Part 4: Prepare a Twist DNA Synthesis Order
Part 4: Prepare a Twist DNA Synthesis Order
Practice exercise — building an sfGFP expression cassette in Benchling, preparing a mock Twist order, and annotating the plasmid.
4.1–4.2 Accounts & Build Your DNA Insert Sequence
Created Twist and Benchling accounts. Built the sfGFP expression cassette in Benchling with annotated parts:
- Promoter (BBa_J23106)
- RBS (BBa_B0034)
- Start codon (ATG)
- Coding sequence (codon-optimized sfGFP from Part 3)
- 7× His tag
- Stop codon (TAA)
- Terminator (BBa_B0015)
Proof of Annotation in Benchling
Benchling sequence link: sfGFP_expression_cassette · Benchling
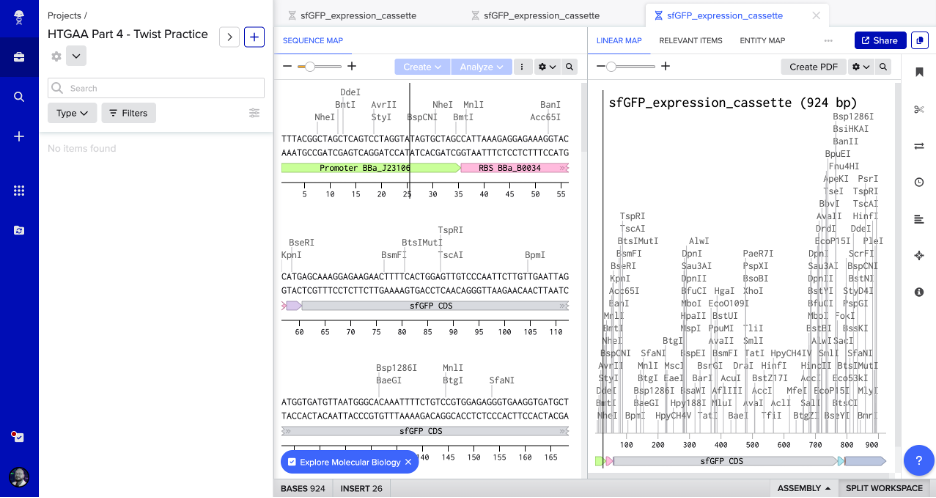
Screenshot: Annotated Sequence Map in Benchling
The sequence map shows the sfGFP expression cassette (924 bp) with promoter, RBS, and sfGFP CDS annotated, plus restriction enzyme cut sites.

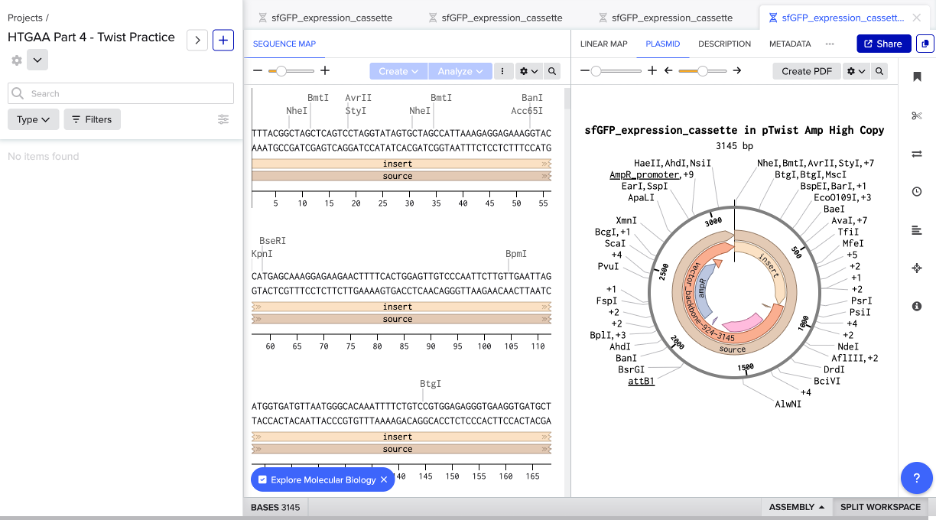
Screenshot: Circular Plasmid Map (sfGFP in pTwist Amp High Copy)
The full construct (3145 bp) in pTwist Amp High Copy, with insert, source, AmpR promoter, and vector backbone annotated.
Note: The color choices for the plasmid annotations are a reflection of my cringe-worthy color skills — consider yourself warned.

4.3–4.6 Twist Order Flow
- Selected Genes → Clonal Genes on Twist
- Uploaded FASTA (sfGFP expression cassette)
- Chose vector: pTwist Amp High Copy from Twist Vector Catalog
- Downloaded GenBank construct and imported into Benchling
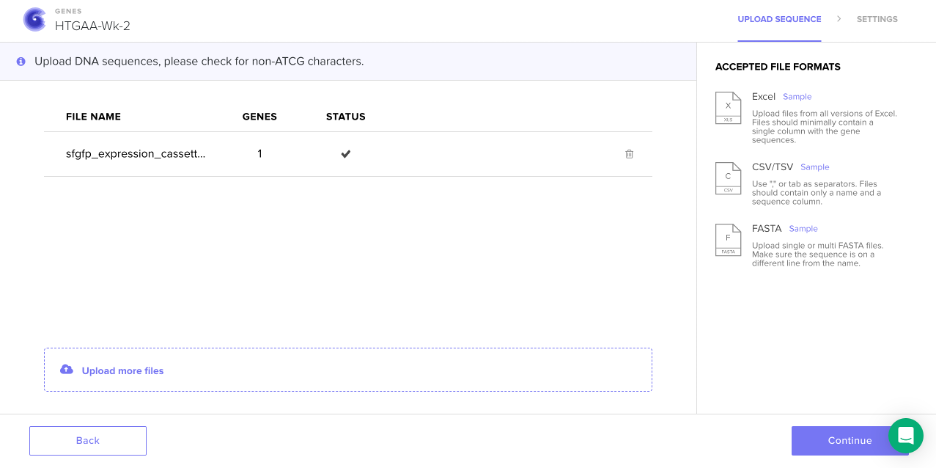
Screenshot: Sequence Upload to Twist

Design Notes: Manual vs. Programmatic
Efficiency: Designing expression cassettes and plasmids can be far more efficient with Python and/or R — tools like DNA Chisel, PyDNA, or SynBioHub enable scripted design, validation, and export. Batch operations, automated codon optimization, and constraint checking become straightforward.
Learning value: Building the construct manually in Benchling — clicking through each part, copying sequences, and annotating by hand — offers a different kind of learning. You develop intuition for how promoters, RBSs, and CDSs fit together, where restriction sites fall, and what the plasmid “looks like” at each step. That tactile understanding is harder to get from a script. For a first expression cassette, the manual approach is worth the extra time.
Documented Deliverables
| Item | Status |
|---|---|
| Desired Twist cloning vector | pTwist Amp High Copy |
| Fully annotated Benchling insert fragment | sfGFP_expression_cassette |
| GenBank construct imported | ✓ |