Week 6 HW: Genetic Circuits Part 1
Protocol Questions
Answer these questions about the protocol in this week’s lab:
-What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
Phusion High-Fidelity PCR Master Mix with HF Buffer is a 2X master mix consisting of Phusion DNA Polymerase, deoxynucleotides and reaction buffer that has been optimized and includes MgCl2. All that is required is the addition of template, primers and water. (New England Biolabs, 2026)
Phusion DNA Polymerase: a highly accurate enzyme that synthesizes new DNA strands with very low error rates due to proofreading activity.
dNTPs (deoxynucleotide triphosphates):the building blocks (A, T, G, C) used to construct new DNA strands.
Reaction Buffer: maintains optimal pH and salt conditions for enzyme activity.
MgCl₂ (magnesium ions): a critical cofactor required for polymerase function.
Stabilizers/additives: enhance enzyme stability and performance during thermal cycling.
-What are some factors that determine primer annealing temperature during PCR?
The primary annealing temperature depends on:
- Primer Melting temperature (usual annealing is 3-5°c below Tm).
- Primer Length (with increasing lenght the Tm also increases).
- GC content (GC increases Tm due to strong H bonding).
- Sequence composition (repetitive or 2nd structure forming sequences affect binding).
- Salt/ion concentration (influences the dna duplex stability).
There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
PCR This method uses primers and DNA polymerase to amplify a specific DNA region. It requires thermal cycling and can introduce custom sequences like overlaps or mutations. It produces DNA fragments with defined ends depending on primer design.
RESTRICTION DIGEST Uses restriction enzymes to cut DNA in specific recognition sites. It is done at a constant temperature and producecs predictable sticky/blunt ends. This methods is limited, as it is required existing restriction sites in the DNA.
PCR vs. RESTRICTION ENZYME DIGEST
When comparing both methods, we can see each one has their specific requirements for protocols and they are tailored for different needs, even if both create linear fragments of DNA. PCR, is used when u need to amplify DNA and there is no restriction sites. Use restriction enzyme digest, when suitable restriction sites exist in the sequence and u want a precise ends. Overall, PCR is a method more flexible while restriciton digestions is simpler but less customizable.
How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
To ensure compatability with Gibson Assembly, we have to sedign fragments with 20-40bp overlapping homologous sequences. We should ensure correct orientation and reading frame, avoid secondary structure or repetitive overlaps, and ensure no unwanted mutations.
How does the plasmid DNA enter the E. coli cells during transformation?
This can be done in 2 ways. Chemical transformation (Heatshock), treats cells with CaCl2 to make them competens and heat shocks them, creating a thermal imbalance that allows the DNA (plasmid) to enter. Electroporation, gives brief pulses to cells to create pores in the mebrane for the DNA to enter. In both cases, the membrane is temporarily permeabilized for the upatake.
Describe another assembly method in detail (such as Golden Gate Assembly)
Golden Gate Assembly, uses restriction enzymes that cut outside their recognition sites, and joins fragments with DNA ligase. This method generates custom overhangs and allows us to create multiple fragments in one reaction.
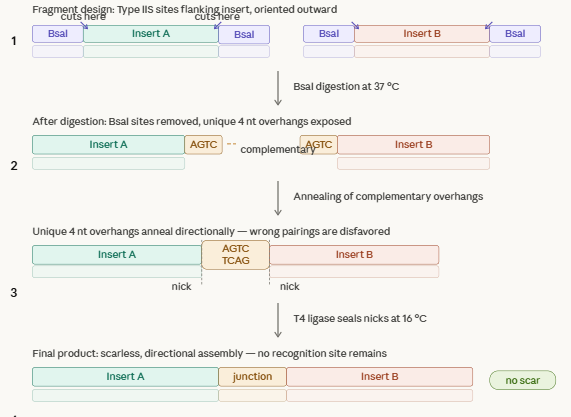
Explain the other method in 5 - 7 sentences plus diagrams (either handmade or online)
This method harnesses restriction enzymes to cut DNA at a defined distance outside their recognition site generating custom 4n overhangs. Each fragment is flanked by typeiis sites oriented so that after digestion, every overhang is exposed and matching its intended neighbor. DNA ligase joins the overhangs thru cycling temperatures. Overhangs are fully defined and at to 50 fragments can be assembled in a reaction, making this method usefult to create gene libraries.

Model this assembly method with Benchling or Asimov Kernel!
PARTS USED:
PROMOTER_J23100:TTTACAGCTAGCTCAGTCCTAGGTACAGTGCTAGC
RBS_B0034: AAAGAGGAGAAACGATG
GFP: ATGGTGAGCAAGGGCGAGGAGCTGTTCACCGGGGTGGTGCC
TERMINATOR_B0015: TTATTAGAGCTAGAAATAGCAAGTTAAAATAAGG
BACKBONE WITH BBSL SITES
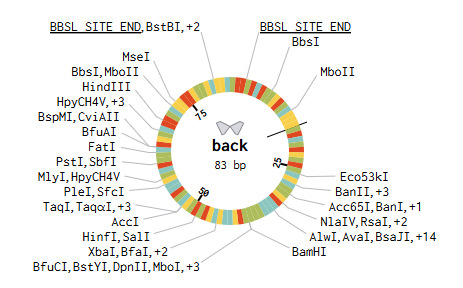

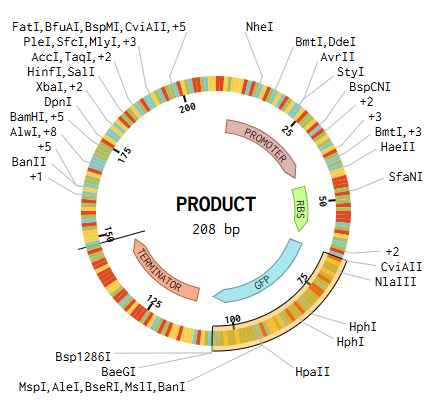
GOLDEN GATE ASSEMBLY IN BENCHLING
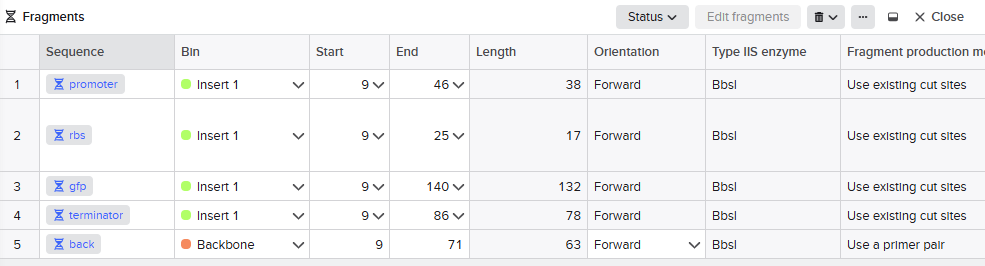

We designed the overhangs and added then to the original dna sequences of the parts we used. This makes possible the annealing and ligation of the final sequence.
Promoter → RBS → GFP → Terminator → Backbone