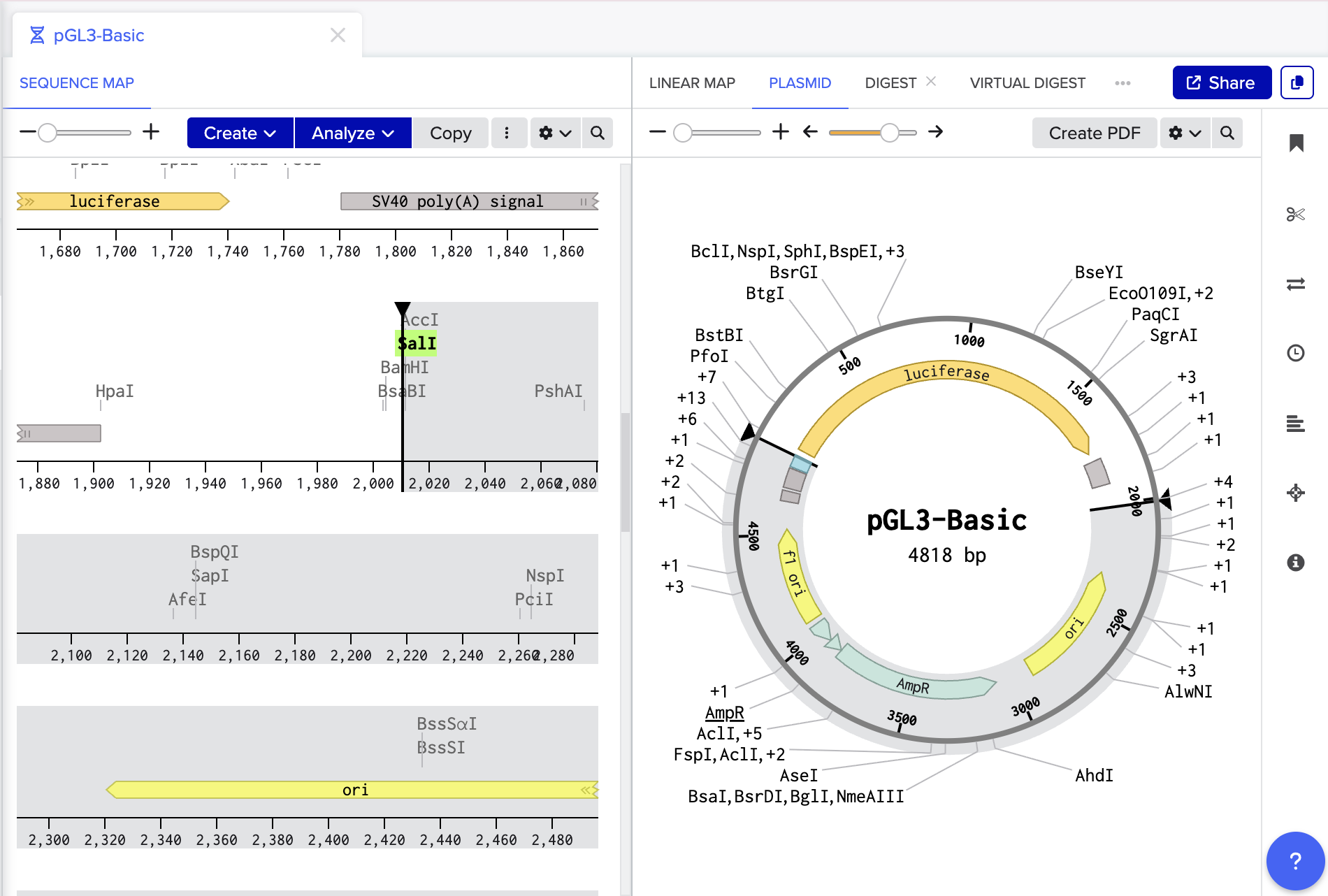
In my attempts to create a similar functioning plasmid in silico I have followed the following workflow and then met some problems!
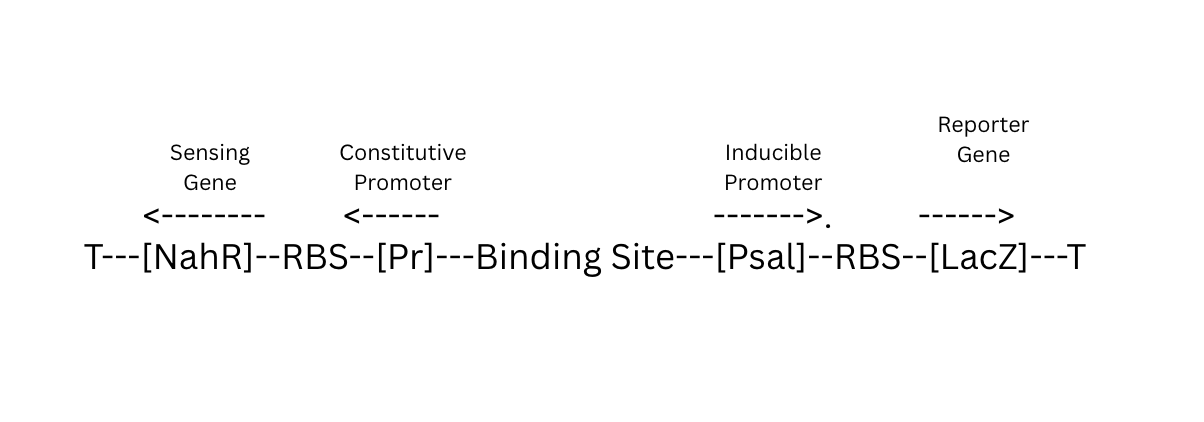


pGL3b basic (2.9kb) digested and linearised with SaII and HindII and Luciferase reporter gene removed.
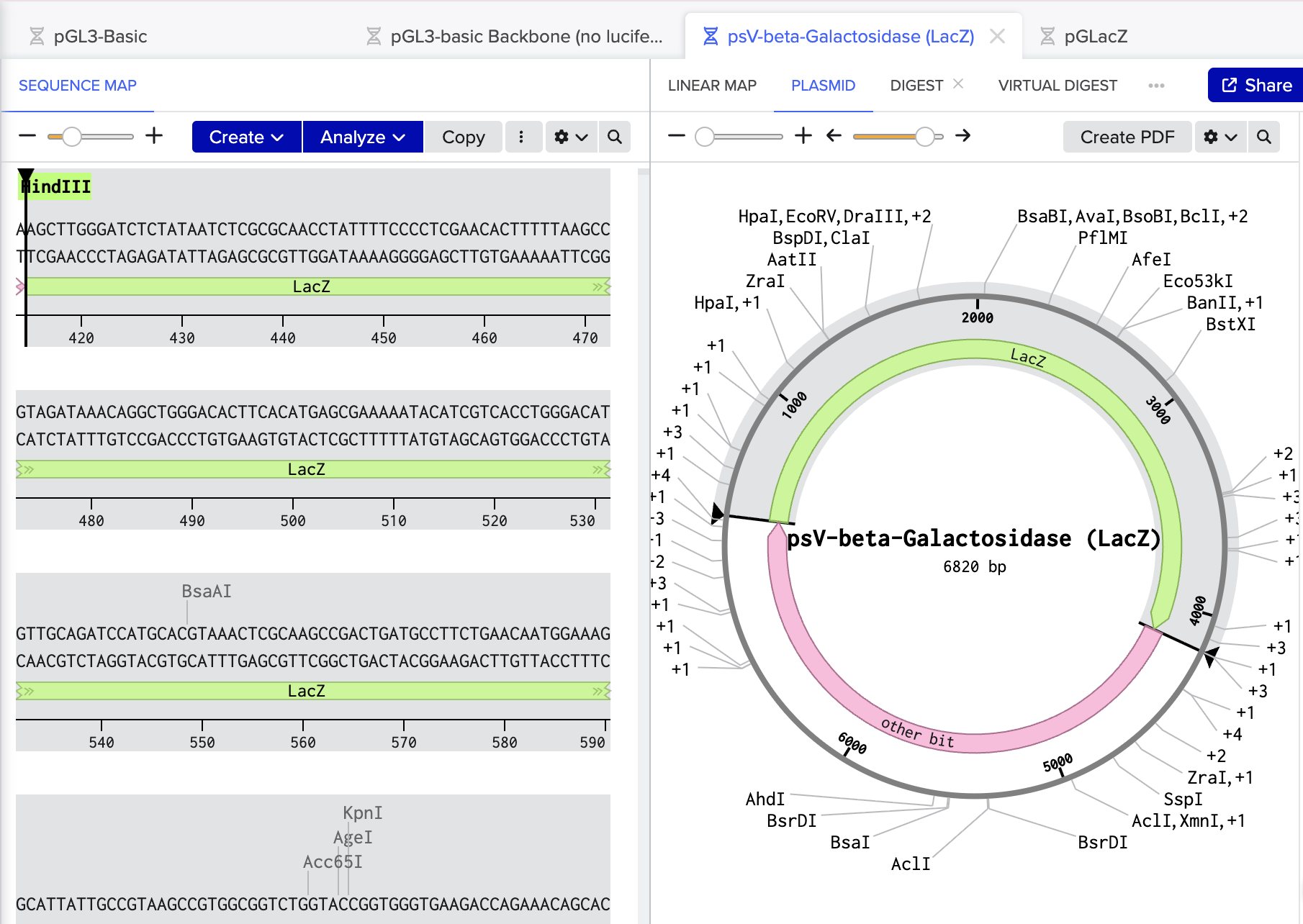
LacZ fragment from psV-beta-Galactosidase. (3.7 kb)
Cut out with restriction enzymes SaII and HindIII and ligated into pGL3b basic to make plasmid PGLacZ.
Containing nahR/Pr/Psal and I have found variants on the sequence located in these places:
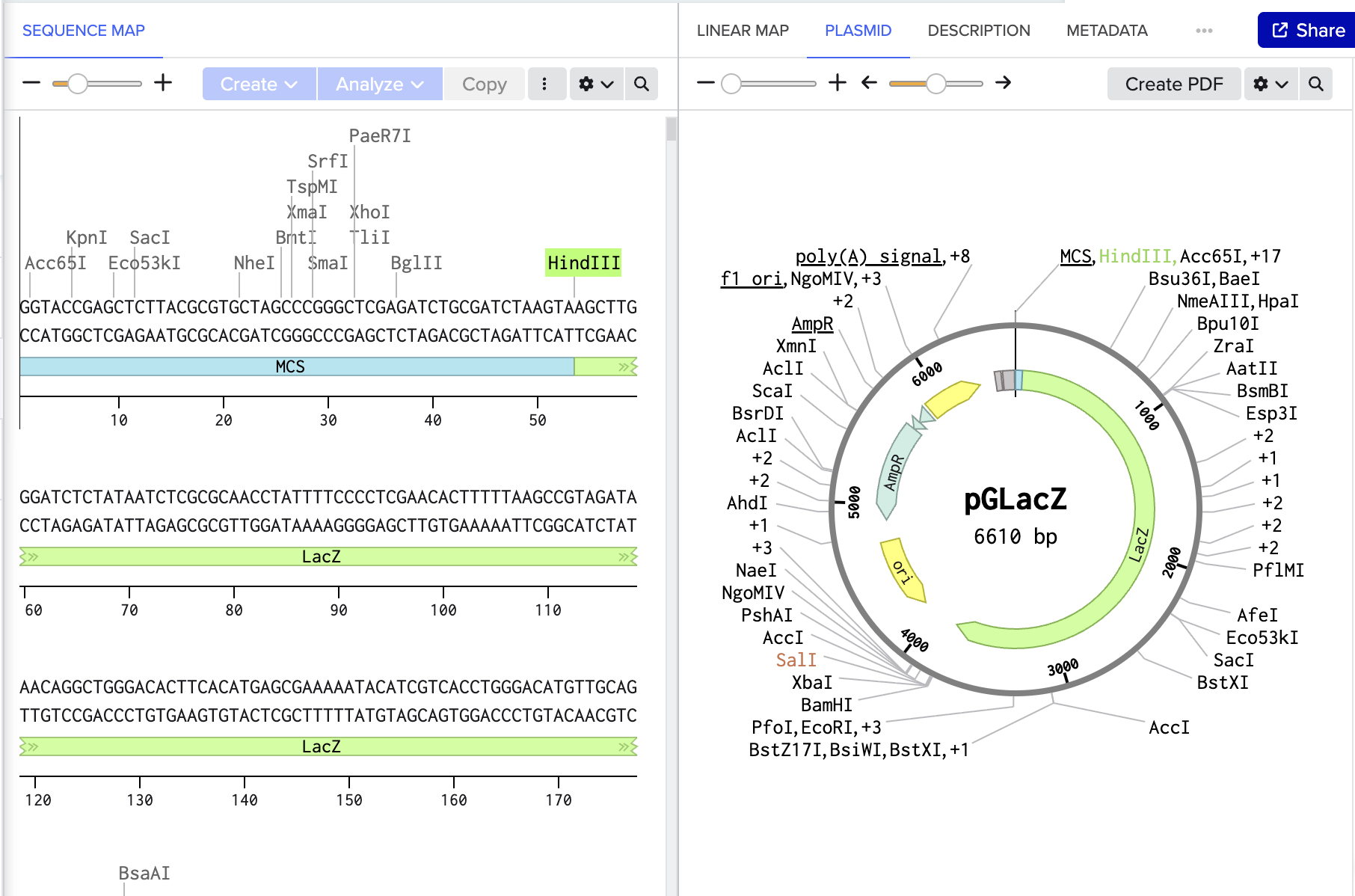
Design and add SacI and Xhol cut sites and ligated into plasmid PGLacZ.

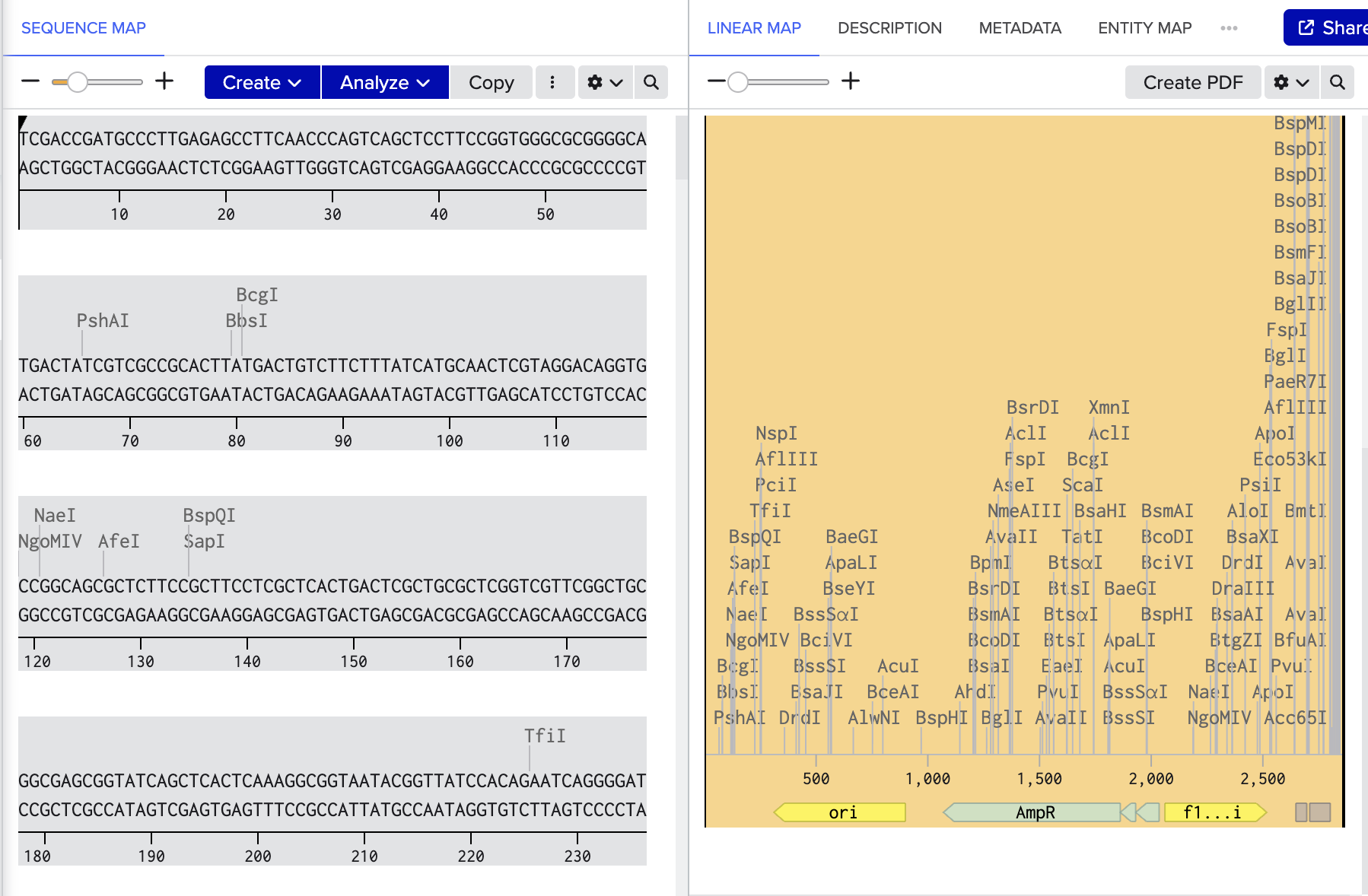

Benchling link: https://benchling.com/s/seq-kf6LTrbodved1t066n13?m=slm-UxQtbtTBRIF5SaInZm2z

Benchling link: https://benchling.com/s/seq-R5yEkXzifvA87rugfx2z?m=slm-7vqIszJaDNRN5P8LbkGW

Benchling link: https://benchling.com/s/seq-VPPhB7LPfBWE65dQCNoa?m=slm-IZFORq41ceeD4uLW8Cra
Versions of sequence I have found:
GenBank: AY294313
Here is where I have hit my problems
As there are no annotated and document sequences for each bit of the NahR/Pr/Psal operon e,g NahR gene, Pr Promoter, Psal promoter and the binding sites used, I don’t know which bit of this gene cluster from GenBank AY294313 is the bit I need.
In addition, I don’t know how to check all the bits I need for a functional sensor e.g RBS, Pr promoter, Psal Promoter are all included in the sequence (which they should be) and if they are the right way round and how to annotate them! I’ve been doing my head in a little trying to work it out!
In the Genbank info it says Gene NahR is 305-1207 and the next gene nahG starts at 1363. Meaning the promoters and binding sites must be between 1207 and 1363 but I am unsure how to identify?
I have tried my best to identify parts of the sequence in this benchling file by cross referencing between different versions of the Nah/Pr/Psal sequence across GenBank and iGEM but I don’t think I’m doing well and if this is a good approach:
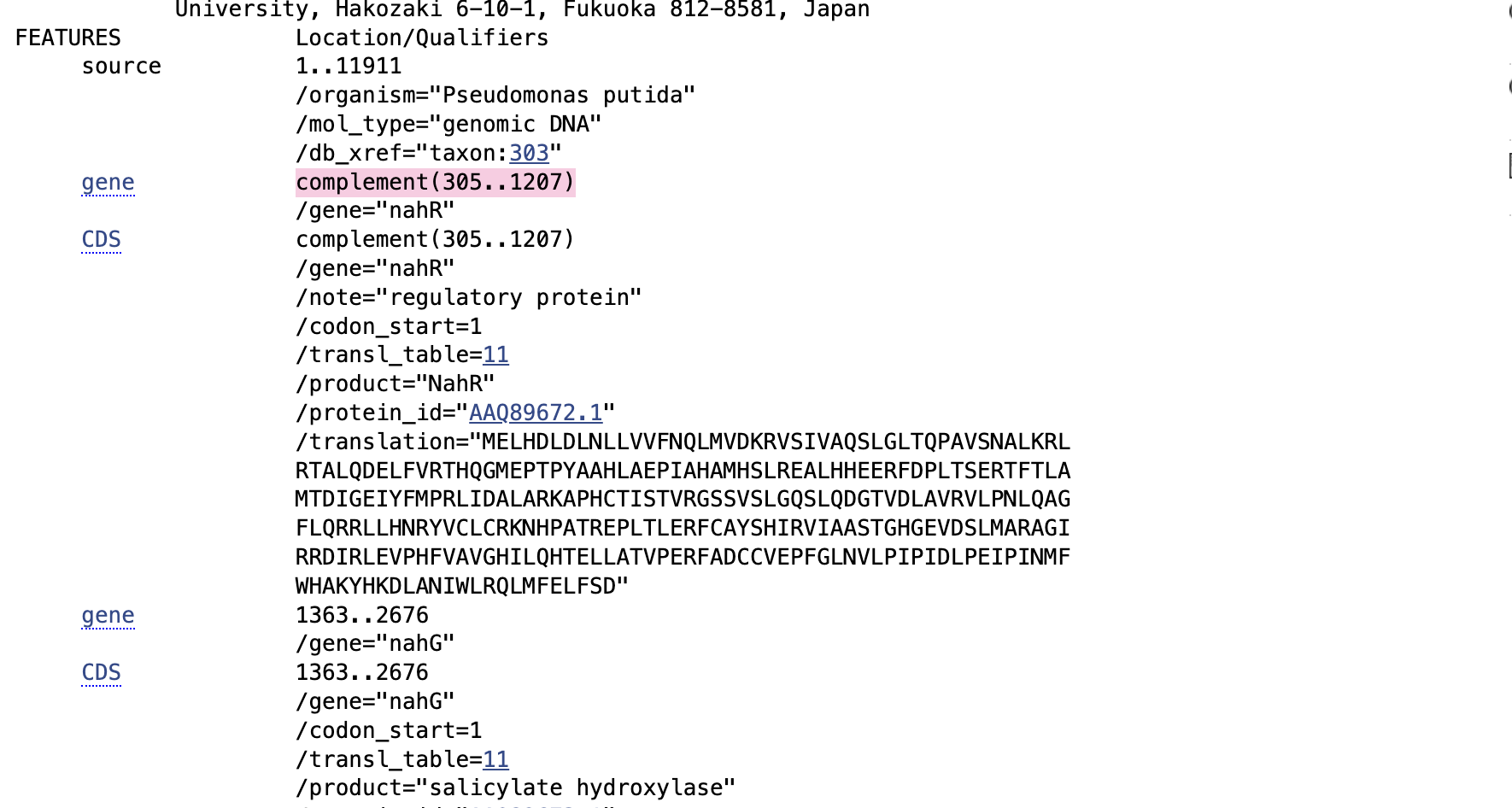

These two iGEM entries give a bit more info but are different from the GenBank sequence and I still don’t know how to tell where everything is??
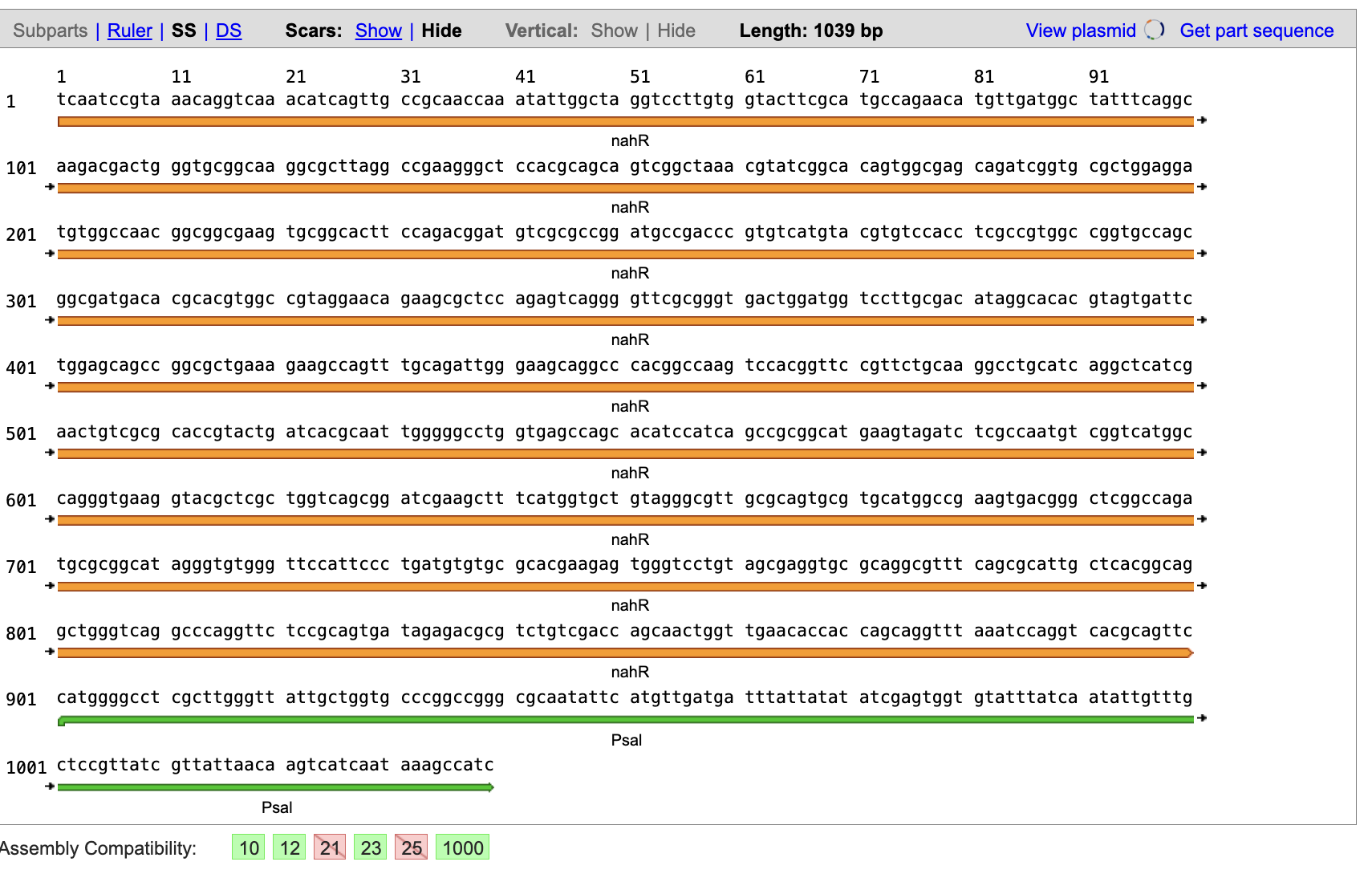



I have the same problem with the LacZ I digested from psV-beta-Galactosidase as although I can see the start and stop codon for the gene, I can’t tell if there is a RBS at the start of the sequence and which one it is?
I don’t know if the problem is that I am taking pieces from different vectors and instead I should try and build the sequence blocks myself with e.g Promoter, RBS, start codon, CDS etc. However, I still have the problem that I am unable to find the sequence for NahR, Pr or Psal promoter within the gene clusters and I wouldn’t know what RBS to use for the LacZ gene.
I have noticed (which is not documented in the paper I am following) that I have 2 SacI cut sites in my new plasmid PGLacZ, because the LacZ I have taken and ligated in has another SacI cut site in it. Meaning that if I were to digest at Xhol and SacI to insert the NahR/Pr/Psal region as described I would get another cut in LacZ.
However, I reason that I can just digest and ligate the pieces in a different order. First digest at Xhol and SacI while the LacZ is not in the backbone. Then digest at HindIII and SalI to remove luciferase and ligate in LacZ.
Is this okay to do??? Or is it better to use Gibson Assembly?? If so how do I design the overlaps for NahR/Pr/Psal so it goes specifically where it needs to go upstream of LacZ.
I am looking into the possibility of innovating with the biosensor by creating a protocol to use the plasmids in a cell free system to create a safer and more stable bio-sensing public sculptures.
I am interested in merging the research I have already done with the research we learnt about in Cell free systems week from these two papers:
Where the possibility of cell free biosensors being 3D printed ina biopolymer matrix for architectural structures is discussed.
Where a cell free system with LacZ and CPRG was successfully designed as a wearable colourmetric bio-sensor.
However, I am a bit stumped on how to create a plasmid design for a cell free system and if the promoters I am using for the NahR operon are compatible with a cell free system as they are from Pseudomonas putida not Ecoli (would it work with Ecoli RNA polymerase or is there such thing as using Pseudomonas putida RNA polymerase in CFPS?). Is this a possible next step or is my goal not compatible?? Is there a way to optimise what I am doing so it would work??
Thank you if you read this far!! :)