Part 1: Benchling & In-silico Gel Art
Part 1: Benchling & In-silico Gel Art
Simulated restriction enzyme digestion with the seven enzymes specified in this week’s lab protocol: SalI, SacI, EcoRV, KpnI, BamHI, HindIII, and EcoRI. Used both the DNA Gel Art Interface (λ DNA) and Benchling (lambda phage genome NC_001416) to visualize digest patterns and verify cut-site predictions.
Lab protocol: Gel Art: Restriction Digests and Gel Electrophoresis
Benchling Digest — NC_001416 (Lambda Phage Genome)
Sequence: NC_001416 — Escherichia phage lambda, 48,502 bp (linear).
Benchling digest link: NC_001416 Digest — Benchling
Proof of Work — Screenshots
1. DNA Gel Art Interface — λ DNA Restriction Digests
Simulated gel electrophoresis using the DNA Gel Art tool. λ DNA was digested with various enzyme combinations (EcoRV + SacI, HindIII + PvuII, NdeI + SalI, etc.) across lanes 2–10. The table documents water, CutSmart buffer, λ DNA, and enzyme volumes per lane.


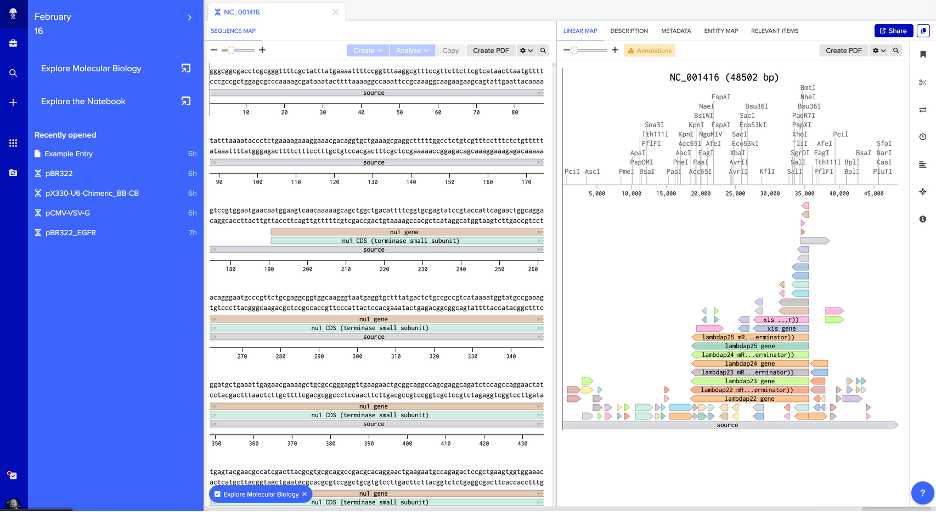
2. Benchling — NC_001416 Sequence Map with Restriction Sites
Linear map of NC_001416 in Benchling showing the raw sequence, annotated genetic features (e.g., xis, nul, lambdap genes), and restriction enzyme cut sites (PciI, AscI, PmeI, BsaI, KpnI, SacI, SalI, and others) along the 48.5 kb genome.

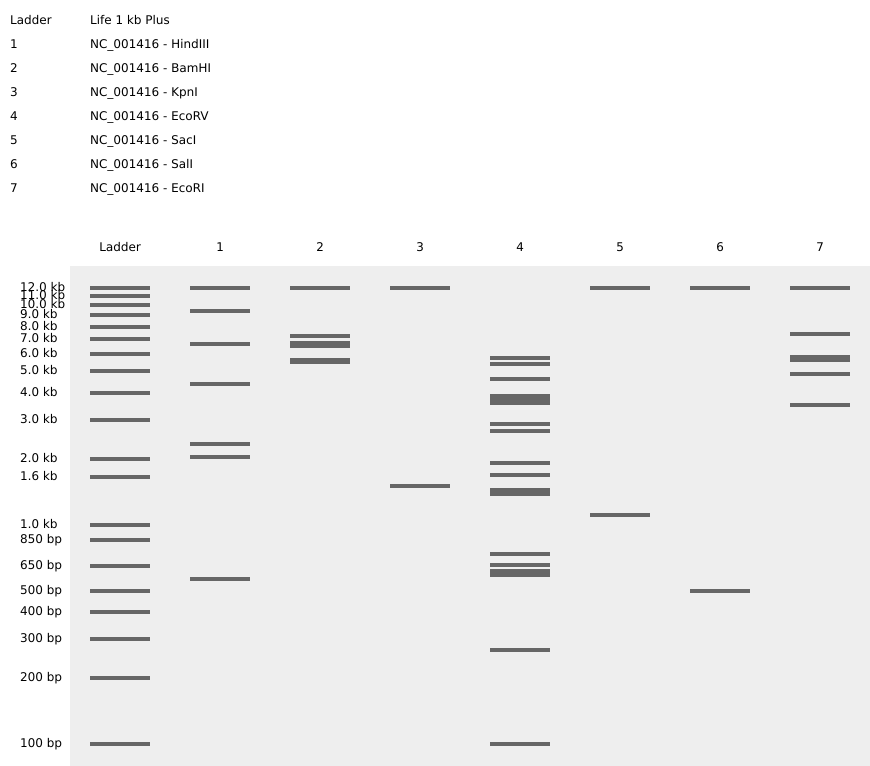
3. Virtual Digest Gel — NC_001416 with All Seven Required Enzymes
Simulated gel (Life 1 kb Plus ladder) showing digest results for NC_001416 with each of the seven required enzymes:
| Lane | Enzyme | Fragment pattern |
|---|---|---|
| 1 | HindIII | 3 bands (~11 kb, ~6.5 kb, ~2.1 kb) |
| 2 | BamHI | 3 bands (~11.5 kb, ~7 kb, ~5.8 kb) |
| 3 | KpnI | 2 bands (~12 kb, ~1.7 kb) |
| 4 | EcoRV | Multiple bands (many cut sites) |
| 5 | SacI | 2 bands (~11.5 kb, ~1 kb) |
| 6 | SalI | 2 bands (~11.5 kb, ~550 bp) |
| 7 | EcoRI | Multiple bands (~12 kb, ~9.5 kb, ~8.5 kb, ~7.5 kb, ~6 kb, ~3.5 kb) |


Enzymes Simulated
| Enzyme | Recognition site | Notes |
|---|---|---|
| SalI | G^TCGAC | 6-cutter |
| SacI | GAGCT^C | 6-cutter |
| EcoRV | GAT^ATC | 6-cutter, blunt |
| KpnI | GGTAC^C | 6-cutter |
| BamHI | G^GATCC | 6-cutter |
| HindIII | A^AGCTT | 6-cutter |
| EcoRI | G^AATTC | 6-cutter |