Week 3 HW: Lab Automation


The Power of Lab Automation
Assignment: Python Script for Opentrons Artwork
- 0: Attended this week’s recitation and reviewed the lab information on programming Opentrons
- 1: Generated an artistic design using Ronan’s Opentrons GUI 1
- 2: Artistic Design Python Script: See script in URL below:
- 3: Listing my sfgfp point coordinates from Ronan’s Opentrons GUI below (the shape is a rightward-facing green arrow):
[(6.6,11), (8.8,11), (11,11), (8.8,8.8), (11,8.8), (13.2,8.8), (11,6.6), (13.2,6.6), (15.4,6.6), (13.2,4.4), (15.4,4.4), (17.6,4.4), (15.4,2.2), (17.6,2.2), (19.8,2.2), (17.6,0), (19.8,0), (22,0), (-22,-2.2), (-19.8,-2.2), (-17.6,-2.2), (-15.4,-2.2), (-13.2,-2.2), (-11,-2.2), (-8.8,-2.2), (-6.6,-2.2), (-4.4,-2.2), (-2.2,-2.2), (0,-2.2), (2.2,-2.2), (4.4,-2.2), (6.6,-2.2), (8.8,-2.2), (11,-2.2), (13.2,-2.2), (15.4,-2.2), (17.6,-2.2), (19.8,-2.2), (22,-2.2), (24.2,-2.2), (-22,-4.4), (-19.8,-4.4), (-17.6,-4.4), (-15.4,-4.4), (-13.2,-4.4), (-11,-4.4), (-8.8,-4.4), (-6.6,-4.4), (-4.4,-4.4), (-2.2,-4.4), (0,-4.4), (2.2,-4.4), (4.4,-4.4), (6.6,-4.4), (8.8,-4.4), (11,-4.4), (13.2,-4.4), (15.4,-4.4), (17.6,-4.4), (19.8,-4.4), (22,-4.4), (24.2,-4.4), (26.4,-4.4), (-22,-6.6), (-19.8,-6.6), (-17.6,-6.6), (-15.4,-6.6), (-13.2,-6.6), (-11,-6.6), (-8.8,-6.6), (-6.6,-6.6), (-4.4,-6.6), (-2.2,-6.6), (0,-6.6), (2.2,-6.6), (4.4,-6.6), (6.6,-6.6), (8.8,-6.6), (11,-6.6), (13.2,-6.6), (15.4,-6.6), (17.6,-6.6), (19.8,-6.6), (22,-6.6), (24.2,-6.6), (17.6,-8.8), (19.8,-8.8), (22,-8.8), (15.4,-11), (17.6,-11), (19.8,-11), (13.2,-13.2), (15.4,-13.2), (17.6,-13.2), (11,-15.4), (13.2,-15.4), (15.4,-15.4), (8.8,-17.6), (11,-17.6), (13.2,-17.6), (6.6,-19.8), (8.8,-19.8), (11,-19.8)]
- 4: Used Gemini 2.5 Flash (built into Google Colab) to assist with completing the coding portion of the homework. I have some rough Python knowledge via a Codecademy course, which helped get things started (i.e., I did do some of the coding for this assignment).
All Gemini 2.5 Flash prompts are listed below:
| Supporting Prompt | Model |
|---|---|
| I want to create some code similar to the code in Examples 1-7. What are the core elements of the code I need to create? | Gemini 2.5 Flash |
| Been working on some code in the ‘Your Code’ module. Have made a single green dot so far. Looking to create a rightward-facing green arrow based on these coordinates in the attached .py file Tell me how the code in the ‘Your Code’ module under the #Aspirate subsection needs to be edited to output the rightward facing green arrow in the attached .py file | Gemini 2.5 Flash |
| Wondering if you could help explain something. Not seeing an actual visualization of a green arrow below. Where is it? Can you give me a picture output of the code similar to the picture output located in the examples in this URL below? https://colab.research.google.com/drive/1VoouRH0nqlk09g50rHxOElaLD-SVknYY#scrollTo=PsOgJ2DndZzt | Gemini 2.5 Flash |
Ensure the ‘run’ function executes all arrow_points | Gemini 2.5 Flash |
| Still getting a single green dot when I run the simulator. Have inputted the coordinates for the rightward-facing green arrow based on the attached file and am aware this will need to likely need a for loop to aspirate the colors. | Gemini 2.5 Flash |
| Still getting a single green dot when I run the simulator. Have inputted the coordinates for the rightward-facing green arrow based on the attached file and am aware this will need to likely need a for loop to aspirate the colors. Recommendations on how to proceed? | Gemini 2.5 Flash |
| Explain why only 1uL was dispensed instead of 20uL | Gemini 2.5 Flash |
| Looking at the code and see that I inputted the points for the rightward-facing arrow under arrow_points. Not understanding what parts of the code need to be changed (if any) so that when the simulation runs, a rightward-facing green arrow is outputted | Gemini 2.5 Flash |
| Thank you. Can you please tell me where cell “pczDLwsq64mk” is located, so I can have an idea of the code used to create the rightward-facing arrow in the simulation? | Gemini 2.5 Flash |
- 5: Coordinating robot time slot with William & Mary node
- 6: Submitted Python file via assignment form (see screenshot below):
 Python Form Submission Confirmation Screenshot
Python Form Submission Confirmation Screenshot
Post-Lab Questions
Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
- The paper An Automated Versatile Diagnostic Workflow for Infectious Disease Detection in Low-Resource Settings was published in the journal Micromachines in 2024 and describes using simple Commercial Off-the-Shelf (COTS) reagents and lab equipment, along with an Opentrons robot, to create an automated workflow for detecting diseases in low-resource settings 2. What’s interesting about the paper is that the workflow the researchers designed reduced the time for detecting a pathogen (in this case meningitis) by approx. 18% (total timne of 118 min.) with an almost 5.8x reduction in cost for sample processing 3. The total cost of to run 8 samples for meningitis detection was approx. $126 USD, a cost savings that matters in a low-resourced environments. The findings and extension of this paper’s workflow shows promise for decentralized disease detection in low-resource settings during future health security incidents. Except for opening and closing of tube lids, the following four steps of the autonated workflow were completed by the Opentrons robot:
- DNA isolation with Dynabeads
- Sample incubation
- Washing step
- DNA resuspension
- DNA amplification
- Recombinase Polymerase Amplification (RPA) mix preparatoon
- RPA amplification
- DNA digestion
- Exonuclease digestion
- DNA dtection
- Preparation of Vertical Flow Microarray (VFM) solutions
- Addition of samples to VFM
- Signal enhancement
- DNA isolation with Dynabeads
Associated AI prompts to address this question included below:
- The paper An Automated Versatile Diagnostic Workflow for Infectious Disease Detection in Low-Resource Settings was published in the journal Micromachines in 2024 and describes using simple Commercial Off-the-Shelf (COTS) reagents and lab equipment, along with an Opentrons robot, to create an automated workflow for detecting diseases in low-resource settings 2. What’s interesting about the paper is that the workflow the researchers designed reduced the time for detecting a pathogen (in this case meningitis) by approx. 18% (total timne of 118 min.) with an almost 5.8x reduction in cost for sample processing 3. The total cost of to run 8 samples for meningitis detection was approx. $126 USD, a cost savings that matters in a low-resourced environments. The findings and extension of this paper’s workflow shows promise for decentralized disease detection in low-resource settings during future health security incidents. Except for opening and closing of tube lids, the following four steps of the autonated workflow were completed by the Opentrons robot:
| Supporting Prompt | Model |
|---|---|
| Can you find me biotechnology related papers from the past 5 years (ideally from Western sources) that incorporate Opentrons or another lab automation tool to create or research a novel biosecurity application | Google Scholar Labs |
| Give me a rundown of what primers are in genomics in simple terms (don’t get overly technical, if possible). Explain this to me as if I was a reasonably educated 15 year-old. Tell me if primers are found in nature or if they’re an artificial construct. Tell me how primers are used in biotechnology Do NOT hallucinate when answering this query | Perplexity |
| In a gene amplification context, what does vortexing mean? | Perplexity |
| What are amplicons? | Perplexity |
| What is a ctrA gene? | Perplexity |
| What is a reagent in a diagnostic or biotechnology context? | Perplexity |
- Write a description about what you intend to do with automation tools for your final project.
- Phage isolation experiments generally require enriching bacterial strains, filtering out bacterial particles, pouring the phage-containing mixture in with fresh bacteria on agar, plating and re-plating phage plaques (areas of phage propagation and bacetrial destruction on agar), and characterizing the resulting phage plaques on the agar. My working thoughts are that I might actually be able to create a somewhat automated workflow a-la the paper referenced in the previous question to help with the filteration through characterization steps of a phage isolation experiment. This might involve using an Opentrons robot and COTS equipment to:
- Handle bacterial liquids
- (Maybe) use an Opentrons module to shake bacterial liquids and reagents
- Operate centrifuge for spinning bacteria
- Pour agar with phage plaques
- Operate micropipettes for phage testing
- Operate Thermocycler module for amplifying specific lysed areas
- Rapidly run software to characterize novel phage DNA
- Phage isolation experiments generally require enriching bacterial strains, filtering out bacterial particles, pouring the phage-containing mixture in with fresh bacteria on agar, plating and re-plating phage plaques (areas of phage propagation and bacetrial destruction on agar), and characterizing the resulting phage plaques on the agar. My working thoughts are that I might actually be able to create a somewhat automated workflow a-la the paper referenced in the previous question to help with the filteration through characterization steps of a phage isolation experiment. This might involve using an Opentrons robot and COTS equipment to:
Note: The maeterial above is not set in stone. It’s an outline of potential automation options for a potential final project (Space (LEO and beyond) or Suborbital Phage Isolation)
Associated AI prompts to address this question included below:
| Supporting Prompt | Model |
|---|---|
| What do phage isolation experiments usually entail? Are any elements of phage isolation experiments dangerous to human health and safety, and if so, why? | Perplexity |
| Take the steps in the “What phage isolation usually involves” section of the last prompt and break down the tools traditionally used for each step. Do NOT hallucinate when answering this question | Perplexity |
| What is supernatant? | Perplexity |
| What are plaques in a phage isolation experiment context? | Perplexity |
| What does it mean to ‘pellet’ bacteria? | Perplexity |
Final Project Ideas
- Submitted 3 Final Project ideas in my node’s section of the slide deck (see screenshot below):
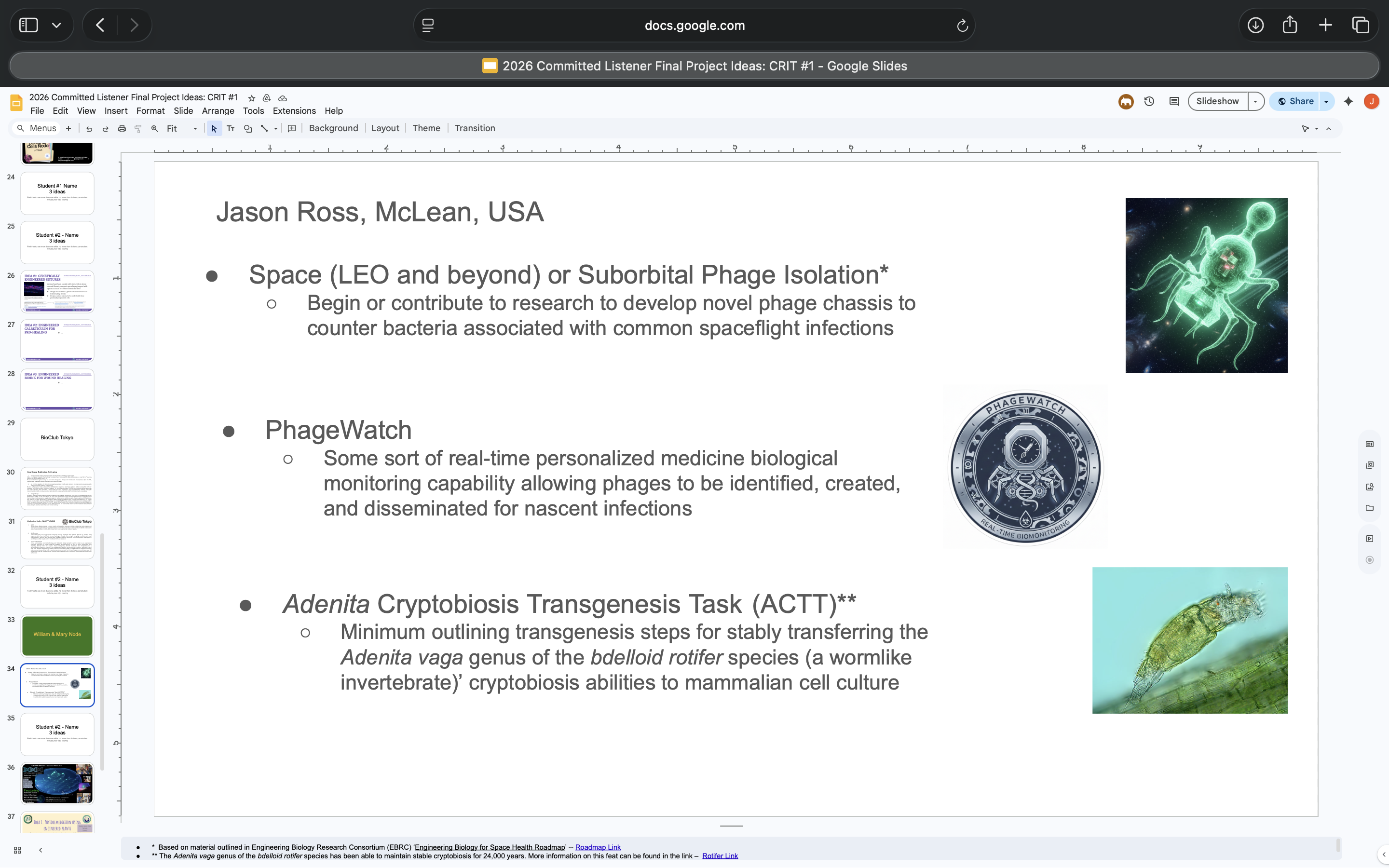

All supporting prompts for Final Project Ideas Slide listed below
| Supporting Prompt | Model |
|---|---|
| What is the microbiome? It’s the gut right? If I’m oversimplifying with the second question, let me know Do NOT hallucinate when answering these questions | Perplexity |
| Does a biotechnological equivalent of ‘Build Your Own Phage’ a-la Build a Bear or Lego exist in the real world? If so, does this capability exist in a personalized medicine context, or is it operable in remote environments? Do NOT hallucinate when answering this question | Perplexity |
| Tell me about the adenita vaga bdelloid rotifer that was able to maintain cryptobiosis for 24,000 years | Google AI Mode |
| Tell me about the adenita vaga bdelloid rotifer that went into cryptobiosis for 24,000 years | Google AI Mode |
| Thinking about creating some sort of real-time personalized medicine biological monitoring capability allowing phages to be identified, created, and disseminated for nascent infections. Create some type of logo image for this concept (the kind you might find on a sticker) that combines the elements of a phage and a time-keeping watch. Let’s not have it be too cartoony or effused with excessive color | Gemini |
| Nice. Change the writing on the top to PhageWatch as opposed to PhageGuard | Gemini |
This is for running samples through the workflow not for equipment like the Opentrons OT-One-Hood ↩︎