Week 2 HW: DNA Read, Write, and Edit


Part 0: Basics of Gel Electrophoresis
- Per instructions for this part, I attended the 02/10 lecture and 02/11. Additionally I attended all 3 Bootcamp sessions.
Part 1: Benchling & In-silico Gel Art
- Make a free account at benchling.com
 Benchling Account Creation Confirmation
Benchling Account Creation Confirmation
- Import the Lambda DNA
 Benchling Phage Lambda DNA Import Confirmation_02.12.26
Benchling Phage Lambda DNA Import Confirmation_02.12.26
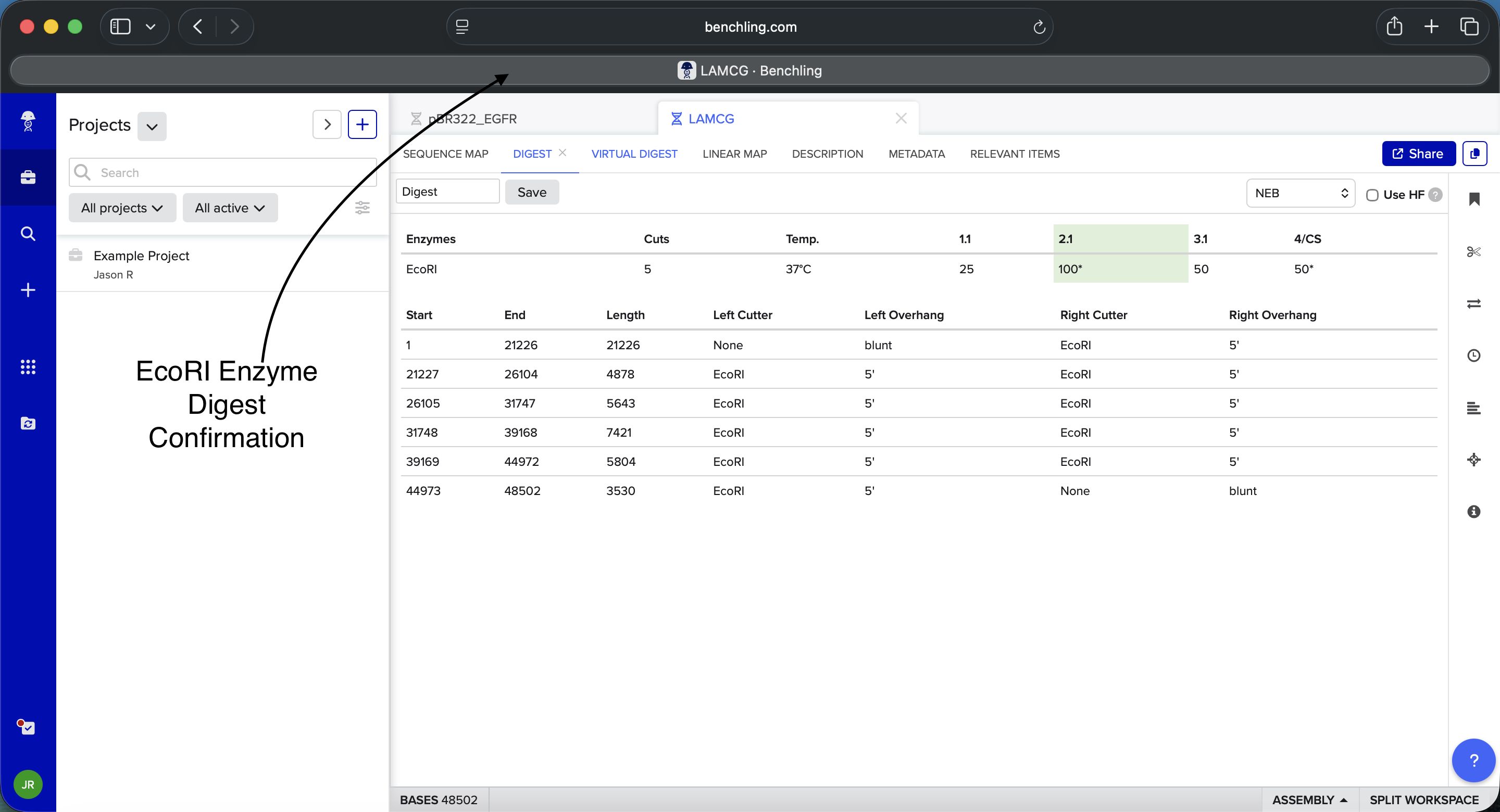
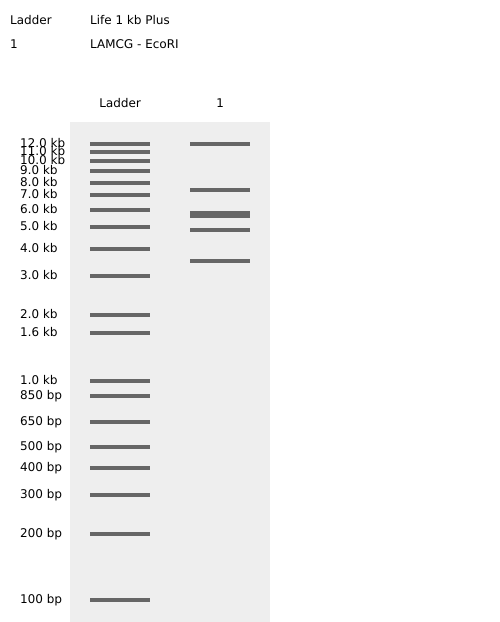
Simulate Restriction Enzyme Digestion with the following Enzymes:
Benchling EcoRI Enzyme Digest Confirmation
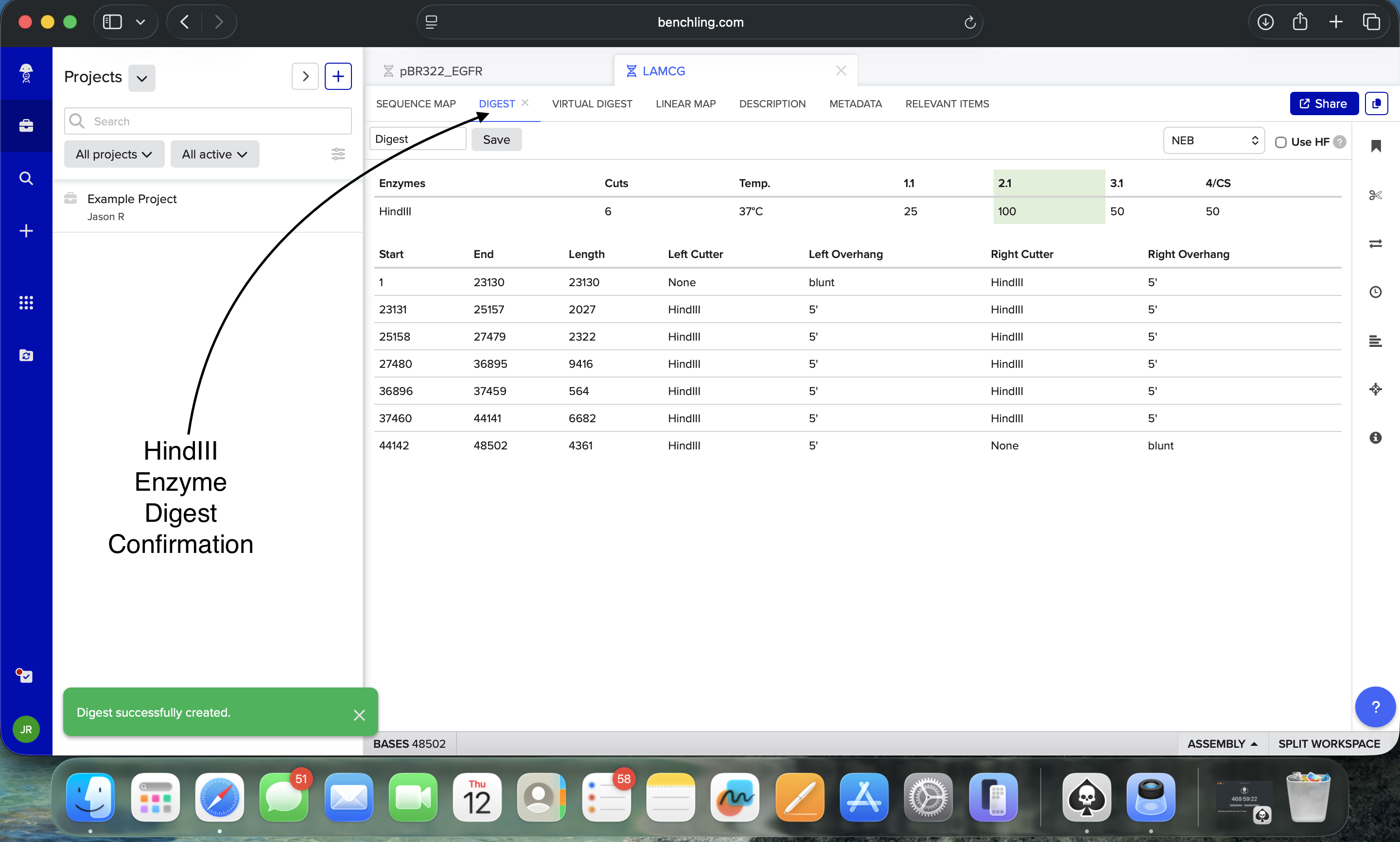
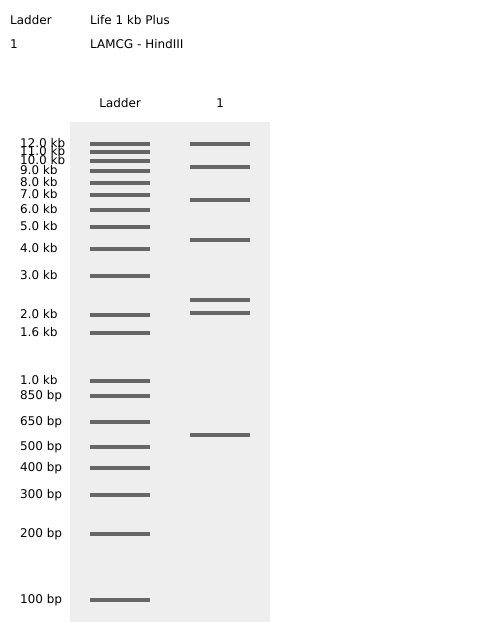
Benchling HindIII Enzyme Digest Confirmation
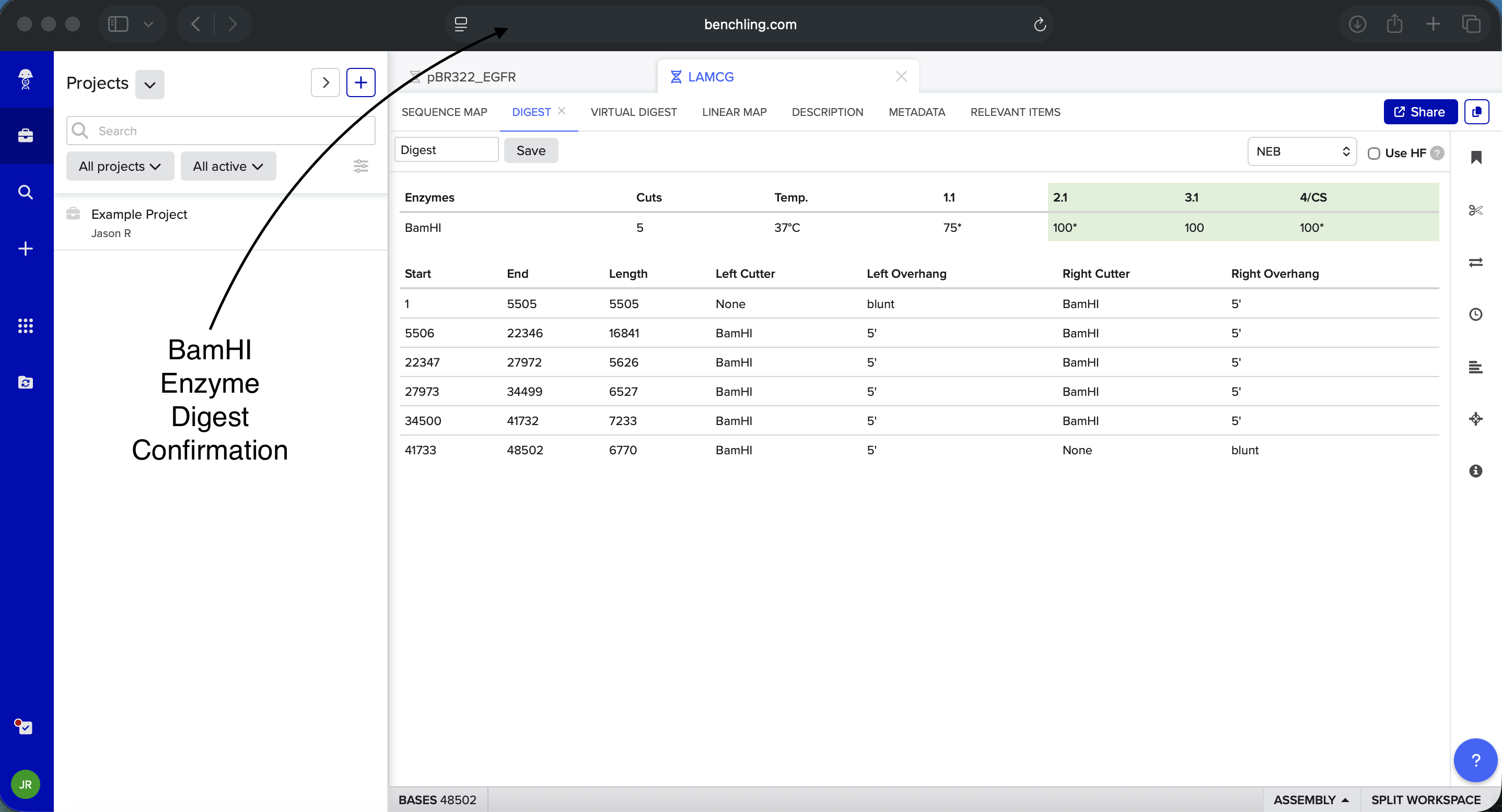
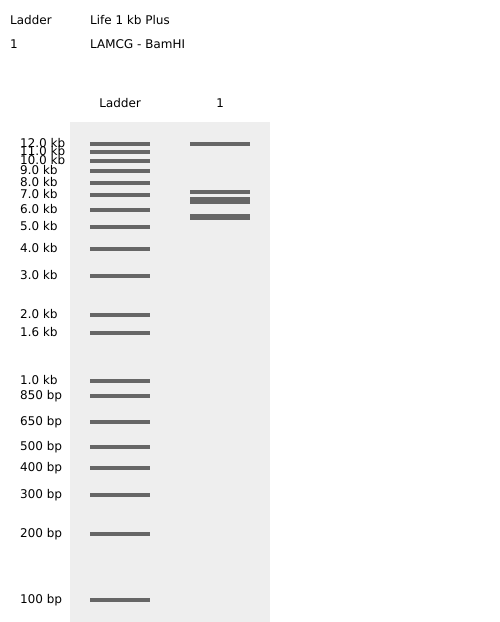
Benchling BamHI Enzyme Digest Confirmation
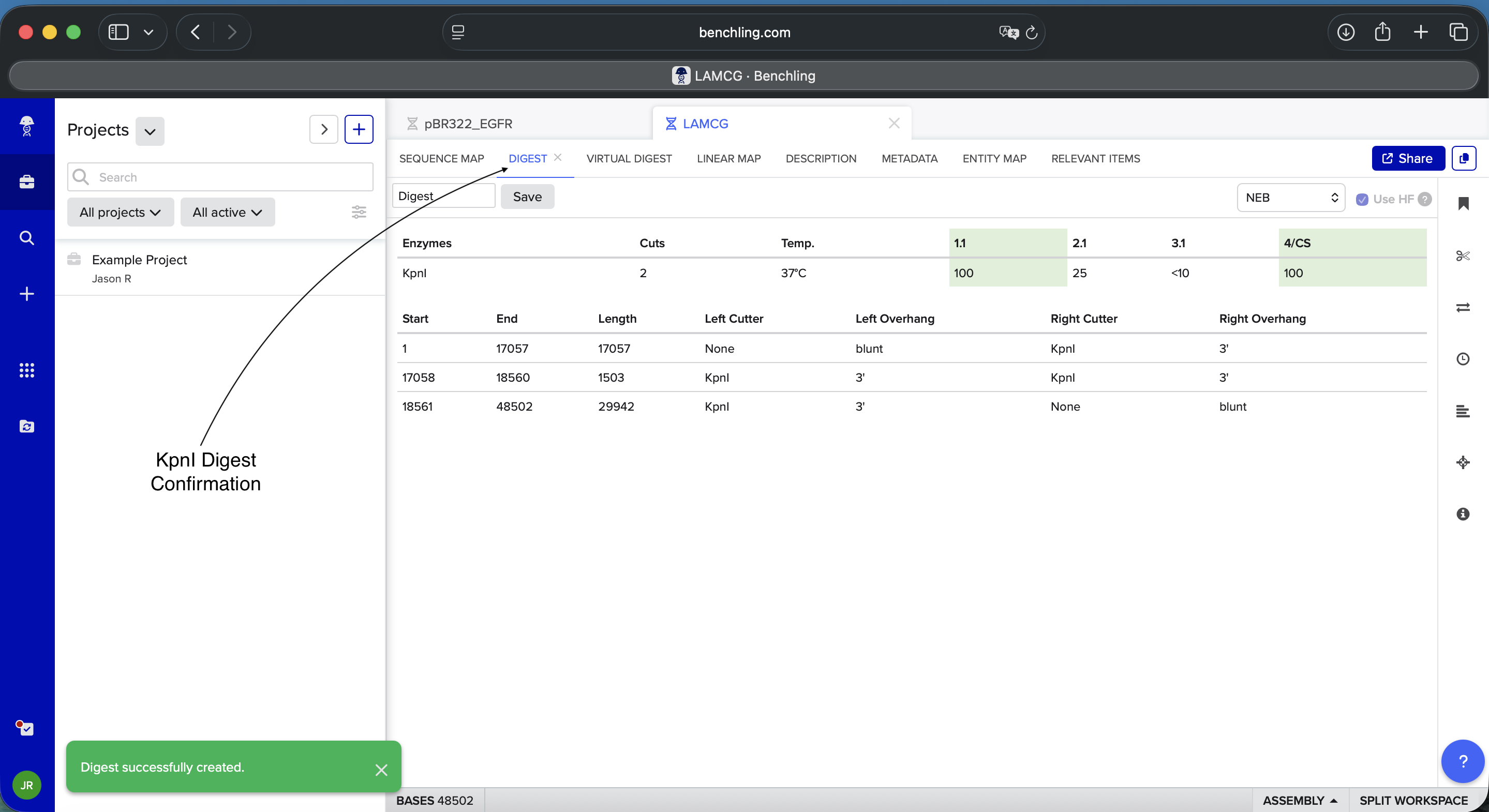
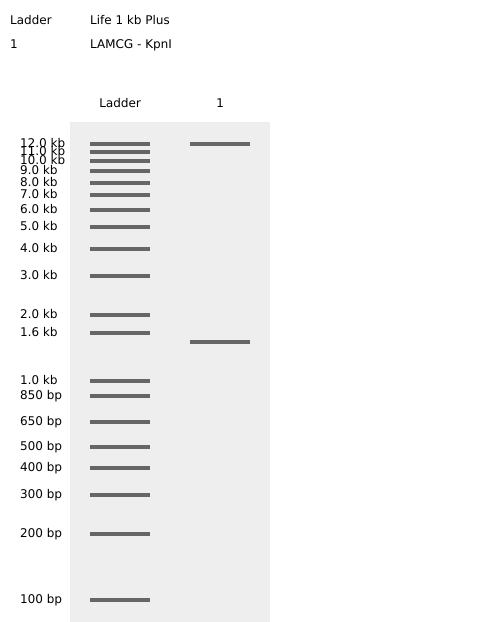
Benchling KpnI Enzyme Digest Confirmation
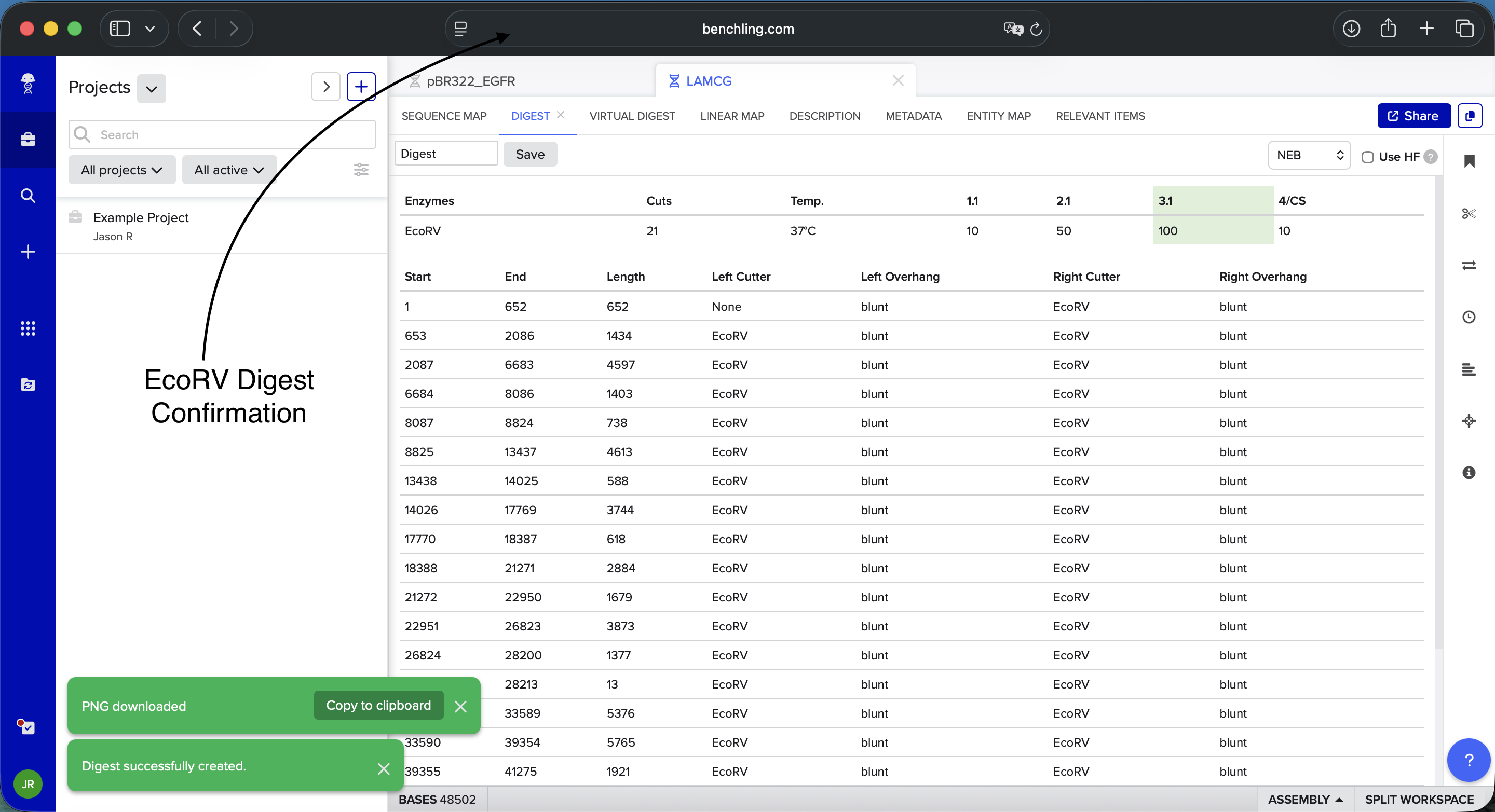
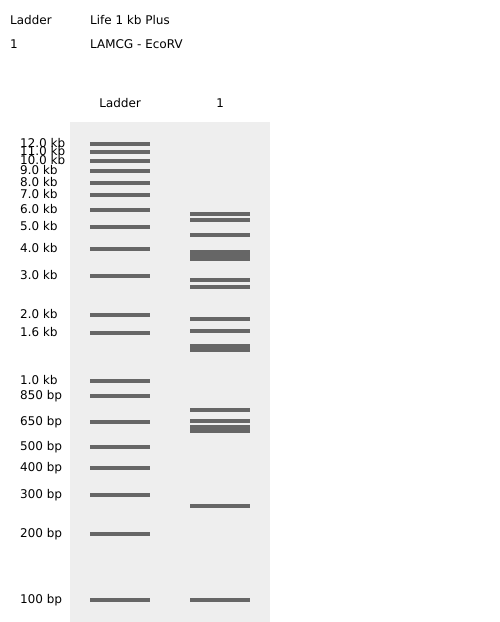
Benchling EcoRV Enzyme Digest Confirmation
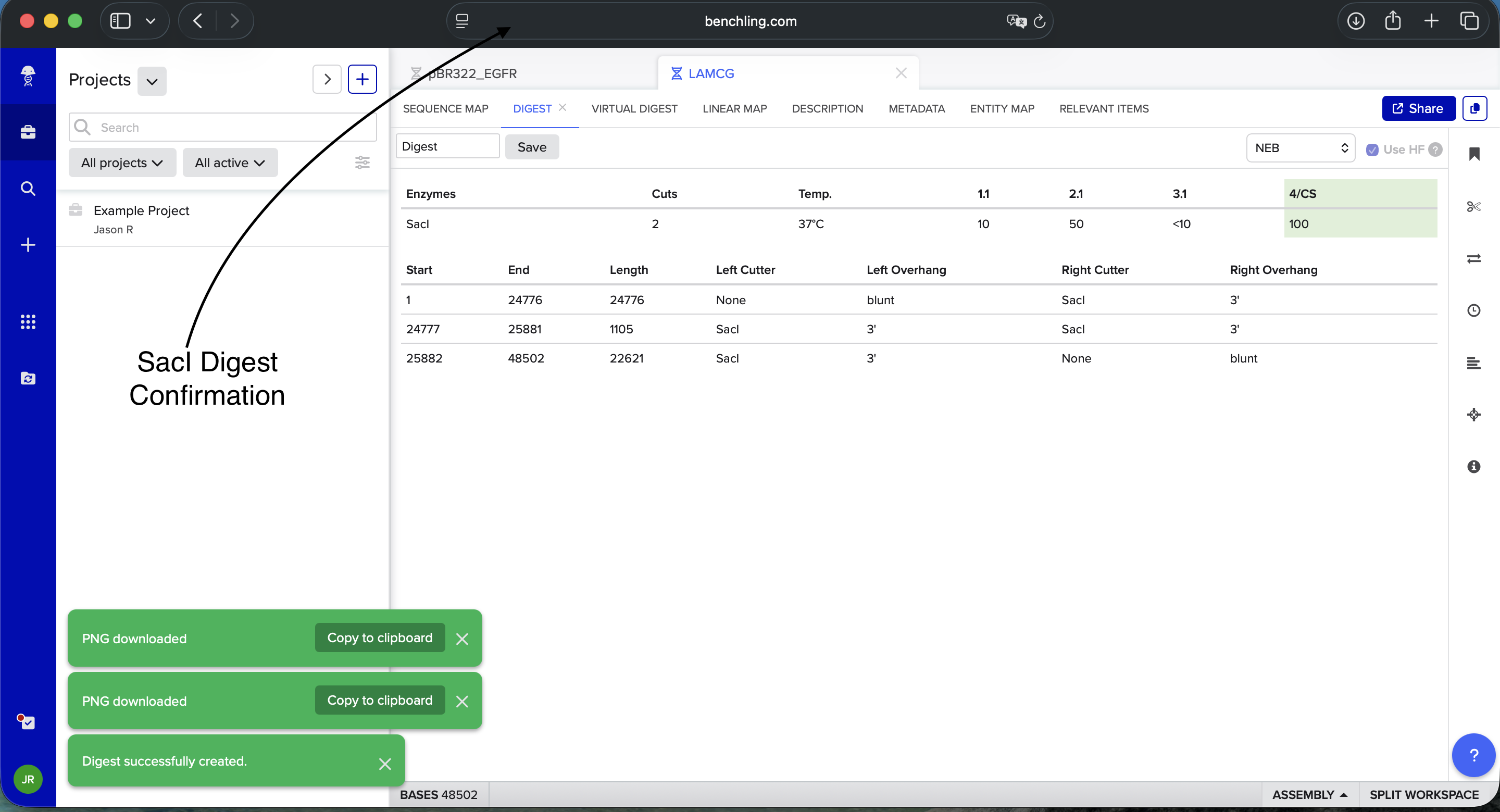
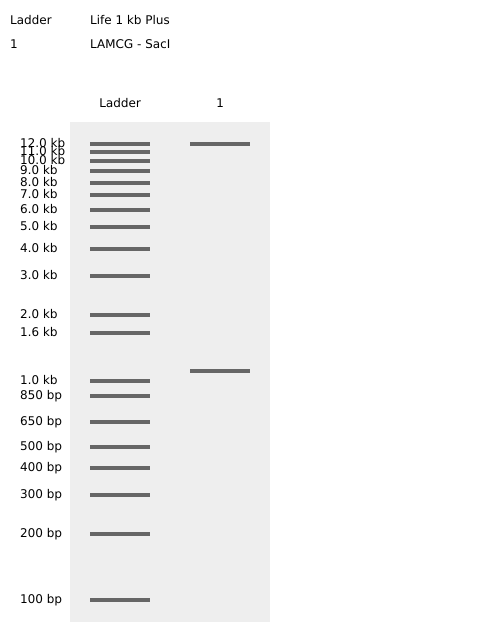
Benchling SacI Enzyme Digest Confirmation
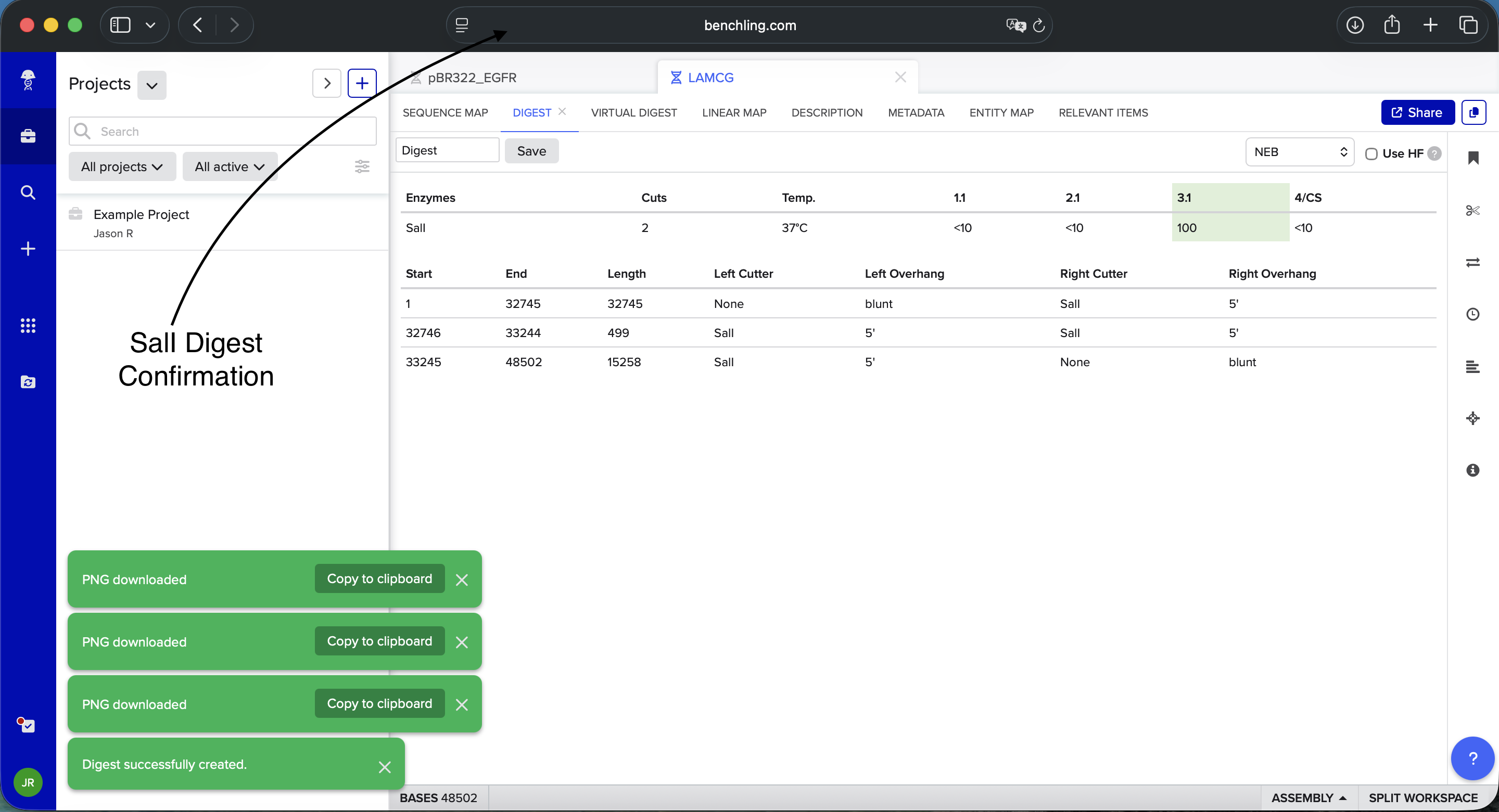
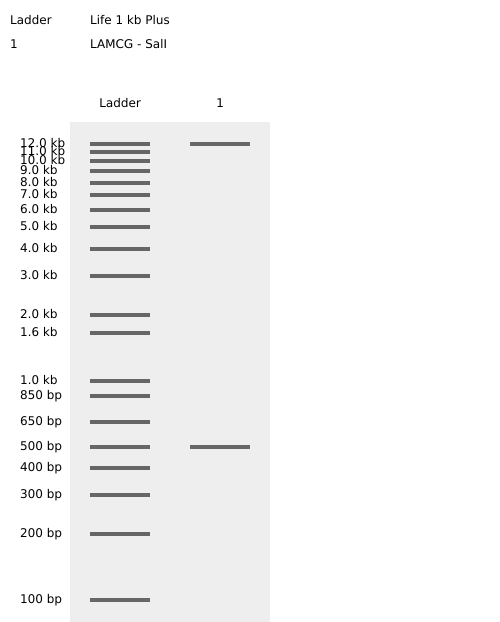
Benchling SalI Enzyme Digest Confirmation
Create a pattern/image in the style of Paul Vanouse’s Latent Figure Protocol artowrks
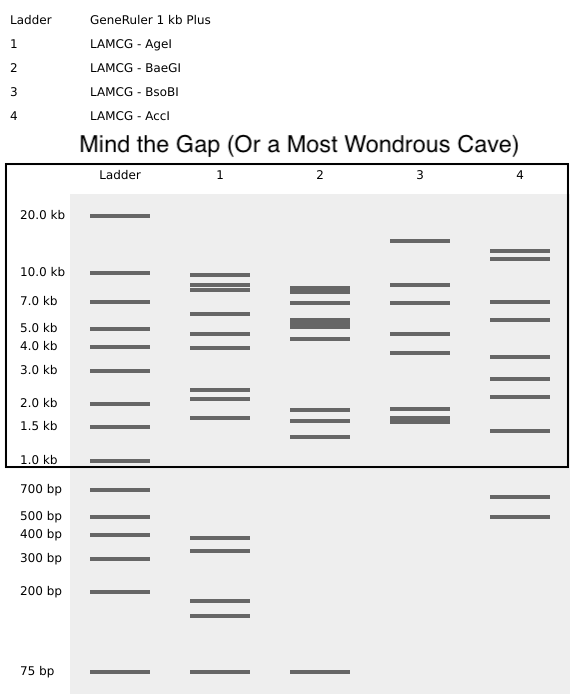

Mind the Gap (Or a Most Wondrous Cave) ➕
Part 2: Gel Art - Restriction Digests and Gel Electrophoresis
Part 3: DNA Design Challenge
3.1 Choose your protein
- I chose the Mantis Fibroin 1 protein because for some reason when I received this assignment, my mind flipped to an insect protein, and then from there, a praying mantis. Upon further research, I was pleased with where my intuition lead me. The Mantis Fibroin 1 protein helps comprise the mantis’ ootheca, otherwise known as its egg casing. What’s fascinating about these proteins is that they create this coiled yet flexible foam-like structure around the mantis’ eggs. This protein piqued my interest, as it might have biomimetic potential. The Mantis Fibroin 1 protein is listed below 12:
tr|I3PM87|I3PM87_9NEOP Mantis fibroin 1 OS=Pseudomantis albofimbriata OX=627833 GN=MF1 PE=2 SV=1 MDSKMLCVSLLLAVFCLWYTEASPLEEKYGEKYGDMEEYQRGTEDSRAVINDHTAKVASQ SARGMVNKAKTTEAAARSNEQLSKDRQYYYREYLKKADYHKKKALEYEQLSAAENAKIAY HESKQKDWETKARESDVQCRDAEAKYEQSYTRSRELKRESIIAYVQAAMHHAEASGDHMK ADRAKDIARDMMRKAESLRGDASNHYQRSEEDKNKARSEKVKAHQNADNSQRHHTACRAY DQEGLKTRLSSKANMMRQIHSSLLAERSHSLAREDGLAADLSHKLAEELARMSEESGAIS KINSGEERGYSNKVRQDEVKAHELAVSKRMMGAEVADNSEMISLAQAKDGSLDEGENYKL STFYADDSTKNMLPDSRGQMSYGDE
3.2 Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
- A translated Mantis Fibroin 1 protein nucleotide sequence of most likely codons is below, as well as evidence showing how I inserted the Mantis Fibroin 1 protein UniProt information into the reverse translation tool
atggatagcaaaatgctgtgcgtgagcctgctgctggcggtgttttgcctgtggtatacc gaagcgagcccgctggaagaaaaatatggcgaaaaatatggcgatatggaagaatatcag cgcggcaccgaagatagccgcgcggtgattaacgatcataccgcgaaagtggcgagccag agcgcgcgcggcatggtgaacaaagcgaaaaccaccgaagcggcggcgcgcagcaacgaa cagctgagcaaagatcgccagtattattatcgcgaatatctgaaaaaagcggattatcat aaaaaaaaagcgctggaatatgaacagctgagcgcggcggaaaacgcgaaaattgcgtat catgaaagcaaacagaaagattgggaaaccaaagcgcgcgaaagcgatgtgcagtgccgc gatgcggaagcgaaatatgaacagagctatacccgcagccgcgaactgaaacgcgaaagc attattgcgtatgtgcaggcggcgatgcatcatgcggaagcgagcggcgatcatatgaaa gcggatcgcgcgaaagatattgcgcgcgatatgatgcgcaaagcggaaagcctgcgcggc gatgcgagcaaccattatcagcgcagcgaagaagataaaaacaaagcgcgcagcgaaaaa gtgaaagcgcatcagaacgcggataacagccagcgccatcataccgcgtgccgcgcgtat gatcaggaaggcctgaaaacccgcctgagcagcaaagcgaacatgatgcgccagattcat agcagcctgctggcggaacgcagccatagcctggcgcgcgaagatggcctggcggcggat ctgagccataaactggcggaagaactggcgcgcatgagcgaagaaagcggcgcgattagc aaaattaacagcggcgaagaacgcggctatagcaacaaagtgcgccaggatgaagtgaaa gcgcatgaactggcggtgagcaaacgcatgatgggcgcggaagtggcggataacagcgaa atgattagcctggcgcaggcgaaagatggcagcctggatgaaggcgaaaactataaactg agcaccttttatgcggatgatagcaccaaaaacatgctgccggatagccgcggccagatg agctatggcgatgaa
 Mantis Fibroin 1 UniProt Information Inserted into Reverse Translation Tool
Mantis Fibroin 1 UniProt Information Inserted into Reverse Translation Tool
3.3 Codon Optimization
- We need to optimize codon usage so a particular sequence can be expressed with greater fidelity, reliability, and efficiency in a host organism. I’ve chosen to optimze the codon sequence for Saccharomyces cerevisiae (baker’s yeast), because it:
- is commonly used as a host in biotechnology applications
- folds in a manner closer to insect protein folding
- is apparently easier to work with than mammalian cells
A codon optimized Mantis Fibroin 1 nucleotide sequence is shown below, as well as evidence showing how I showing how I inserted the Mantis Fibroin 1 protein nucleotide sequence information into the codon optimization tool
ATGGACAGTAAGATGTTATGTGTCTCCTTATTGTTGGCTGTTTTTTGTTTATGGTATACTGAAGCTTCCCCATTAGAAGAAAAGTATGGTGAAAAGTACGGTGACATGGAAGAGTACCAAAGAGGTACTGAAGATTCAAGAGCAGTTATTAACGATCATACTGCTAAAGTTGCTTCCCAATCCGCCAGAGGTATGGTTAATAAGGCTAAGACTACAGAAGCTGCTGCTAGAAGTAATGAACAATTATCTAAAGATAGACAATACTATTACAGAGAATATTTGAAAAAGGCTGATTATCATAAGAAGAAAGCTTTGGAATATGAACAGCTTTCAGCTGCTGAAAATGCAAAAATTGCTTATCATGAATCTAAACAAAAAGACTGGGAAACGAAAGCCAGAGAATCCGATGTTCAATGTCGTGATGCTGAAGCAAAATATGAACAATCTTACACAAGGTCCAGAGAACTGAAAAGGGAATCTATTATTGCTTATGTTCAAGCTGCTATGCATCATGCTGAAGCTAGCGGTGATCACATGAAAGCTGATAGAGCTAAAGATATCGCTAGAGATATGATGAGAAAGGCAGAATCCTTAAGGGGTGACGCTAGCAACCATTATCAGAGATCCGAAGAAGATAAGAATAAGGCCAGATCTGAAAAGGTTAAAGCTCATCAAAACGCTGATAATTCTCAAAGACATCATACTGCATGCAGAGCGTATGACCAAGAAGGTTTAAAGACGAGATTGAGCTCAAAAGCCAACATGATGAGACAAATTCACTCCTCACTACTGGCTGAAAGATCTCATTCATTAGCAAGAGAAGACGGTCTTGCGGCCGATTTATCACATAAGTTGGCTGAAGAATTAGCTAGAATGTCCGAAGAATCAGGTGCTATATCTAAAATAAACTCAGGTGAAGAAAGAGGCTATTCGAATAAAGTGAGACAAGATGAGGTTAAAGCACATGAATTGGCTGTTAGCAAAAGAATGATGGGTGCTGAAGTTGCTGATAATTCGGAGATGATTAGTTTGGCACAAGCTAAAGACGGTTCTTTAGATGAAGGTGAGAACTATAAATTATCCACTTTTTATGCAGACGATTCTACAAAAAATATGCTACCAGATTCTAGGGGTCAAATGTCTTACGGTGATGAA
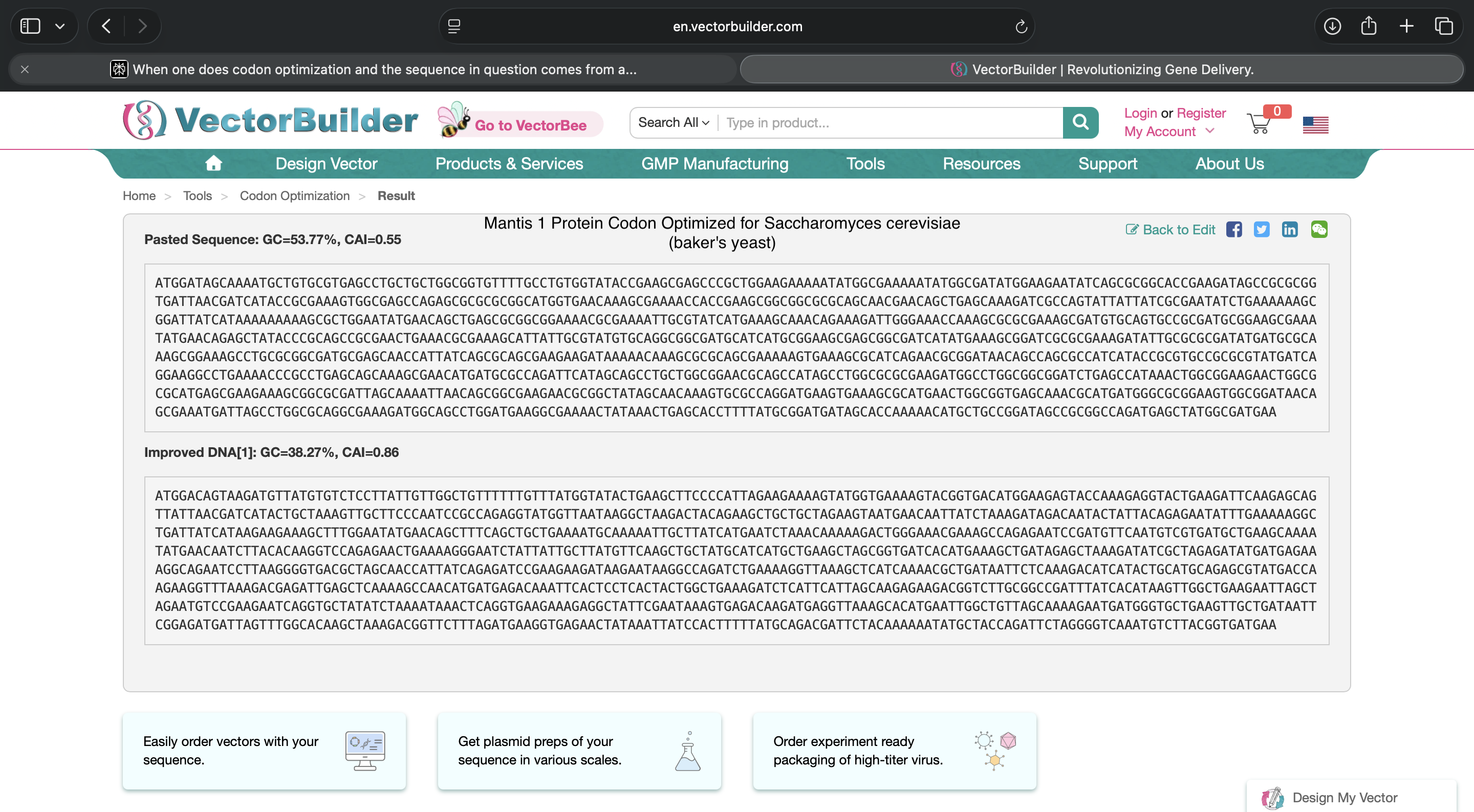
 Mantis Fibroin 1 nucleotide sequence codon optimized for Saccharomyces cerevisiae (baker’s yeast)
Mantis Fibroin 1 nucleotide sequence codon optimized for Saccharomyces cerevisiae (baker’s yeast)
3.4: You have a sequence! Now what?
- Several cell-dependent technologies could be used for producing the codon-optimized Mantis Fibroin 1 protein. One such technology, a yeast system, has already been pursued in the previous steps of this section, as Saccharomyces cerevisiae (baker’s yeast) is a form of yeast. Bacterial systems, such as E. coli could also be used for producing the protein in host cell culture, although this would require different codon optimization. Other cell-dependent technologies could have included insect or mammalian-based systems, although I’m not sure of the value of expressing an insect-associated protein in another insect-based host (although this may be a failure of imagination on my part). As mentioned previously, mammalian systems could also be used, but apparently mammalian cells are more difficult to work with than bacterial or yeast-based hosts. Cell-free methods for producing the codon-optimized Mantis Fibroin 1 protein would involve breaking open a cell, extracting relevant ribosomes, enzymes, tRNAs, etc., and then taking these contents and combinining them with a DNA template (in this case our Mantis Fibroin 1 protein nucleotide sequence), energy sourcesm relevant amino acids, and a reaction buffer. There’s a time advantage of cell-free methods over cell-dependent methods of protein expression.
3.5 Optional: How does it work in nature/biological systems?
- From my research in answering this question, I think the answer is that post-transcriptional process called alternative splicing occurs, where non-coding mRNA (introns) are cut and removed, while coding regions (exons) remain 3. It’s pretty fascinating because this splicing can create several different types of mRNA molecules, and therefore different proteins. This increases the efficiency with which different proteins can be expressed within a particular organism.
- See below
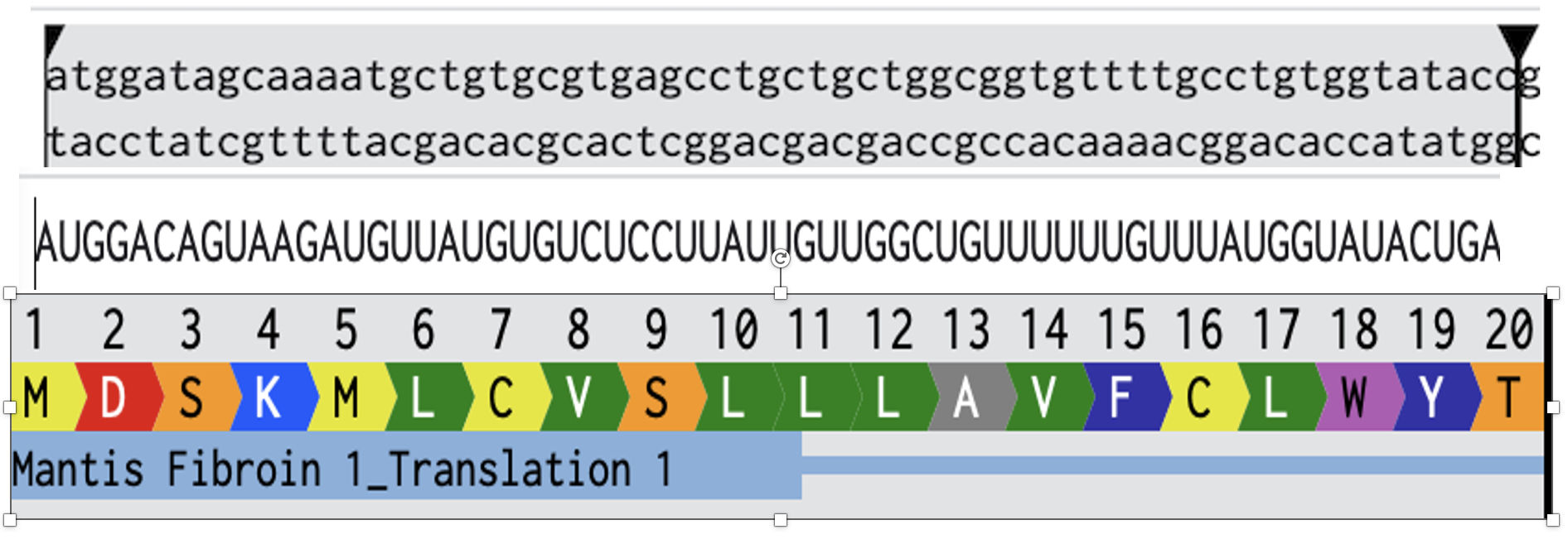
 Attempted Mantis Fibroin 1 Alignment
Attempted Mantis Fibroin 1 Alignment
All Section 3 Prompts Listed Below
| Supporting Prompt | Source |
|---|---|
| I want to make DNA gel art in the style of Paul Venouse’s gel electrophoresis works. I want to use the website below to help me:https://rcdonovan.com/gel-art How should I go about making this art virtually? I have a Benchling account, an Internet connection, and no access to a wet lab | Perplexity |
| Help me design a specific lane‑by‑lane “band plan” for your first virtual gel artwork. My knowledge of Benchling is very minimal and my desired art is a right-facing arrow | Perplexity |
| Not understanding the 3rd Step under Step 3. How do I run any digest that gives me 2 fragments? How do I know my target size for each copy of the Arrow_Base sequence? | Perplexity |
| I see a scissors icon. Write out a literal “click this, then this” sequence for one lane so I can duplicate it for the others | Perplexity |
| Ok I’ve found and made 5 digests of the requested bp sizes. When I’m seeing the combined virtual digest I’m not seeing an arrow. What are next steps I should take? | Perplexity |
| Ok – let’s take a step back. I’ve tried what you’ve said and am not seeing a right facing arrow. Think I need to learn how to do this from the ground up with some more tweaking. I know how to select the Digests option in Benching, I know how to save a Digest, and I know how to click on ‘Virtual Digest’ and select different kinds of ladders. What type of ‘Cut sites visible on maps’ should I select to begin compiling the right facing arrow? Under ‘Show enzymes that cut’ is there anything in particular I should select? Should I select ‘Highlight enzymes with compatible sticky ends’? How do I actually learn how to make the right facing arrow? | Perplexity |
| This seems like a good workflow. How do I create a new DNA with a 5000bp length? I assume I have to import a sequence of that length from an NCBI accession correct? | Perplexity |
| When I type in HindIII I see 0 cuts. I can’t make a ~4000bp band based on that, right? | Perplexity |
| Everything is coming up a 0 when I try to put in an enzyme. I think there’s hallucinating going on or there is something wrong when you started having me make the 5kb sequence | Perplexity |
| When I select an enzyme for a digest, do I then need to select a cut site to make the bio art or no? | Perplexity |
| Dumb question: How do I see the length of a digest for a given enzyme in a sequence? | Perplexity |
| Tell me how to search NCBI for an accession | Perplexity |
| When I look inside the Digest feature in Benchling, how do I find an enzyme that can give me a cut of a certain length? I see Name in one column, followed by Cuts, but am not exactly understanding what I’m seeing here | Perplexity |
| wondering if there are any special proteins found in the praying mantis insect and what exactly about these proteins make them special | Google AI Mode |
| Are there any proteins in nature that have unbelievable economy of space that would make them particularly useful for data storage? | Google AI Mode |
| Are there any proteins in nature that have unbelievable economy of space that would make them particularly useful for data storage? | Google AI Mode |
| i want to make a box where I can put some text in in markdown hugo relearn theme. I don’t want to create a table. What should i create? | Google AI Mode |
| When one does codon optimization and the sequence in question comes from a protein traditionally associated with a given species (let’s say an insect), does one traditionally optimize the codon sequence for that same species or its genus/family of species? How does this work given standard codon optimization practices? | Perplexity |
| So in essence, we perform codon optimization so our sequence in question can be expressed with greater fidelity or reliability in the host? Or so the host can receive or incorporate the sequence as efficiently as possible? Let me know if my thinking or terminology is off here | Perplexity |
| I have a nucleotide sequence for the Mantis Fibroin 1 protein. Have learned about some of the Mantis Fibroin 1 protein’s interesting properties, namely how it helps create a coiled yet flexible casing around Praying Mantis eggs. At this point, I think this protein might have some biomimetic potential, but am not sure what organism I should optimize the sequence for. Traditionally I know E. coli and Baker’s yeast are used a lot in synthetic biology applications, and I know mammalian cells are apparently more challenging to work with. For a use case like this one, where I have an insect-associated protein that may have biomimetic properties, let me know some traditional host organisms in biotechnology that are used for codon optimization in cases like this one. Do NOT hallucinate. Use existing sources. If you cannot provide anything whatsoever, say so | Perplexity |
| When we have a protein we find in the wild, and then we codon optimize its nucleotide sequence for expression in a host organism, what do cell-dependent or cell-free methods to produce this codon optimized protein from the sequence mean in a biotechnological context? What exactly are we talking about? | Perplexity |
| Based on the answer to the previous prompt, what is a promoter? What is a lysate mix? Do NOT hallucinate when answering these questions | Perplexity |
| What are some cell-dependent methods of producing proteins from DNA in biotechnology? Are there multiple types of cell-dependent methods? Do NOT hallucinate when answering these questions | Perplexity |
| Are cell-dependent methods for producing proteins from DNA in biotechnology distinct from cell-dependent technologies for producing proteins from DNA or are the terms ‘methods’ and ’technologies’ essentially interchangeable in a biotechnology context? If they’re not, describe some cell-dependent technologies for producing proteins from DNA. Do NOT hallucinate when addressing this query. If you cannot answer this question, say so | Perplexity |
| explain to me how cell-free expression of a protein works | Google AI Mode |
| Does the histone code have anything to do with the ability for a single gene in nature to code for multiple proteins at the transcriptional level? How does the histone code relate to the transcriptome? Are they one and the same?. Do NOT hallucinate when answering these questions. If you don’t know the answers to these questions, say so | Perplexity |
| Based on the answer to the last prompt, then what does allow for a single gene in nature to code for multiple proteins at the transcriptional level? In plain terms, what does “…different exon combinations produce distinct mRNA isoforms from one gene, which then translate into varied proteins” mean? What are exons again, and how do they produce different combinations? What is an mRNA isoform? Do NOT hallucinate when answering this query | Perplexity |
| Thank you for the answer to the last query. What exactly is being sliced (the gene itself or something else), and why is it being spliced? Why would nature/evolution create this ability? Do NOT hallucinate when answering this query. Go off existing literature | Perplexity |
| Does alternative splicing operate at the transcriptional level? From what I can see in the link below, it operates on the translational level. https://www.yourgenome.org/theme/what-is-rna-splicing/. If there was hallucination, or an error in answering the previous prompts, say so. Or, if I’m misreading or misunderstanding things, say so. Just wondering what at the transcriptional level allows for a single gene to code for multiple proteins | Perplexity |
| how does a single gene in nature code for multiple proteins at the transcriptional level? | Google AI Mode |
| I want to align a DNA sequence, its transcribed RNA, and a resulting translated protein. I believe I can capture the separate pieces (the DNA sequence, the transcribed RNA, and the resulting translated protein) in Benchling. If this is true, how can I go about doing this? | Perplexity |
| I have a codon optimized sequence for the Mantis Fibroin 1 protein in a Saccharomyces cerevisiae (baker’s yeast) host. I want to produce the RNA sequence and the final translated protein. What services online can I use to do this? | Perplexity |
| What does forward and reverse translation of a DNA sequence in Benchling mean? | Perplexity |
| I have a codon optimized nucleotide sequence (DNA). How can I find what the RNA sequence and translated protein look like for this sequence? What services can I use to see these items? | Google AI Mode |
Part 4: Prepare a Twist DNA Synthesis Order
4.1: Create a Twist account, and Benchling account

 Benchling Account Creation Confirmation
Benchling Account Creation Confirmation
4.2: Build Your DNA Insert Sequence
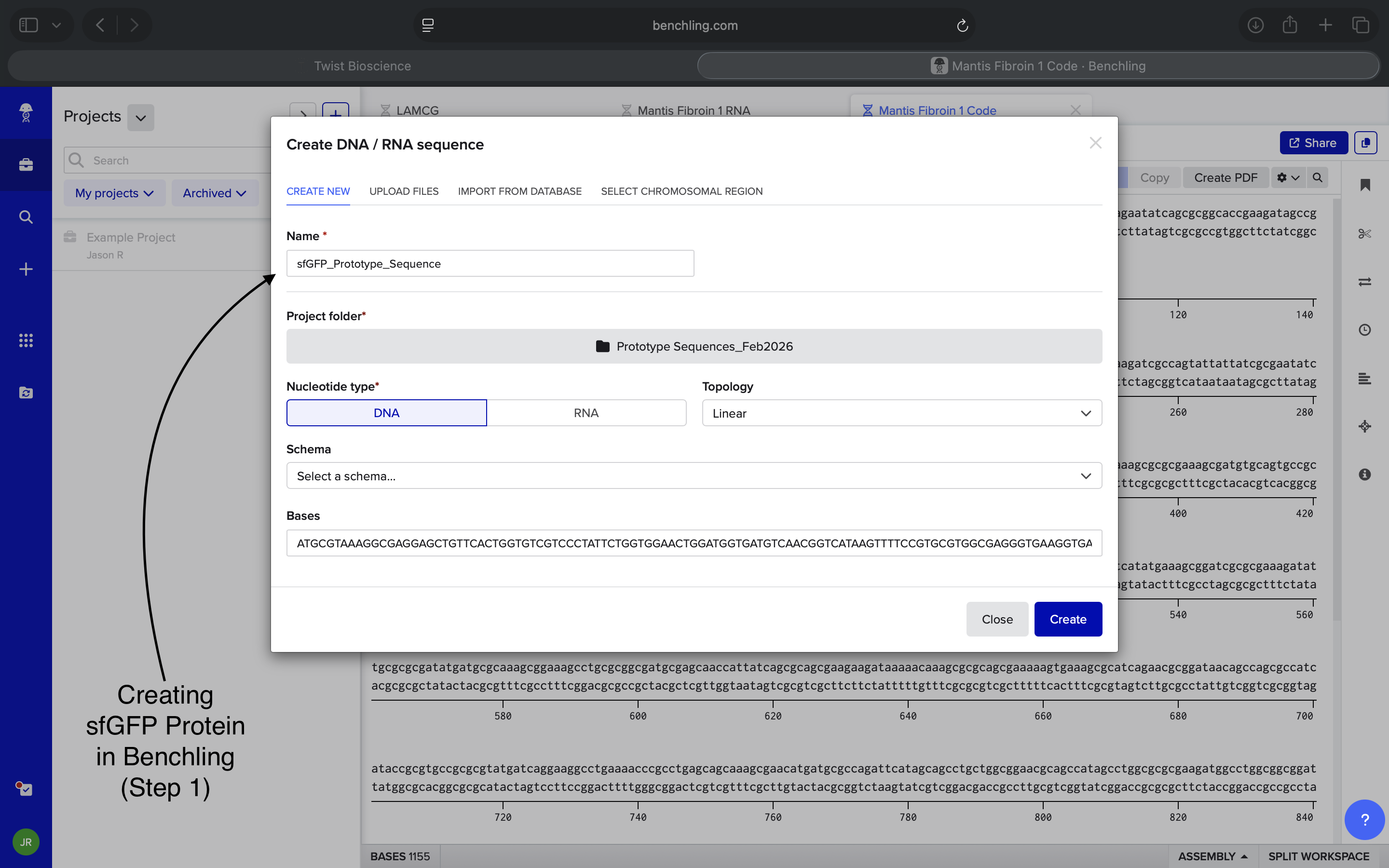
 Original Sequence Insertion NOTE: Think I may’ve started off inserting the wrong sequence. This may have been potentially fixed when I inserted a sfGFP sequence from NCBI*
Original Sequence Insertion NOTE: Think I may’ve started off inserting the wrong sequence. This may have been potentially fixed when I inserted a sfGFP sequence from NCBI*
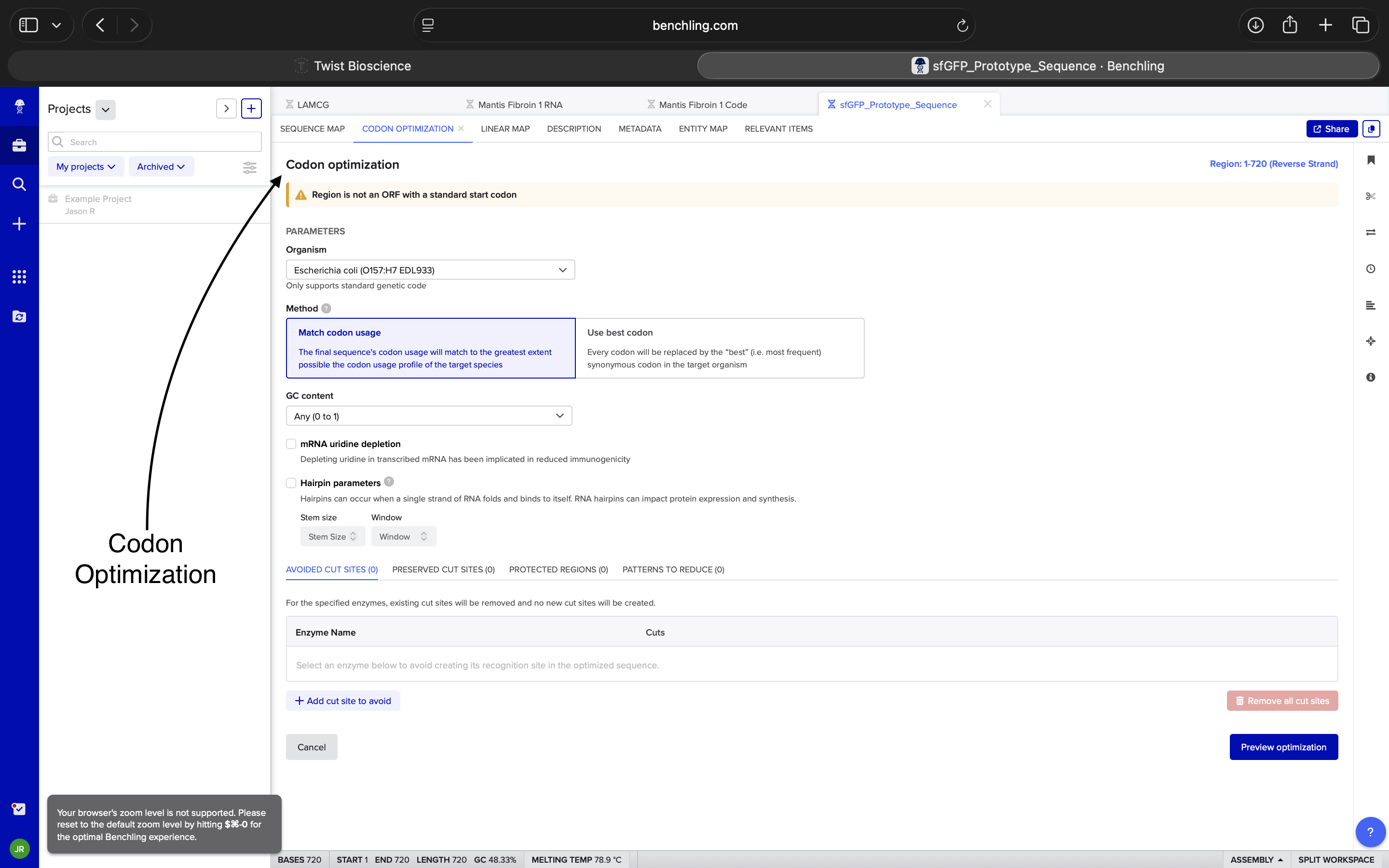
 Codon Optimization NOTE: Think I may’ve started off inserting the wrong sequence. This may have been potentially fixed when I inserted a sfGFP sequence from NCBI*
Codon Optimization NOTE: Think I may’ve started off inserting the wrong sequence. This may have been potentially fixed when I inserted a sfGFP sequence from NCBI*
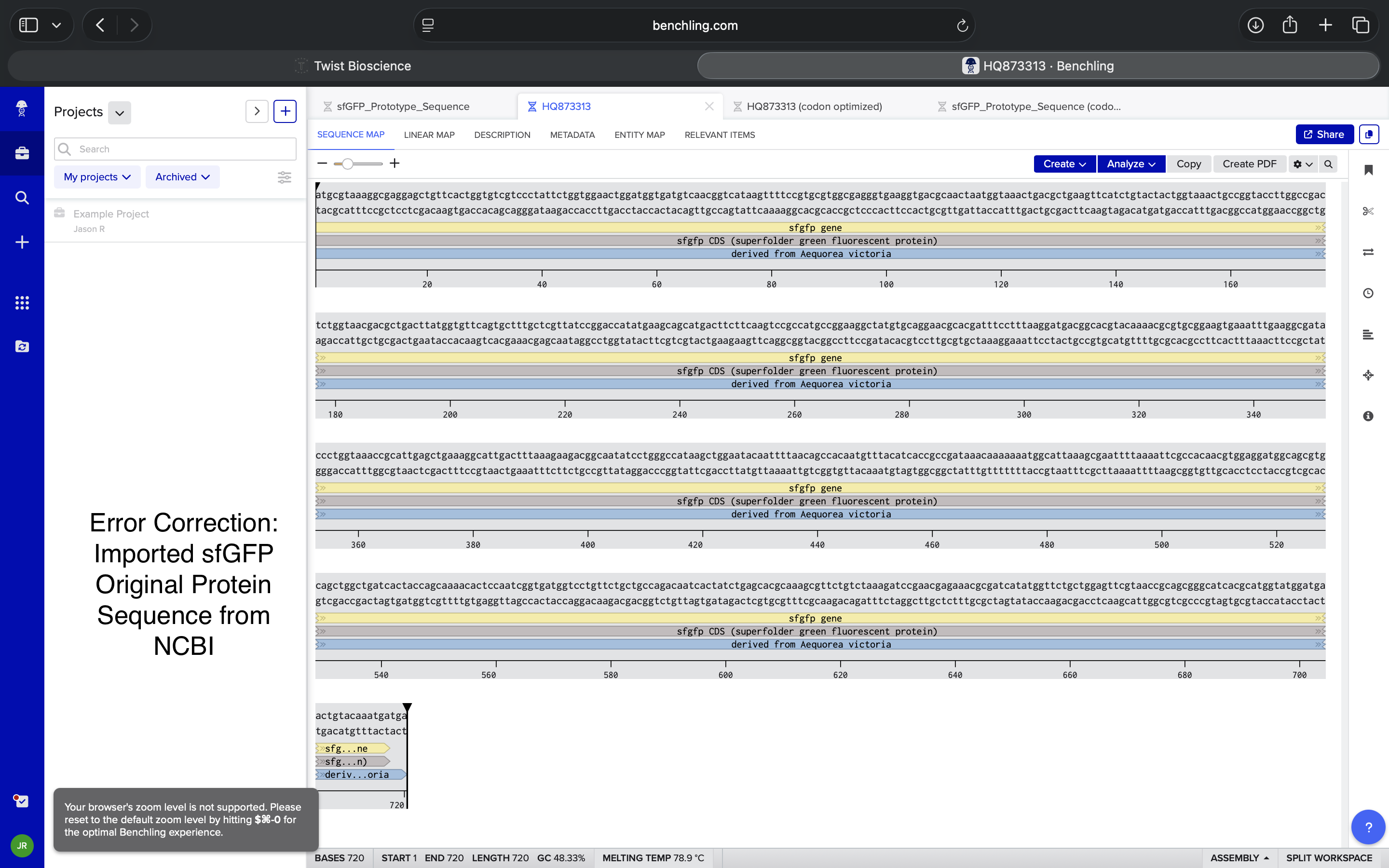
 Corrected NCBI Sequence Insertion
Corrected NCBI Sequence Insertion
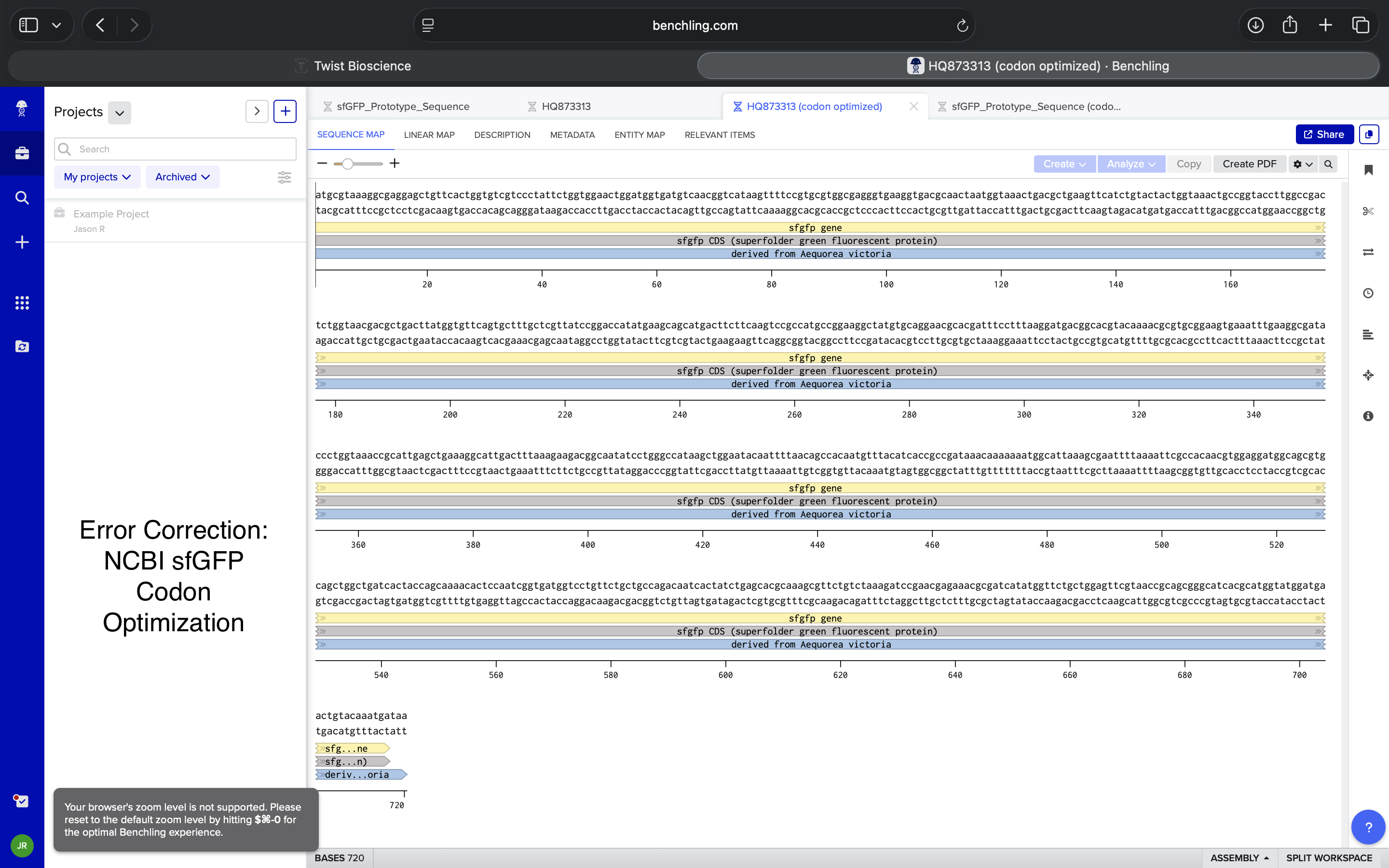
 Corrected NCBI Sequence Codon Optimization
Corrected NCBI Sequence Codon Optimization
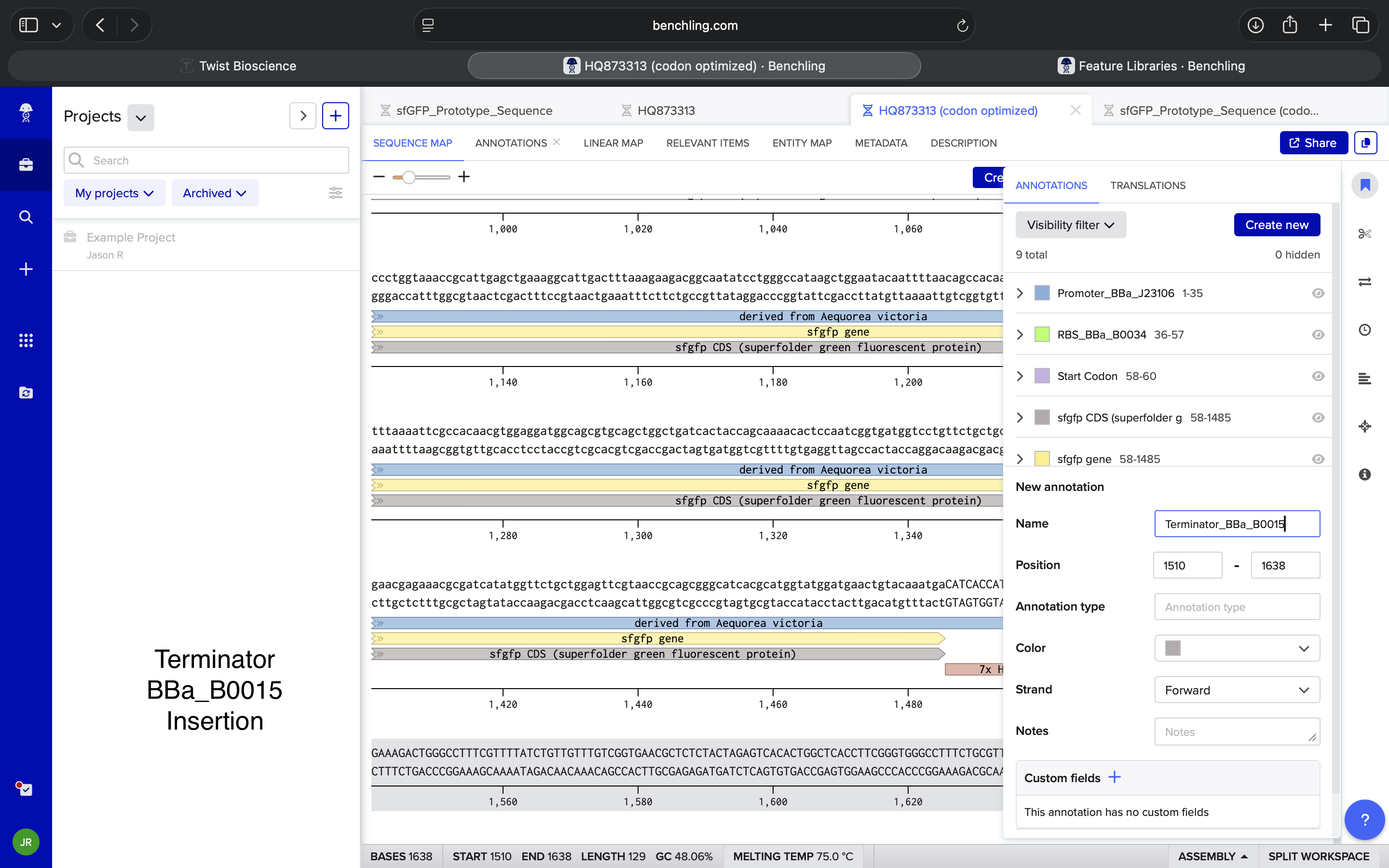
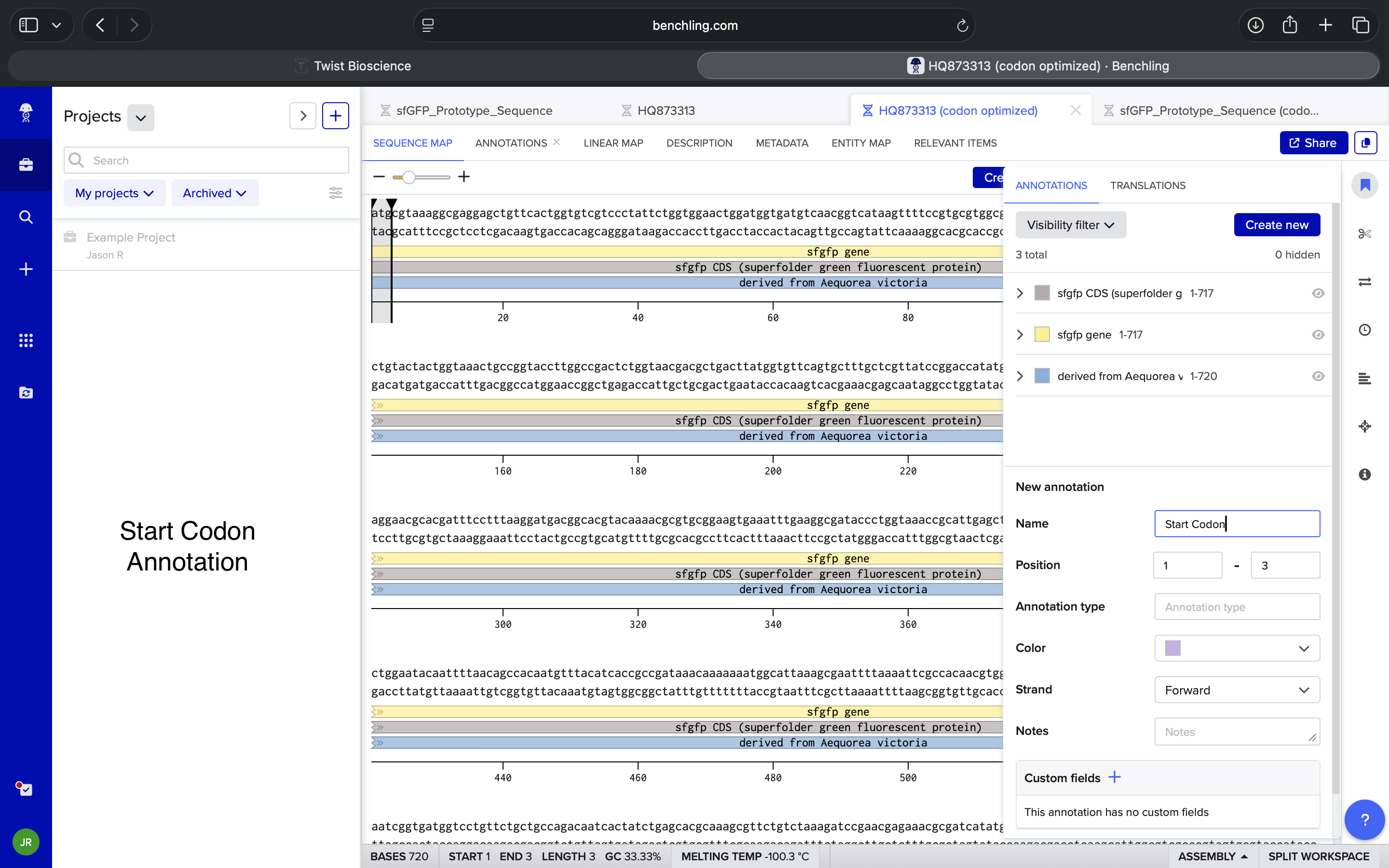
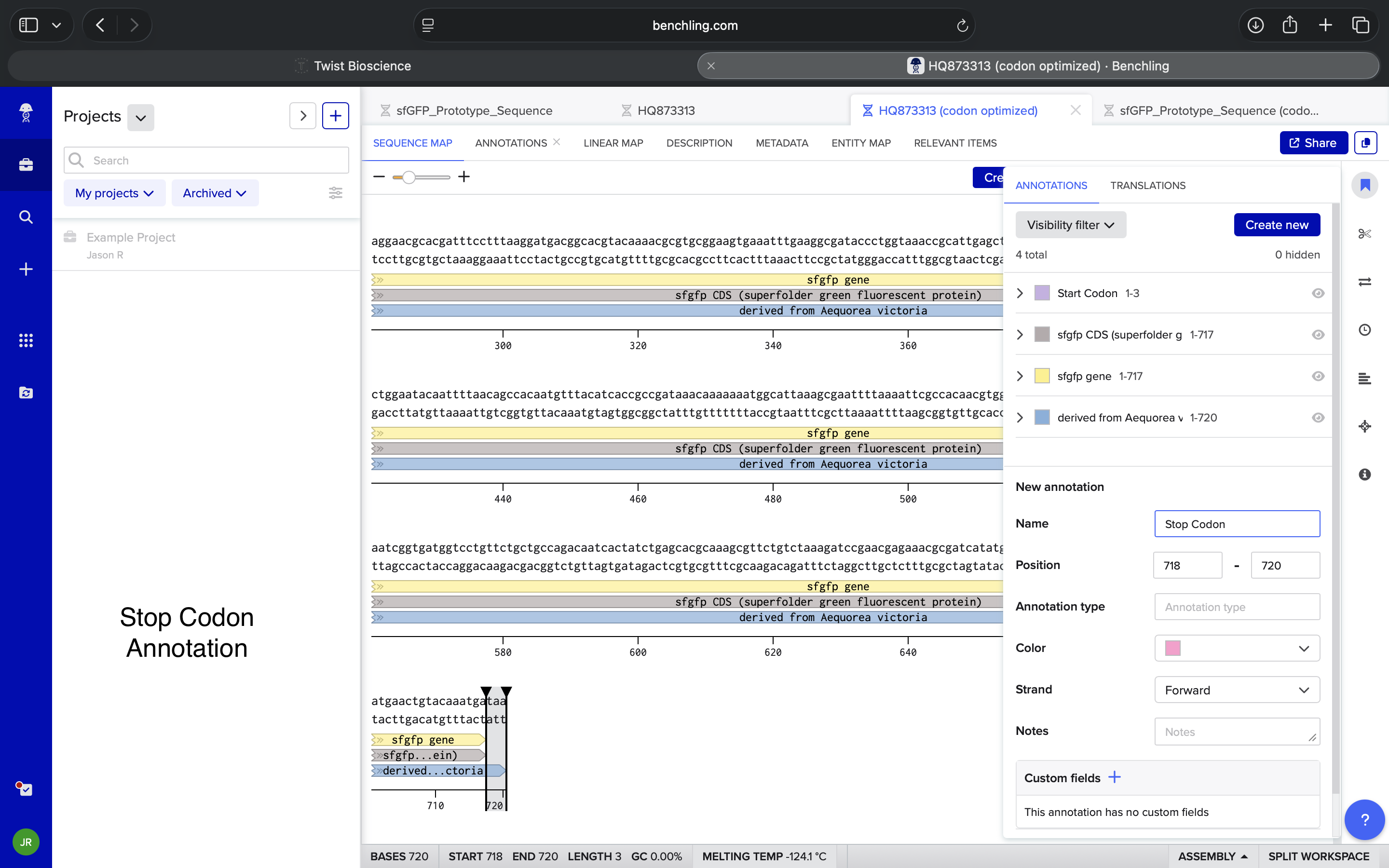
 Terminator BBa_B0015 Insertion
Terminator BBa_B0015 Insertion
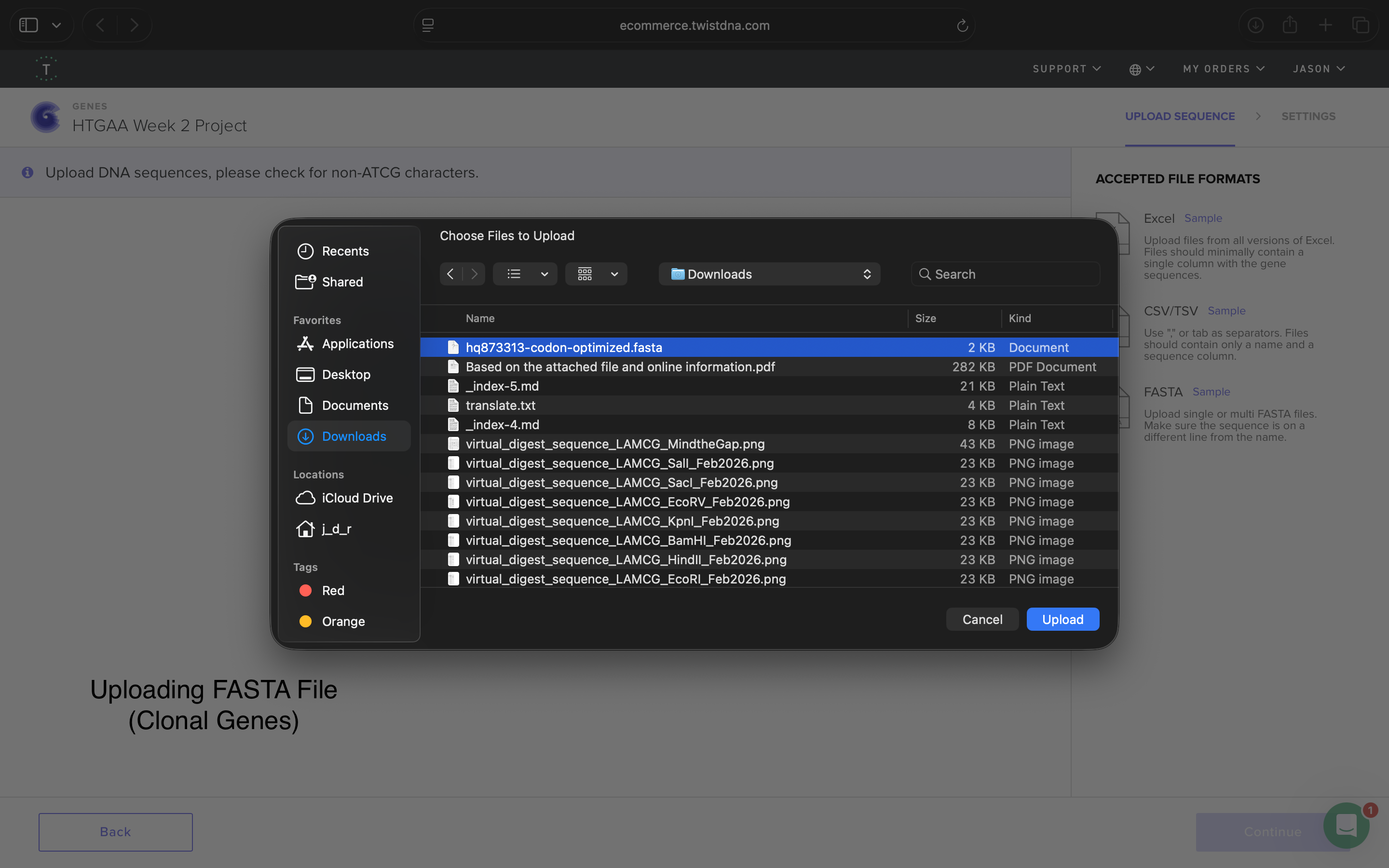
- Downloaded Sequence FASTA file (via Mac OS TextEdit). See below:
HQ873313 (codon optimized) TTTACGGCTAGCTCAGTCCTAGGTATAGTGCTAGCCATTAAAGAGGAGAAAGGTACCatgAGCAAAGGAGAAGAACTTT TCACTGGAGTTGTCCCAATTCTTGTTGAATTAGATGGTGATGTTAATGGGCACAAATTTTCTGTCCGTGGAGAGGGTGA AGGTGATGCTACAAACGGAAAACTCACCCTTAAATTTATTTGCACTACTGGAAAACTACCTGTTCCGTGGCCAACACTT GTCACTACTCTGACCTATGGTGTTCAATGCTTTTCCCGTTATCCGGATCACATGAAACGGCATGACTTTTTCAAGAGTG CCATGCCCGAAGGTTATGTACAGGAACGCACTATATCTTTCAAAGATGACGGGACCTACAAGACGCGTGCTGAAGTCAA GTTTGAAGGTGATACCCTTGTTAATCGTATCGAGTTAAAGGGTATTGATTTTAAAGAAGATGGAAACATTCTTGGACAC AAACTCGAGTACAACTTTAACTCACACAATGTATACATCACGGCAGACAAACAAAAGAATGGAATCAAAGCTAACTTCA AAATTCGCCACAACGTTGAAGATGGTTCCGTTCAACTAGCAGACCATTATCAACAAAATACTCCAATTGGCGATGGCCC TGTCCTTTTACCAGACAACCATTACCTGTCGACACAATCTGTCCTTTCGAAAGATCCCAACGAAAAGCGTGACCACATG GTCCTTCTTGAGTTTGTAACTGCTGCTGGGATTACACATGGCATGGATGAGCTCTACAAAcgtaaaggcgaggagctgt tcactggtgtcgtccctattctggtggaactggatggtgatgtcaacggtcataagttttccgtgcgtggcgagggtga aggtgacgcaactaatggtaaactgacgctgaagttcatctgtactactggtaaactgccggtaccttggccgactctg gtaacgacgctgacttatggtgttcagtgctttgctcgttatccggaccatatgaagcagcatgacttcttcaagtccg ccatgccggaaggctatgtgcaggaacgcacgatttcctttaaggatgacggcacgtacaaaacgcgtgcggaagtgaa atttgaaggcgataccctggtaaaccgcattgagctgaaaggcattgactttaaagaagacggcaatatcctgggccat aagctggaatacaattttaacagccacaatgtttacatcaccgccgataaacaaaaaaatggcattaaagcgaatttta aaattcgccacaacgtggaggatggcagcgtgcagctggctgatcactaccagcaaaacactccaatcggtgatggtcc tgttctgctgccagacaatcactatctgagcacgcaaagcgttctgtctaaagatccgaacgagaaacgcgatcatatg gttctgctggagttcgtaaccgcagcgggcatcacgcatggtatggatgaactgtacaaatgaCATCACCATCACCATC ATCACtaaCCAGGCATCAAATAAAACGAAAGGCTCAGTCGAAAGACTGGGCCTTTCGTTTTATCTGTTGTTTGTCGGTG AACGCTCTCTACTAGAGTCACACTGGCTCACCTTCGGGTGGGCCTTTCTGCGTTTATA
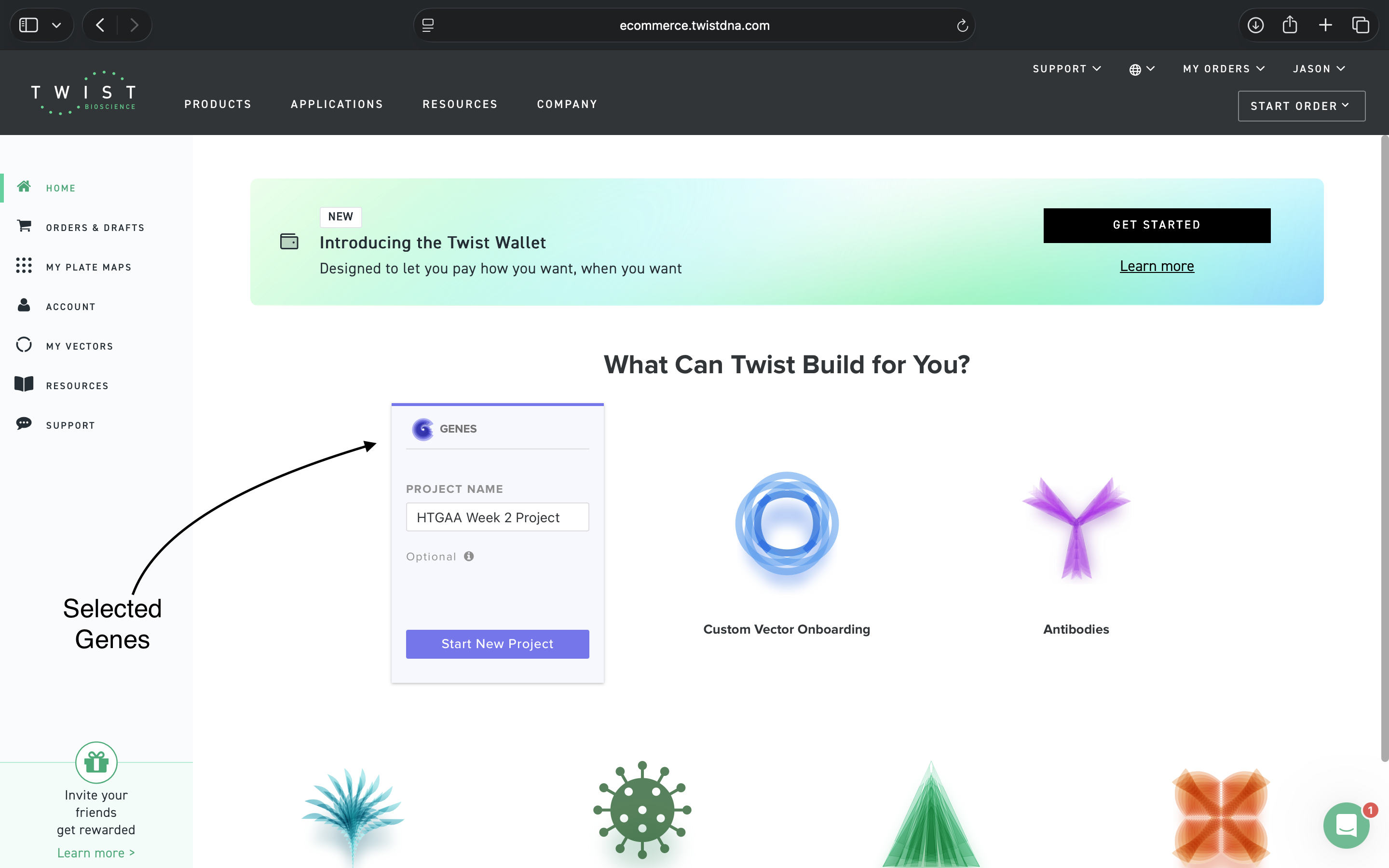
4.3: On Twist, Select the Genes Option
4.4: Select the ‘Clonal Genes’ Option
Clonal Genes Selected see results in subsections 4.5 and 4.6 below
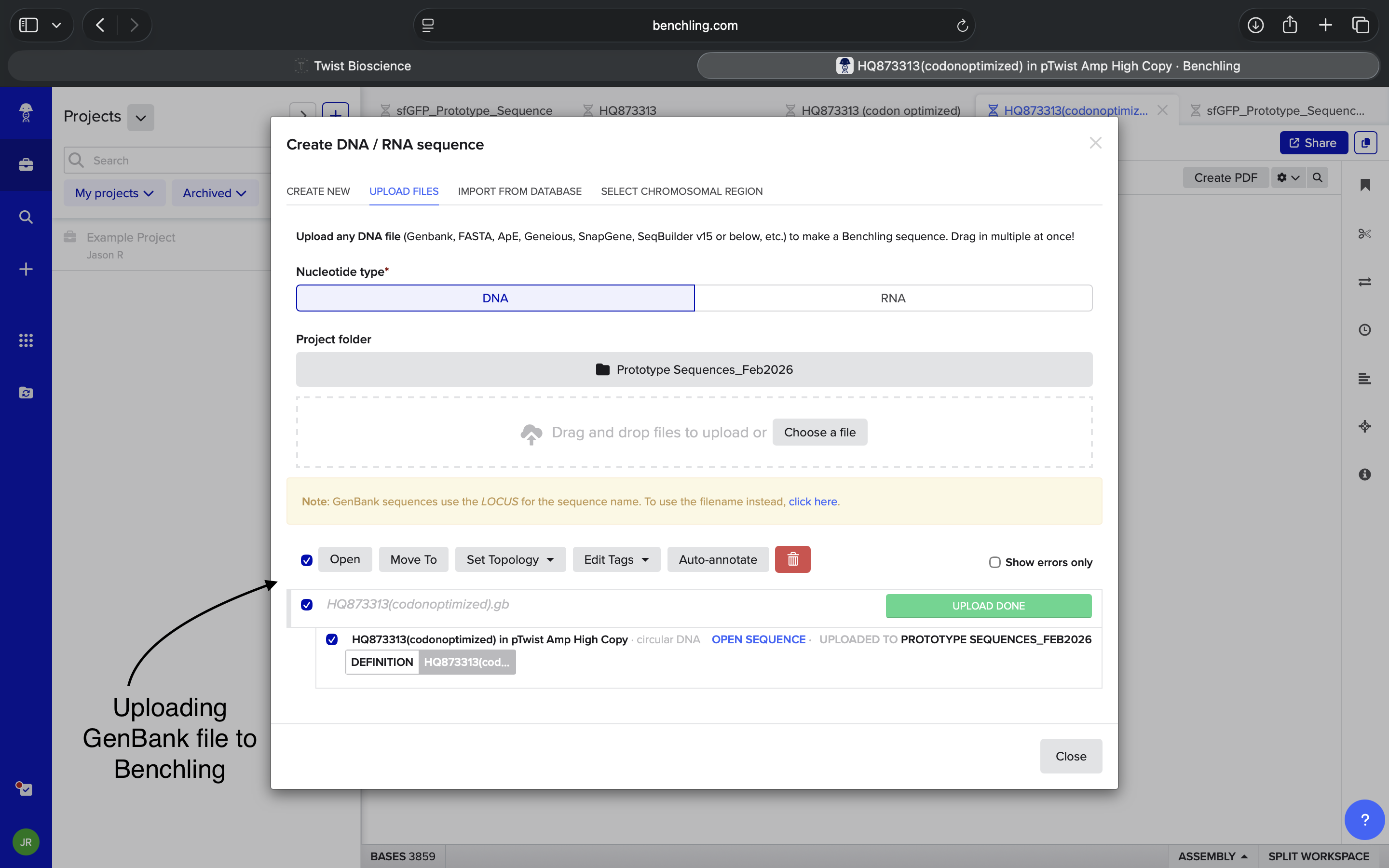
4.5: Import your sequence
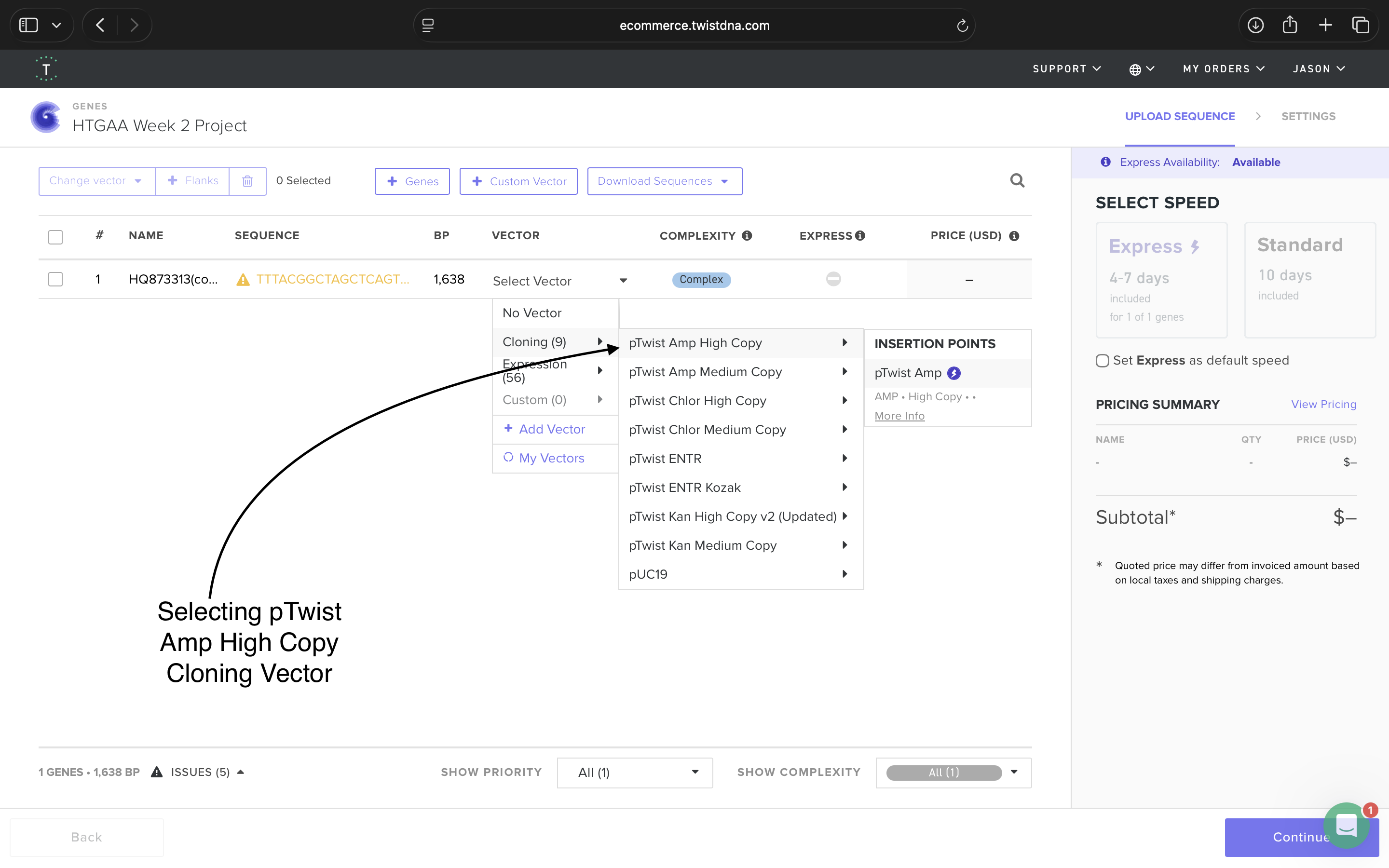
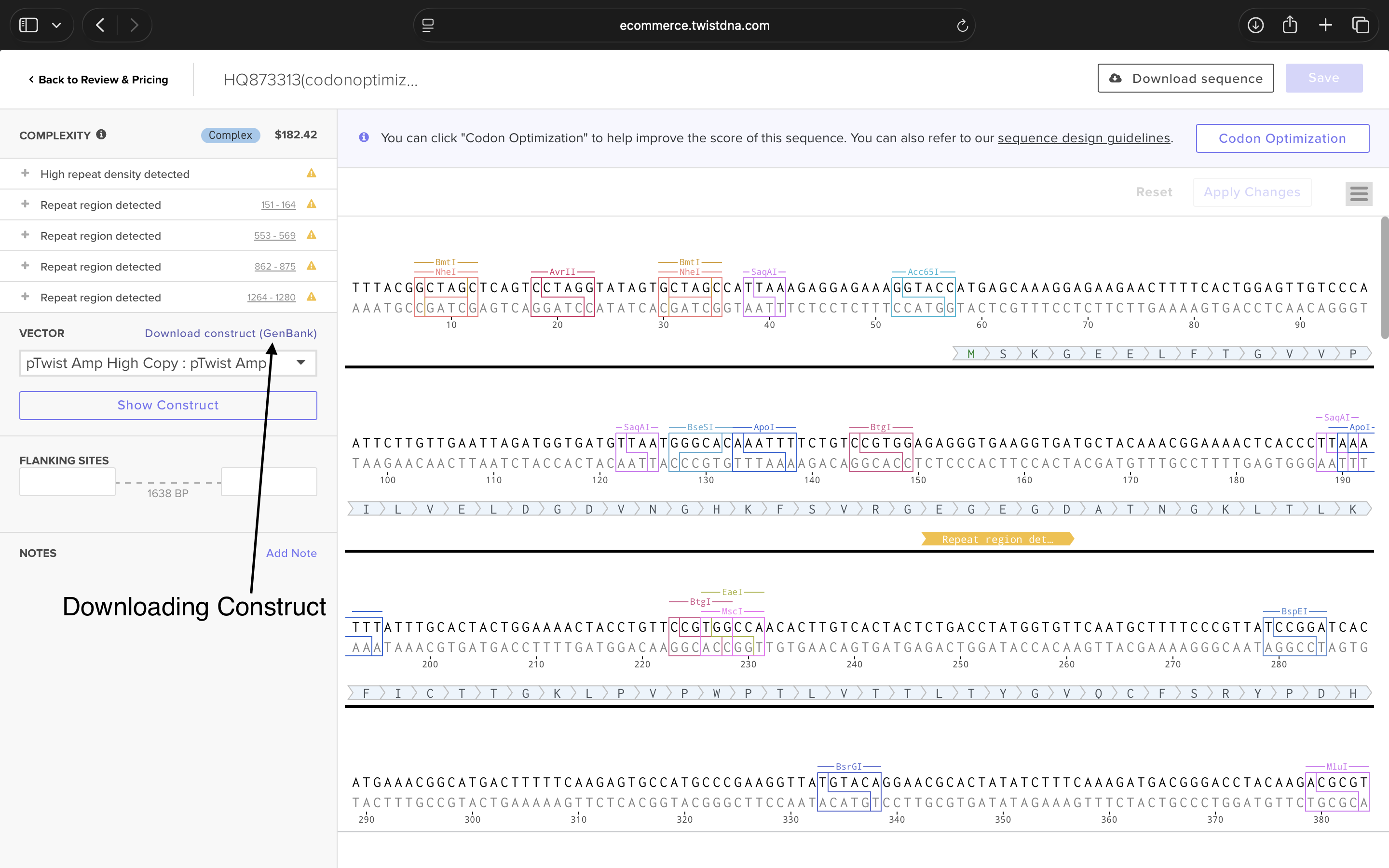
4.6: Choose Your Vector
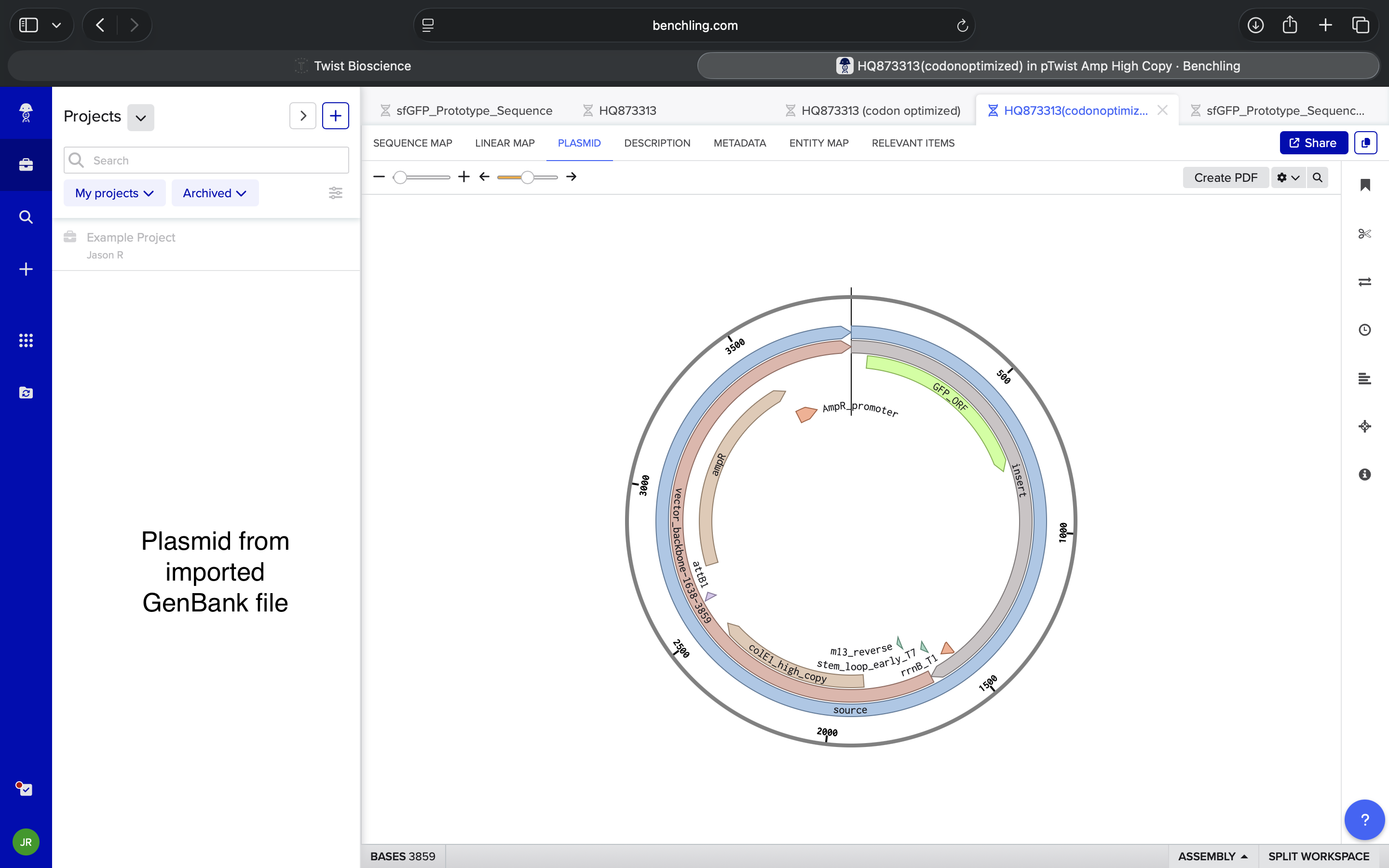
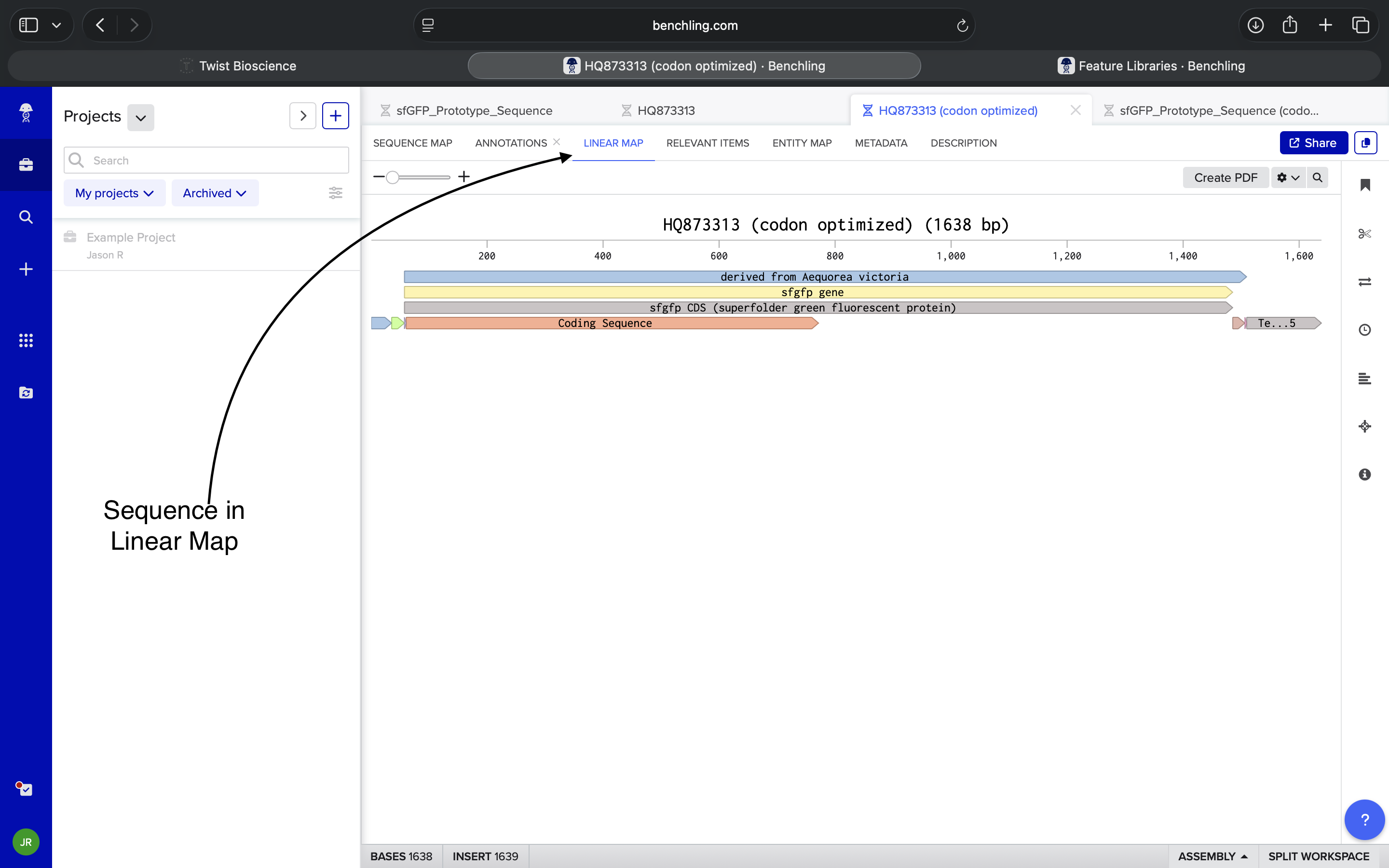
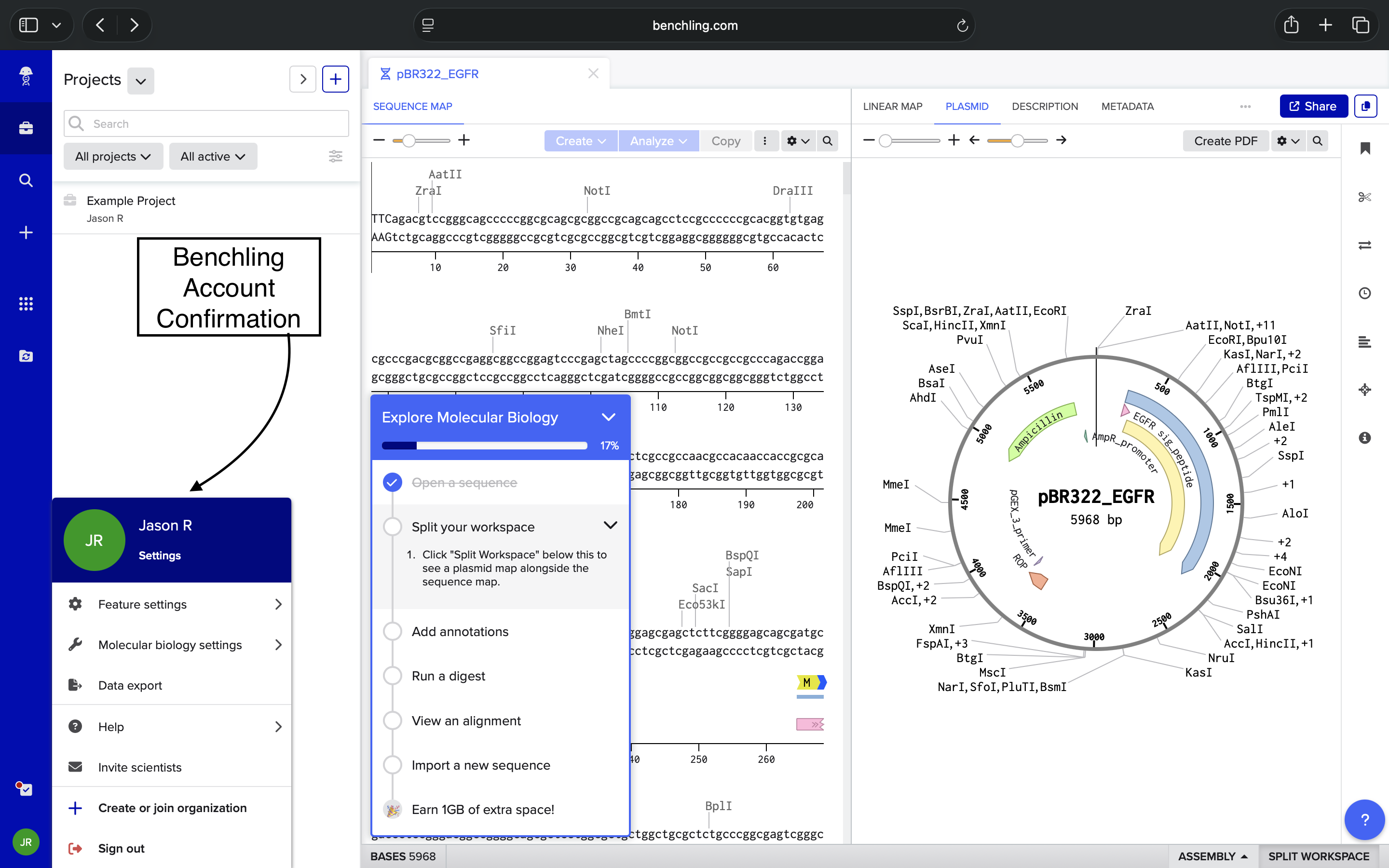
 Imported sequence into Benchling and viewed resulting plasmid
Imported sequence into Benchling and viewed resulting plasmid
All Section 4 Prompts Listed Below
| Supporting Prompt | Source |
|---|---|
| Tell me how to add a Promoter to a sequence in Benchling | Perplexity |
| Found this information from the Registry of Standard Biological Parts: BBa_J23106 Can you break down what this naming convention means and how I can find the relevant Promoter information in a sequence based on this naming convention? Do NOT hallucinate. If you don’t know the answer, say so | Perplexity |
| What is an alignment in Benchling? In Benchling, how do I put a codon optimized sequence under or next to a sequence I originally imported? Do NOT hallucinate when answering this question | Perplexity |
| How do I replace a sequence in Benchling with a codon-optimized sequence? | Perplexity |
| Bit confused regarding how to find a Promoter in a sequence in Benchling. I tried Auto-Annotate and it doesn’t seem to be working. Where should I go from here? | Perplexity |
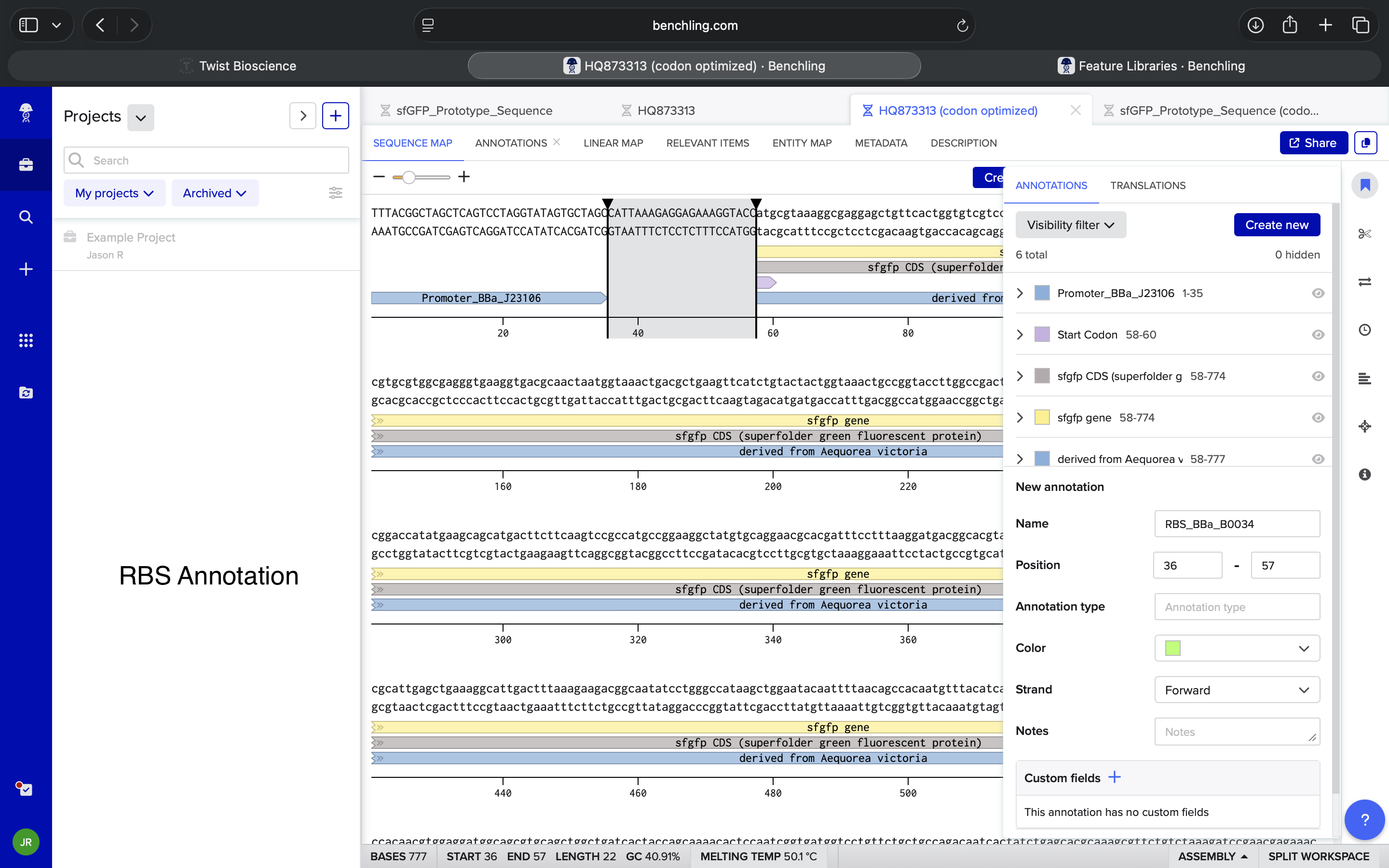
| What is an RBS in Benchling? | Perplexity |
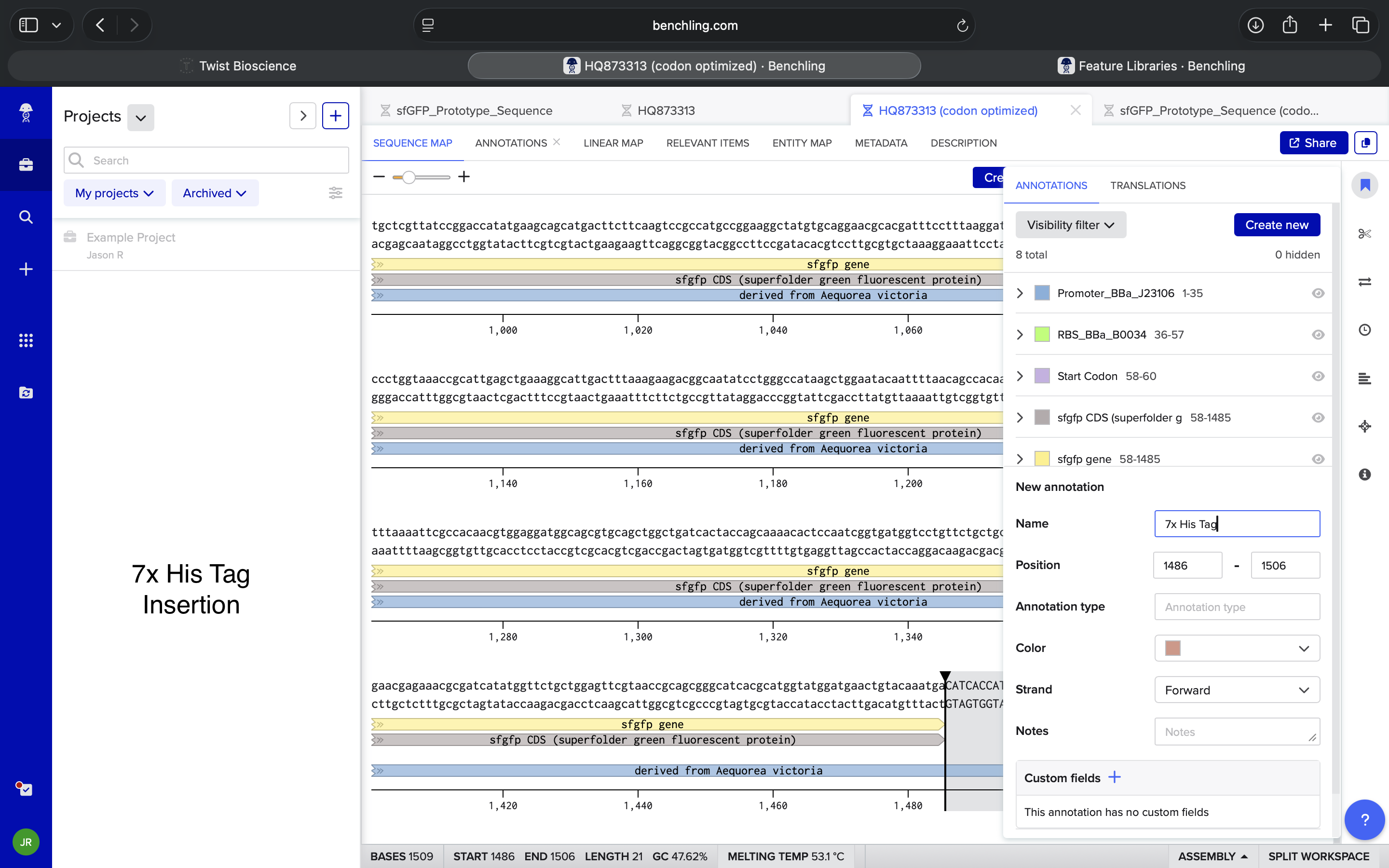
| What is a 7x His Tag? What is a Terminator? How do I find these in Benchling? Where are these traditionally inserted into a sequence in Benchling? | Perplexity |
| How do I paste sequences into a Benchling file? | Perplexity |
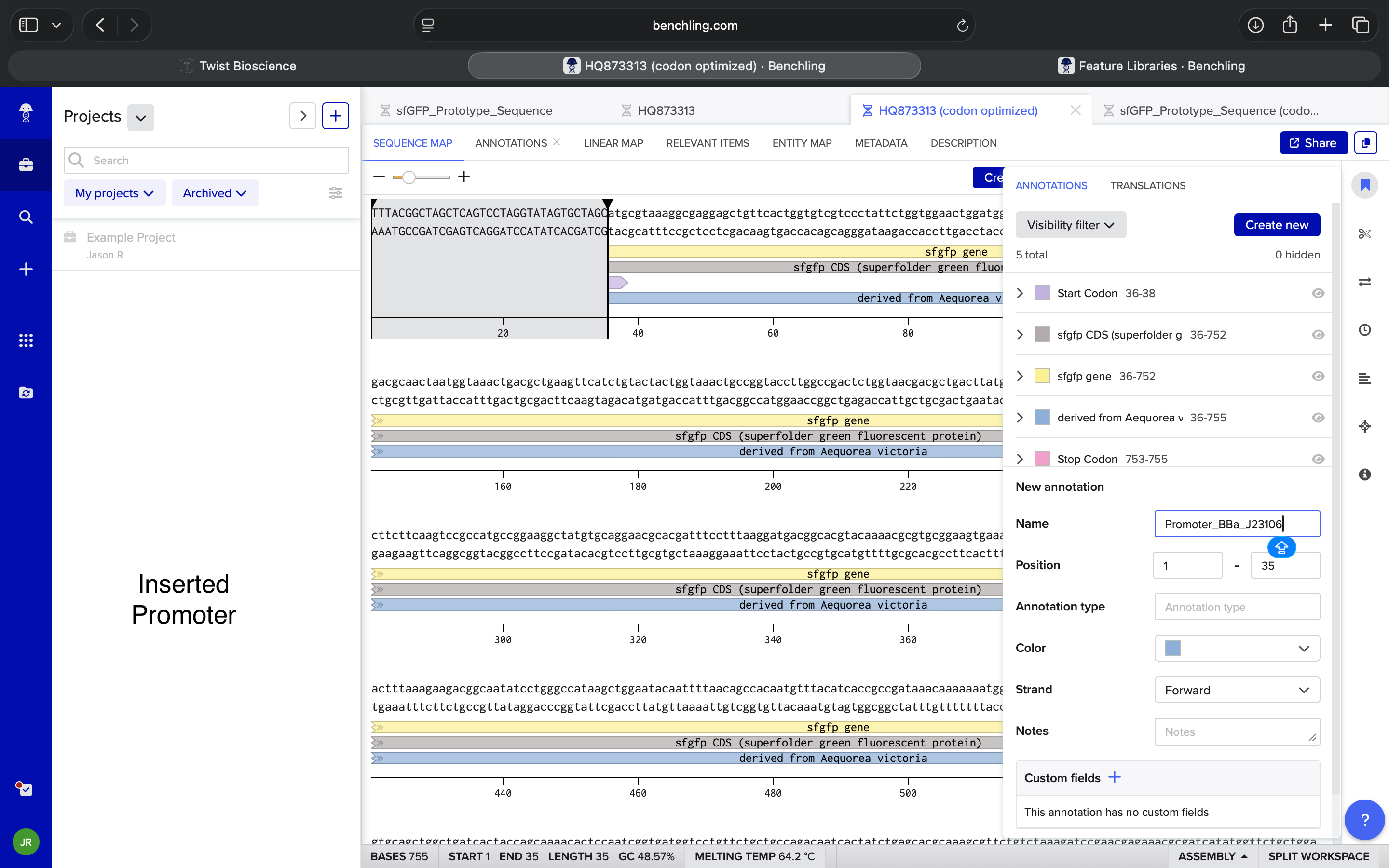
| How do I know where to insert a Promoter into a given sequence in Benchling? | Perplexity |
| Not totally understanding. If the start codon (the ATG) represents the start of the sequence, how do I insert something before that in Benchling? | Perplexity |
| What is an RBS? Where are they traditionally inserted into a sequence? | Perplexity |
| What do spacers look like in Benchling? Is it literally just empty space with no letters/codons? Something tells? | Perplexity |
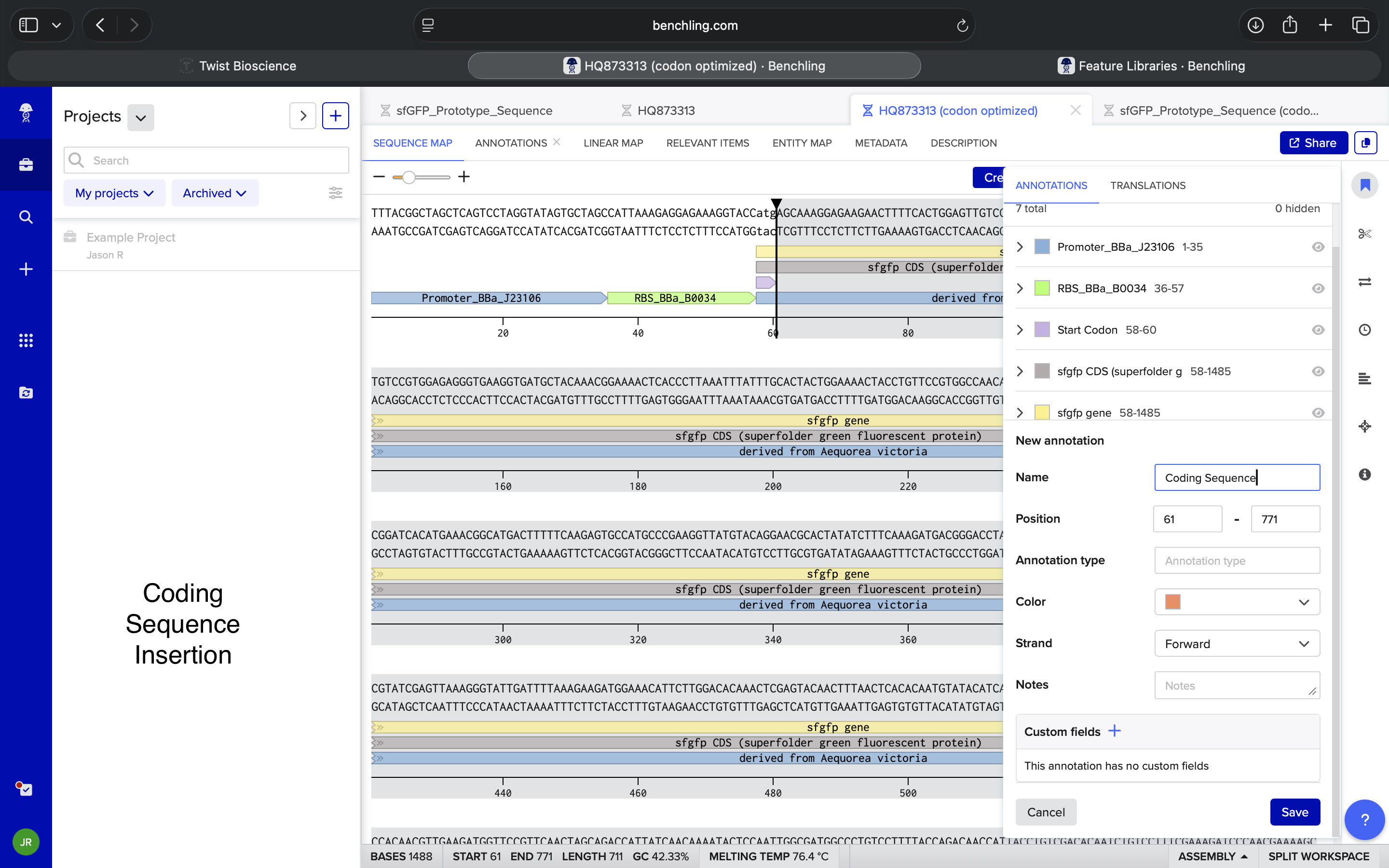
| Where is a coding sequence traditionally inserted in a codon optimized sequence in Benchling? If there’s something off in what I’m saying, let me know | Perplexity |
| Where is a C-terminus in a protein in Benchling? | Perplexity |
| How do I find an amino acid view for a sequence in Benchling? | Perplexity |
| In Benchling, if I’m inserting a 7x His Tag and a Terminator, and I have a stop codon in my sequence, what is the traditional sequence? Is it 7x His Tag, stop codon, Terminator? Something else? | Perplexity |
| Any way I can add a Schema to a sequence after the fact in Benchling? | Perplexity |
Part 5: DNA Read/Write/Edit
5.1 DNA Read
(i) What DNA would you want to sequence (e.g., read) and why?
I want to sequence the DNA of the bdelloid rotifer Adineta genus, a microscopic-sized invertebrate that kind of looks like a worm. I’m fascinated by its ability to sustain cryptobiosis for thousands (in the case of a bdelloid rotifer thawed out in Russia in 2015, more than 24,000!!) of years. Transgenesis of the bdelloid rotifer’s cryptobiotic abilities in mammalian organisms could have profound impacts on the future of the species, specifically the ability for groups of homo sapiens or other future sapiens forms to engage in interstellar travel over large durations of time and space. More information on the bdelloid rotifer Adineta genus and its cryptobiotic abilities can be found in the footnote at the end of this sentence 4.
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
- I’d use Next-Generation Sequencing (NGS) on my DNA because it’s well-suited for transgenesis. It’s fast, high-resolution, and allows for massively parallel sequencing (if desired). More importantly, it’s highly precise, meaning it can pinpoint transgene locations within a host genome.
Is your method first-, second- or third-generation or other? How so?
- It’s a second-generation sequencing method, as it emerged in the 2000’s after the advent of Sanger sequencing in the 1970’s and before the advent of single-molecule sequencing in the 2010’s.
What is your input? How do you prepare your input (e.g. fragmentation, adapter ligation, PCR)? List the essential steps.
- If the gene exists in an organism, the initial input is extracted genomic DNA from that particular organism. Otherwise, the initial input can be a plasmid (if the DNA’s already cloned) or complementary DNA (cDNA) if only mRNA is available. Essential preparation steps listed below (assuming gene exists in an organism):
- Isolate or extract the gene; lyse cells or tissues from donor, remove proteins and contaminants
- Fragment the isolated or extracted gene into fragments of apprxo. ~200-600bp
- Convert ragged ends into blunt or sticky ends
- Attach adapter sequences to ends of each fragment
- Enrich fragments of the intended size (i.e., the size you want), removing any remaining small artifacts and excess adapters
- Review fragement size distribution
- Convert double-stranded library into single strands if necessary
- Load into sequencing instrument
- If the gene exists in an organism, the initial input is extracted genomic DNA from that particular organism. Otherwise, the initial input can be a plasmid (if the DNA’s already cloned) or complementary DNA (cDNA) if only mRNA is available. Essential preparation steps listed below (assuming gene exists in an organism):
What are the essential steps of your chosen sequencing technology, how does it decode the bases of your DNA sample (base calling)?
- The essential NGS steps are listed below:
- Preparation (see above)
- Amplification: Many copies of each DNA fragment are created on flow cells. One end of each fragment sticks to a primer, gets copied to a complementary strand, and bends over to stick to a primer. This bridging repeats nth times, forming amplified (hence the name) clusters of these identical grouped fragments
- Sequencing: A polymerase enzymes adds 1 colored flourescent nucleotide to each strand in each cluster of grouped fragments. A camera takes a picture of recording the color for the given nucleotide (revealing its A, T, G, or C base) then chemicals are used to wash away any free-floating nucleotides that aren’t part of a given cluster.
- Base Decoding/Base Calling: A computer analyzes these colored image clusters, assigning each cluster a sequence of bases based on its colors. This analysis then becomes a text string of DNA code, with confidence ratings per base sequence based on the resolution of the read 5
- The essential NGS steps are listed below:
What is the output of your chosen sequencing technology?
- See above. NGS outputs a text string of DNA code, with confidence ratings per base sequence
5.2 DNA Write
(i) What DNA would you want to synthesize (e.g., write) and why?)
- See answer to 5.1 (i). I’m fascinated by the potential of transgenesis of the bdelloid rotifer Adineta’s cryptobiotic abilities for interstellar travel, and therefore, am very interested in reading and editing its sequence.
NOTE: I found a bdelloid rotifer Adineta sequence in the link in the footnote. However, the raw FASTA information is so long that upon insertion into this webpage, it seemingly broke the webpage, or caused it to freeze up (pardon the unintentional cryptobiosis pun) 6
(ii) What technology or technologies would you use to perform this DNA synthesis and why?
- I’d use PCR amplification, chemical synthesis, and restriction enzymes and ligation to perform this DNA synthesis. I’d use PCR so I can make many copies of the original DNA, I’d use chemical synthesis so I can clone the DNA into a given plasmid, encapsulating the DNA for desired level expression in an appropiate vehicle, and I’d use restriction enzymes and ligation for precise synthesis.
- What are the essential steps of your chosen sequencing methods?
- PCR amplification:
- Denaturation: Heat DNA so it breaks into stingle strands
- Annealing: Cool the DNA, allowing primers to bind to sites in target gene
- Extension: DNA polymerase adds nucleotides from the primer starting point, allowing each strand to fully copy
- Chemical synthesis:
- Deprotection: Remove protecting DMT group
- Base Coupling: Add protected phosphoramidite nucleotide for phosphate linkage
- Capping: Cap the chain to prevent errors
- Oxidation: Stabilize phosphate triester bonds
- Restriction enzymes and ligation:
- Stitch oligos from prior chemical synthesis step into a complete gene for insertion into plasmid
- Add flanking restriction enzymes at ends of oligos as needed
- Clean the isolated DNA via gel extract
- Use matching restriction enzymes to incubate the gene insert and its plasmid vector. This incubation process recognizes specific sequences in the gene insert and cut, creating blunt and sticky ends
- Use DNA ligase enzyme to mix compatible ends
- PCR amplification:
- What are the limitations of your sequencing method (if any) in terms of speed, accuracy, scalability?
- A scalability challenge is that it’s difficult to synthesize a sequence of more than 200nt via direct synthesis methods because at a certain point, too many errors accumulate in the synthesized sequence
5.2 DNA Edit
- (i) What DNA would you want to edit and why?
- See answer to 5.1 (i). I’m fascinated by the potential of transgenesis of the bdelloid rotifer Adineta’s cryptobiotic abilities for interstellar travel, and therefore, am very interested in reading and editing its sequence.
- (ii) What technology or technologies would you use to perform these DNA edits and why?
- I’d use CRISPR-Cas9 to perform these DNA edits because when performing transgenesis from an invertebrate to a mammalian vertebrate, it allows for non-random, precise insertion of large amounts of genetic data with reduced risk of unintended or off-target effects.
- How does your technology of choice edit DNA? What are the essential steps?
- CRISPR-Cas9 edits DNA through a multi-stage mechanism. This mechanism is broken down below:
- Recognition: A single guide RNA (sgRNA) pairs with a Cas9 protein
- This pair scans the genome for a 20bp DNA sequence if the sequence is next to a protospacer adjacent motif (PAM)
- If there is a PAM next to the desired 20bp DNA sequence, the Cas9 makes a dobule-stranded break (DSB)
- The DSB then triggers repair mechanisms (either non-homolgous end joining [NHEJ] or homology-directed repair [HDR]). These repair mechanisms allow the desired edited DNA to be incorporated into the sequence
- CRISPR-Cas9 edits DNA through a multi-stage mechanism. This mechanism is broken down below:
- What preparation do you need to do (e.g. design steps) and what is the input (e.g. DNA template, enzymes, plasmids, primers, guides, cells) for the editing?
- Preparation:
- Select target site(s)
- Design and synthesize gRNA
- Build DNA donor templates
- Create mixture of recombinant Cas9 protein and purified gRNA
- Preapre cells/embryos
- Inputs to this process are gRNDA,Cas9 protein, donor DNA (usually linear templates), and a delivery vehicle (usually an injection buffer)
- Preparation:
- What are the limitations of your editing methods (if any) in terms of efficiency or precision?
- HDR can have low efficiency in a transgenesis context
- Off-target or unintended consequences can still occur
- Need PAMs near target sequences for precise DSBs
All Section 5 Prompts Listed Below
| Supporting Prompt | Source |
|---|---|
| Remind me what a digest is in a biotechnological context. I know it has something to do with subdividing DNA sequences into fragments based on enzymes, but there’s some additional information I know I’m missing. Do NOT hallucinate when answering this question | Perplexity |
| What is horizontal gene transfer? A separate question (perhaps): What is the technical term in biotechnology for transferring the abilities of one organism to another (ex. if I wanted to actually give a lizard the ability to fly like a bird by importing genetic properties that allow for the creation of wings for example)? | Perplexity |
| What is it called in biotechnology when traits from one organism are transferred or conferred to another via an engineered process or processes? | Google AI Mode |
| If I want to perform transgenesis in a biotechnological context (i.e., introduce a foreign gene into a new organism to confer a desired trait), and I want to start this process by sequencing the original foreign gene, what is considered the best practice in modern biotechnology for sequencing this original foreign gene? Is this sequencing method first, second, or third generation in the history of biotechnology? From some other period? What essential steps does it involve and how does it decode the bases of the original foreign gene? What is its output? Do NOT hallucinate when answering these questions. If you don’t know the answer to any of these questions, say so | Perplexity |
| How is Next-Generation Sequencing (NGS) considered second generation? Do NOT hallucinate when answering this question | Perplexity |
| If I want to perform transgenesis in a biotechnological context (i.e., introduce a foreign gene into a new organism to confer a desired trait), and I want to start this process by sequencing the original foreign gene via Next-Generation Sequencing (NGS), what is my input at the very beginning of the sequencing process? How is that input prepared for sequencing? Do NOT hallucinate when answering these questions. If you don’t know the answer to any of these questions, say so | Perplexity |
| What is cDNA? | Perplexity |
| I have the following information describing and Illumina Next-Generation Sequencing (NGS workflow: –Library Prep: Extract/isolate the gene (PCR amplify if needed), fragment to ~200-500 bp, add adapters/barcodes.–Amplification: Bridge amplification on flow cell creates clusters of identical fragments.–Sequencing: DNA polymerase incorporates fluorescent reversible terminator nucleotides (A/T/G/C); camera captures color/emission per base, cleaving terminator for next cycle.–Imaging/Analysis: Base calling from images, alignment to reference or de novo assembly. Explain the amplification through imaging/analysis steps to me as if I was a reasonably educated 16 year old without an advanced biotechnology background. Tell me what the terms in the amplification through imaging/analysis steps mean. Do NOT hallucinate when addressing this query | Perplexity |
| What is a polymerase enzyme? Explain this to me as if I was a reasonably educated 16 year old without an advanced biotechnology background | Perplexity |
| Found the following on NCBI: “Uncultured bdelloid rotifer isolate Undet.AN.1.3 cytochrome oxidase subunit I gene, partial cds; mitochondrial"Is that the same as the full bdelloid rotifer genome? Bit confused and am unsure whether or not it isDo NOT hallucinate. If you don’t know the answer to this question, say so | Google AI Mode |
| Yes, I’m looking for a complete nuclear genome of a bdelloid rotifer, although I’m not sure which species of bdelloid rotifer I’m looking for. I thought bdelloid rotifer was its own species. Open to having any misconceptions cleared up. If you can provide me links where I can find a complete bdelloid rotifer nuclear genome, that would be greatly appreciated | Google AI Mode |
| Am aware that there was a bdelloid rotifer that came out of long-term cryptobiosis in Russia back in 2015. Any chance we know/you can find the specific species of this rotifer that had this cryptobiotic ability? If so, I’d like the complete nuclear genome for that rotifer species | Google AI Mode |
| In the answer two prompts ago, there was a mention of “chemicals wash away extras” in the ‘Sequencing by synthesis’ step. What are ’extras’ in this context? Do NOT hallucinate when answering this question | Perplexity |
| Is the workflow sequence described in the answer to the prompt 2 prompts ago Polymerase Chain Reaction (PCR)? Believe so, but am not sure | Perplexity |
| If I want to perform transgenesis and I start by Next-Generation Sequencing (NGS) to read the DNA, what comes after (i.e., what technologies are traditionally used for writing and then editing the DNA or the original organism that has the abilities I want to confer in a host)? Do NOT hallucinate when answering this question | Perplexity |
| In the answer to the last prompt, can you give me the ‘So What?’ or ‘So What’? behind the ‘Writing the DNA (Gene Preparation & Cloning)’ section from a transgenesis perspective? What do gene synthesis steps like “clone it into a plasmid vector with promoter, terminator, and selection marker.” allow one to do in a transgenesis context? What does restriction enzyme digestion and ligation allow one to do? Do NOT hallucinate. If you don’t know the answers to these questions, say so | Perplexity |
| I have a DNA writing workflow consisting of the following steps:–PCR amplification: Use the sequenced gene info to design primers and amplify the gene from the original source DNA.–Gene synthesis: Chemically synthesize the DNA sequence (especially if optimized for the host), then clone it into a plasmid vector with promoter, terminator, and selection marker.–Restriction enzyme digestion & ligation: Cut the vector and gene insert with enzymes, then join (ligate) them using DNA ligase to create a recombinant plasmidCan you give me the ‘So What?’ or ‘So What’? behind this workflow from the perspective of someone who wants to perform transgenesis from one organism to another? What do gene synthesis steps like “clone it into a plasmid vector with promoter, terminator, and selection marker.” allow one to do in a transgenesis context? What does restriction enzyme digestion and ligation allow one to do?Do NOT hallucinate. If you don’t know the answers to these questions, say so | Google AI Mode |
| How does PCR amplification and gene synthesis actually work? What are the essential steps? | Perplexity |
| How does the content in the last prompt related to the Phosphoramidite DNA Synthesis Cycle? Do NOT hallucinate when answering this question | Perplexity |
| What is a dNTP in genomics? | Perplexity |
| What is a ‘phosphate triester’? Explain this to me in simple terms. Do NOT hallucinate | Perplexity |
| Understanding the phosphoramidite DNA synthesis cycle. How does this transition to the essential steps of restriction enzyme digestion and ligation? Do NOT hallucinate when answering this question | Perplexity |
| If I want to perform transgenesis from one organism to another in a biotechnological context, particularly if I want to confer a trait from an invertebrate organism to a vertebrate, mammalian organism, what is/are the recommended DNA technology or technologies for accomplishing this task? What are the benefits and drawbacks, and respective workflows of each ot these technologies? Where does CRISPR fit into the mix? | Perplexity |
| Can you elaborate on the CRISPR-Cas9 workflow from the answer to the last prompt, specifically describing how it edits DNA? Can you make a workflow just focused on that? Explain this workflow to me as if I was a reasonably educated person with some (not extensive) biology and biotechnology knowledge. Do NOT hallucinate when answering this query | Perplexity |
Excerpt from “The Great Siberian Thaw” (New Yorker Magazine; 2022-01-17): “Permafrost thaw has brought to the surface all sorts of mysteries from millennia past. In 2015, scientists from a Russian biology institute in Pushchino, a Soviet-era research cluster outside Moscow, extracted a sample of yedoma from a borehole in Yakutia. Back at their lab, they placed the piece of frozen sediment in a sterilized culture box. A month later, a microscopic, wormlike invertebrate known as a bdelloid rotifer was crawling around inside. Radiocarbon dating revealed the rotifer to be twenty-four thousand years old. In August, I drove out to Pushchino, where I was met by Stas Malavin, a researcher at the laboratory. “It’s one thing for a simple bacterium to come back to life after being buried in the permafrost,” he said. “But this creature has intestines, a brain, nervous cells, reproductive organs. We’re clearly dealing with a higher order.” The rotifer had survived the intervening years in a state of “cryptobiosis,” Malavin explained, “a kind of hidden life, where metabolism effectively slows down to zero.” The animal emerged from this geological “time machine,” as he put it, not just alive but able to reproduce. A rotifer lives for only a few weeks, but replicates itself multiple times through parthenogenesis, a type of asexual reproduction. Malavin removed from the lab fridge a direct descendant of the rotifer that had crawled out of the permafrost and placed it under a microscope. An oval-shaped plankton squirmed around; I imagined this blob, two-tenths of a millimetre in size, as a nervous explorer who awoke to find itself in a strange and unexpected future. “Why be modest?” Malavin asked. Unlocking the secret of how an animal with a complex anatomy was able to shut down for tens of thousands of years and then turn itself back on might, for example, offer hints for using cryogenic conditions to store organs for donation. Neuroscientists at M.I.T. have been in touch. “I’m obviously not saying our findings will lead to people being put into long-term cryogenic slumber tomorrow,” Malavin said. “But it’s a step in that direction.”” ↩︎
I think this might mean the resolution of the image of the cluster ↩︎
https://www.ncbi.nlm.nih.gov/datasets/genome/GCA_021613535.1/ ↩︎