Final Project Selected: Boltr
Final Project Selected: Boltr
In short: I am building a structure predictor aimed at outperforming AlphaFold on both speed and accuracy in dynamic regions—loops, hinges, and other flexible or poorly ordered stretches where rigid single-state models weaken.
Boltr is a Rust-native biomolecular inference platform for reproducible structure prediction—a Rust implementation of Boltz2-style workflows. It targets transparent, modular pipelines with YAML inputs, Boltz-compatible preprocessing, and LibTorch-backed inference, producing structural predictions in mmCIF and PDB formats.
Developed to support reproducible protein structure prediction in synthetic biology, Boltr addresses limitations of complex Python-heavy modeling environments by asking whether a Rust-based system can deliver useful structural predictions while improving transparency and maintainability.
Repository: github.com/SampleBias/Boltr


Project aims
Aim 1 — Experimental aim: Validate Boltr for biomolecular prediction using YAML inputs, Boltz-compatible preprocessing, and LibTorch-backed inference; generate usable structural predictions for protein or protein–peptide systems in mmCIF and PDB.
Aim 2 — Development aim: Expand beyond initial validation toward stronger numerical parity, reliability, and usability—preprocessing, confidence scoring, and benchmarking so Boltr can serve as a robust platform for protein design.
Aim 3 — Vision aim: Evolve toward a fully independent, Rust-native prediction engine using the Burn ML library, replacing Boltz-derived dependencies with a standalone architecture for transparent, scalable scientific inference.
James Utley, PhD — Quito Node, Ecuador (Committed Listener) · james.utley@syndicate-labs.io
Not selected for final project — Gumol Microdrop (click to expand)
Gumol Microdrop (archived draft)
Integrating molecular dynamics simulation, engineered MSC antioxidant systems, and programmable microfluidic experimentation for oxidative stress validation.
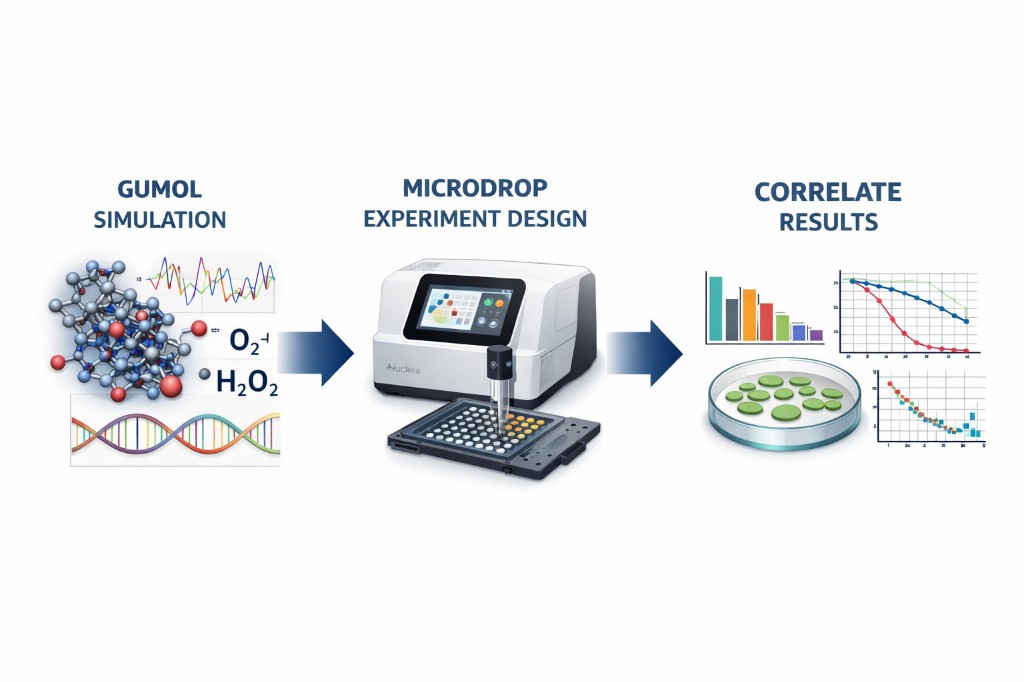

MicroDrop is a design companion application written entirely in Rust that bridges the Gumol Simulation engine with the Nuclera digital microfluidic platform. It enables researchers to design, configure, and execute validation wet-lab experiments that directly test computational predictions from Gumol molecular dynamics simulations.
📱 Note: The video renders better on mobile. Draft 2 will be improved.
MicroDrop application workflow
The application guides users through a seven-step workflow from experiment design to data correlation. Key screens include:
2. Card Editor — Experiment Design — Map experimental parameters (oxidant concentrations, antioxidant types, exposure times) onto a physical 96-well layout for microfluidic testing.
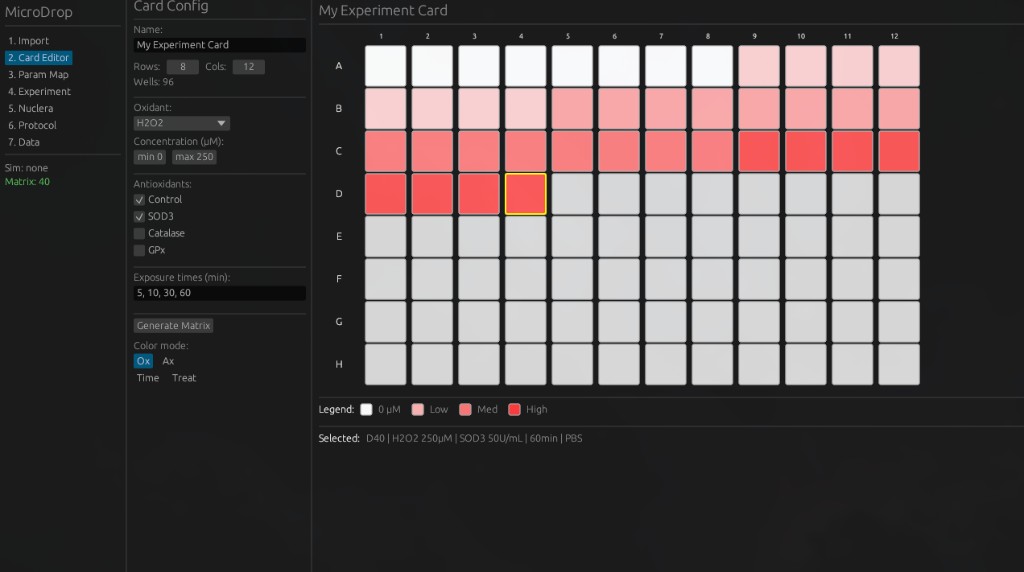

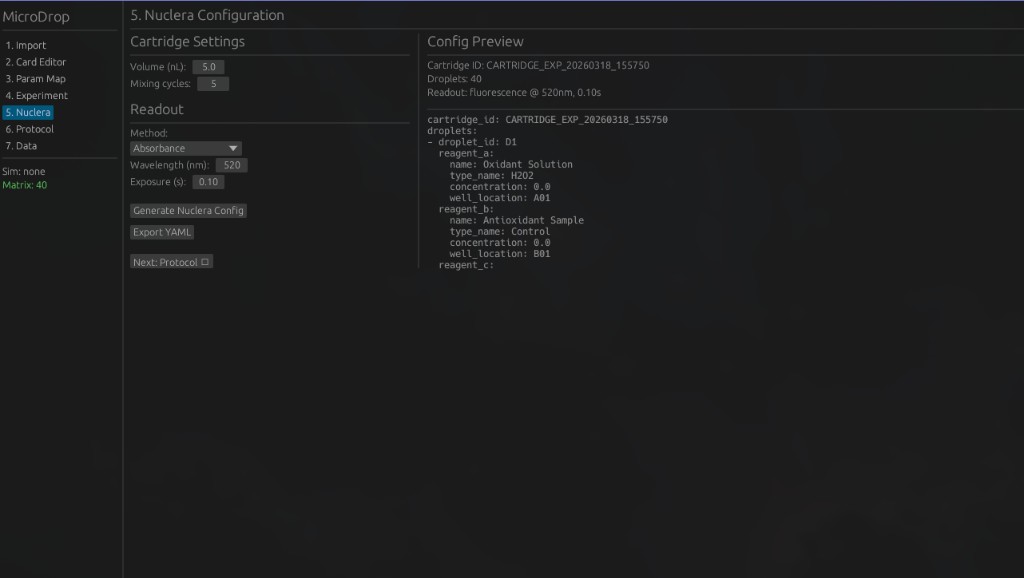
5. Nuclera Configuration — Configure cartridge settings and readout parameters for the Nuclera eProtein Discovery system.

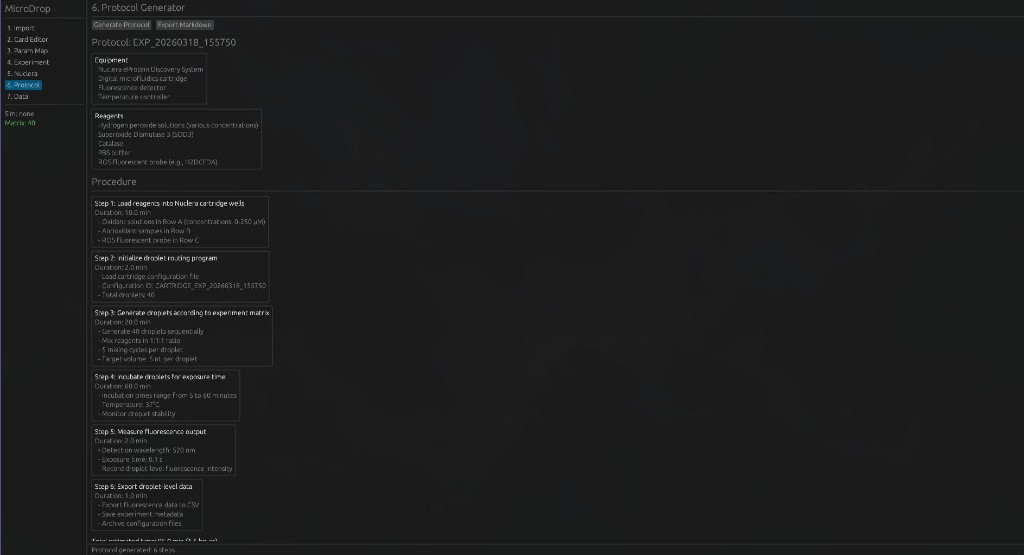
6. Protocol Generator — Generate executable wet-lab protocols with equipment lists and step-by-step procedures.

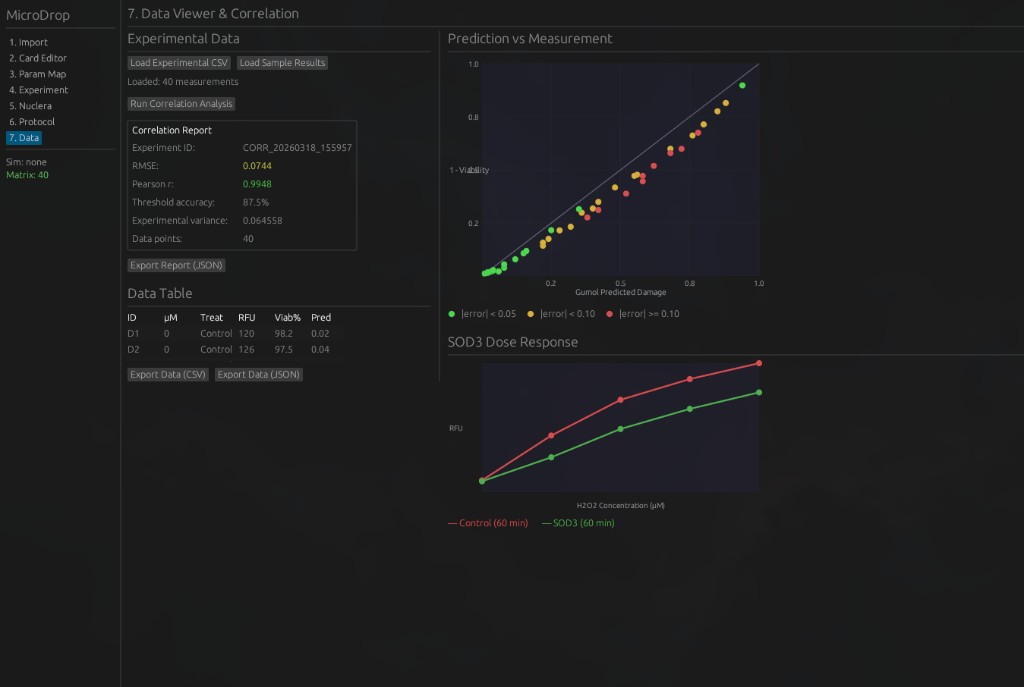
7. Data Viewer & Correlation — Load experimental CSV data, run correlation analysis against Gumol predictions, and visualize results.

Section 1: Background & rationale
Human exploration of deep space presents substantial biological challenges, particularly due to prolonged exposure to ionizing radiation and the resulting oxidative stress experienced by biological tissues. Outside Earth’s magnetosphere, astronauts are exposed to galactic cosmic radiation (GCR) and solar particle events (SPE), which generate high levels of reactive oxygen species (ROS) within biological systems. These reactive molecules—including superoxide radicals (O₂⁻), hydrogen peroxide (H₂O₂), and hydroxyl radicals—can damage DNA, proteins, lipids, and mitochondrial structures, ultimately leading to cellular dysfunction, senescence, and increased cancer risk. The mitigation of oxidative damage therefore represents a critical problem in human spaceflight and long-duration missions to destinations such as Mars.
Cells possess endogenous antioxidant defense systems that neutralize reactive oxygen species and maintain redox homeostasis. One of the key enzymes involved in this defense is extracellular superoxide dismutase (ECSOD), also known as SOD3, which catalyzes the conversion of superoxide radicals into hydrogen peroxide and molecular oxygen in extracellular environments. ECSOD plays an important role in protecting tissues from oxidative injury by regulating extracellular redox balance and preventing radical propagation. Studies have demonstrated that increased expression of ECSOD can significantly reduce oxidative damage in multiple tissues and disease models associated with inflammation, ischemia, and radiation exposure.
Mesenchymal stem cells (MSCs) represent a promising biological platform for delivering therapeutic antioxidant activity because of their capacity for secretion of protective factors and their ability to modulate tissue microenvironments. Engineering MSCs to overexpress ECSOD could potentially enhance extracellular antioxidant capacity and create a biologically active defense against ROS accumulation. Such engineered MSC systems could serve as a biological countermeasure against oxidative stress encountered in extreme environments such as deep-space radiation fields.
While biological antioxidant strategies are promising, predicting how these systems behave under complex oxidative environments remains challenging. Reactive oxygen species interact through nonlinear reaction–diffusion processes that depend on radical concentrations, diffusion coefficients, enzymatic kinetics, and environmental conditions. Molecular dynamics (MD) simulations offer a powerful computational approach to modeling these processes at high temporal and spatial resolution. By simulating radical diffusion, reaction kinetics, and antioxidant enzyme interactions, MD systems can generate predictions regarding oxidative burden and the effectiveness of ROS neutralization mechanisms.
However, computational predictions must ultimately be validated experimentally. Traditional biochemical assays often lack the throughput and environmental control required to systematically test large numbers of oxidative stress conditions. Microfluidic systems provide a promising alternative by enabling the creation of precisely controlled microscale reaction environments. Digital microfluidic platforms, such as the Nuclera eProtein Discovery system, allow programmable generation and manipulation of microdroplets containing defined reagent mixtures. These droplets can function as discrete experimental microreactors where oxidative stress reactions and antioxidant activity can be quantified.
Recent research demonstrates that microfluidic droplet systems can be used to study biochemical reactions, enzyme kinetics, and cellular stress responses in highly parallelized experimental formats. Because each droplet can represent a unique experimental condition, microfluidics enables systematic exploration of large parameter spaces with minimal reagent consumption. When combined with fluorescence-based reporter systems for detecting reactive oxygen species, microdroplet assays can provide quantitative measurements of oxidative dynamics in controlled environments.
The integration of molecular simulation with programmable microfluidic experimentation represents an emerging paradigm in synthetic biology and systems biology. In such frameworks, computational models generate predictions regarding biological behavior, which are then rapidly tested using automated experimental platforms. The resulting data can be used to refine simulation parameters and improve predictive accuracy, creating a closed loop between modeling and experimentation.
The present project aims to establish such a pipeline by integrating Gumol molecular dynamics simulations, engineered MSC antioxidant biology, and Nuclera microdroplet assays. By modeling ROS behavior computationally and experimentally testing those predictions using high-throughput microfluidic experiments, the project seeks to create a framework for evaluating antioxidant countermeasures relevant to space radiation environments.
Novelty and innovation: This project introduces a novel integration of molecular dynamics modeling, engineered stem cell antioxidant systems, and programmable microfluidic experimentation. While oxidative stress biology and ROS chemistry are well studied independently, the direct coupling of molecular simulations with automated microdroplet validation experiments remains relatively unexplored.
Why this project matters: Understanding how biological systems respond to oxidative stress is critical for medicine, aging research, and space exploration. Microfluidic technologies enable highly parallelized experiments at small scales; validating computational predictions accelerates systems biology.
Section 2: Project aims (Gumol Microdrop)
Aim 1 — Experimental aim: Experimentally validate molecular dynamics simulations of ROS neutralization mediated by ECSOD/SOD3 using a programmable microdroplet assay on the Nuclera digital microfluidic platform.
Aim 2 — Development aim: Develop an automated pipeline that converts molecular dynamics simulation outputs into experimentally executable microdroplet assay designs for high-throughput oxidative stress testing.
Aim 3 — Visionary aim: Establish a simulation-driven experimental platform for evaluating biological countermeasures against oxidative damage in extreme radiation environments, including deep-space missions to Mars and beyond.
Archived draft — not selected as final project.