Week 6 HW: Genetic Circuits Part 1


- Robot Crafting Genetic Circuit (Stylized)
DNA Assembly
- What are some components in the Phusion High-Fidelity (HF) PCR Master Mix and what is their purpose?
- HF DNA Polymerase: This is the enzyme responsible for copying DNA as it moves from the 5’ to the 3’ position across the DNA
- Deoxynucleotide triphosphates (dNTPs): These are the DNA molecular building blocks, consisting of Adenine (A), Thymine (T), Cytosine (C), and Guanine (G) variants
- HF Buffer: This consists of magnesium chloride, which is salt added to the reaction. It matters because it dissolves into Mg²⁺, which helps nucleotides bond during the reaction
- What are some factors that determine primer annealing temperature during PCR?
- Some factors that determine primer annealing tempeature during PCR include:
- Primer lengths
- Primer melting tempratures
- GC content/sequence content
- Buffer components
- Some factors that determine primer annealing tempeature during PCR include:
- There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
- PCR: PCR creates new linear DNA fragments by via enzymatic amplification of a given region nth number of times. The PCR protocol essentially consists of setting up reaction mixes, denaturating the DNA into single strends, annealing so primers can anneal to specific complementary sequences, extension so the polymerase can syntehsize a new strand, and then repeating this as many times as neccessary. This method might be more useful when there is a specific fragment of DNA one wants to amplify for further use.
- Restriction Enzyme Digests: Restriction Enzyme Digests create new linear DNA fragments by cutting DNA at specific points/recognition sites. The Restriction Enzyme Digest protocol consists of setting up a reaction mix, incubation, and then stopping the reaction. This method might be more useful when there is a specific fragment of DNA one wants to isolate for further analysis.
- How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
- You can ensure the DNA sequences have appropriate 5’ –> 3’ orientation with corresponding overlaps. Fragments salso need to cover the relevant region for cloning, and also need to be inserted at the appropriate molar ratio relative to the plasmid backbone (vector). This is usually a 2:1 ratio.
- How does the plasmid DNA enter the E. coli cells during transformation?
- The plasmid DNA enters the E. coli either via heat shock (temperature change) or electroporation (high electrical voltage). Both methods shock the E. coli cell, causing its cell membrane to open for the plasmid DNA to enter.
- Describe another assembly method in detail (such as Golden Gate Assembly)
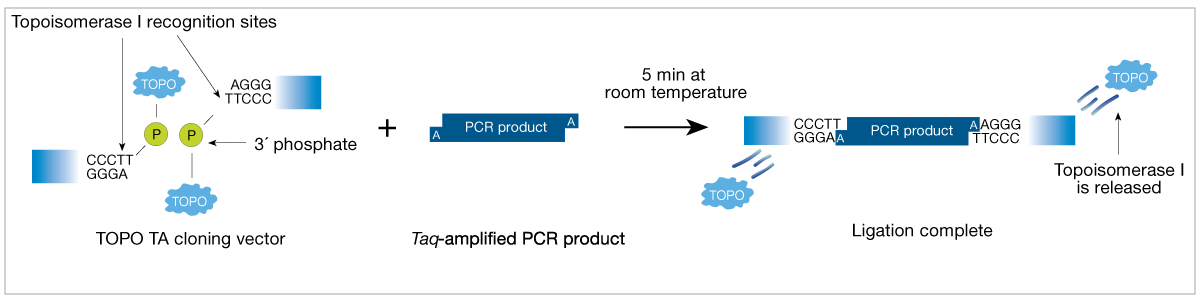
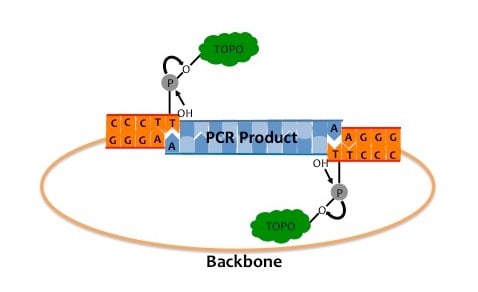
- DNA topoisomerase I (TOPO) Cloning: TOPO cloning’s traditionally used, as it’s a fast, reliable method for cloning products from PCR for later sequencing, etc. The first step in TOPO cloning is generating an insert with Taq polymerase via PCR. This creates inserts with an A-overhang, which can then help address the second step. The second step is to combine this PCR product with the TOPO vector. This is usually done for a couple of minutes. The insert’s 5’ OH/hydroxyl interacts with the TOPO DNA at its end, and as part of this process A and T base pairing occurs between the respective insert and the vector . Then the TOPO religates the strangs and dissociates, creating a closed circular plasmid with the given insert. See diagrams below:
Supporting prompts for this section listed below:
| Supporting Prompt | Model |
|---|---|
| Within the context of Gibson Assembly (biotechnology DNA assembly method), why exactly are molar ratios (apparently they need to be 2:1, insert:vector) important? What are molar ratios? Do NOT hallucinate when replying to this prompt | Perplexity |
| What exactly is the insert and what exactly is the vector within the context of the Gibson Assembly DNA Assembly method? Do NOT hallucinate when replying to this prompt | Perplexity |
| In the context of biotechnology and synthetic biology, what exactly is a plasmid backbone? Explain this to me as if I were a reasonably educated 16-year old. Do NOT hallucinate when addressing this prompt | Perplexity |
| Tell me about the Phusion High-Fidelity (HF) Polymerase Chain Reaction (PCR) Master Mix. What is it? What are its subcomponents and what do they do? Do NOT hallucinate when addressing this prompt | Perplexity |
| The 4 dNTPs referenced in the answer to the last prompt are essentially chemical mixtures, correct? If that is incorrect, what are they? Do NOT hallucinate when answering this prompt | Perplexity |
| In the answer to the prompt before the previous prompt, there was reference to MgCl₂, and Mg²⁺. What are MgCl₂, and Mg²⁺ respectively? Are they chemicals? Something else? Why do they matter? Do NOT hallucinate when addressing this prompt | Perplexity |
| Within the context of a Polymerase Chain Reaction (PCR), I believe primers are the pieces of DNA that get copied nth number of times, correct? If I’m mistaken, indicate as such, and the error in the initial reasoning. Do NOT hallucinate when addressing this prompt | Perplexity |
| So based on the answer to the last prompt: –Primers essentially define the space in the DNA sequence that will be copied? –What is a free 3′‑OH end? Explain this to me as if I were a relatively educated 16-year old Do NOT hallucinate when answering this prompt | Perplexity |
| Do primer pairs always need to have a temperature difference of 5°C from each other? If so, why? Do primer pairs always need to at a temperature of between 52–58°C before annealing? If so, why? What factors determine ideal primer annealing temperatures, and why? Do NOT hallucinate when addressing these prompts | Perplexity |
| Both Polymerase Chain Reaction (PCR) and Restriction Enzyme Digests create linear DNA fragments. PCR creates these linear DNA fragments via enzymatic amplification and Restriction Enzyme Digests create these by essentially cutting the DNA, correct? What do the basic steps of each look like (in some simple broken down steps)? Do NOT hallucinate when addressing this prompt | Perplexity |
| Other than PCR, Restriction Enzyme Digest, and Golden Gate Assembly, what other DNA assembly methods exist? Do NOT hallucinate when addressing this prompt | Perplexity |
| Which of the following results from the answer to the previous prompt is easiest to model in Benching? EXCLUDE Gibson Assembly from the selection and do NOT hallucinate when addressing this prompt | Perplexity |
| Ok. Based on the content in the answer to the prompt before the last prompt, explain to me what TOPO cloning is. What is a TOPO? What does it consist of/what are the basic steps? Do NOT hallucinate when addressing this prompt | Perplexity |
| Based on the answer to the last prompt, what exactly is Taq polymerase again? When we say TOPO cloning is ’ligase-free’, what do we mean when we say that? Why is TOPO cloning traditionally used? | Perplexity |
| In the context of the answer to the last prompt, why would one clone a PCR product using TOPO cloning? An insert is just the piece/fragment of DNA being inserted into the vector (usually a bacterial plasmid), correct? Do NOT hallucinate when addressing this prompt | Perplexity |
| Break down the basic steps of how I would model basic (i.e., not complicated TOPO Cloning) in Benchling. Do NOT hallucinate when addressing this prompt | Perplexity |
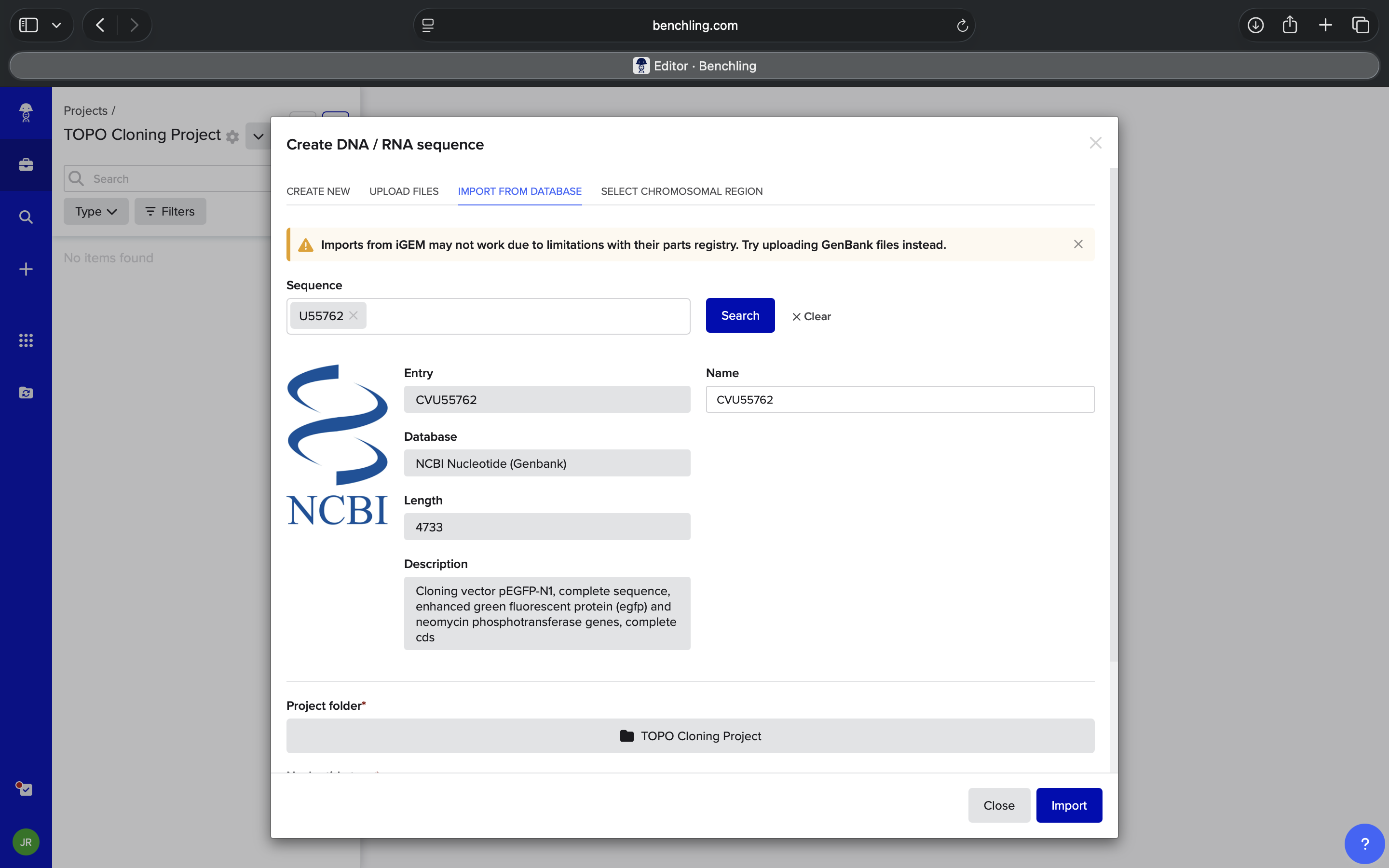
| Where can I find a PCR insert to insert into Benchling for TOPO Cloning? Can I basically choose anything from GenBank or UniProt? What should I be looking for in the context of TOPO Cloning? Do NOT hallucinate when addressing this prompt | Perplexity |
| Thanks. Based on the answer to the last prompt, EGFP is a green fluorescent protein, correct? Do NOT hallucinate when addressing this prompt | Perplexity |
| Is TOPO Cloning a form of homologous, homology-based cloning? Do NOT hallucinate when addressing this prompt | Perplexity |
| Would Benchling’s ‘Concantenate sequences’ feature work/be suitable if one was trying to model TOPO Cloning within Benchling? Why or why not? Do NOT hallucinate when addressing this prompt | Perplexity |
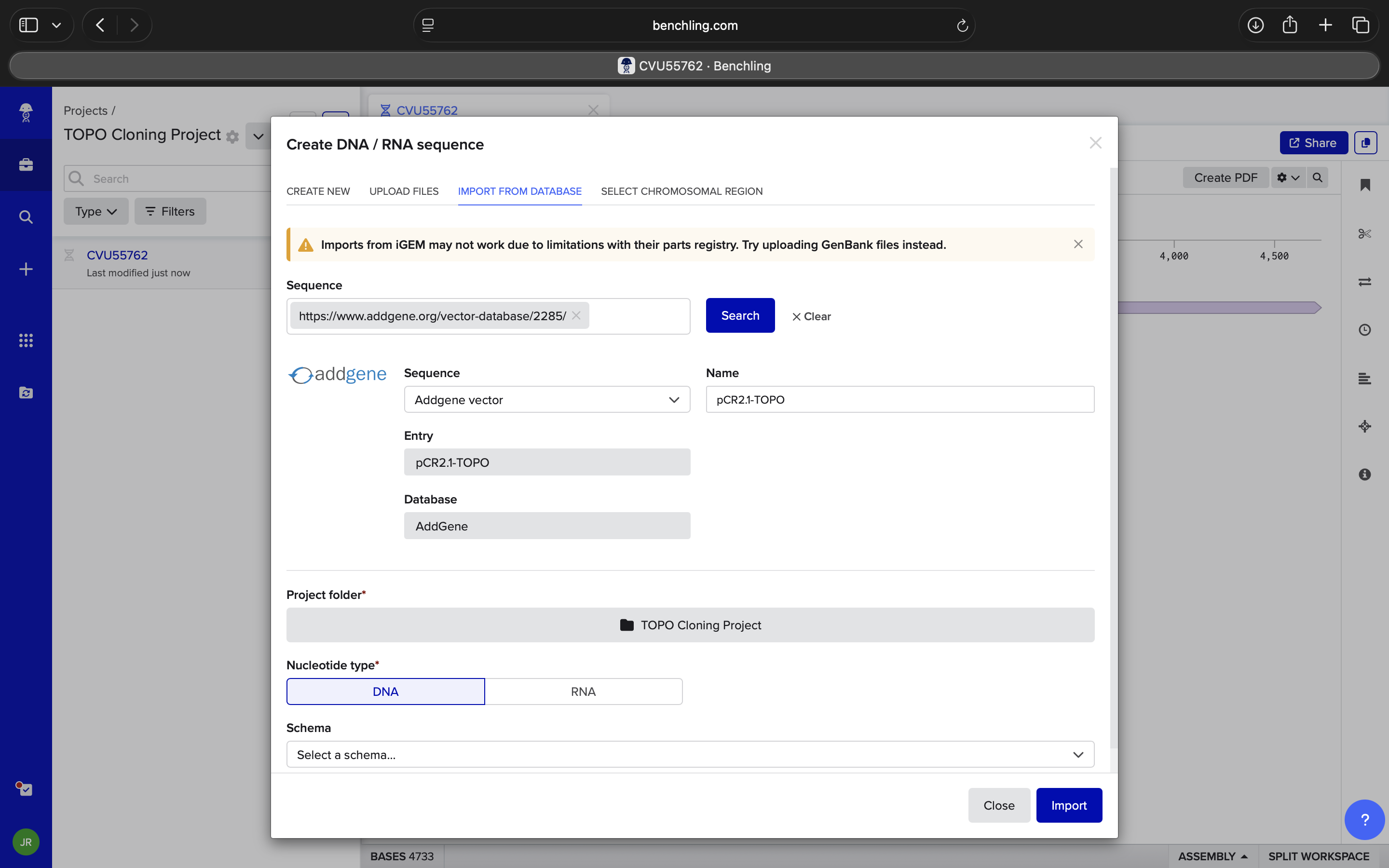
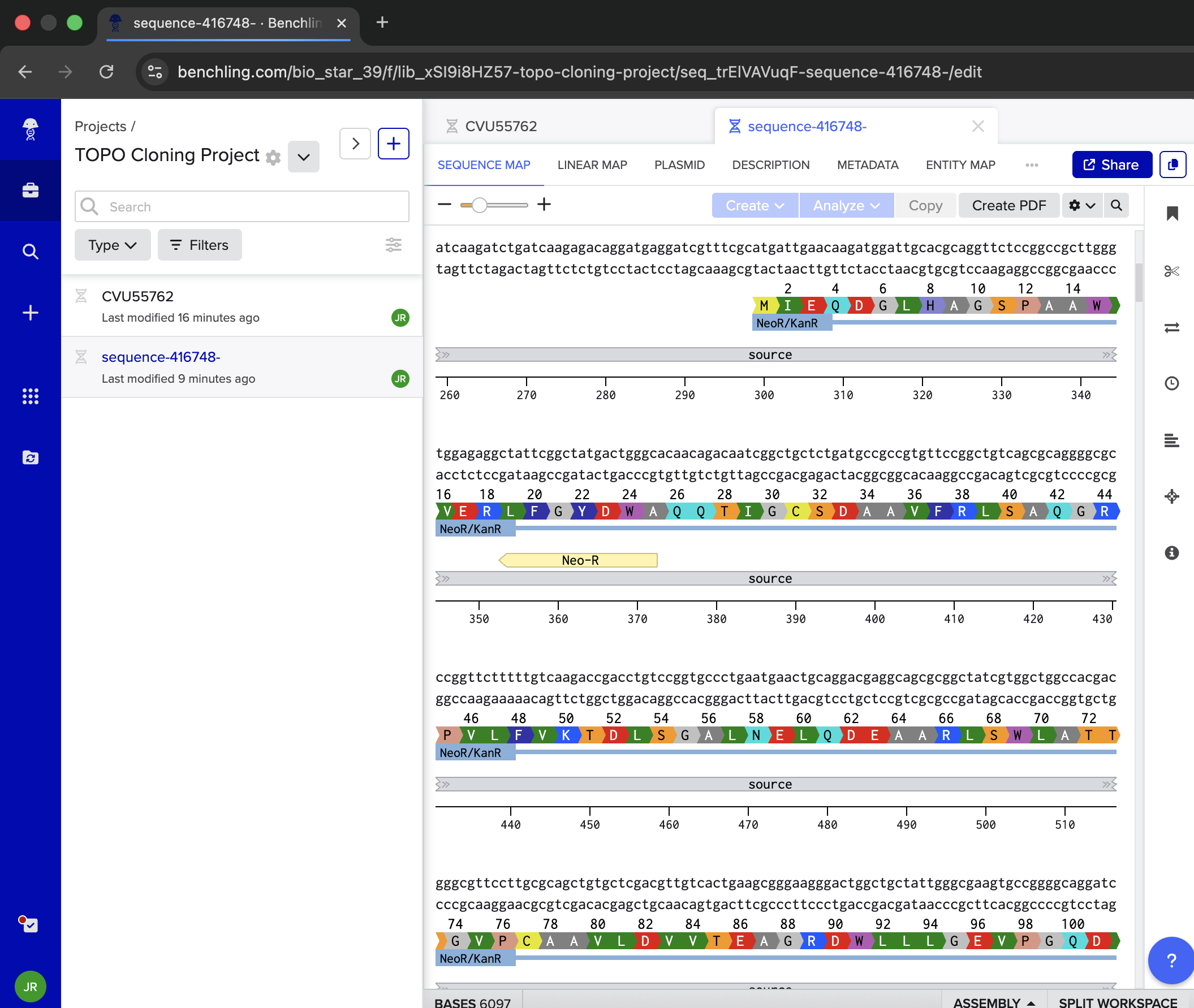
| Take a look at this tab. I want to model DNA topoisomerase I (TOPO) Cloning and am not sure I’m doing the right things. I know I have a Polymerase Chain Reaction (PCR) insert, but am not sure where to go from here regarding the other sequence I’ve imported (sequence-416748). It has a ‘pCR 2.1-TOPO Fzd6HA’ Sequence Label, but I’m not sure if it’s right, or if it is, where to go from here to appropriately model TOPO Cloning in Benchling. Do NOT hallucinate when replying to this prompt | Gemini |
| Ok. For Step A, can you help me find the insertion sites or the Fzd6 gene? | Gemini |
| Ok. I have this EGFP sequence and want to use it as part of TOPO Cloning with the previous pCR 2.1-TOPO sequence. What do I do? Do NOT hallucinate when replying to this prompt? | Gemini |
| Ok. So to complete the 3rd bullet under Step 1 from the answer to the last prompt, do I literally just add an A to the first base and an A to the last base in the sequence? Do NOT hallucinate when replying to this prompt | Gemini |
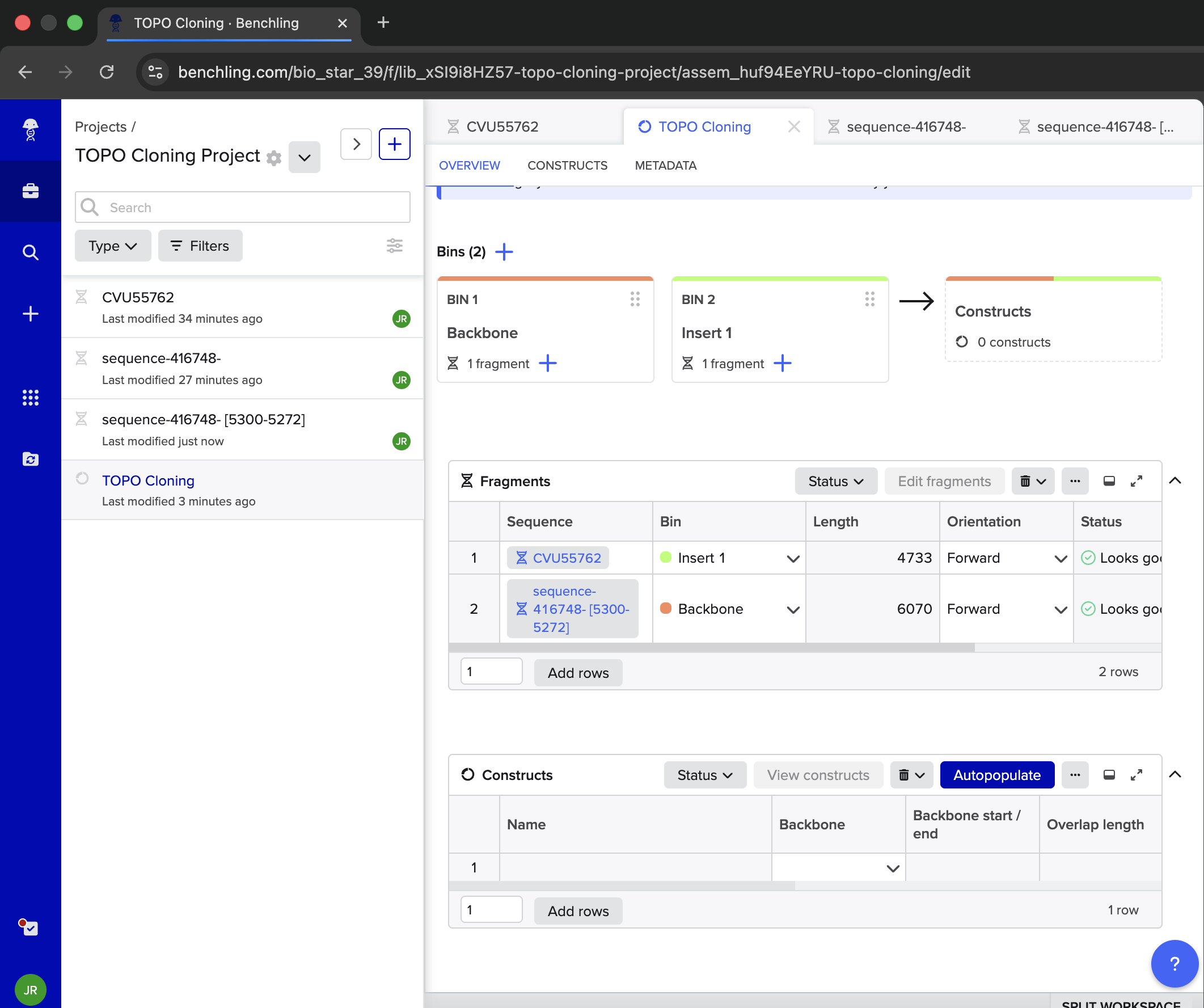
| When I go to Asembly Wizard, I can only do ‘Golden Gate’, ‘Gibson’ or ‘Homology’. How should I proceed? | Gemini |
| When we say ‘select the part of the vector excluding the Fzd6HA gene’, that can mean just copy the entire sequence EXCEPT for the Fzd6HA gene in a new sequence and then including that or no? | Gemini |
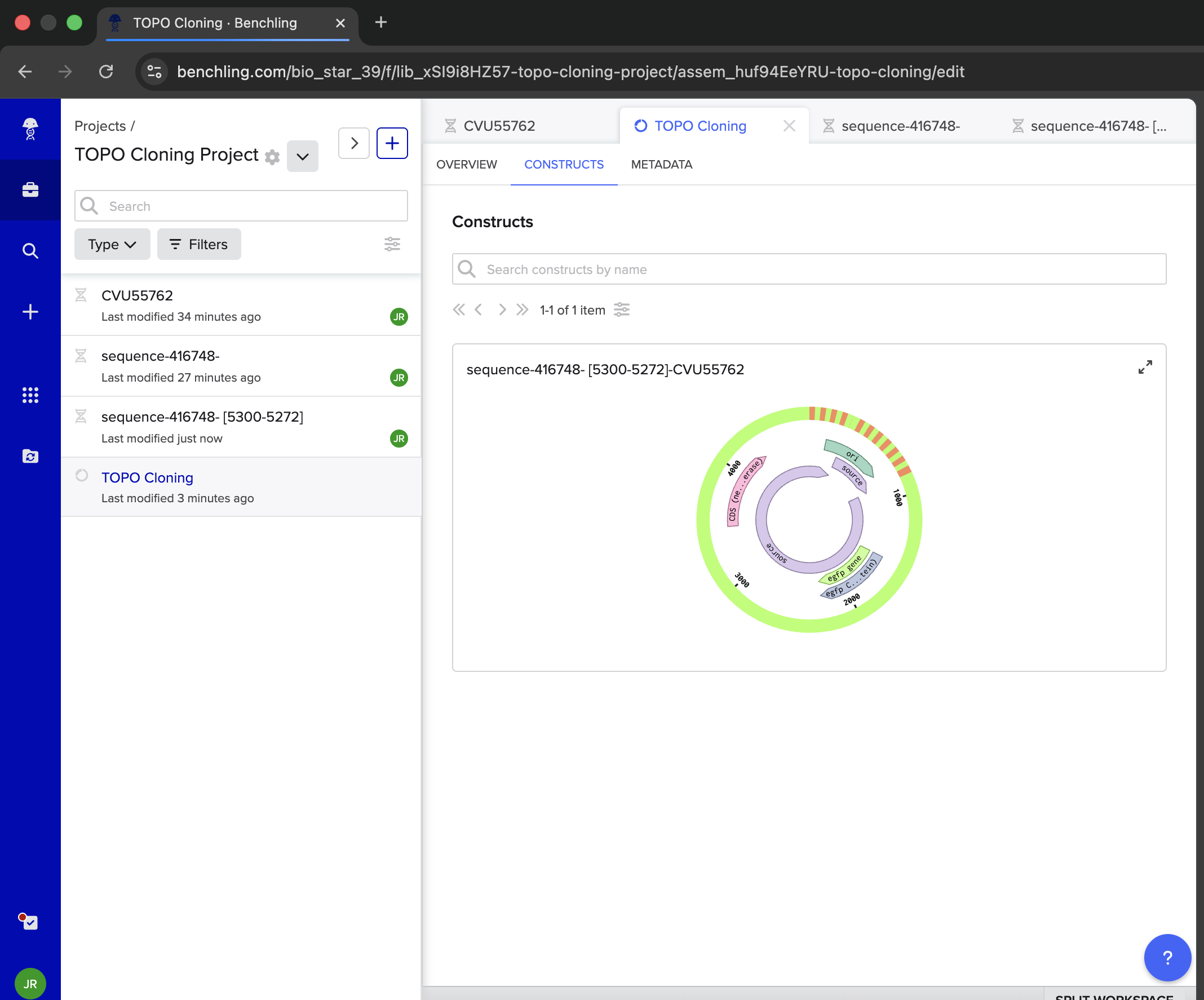
| Ok. Take a look at the construct I made. Does this depict TOPO Cloning in a reasonable way based on the insert and the vector? Do NOT hallucinate when replying to this prompt | Gemini |
| Getting an error that one of my constructs is invalid. Show me how to fix this | Gemini |
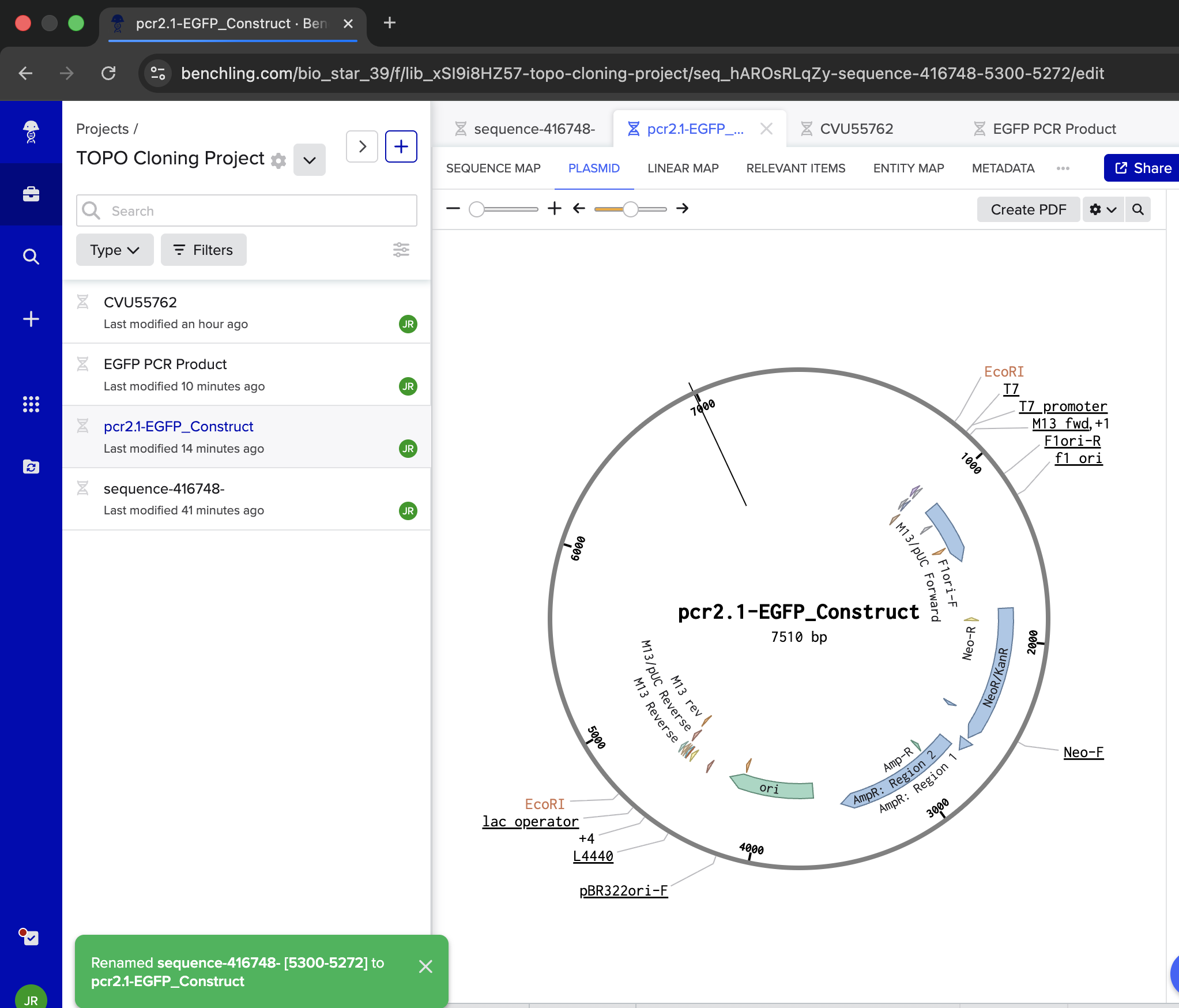
| Confused. Want to do this manually WITHOUT the Assembly Wizard. Not seeing where to add the EGFP PCR Product. Also not seeing how I can see the plasmid | Gemini |
Asimov Kernel
See Repository below
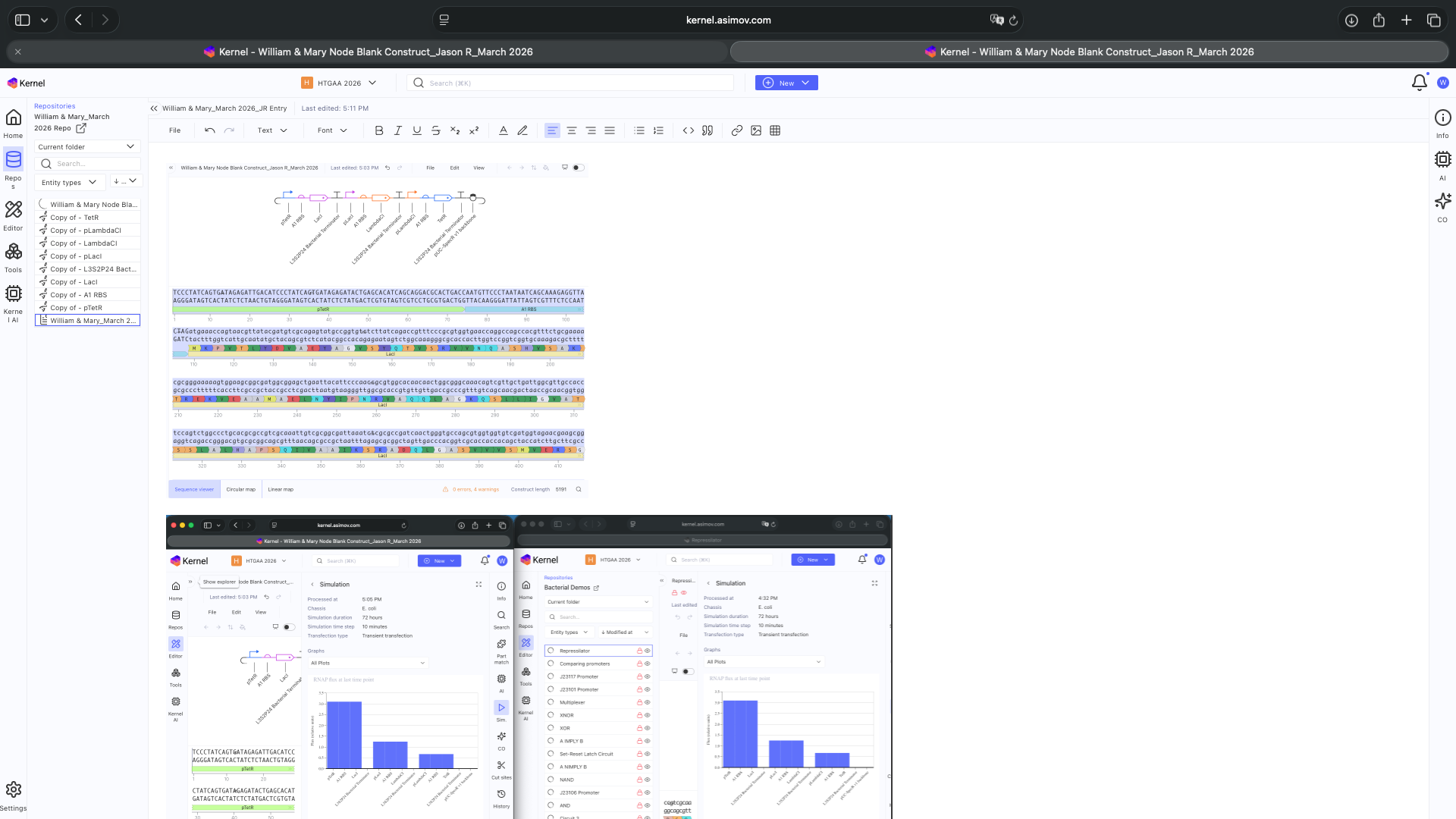
See blank Notebook entry below
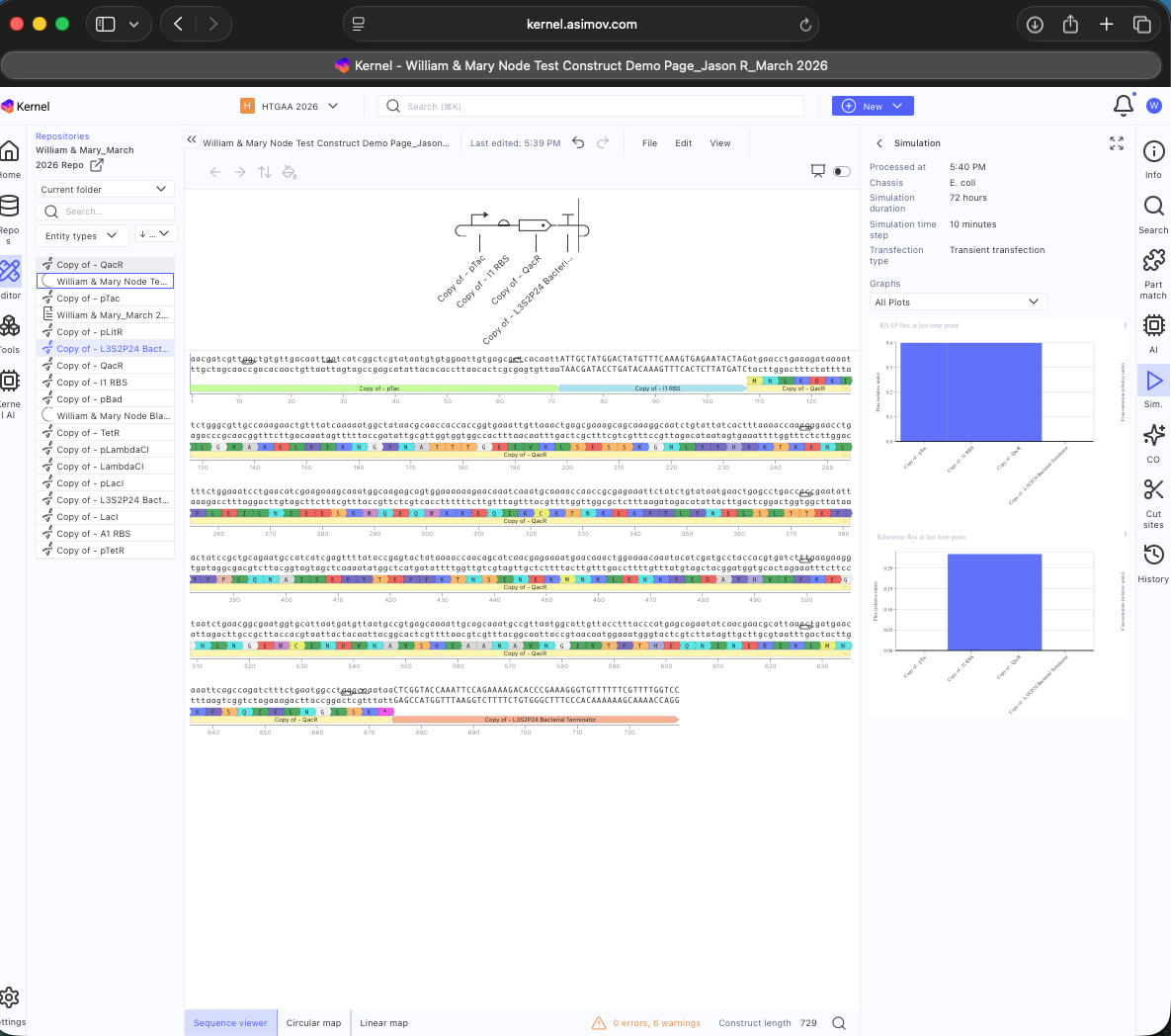
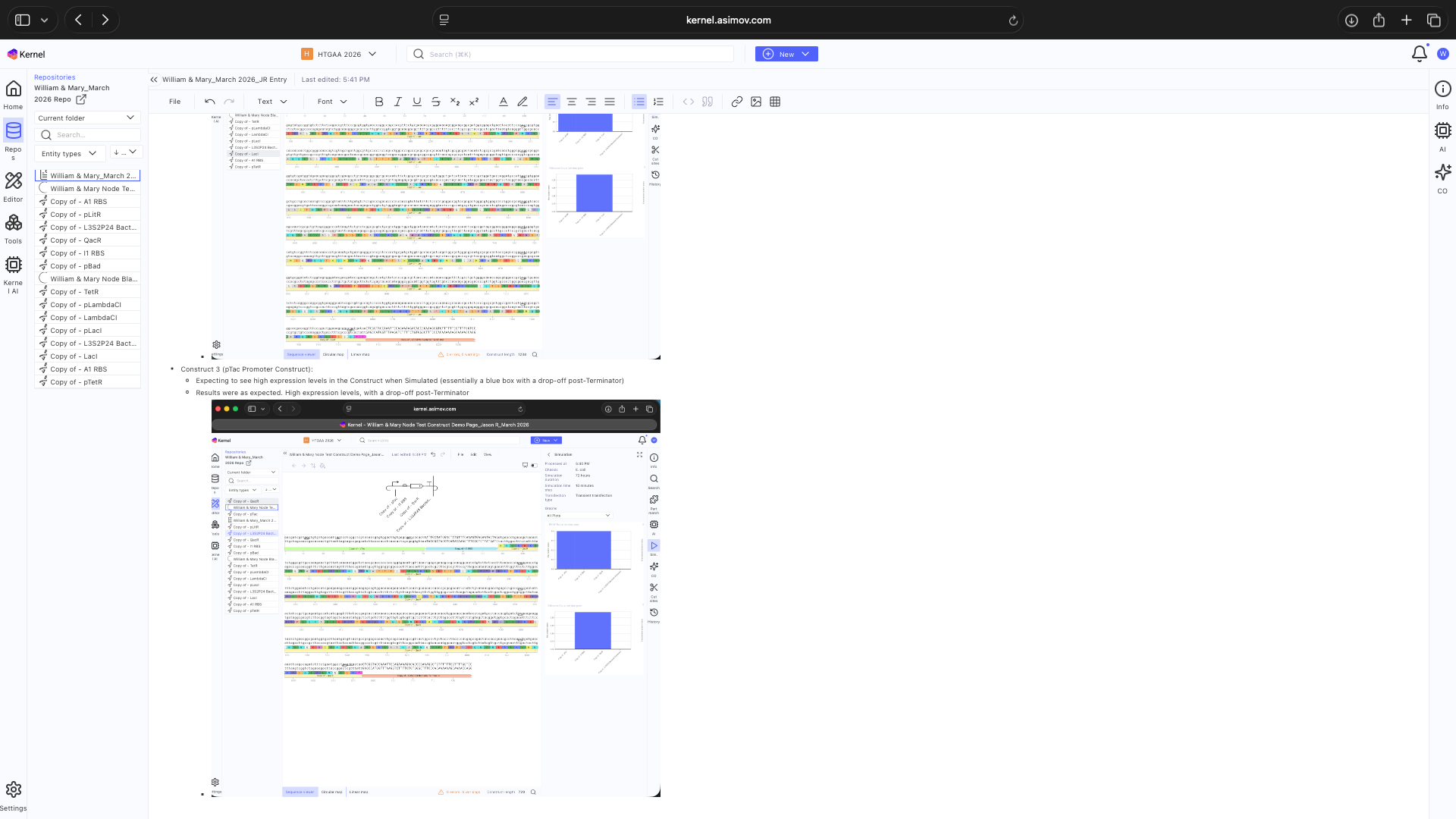
See results below
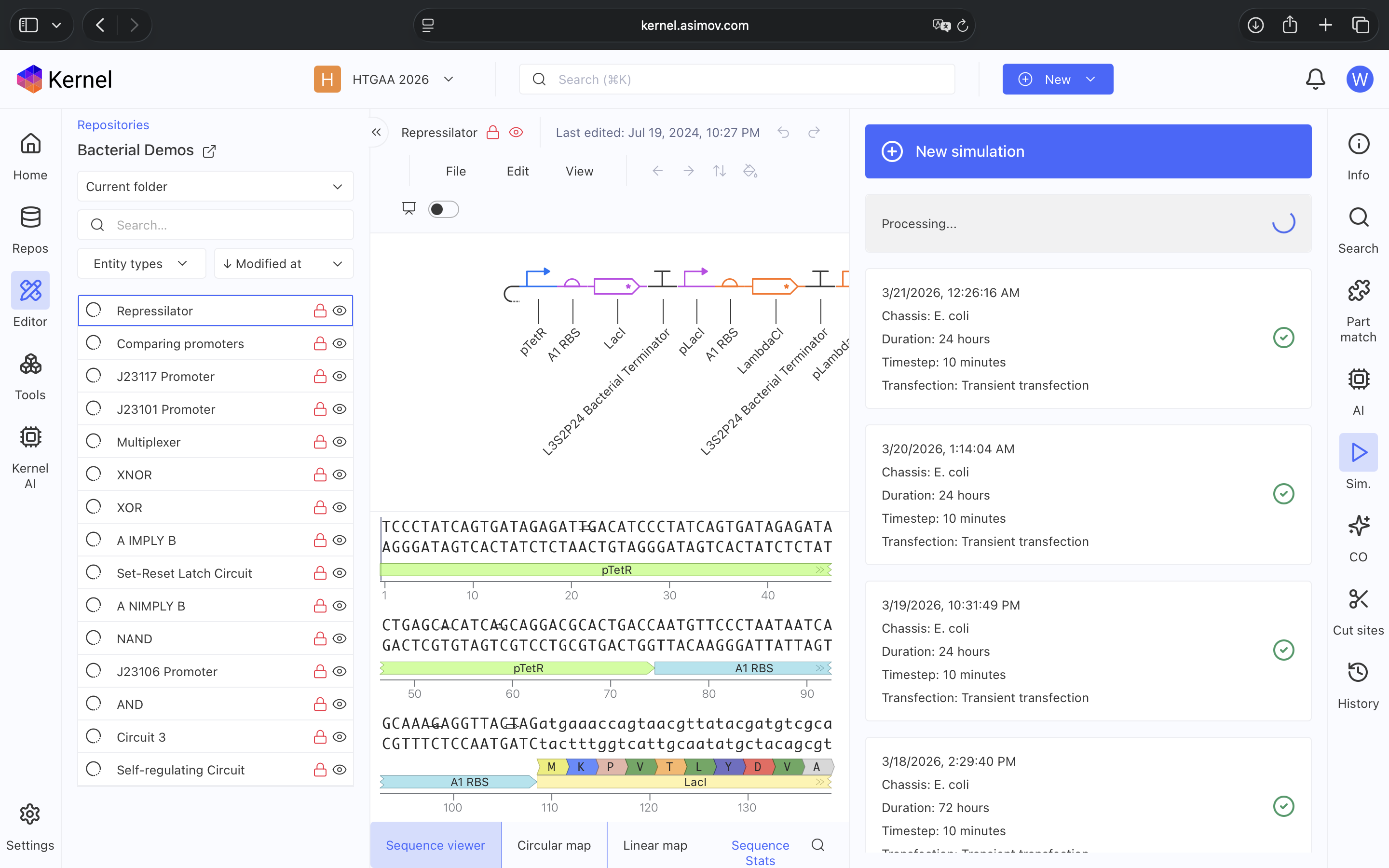
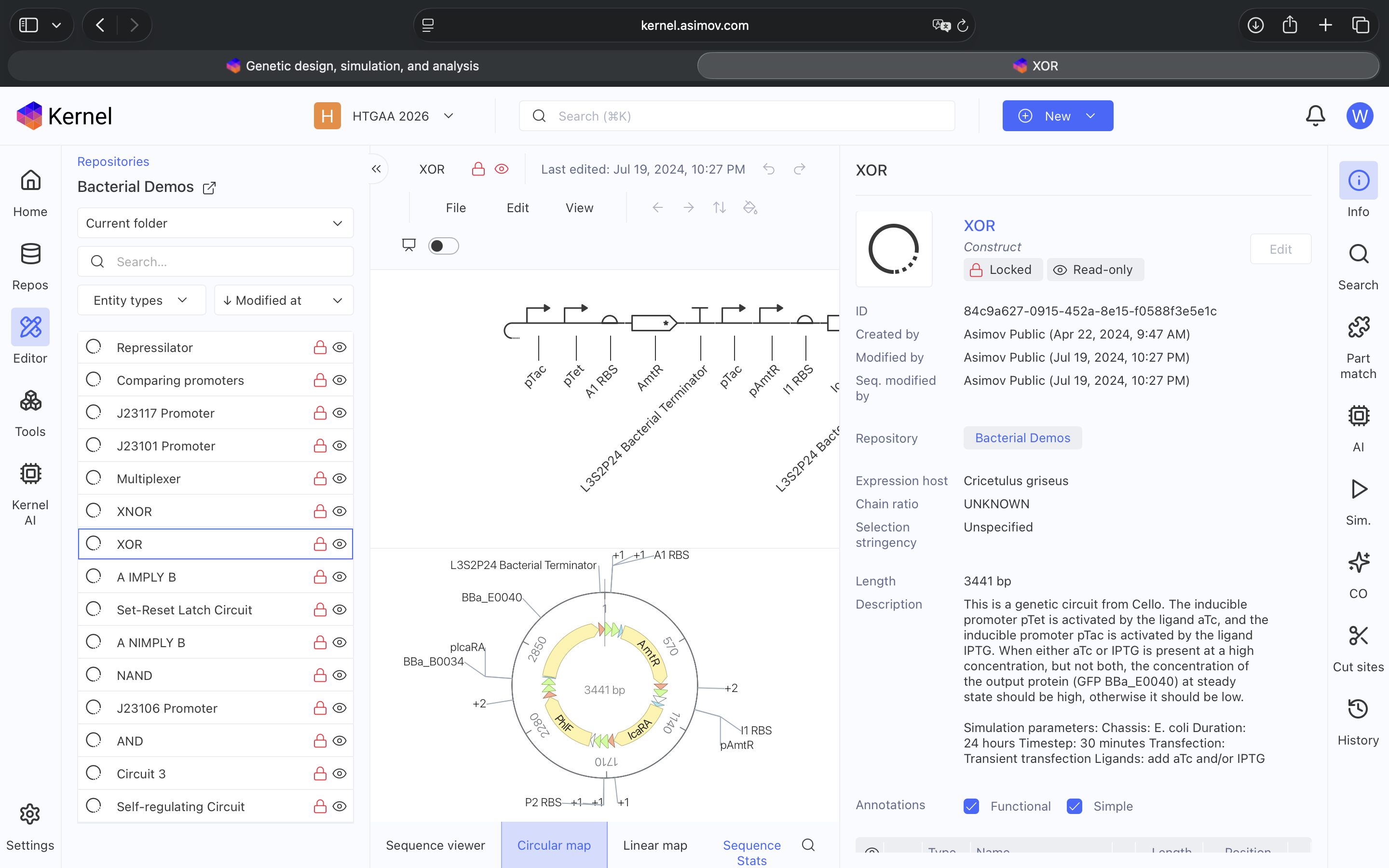
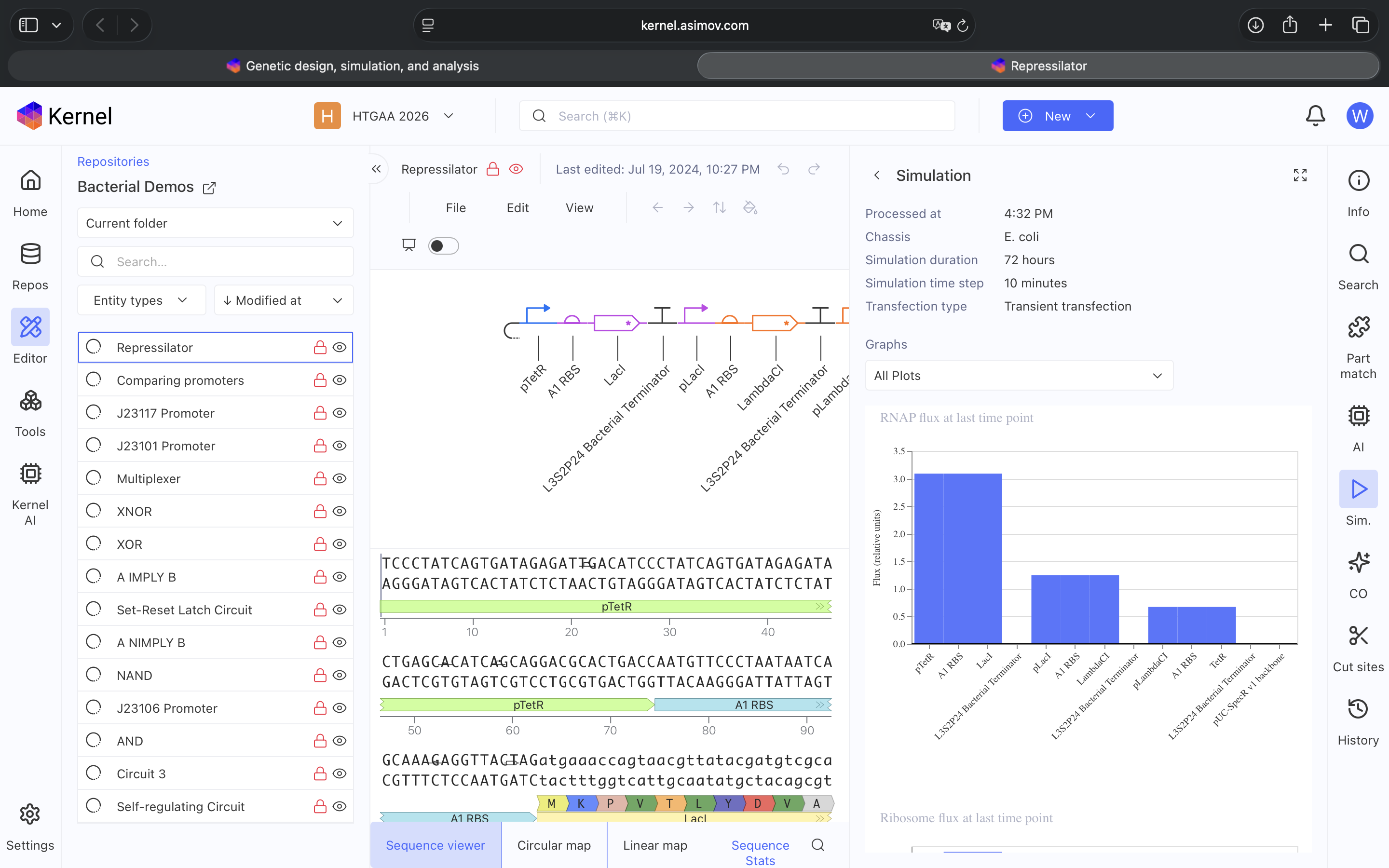
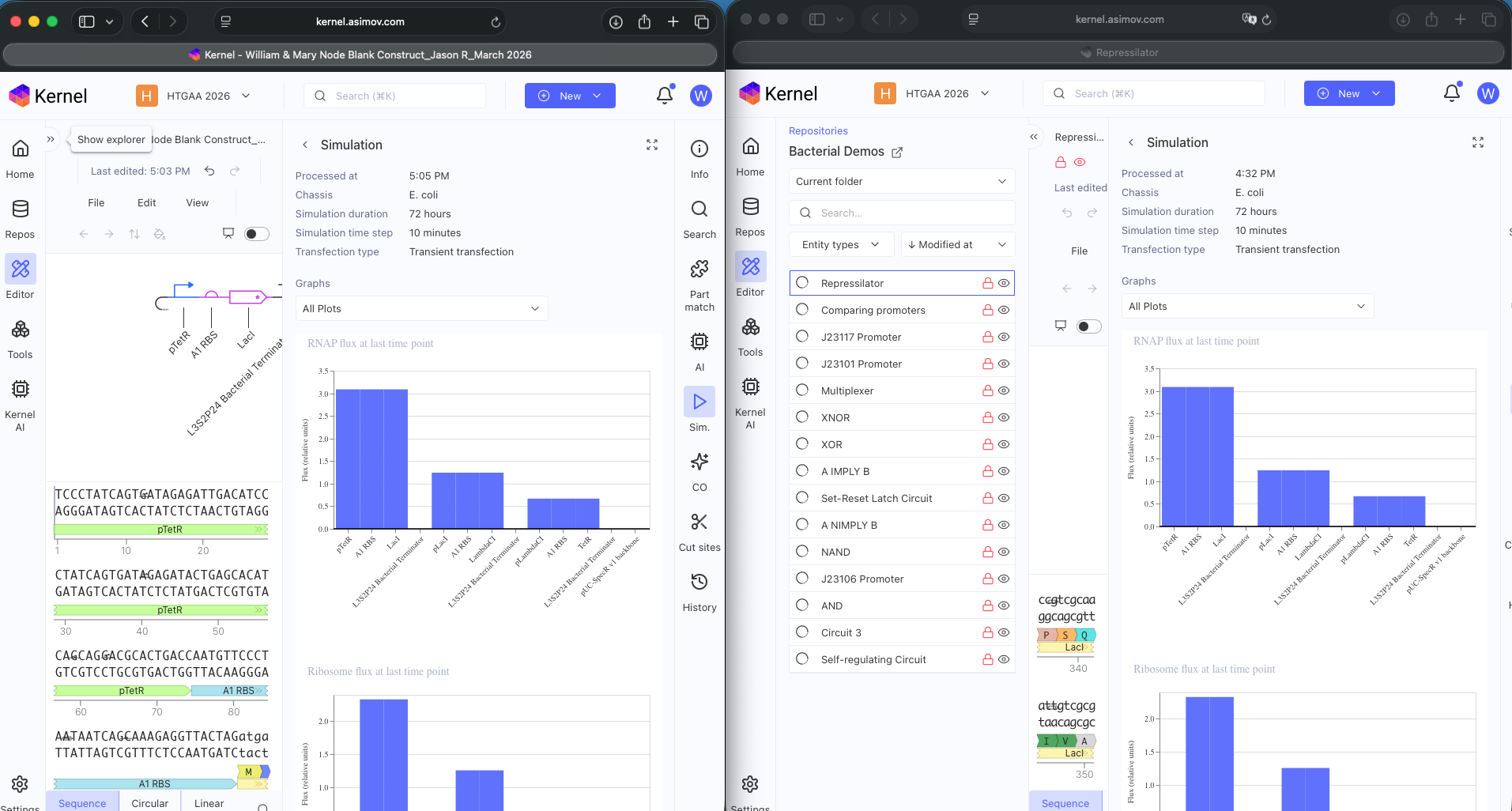
Explored Bacterial Repos Devices and ran Simulator on various examples (see above)
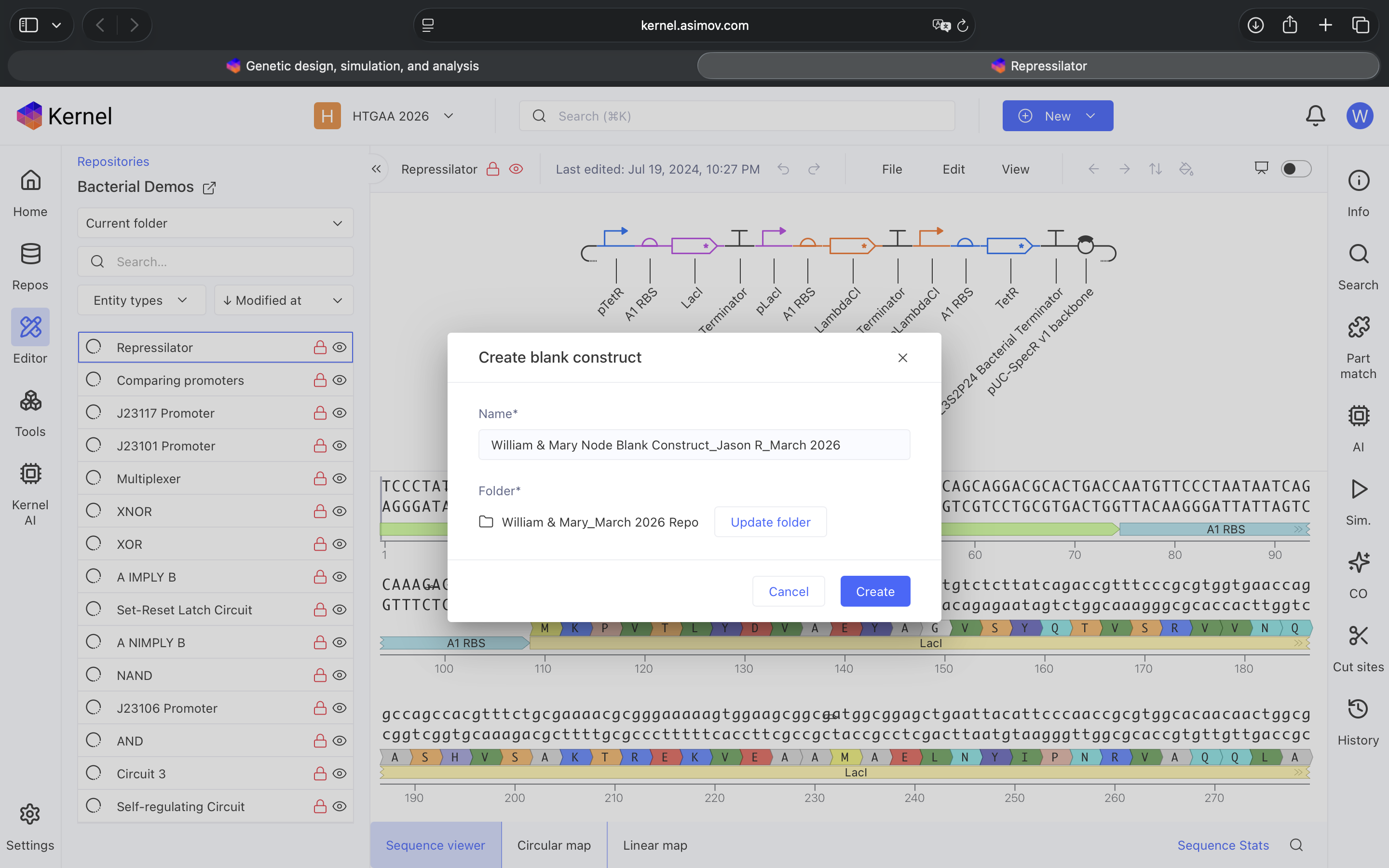
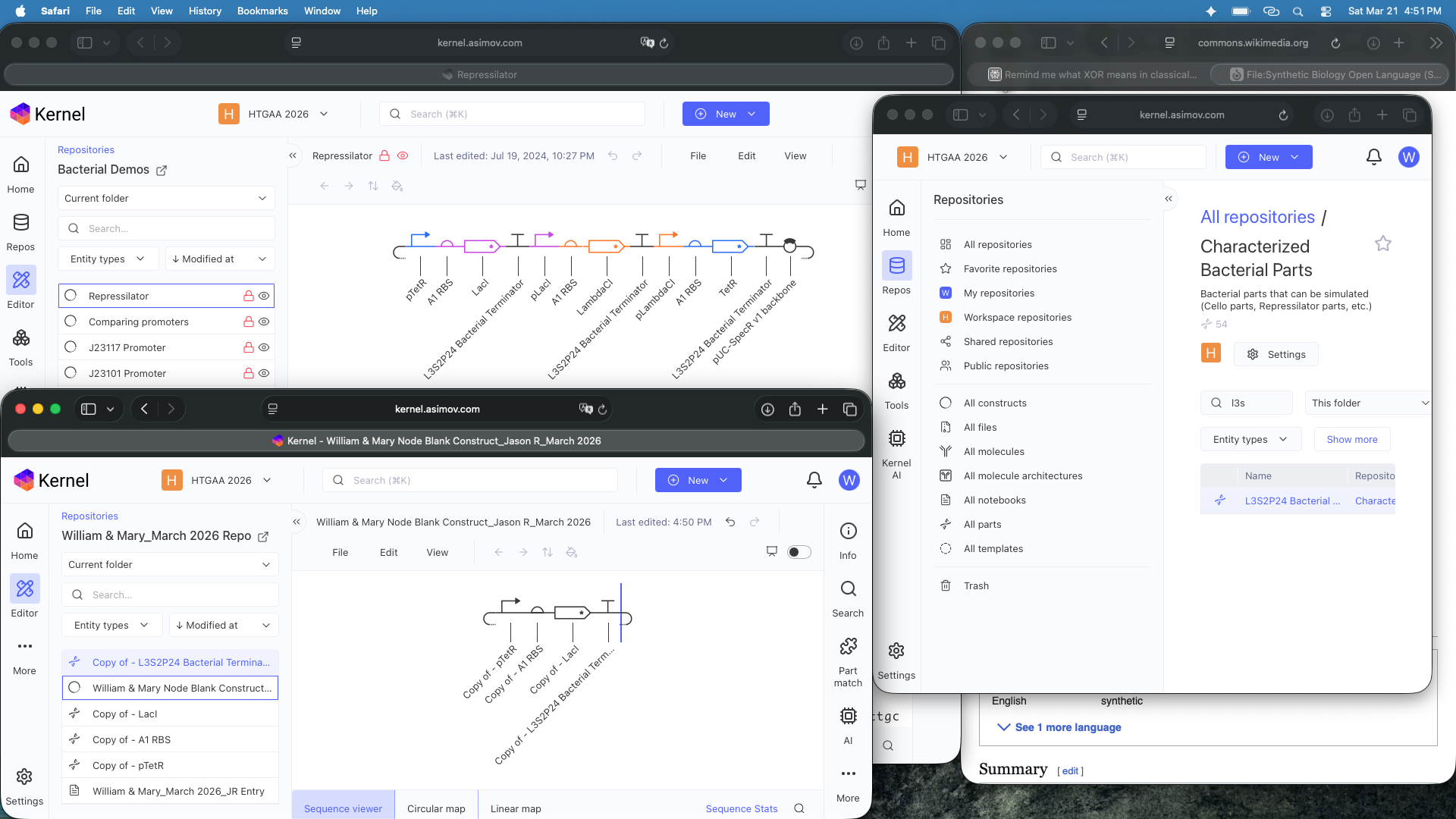
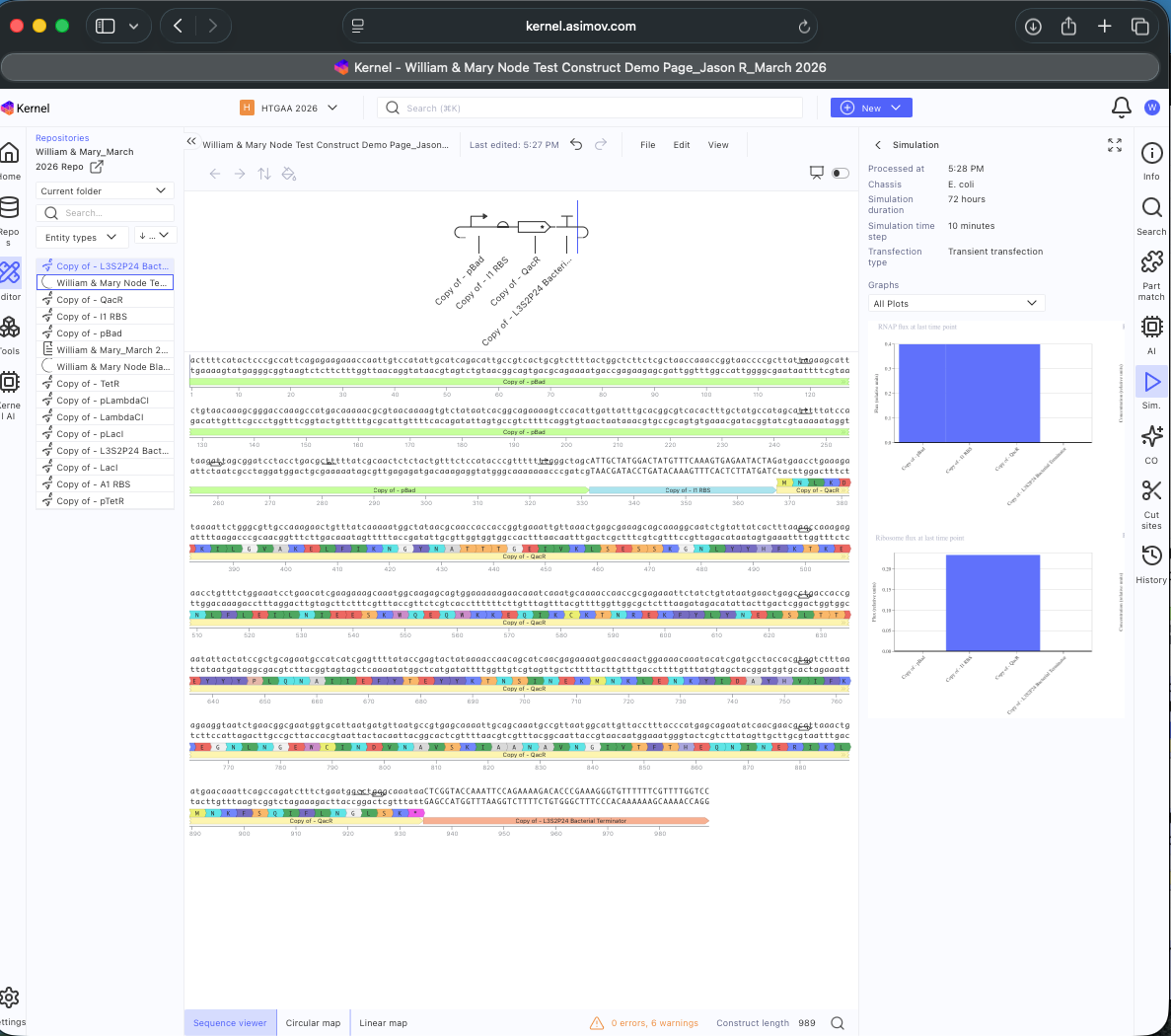
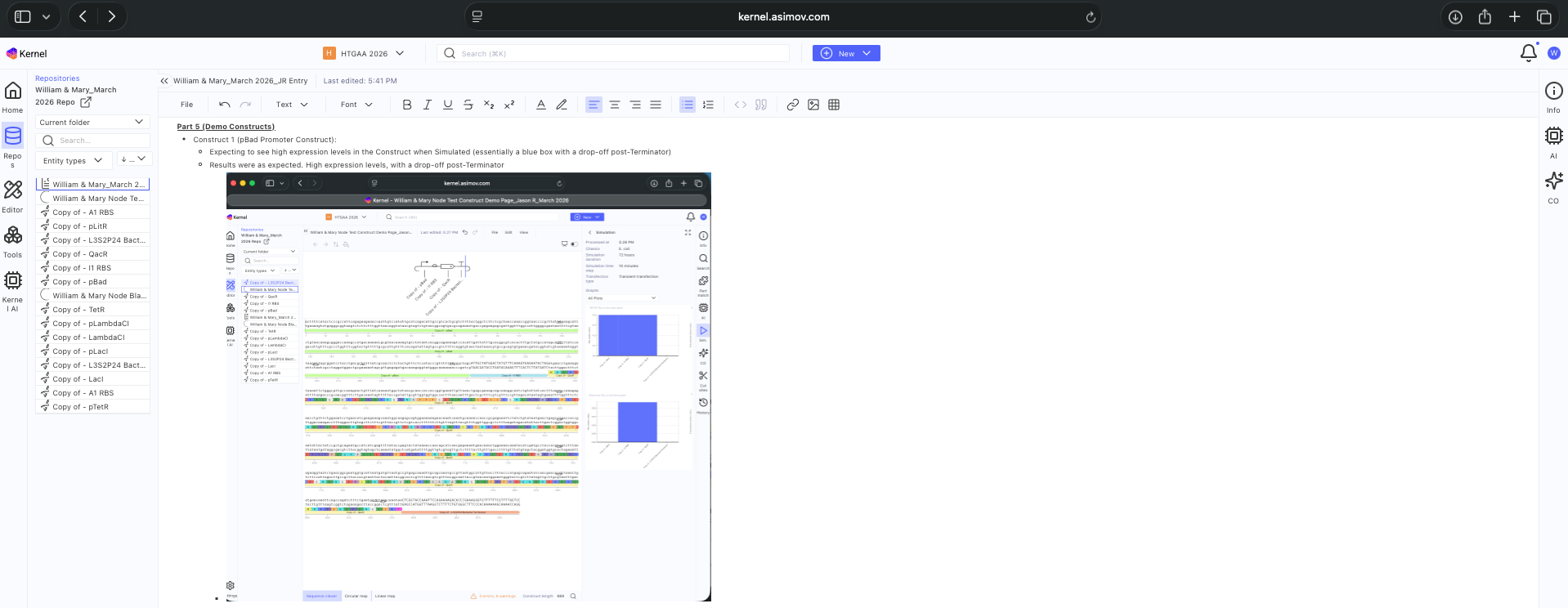
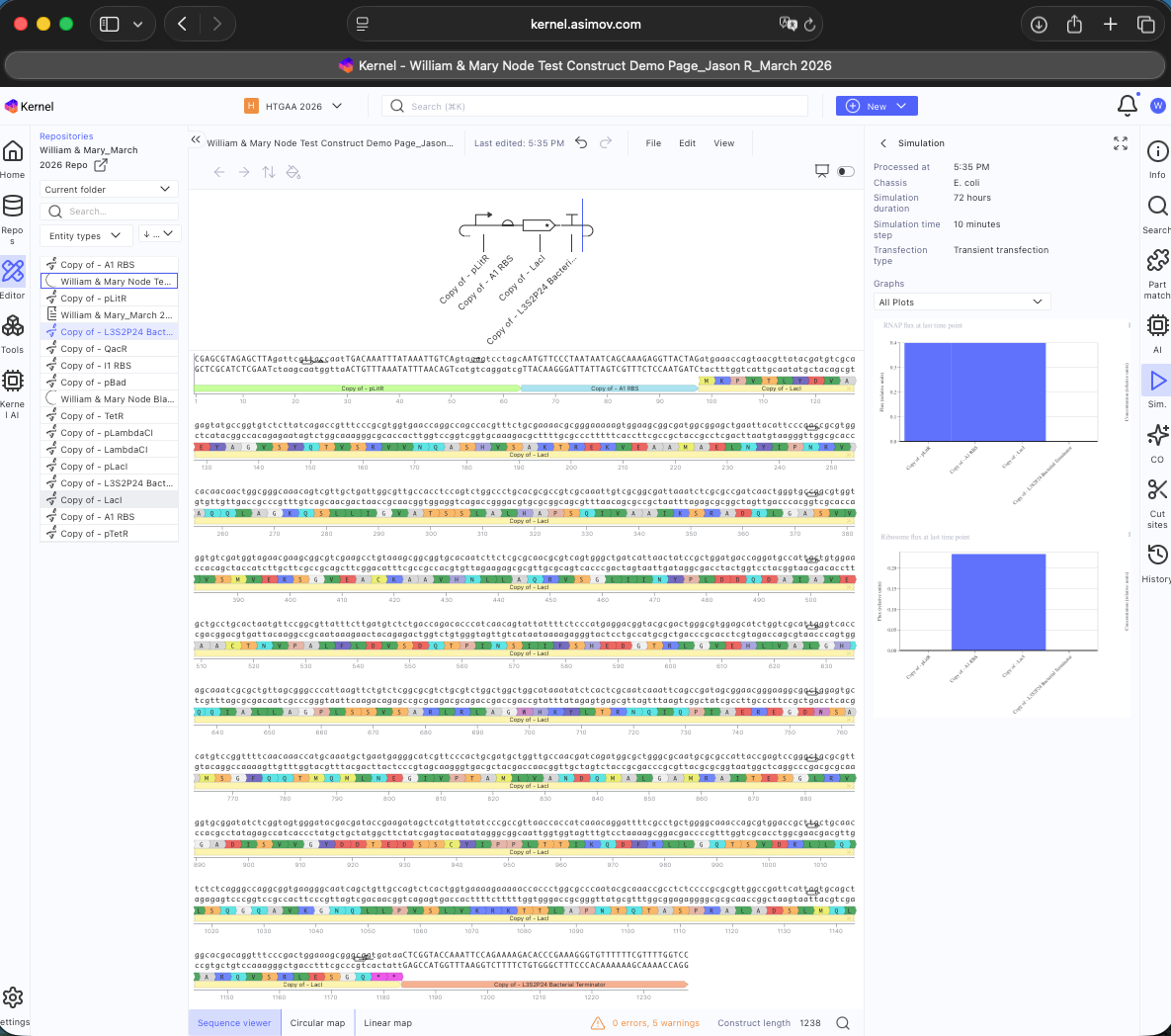
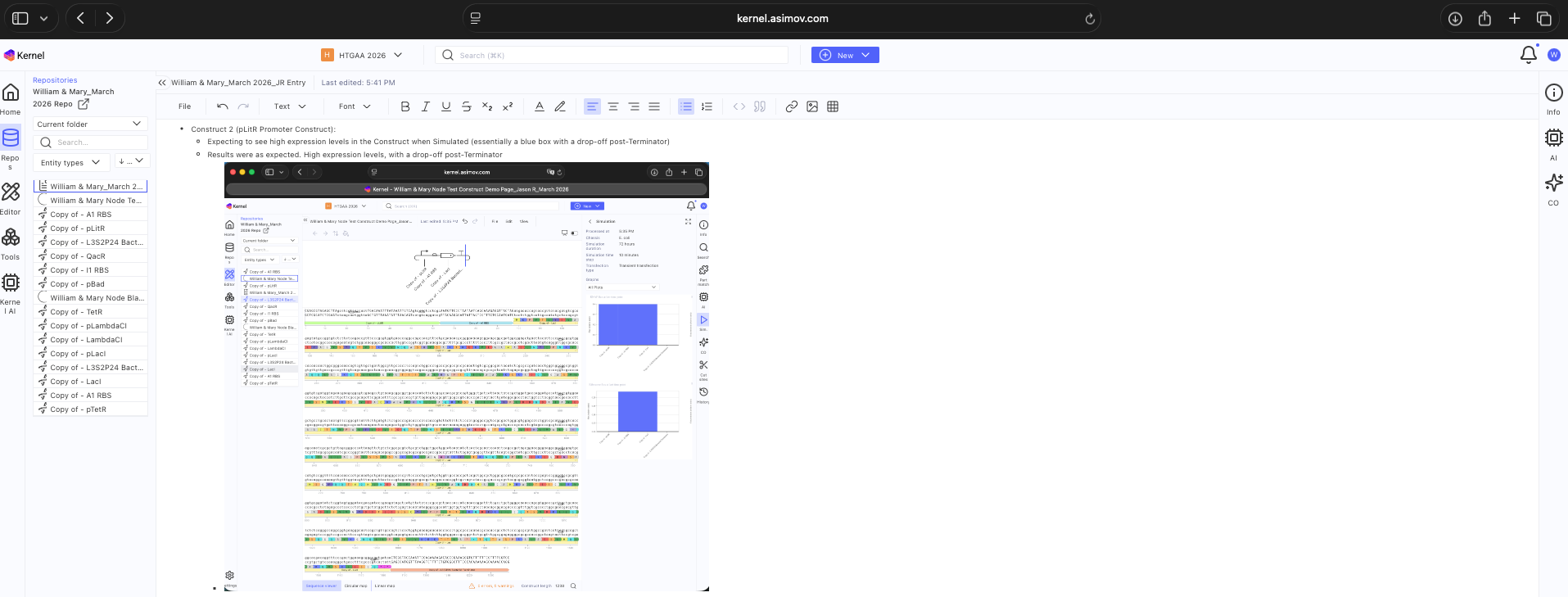
See Construct creation results below
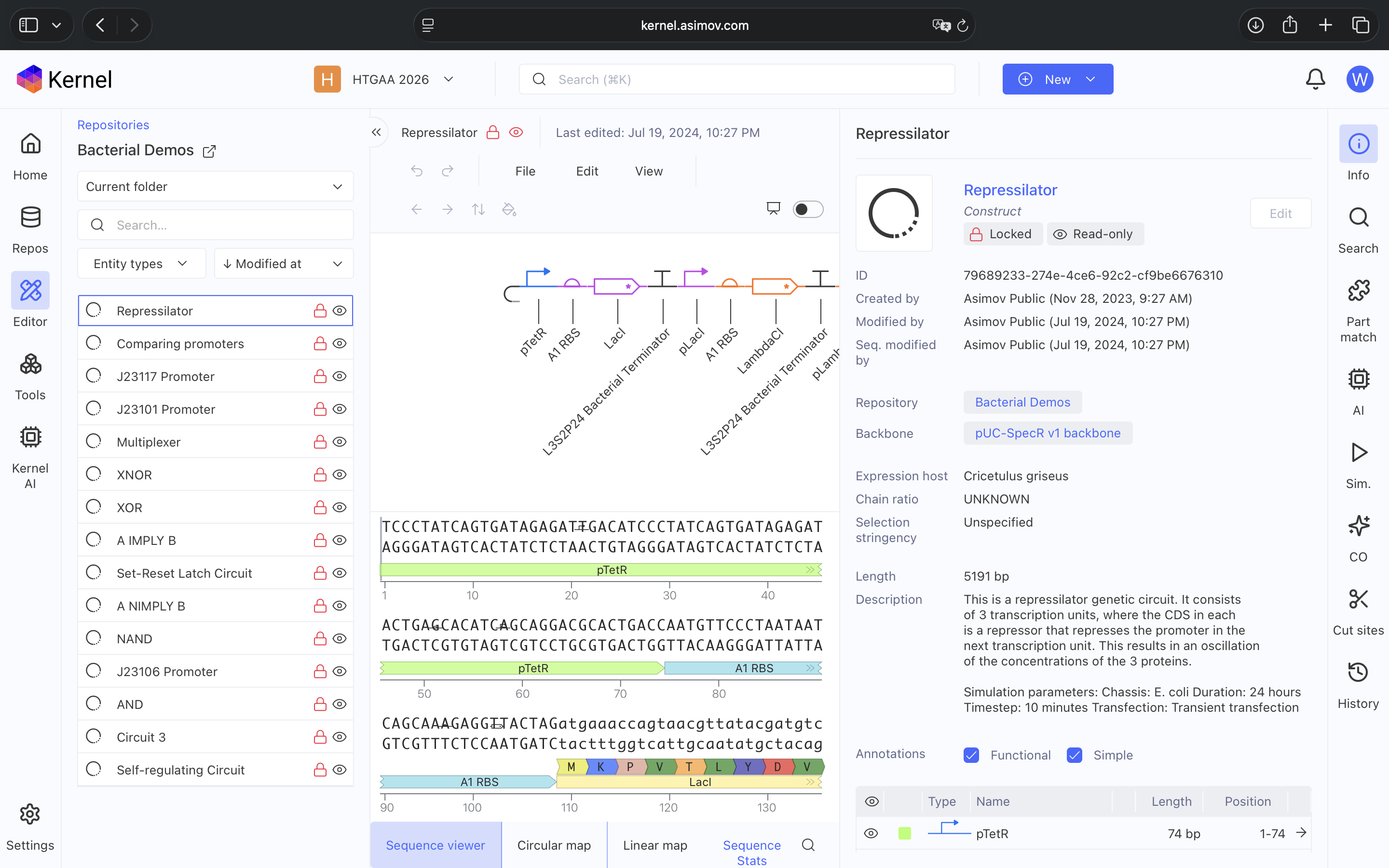
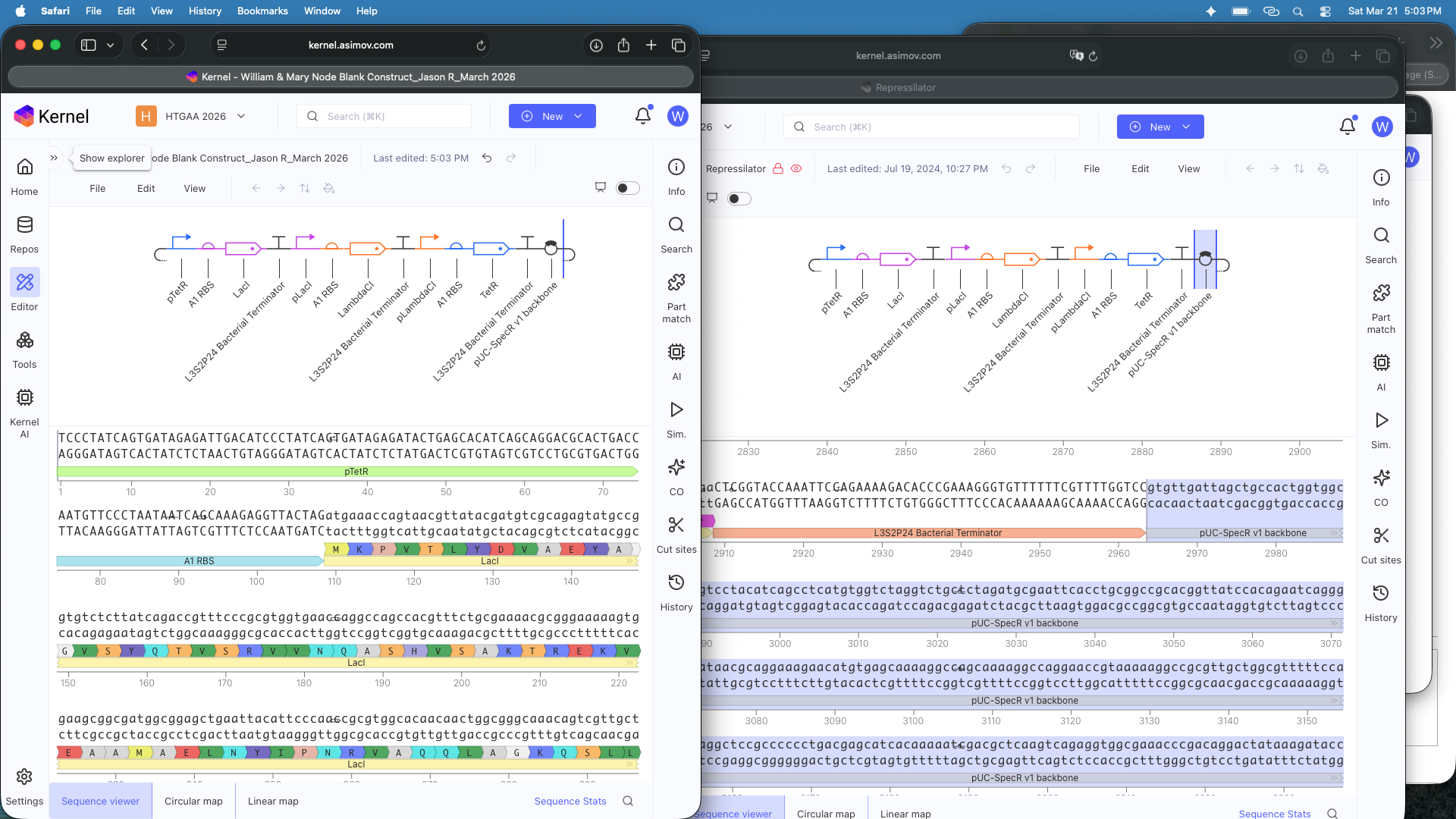
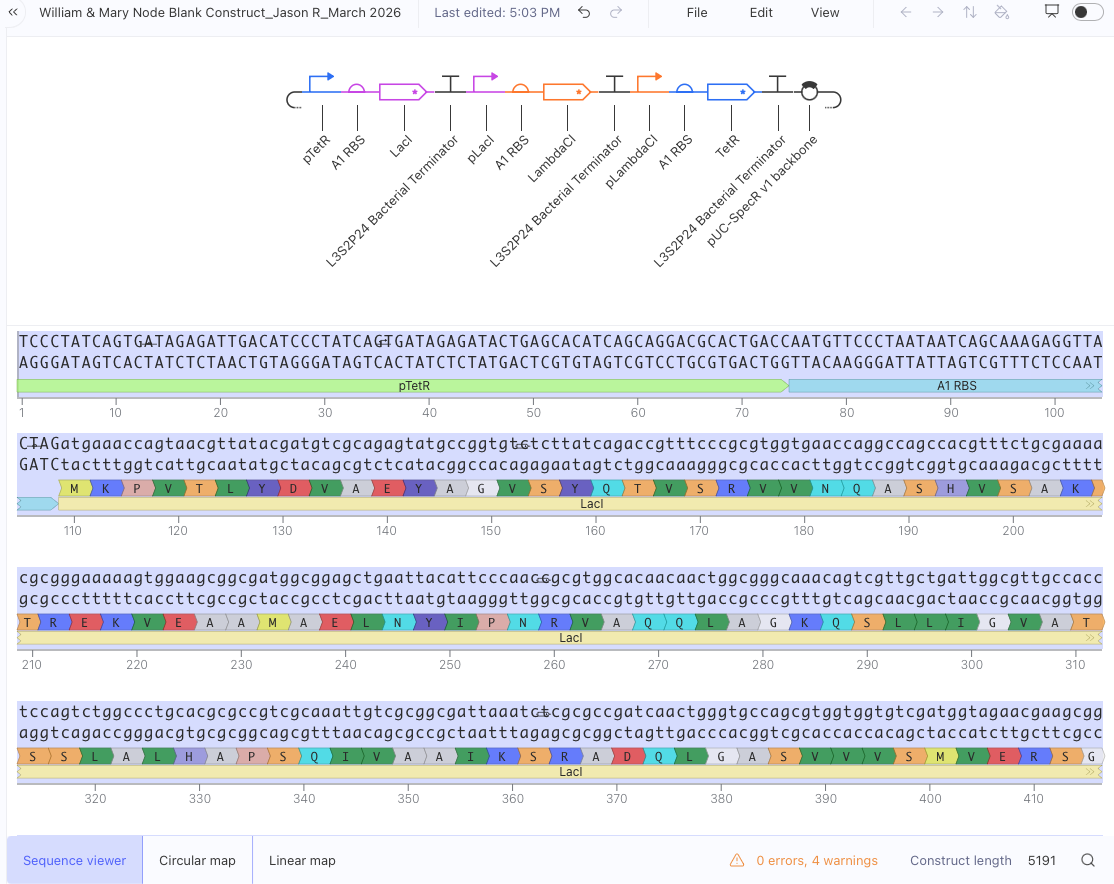
- Question 1-3 Results: Recreated Repressilator in empty Construct using Characterized Bacterial Parts repository parts, searched and selected parts using the Search function, and dragged and dropped parts into Construct (see photos below)
- Question 4 Results: The Repressilator wasn’t running as expected so I re-made it and ensured I directlty copied and pasted everything. Then I re-ran the Simulation and the Repressilator operated as expected (see photos below)
- Question 5 Results: Documented results in Notebook (see below)
See 3 Construct creation results below
Supporting prompts for this section listed below:
| Supporting Prompt | Model |
|---|---|
| Remind me what XOR means in classical computation (including, but not limited to its instantiation on digital computers). Do NOT hallucinate when addressing this prompt | Perplexity |
| Share any links that break down the basic symbology/legend of Synthetic Biology Open Language (SBOL). Do NOT hallucinate when replying to this prompt | Perplexity |
| Little bit confused here. Copied all the Repressilator parts from the ‘Bacterial Demos’ repo, but am getting different results when running the Simulation. What could be going on? | Kernel AI |