Week 2 HW: DNA- Read, Write and Edit


Part 1: In Silico Gel Art
-Import the Lambda DNA.
https://www.neb.com/en/products/n3011-lambda-dna
-Simulate Restriction Enzyme Digestion
-Create a pattern/image in the style of Paul Vanouse’s Latent Figure Protocol artworks
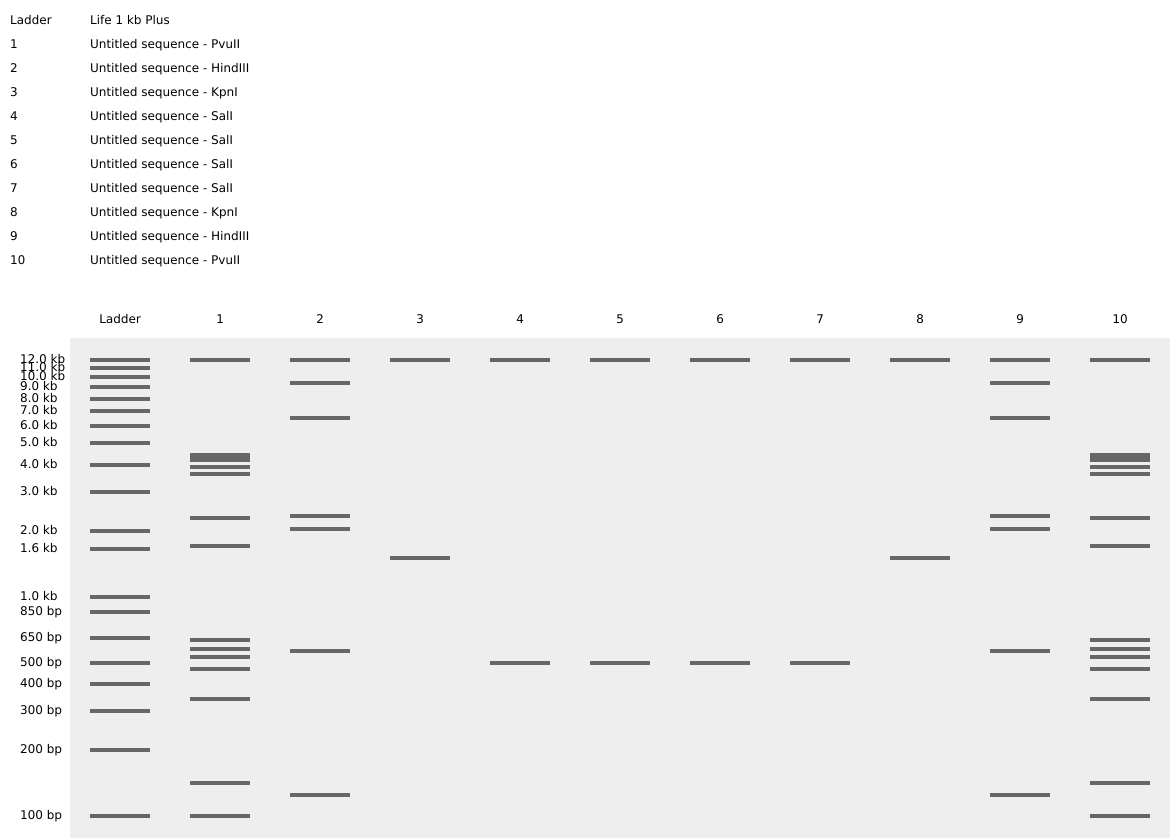


While looking the patterns with the different restriction enzymes, I try to create the top part of a cats siluette, basically just its ears. I used the patterns of the digestion of enzymes: Pvull, Hindll, kpnl, SalI.
Part 3: DNA Design Challenge
Choose a protein: INTERLEUKIN-10
MHSSALLCCLVLLTGVRASPGQGTQSENSCTHFPGNLPNMLRDLRDAFSRVKTFFQMKDQLDNLLLKESLLEDFKGYLGCQALSEMIQFYLEEVMPQAENQDPDIKAHVNSLGENLKTLRLRLRRCHRFLPCENKSKAVEQVKNAFNKLQEKGIYKAMSEFDIFINYIEAYMTMKIRN
Sequence obtain from UniProt: https://www.uniprot.org/uniprotkb/P22301/entry#sequences
atgcatagcagcgcgctgctgtgctgcctggtgctgctgaccggcgtgcgcgcgagcccg ggccagggcacccagagcgaaaacagctgcacccattttccgggcaacctgccgaacatg ctgcgcgatctgcgcgatgcgtttagccgcgtgaaaaccttttttcagatgaaagatcag ctggataacctgctgctgaaagaaagcctgctggaagattttaaaggctatctgggctgc caggcgctgagcgaaatgattcagttttatctggaagaagtgatgccgcaggcggaaaac caggatccggatattaaagcgcatgtgaacagcctgggcgaaaacctgaaaaccctgcgc ctgcgcctgcgccgctgccatcgctttctgccgtgcgaaaacaaaagcaaagcggtggaa caggtgaaaaacgcgtttaacaaactgcaggaaaaaggcatttataaagcgatgagcgaa tttgatatttttattaactatattgaagcgtatatgaccatgaaaattcgcaac
Tool used: https://www.genecorner.ugent.be/rev_trans.html
ATGCATTCGAGCGCGCTGCTGTGCTGCCTGGTGCTGCTGACCGGCGTGCGCGCATCTCCGGGCCAGGGTACCCAGAGTGAAAACAGCTGCACCCATTTTCCGGGCAATCTGCCGAACATGCTGCGCGATCTGCGTGATGCCTTTAGCCGTGTGAAAACCTTCTTTCAGATGAAAGATCAGCTGGATAACCTGCTGCTGAAAGAAAGCCTGCTGGAAGATTTTAAAGGCTACCTGGGCTGCCAGGCCCTGAGCGAAATGATTCAATTTTATCTGGAAGAAGTGATGCCGCAGGCCGAAAATCAGGATCCGGACATTAAAGCGCATGTGAACAGCCTGGGCGAAAACCTGAAAACCCTGCGCCTGCGCCTGCGCCGTTGCCATCGCTTTCTGCCGTGCGAAAACAAAAGCAAAGCCGTGGAACAGGTGAAAAACGCGTTTAACAAACTGCAGGAAAAAGGCATTTACAAAGCGATGAGCGAATTTGATATTTTTATTAATTACATTGAAGCGTATATGACCATGAAAATTCGCAAC
Tool used:https://en.vectorbuilder.com/tool/codon-optimization.html
Which protein have you chosen and why?
While looking into the possibilites of the use of bacteria as drug delivery systems, I saw the concept of designing bacteria with the capability of secreting anti-inflammatory proteins. I chose Interleukin-10, as its a anti-inflammatory cytokine that usually is secreted by immune cells and has a role in reducing the inflammation in tissues. I used this week assignment as a way to explore the possiblity of engineering bacteria to deliver this therapeutic protein, choosing this protein sequence and optimizing it to E.coli as a trial to see the efficacy of this concept.
In your own words, describe why you need to optimize codon usage. Which organism have you chosen to optimize the codon sequence for and why?
The genetic code is degenerate, which means most aminoacids could be encoded by more than one codon. This makes it possible for organisms to have preferences with codon for the expression of aminoacids, creating a bias. To maximaize the protein production, we need to work according to what the host prefers, improving the trnaslation efficiency and reducing errors. I chose to directly optimized for E.coli, as sequence is more a trial to see if the expression of the protein through out the bacteria is possible. Choosing E.coli simplifies the design and clonign workflow, letting us to conceptially see if the therapeutic delivery is possible.
What technologies could be used to produce this protein from your DNA? Describe in your words the DNA sequence can be transcribed and translated into your protein. You may describe either cell-dependent or cell-free methods, or both.
Some tehcnologies we could use to optain the protein from the DNA, is cell dependent expression. With this technique the IL-10 gene optimized for E.coli, gets used by the bacterial transcription machinery, producing mRNA that gets bound and translated giving us the end product of IL-10.
Part 4. Build Your DNA Sequence
https://benchling.com/s/seq-5lw1UIOlQAZWLoQm6VbH?m=slm-SjnrMfysVLzz1ixK6UM9
BBa_J23106
tttacggctagctcagtcctaggtatagtgctagc
https://parts.igem.org/wiki/index.php?title=Part:BBa_J23106
BBa_B0034
aaagaggagaaa
https://parts.igem.org/Part:BBa_B0034
pelB Signal
ATGAAATACCTATTGCCTACGGCAGCCGCTGGATTGTTATTACTCGCGGCCCAGCCGGCCATGGCG
https://parts.igem.org/wiki/index.php?title=Part:BBa_J32015
Why?
I decided to include the pelB signal in the Il-10 seuqnece so the protein when it is translated, it can be directed to the periplasmic space where it can be more easily secreted.
ATGCATTCGAGCGCGCTGCTGTGCTGCCTGGTGCTGCTGACCGGCGTGCGCGCATCTCCGGGCCAGGGTACCCAGAGTGAAAACAGCTGCACCCATTTTCCGGGCAATCTGCCGAACATGCTGCGCGATCTGCGTGATGCCTTTAGCCGTGTGAAAACCTTCTTTCAGATGAAAGATCAGCTGGATAACCTGCTGCTGAAAGAAAGCCTGCTGGAAGATTTTAAAGGCTACCTGGGCTGCCAGGCCCTGAGCGAAATGATTCAATTTTATCTGGAAGAAGTGATGCCGCAGGCCGAAAATCAGGATCCGGACATTAAAGCGCATGTGAACAGCCTGGGCGAAAACCTGAAAACCCTGCGCCTGCGCCTGCGCCGTTGCCATCGCTTTCTGCCGTGCGAAAACAAAAGCAAAGCCGTGGAACAGGTGAAAAACGCGTTTAACAAACTGCAGGAAAAAGGCATTTACAAAGCGATGAGCGAATTTGATATTTTTATTAATTACATTGAAGCGTATATGACCATGAAAATTCGCAAC
Tool used:https://en.vectorbuilder.com/tool/codon-optimization.html
CATCACCATCACCATCATCAC
Why?
The His-tag gets add into the c terminus of the protein, and it helps if u want to purify the protein when we are testing the overall sequence.
BBa_B0015
ccaggcatcaaataaaacgaaaggctcagtcgaaagactgggcctttcgttttatctgttgtttgtcggtgaacgctctctactagagtcacactggctcaccttcgggtgggcctttctgcgtttata
https://parts.igem.org/Part:BBa_B001
DNA SEQ:
[Promoter] → [RBS] → ATG (Start Codon) → IL-10 Coding Sequence → Stop Codon (TAA) → [Terminator]
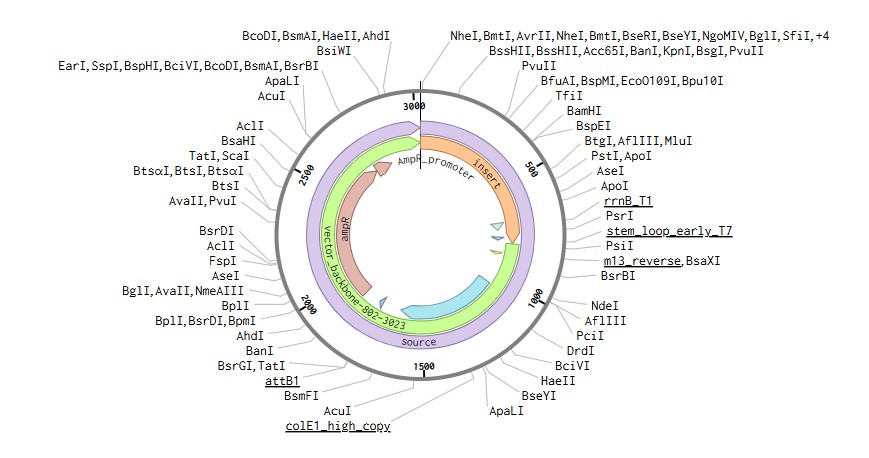

To express IL-1O in E.coli, I designed a expression cassette consisting of a constitutive promoter, a strong RBS, and a start codon. I added a pelB signal sequence that makes possible for the bacteria to secrete the protein. We added a HISX7 TAG, to make it easier to purify the proteins, a stop codon and finally a terminator. We choose clonal genes so it can be delivered as a circular plasmid and direclty transformed into the E.coli, and for vector the pTwist Amp High Copy, that gives us a high plasmid copy number resulting into a strong protein expression.
PART 5: DNA READ/WRITE/EDIT
5.1 READ
-What DNA would you want to sequence (e.g., read) and why?
Even if it isnt strictly related to my project, i would like to sequence genes involved in collagen production and extrecellular matrix stability, specially in patients with EDS. Ehler Danlos Syndrome is a group of inherited connective tissue disorders, with a combined stimated prevelance of at least 1 in 5000, even with it being underdiagnosed, where collagen sequences are mutated leading to misfolded collagen and connective tissue fragility. Because of collagen structure being complex and relying in the triple helix assemblance, small mutations can cause big consequences. Studying genes such as COL1A1, COL1A2, COL3A1 in patients with the disorder, it would give insight into how specific mutaitons lead to misfolded colalgen and connective tissue fragility and give better understanding of genotype/phenotype of variants and a lead to develop targeted therapies.
-What technology or technologies would you use to perform sequencing on your DNA and why?
I think the best choice would be a second generation sequencing, like Ilumina NGS. It’s considered 2nd gen because of the ability of sequencing millions of short dna fragments in parallel and requires the amplification of dna before the sequencing. The input could be be genomic dna extracted from the patients blood, which would have to undergo certain preparation steps. After DNA is extracted, it’s mechanically brokendown into short fragments. Synthetic adapters are lgiated to both ends of each fragments, which allows the binding to flow cells. After the fragments are amplified with pcr and then denatured to a single strand to sequence. For Illumina, the sequencing is done by synthesis. It creates cluster generation by binding the dna fragments into complementary oligos onf the flowcells and thru bridge amplificaiton creating the clusters of identical copies. Labeled nucleotides with a flurescent tag and reversible terminators are added and are incorporeated by polymerase 1 at time per cycle. As each nucleotide has a different signal, we can detect which base is being used each cycle. The output is millions of short dna reads, each giving us the nucleotide sequence and the a quality score of each base. It also makes it possible to identify any variants like deletions or insertions, which woulds allow the detection of any mutation for EDS in collagen genes.
5.2 WRITE
What DNA would you want to synthesize (e.g., write) and why?
I would like to synthesize a correct repair template for the pathogenic mutations in the collagen genes. The many of the mutations associated with EDS, involve the susbtitution of glycine residues in the repeating Gly–X–Y motif that stabilizes the collagen triple helix. If glycine is substituted for a bulkier aminoacid it disrupts the folding of the protein and weakes the connective tissues. If we could synthetize a fragment with the proper repreats and a way to flank the mutation site, it could be used to investigate the possiblities to correct the misfolded collagen and help compared stability of proteins.
What technology or technologies would you use to perform this DNA synthesis and why?
I would use the synthesis thru phosphoramidite chemistry. This methods chemically synthetizes base by base short dna oligonucleotides using protected nucleotides. The overlappping oligos are assembled into a full fragment that its cloned to a plasmid, and later the sequence is verified. This tecnique gives me a high fidelity which is needed for the repeated collagen sequences, and enables precise contorl over mutation correction.
5.3 EDIT
What DNA would you want to edit and why? What technology or technologies would you use to perform these DNA edits and why? VEDS, is associated with pathogenic mutations in the COL3A1 gene, arising as disruptions with the glycine residues needed for the correct formation of collagen. You could use CRISPR CAS9, to design an RNA capable to target the mutated region and introduce a donor repair template with the correct sequence. Alternatively, base editing could be use to change the erroneous base.