Week 12 HW: Bioproduction & Cloud Labs Part 2
Part A: The 1,536 Pixel Artwork Canvas | Collective Artwork


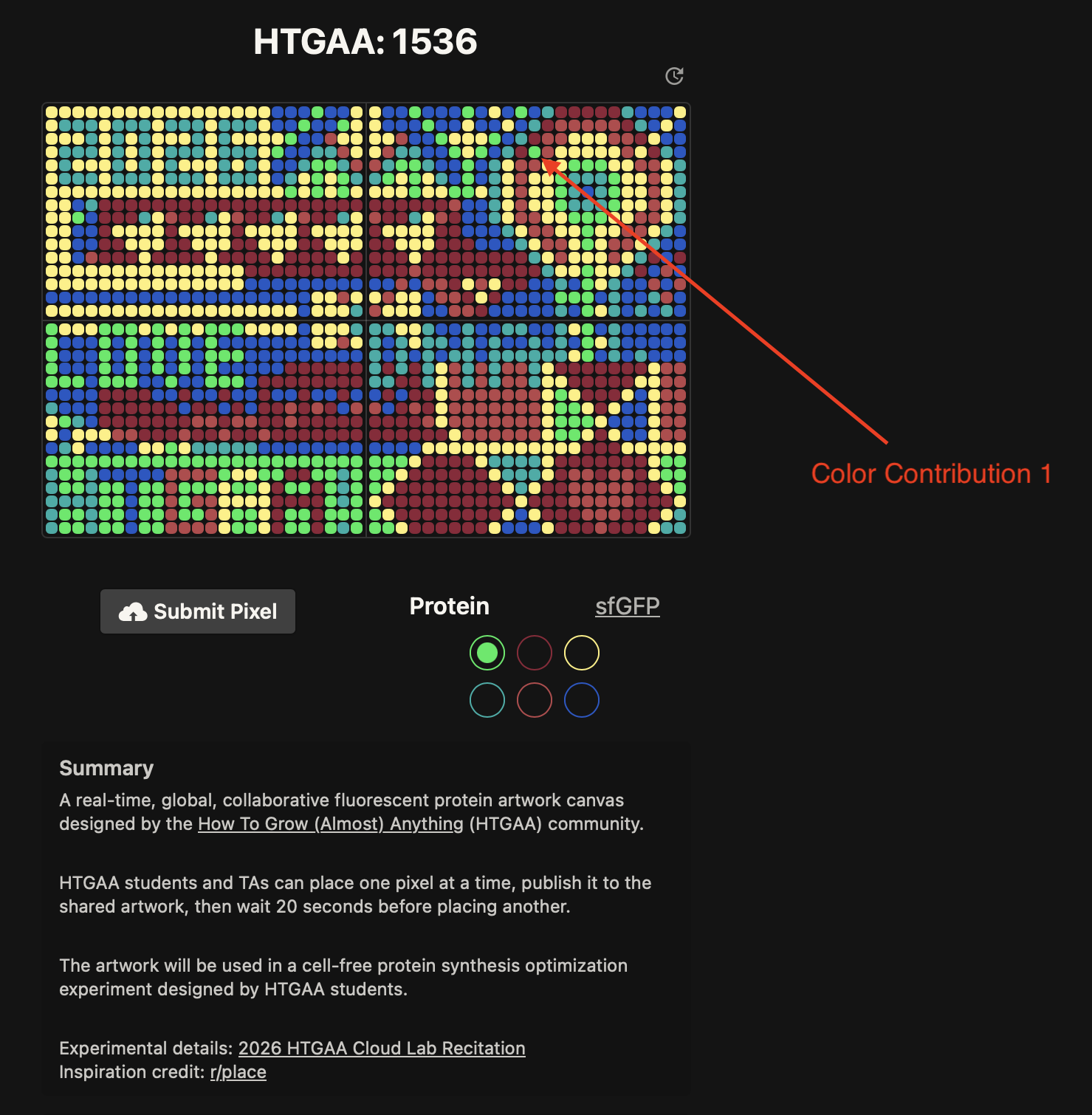
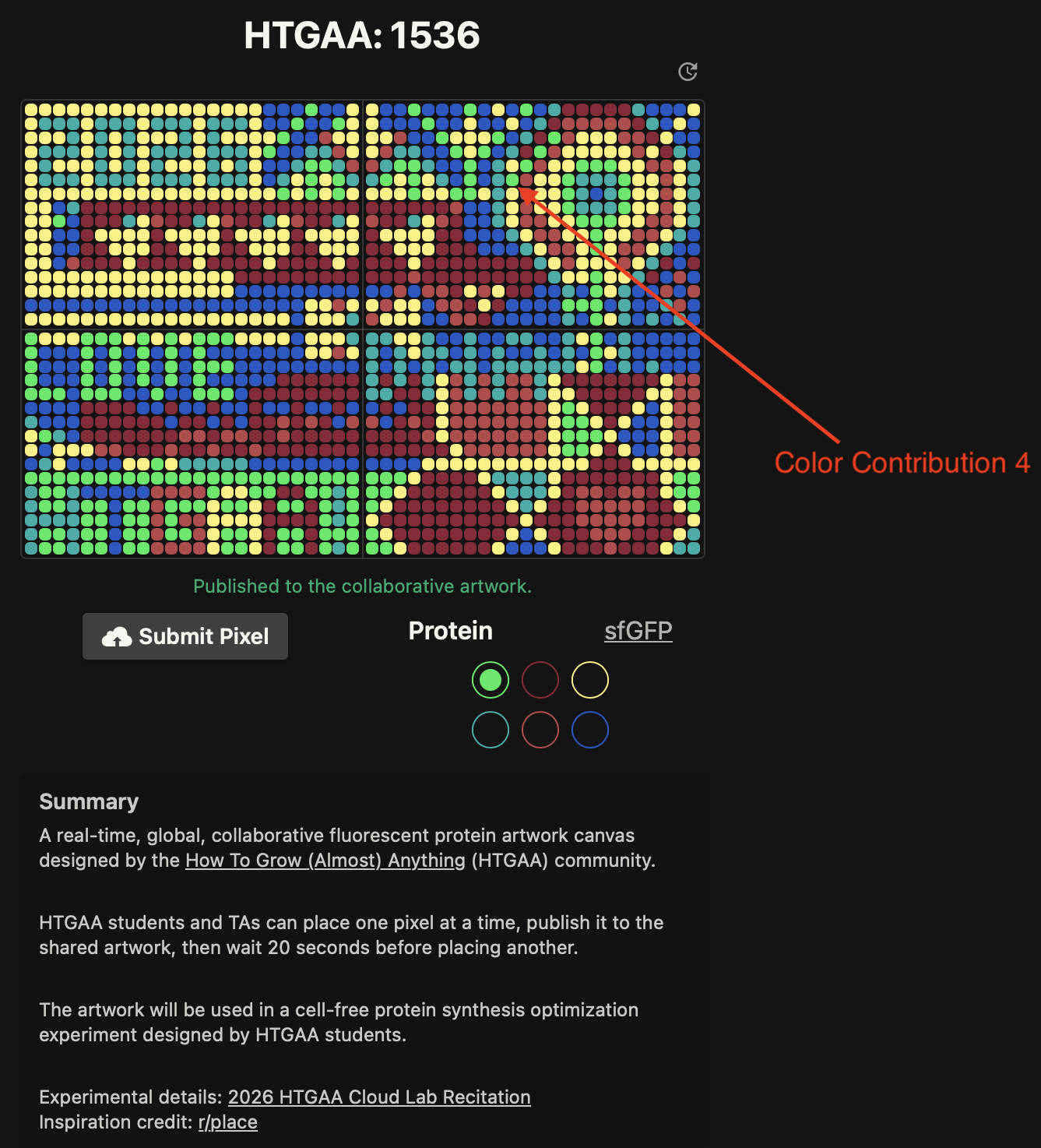
Contribute at least one pixel to this global artowrk experiment before the editing ends on Sunday 4/19 at 11:59 PM EST.
- Contributed 4 pixels to the global artwork experiment on Saturday 4/18
Make a note on your HTGAA webpages including:
what you contributed to the community bioart project
what you liked about the project
- The project was a nice opportunity to contribute to a larger HTGAA effort. It was nice to see the creativity of the community at play! I also appreciated Ronan’s page for contributing pixels – very intuitive and easy to understand
what about this collaborative art experiment could be made better for next year
- I’d probably say a bit more advance notice might have been useful. Perhaps a bit more clarity on ground rules. These are relatively minor nitpicks in the grand scheme of things
Part B: Cell-Free Protein Synthesis | Cell-Free Reagents
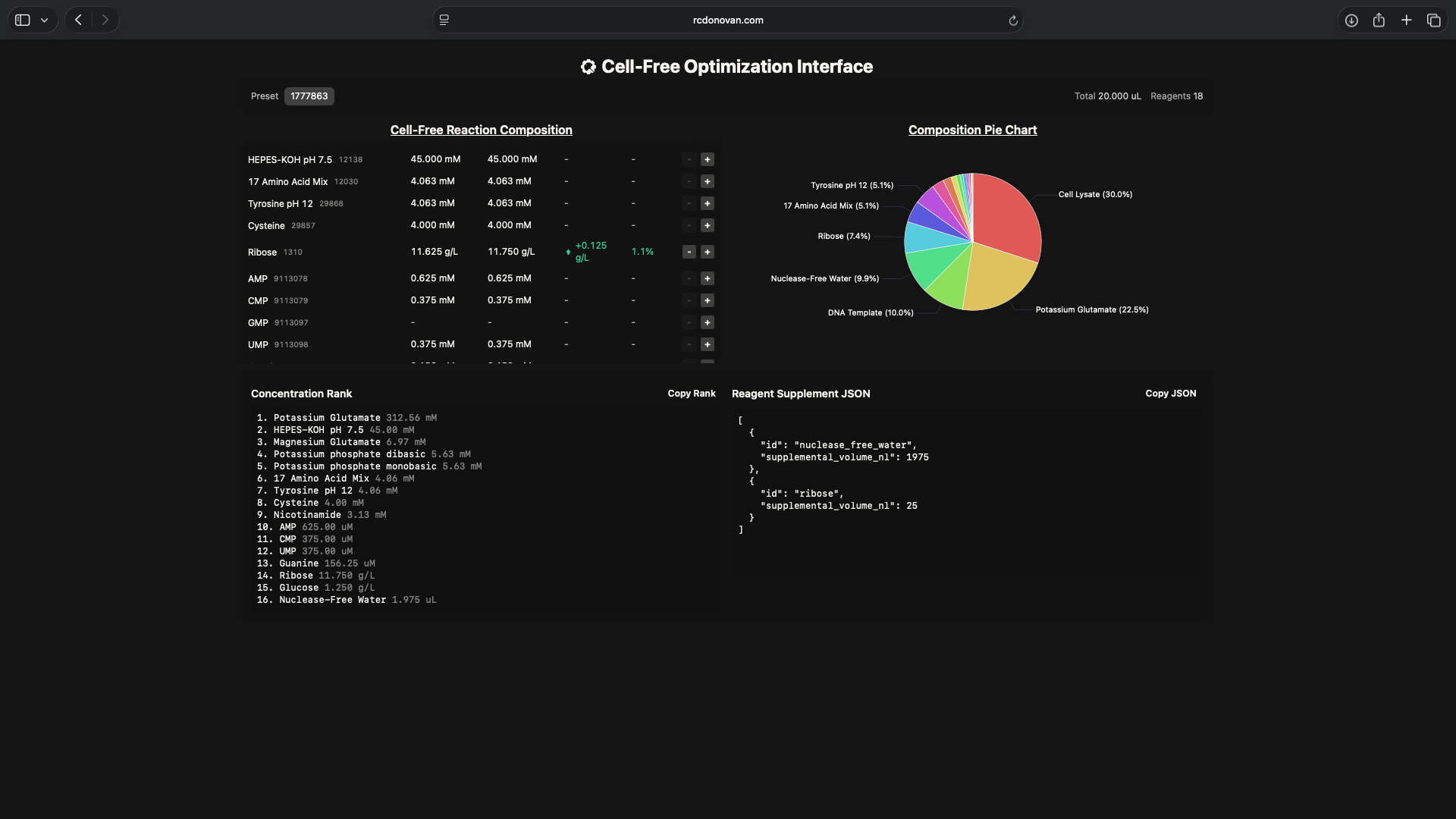
- Referencing the cell-free protein synthesis reaction composition (the middle box outlined in yellow on the image above, also listed below), provide a 1-2 sentence description of what each component’s role is in the cell-free reaction.
- E. coli Lysate
- BL21 (DE3) Star Lysate (includes T7 RNA Polymerase)
- I think BL21 (DE3) Star Lysate’s role is to provide the E. coli bacteria necessary to be synthesized into fluorescent proteins. Basically it seems to serve as a starting ingredient/necessary component, for lack of a better word
- BL21 (DE3) Star Lysate (includes T7 RNA Polymerase)
- E. coli Lysate
- Salts/Buffer
- Potassium Glutamate
- Helps aid in the elongation portion of the RNA –> protein translation process. In this context, I think elongation refers to the reaction timeframe
- HEPES-KOH pH 7.5
- Important cell buffer. It allows for extra buffering capacity if cell culture manipulation occurs for prolonged/longer than normal time period
- Magnesium Glutamate
- Helps stablizie ribosome construction. Ir also helps neutralize mRNA and DNA backbone negative charge
- Potassium phosphate monobasic
- Potassium source and buffer for the reaction, helping stabilize pH levels/keep them nominal during the reaction. Contains 1 replaceable atom
- Potassium phosphate dibasic
- Another potassium source and buffer for the reaction, helping stabilize pH levels/keep them nominal during the reaction. Contains 2 replaceable atoms
- Potassium Glutamate
- Energy / Nucleotide System
- Ribose
- Key cell energy source. It’s crucial for creating adenosine triphosphate (ATP) the primary form of energy within cells
- Glucose
- Another essential energy source for cell processes. Also helps produce ATP
- AMP
- Adenosine Monophosphate (AMP) is a metabolite helping regulate energy levels. It acts as a form of an ATP sensor/response mechanism
- CMP
- Cytidine monophosphate (CMP) assists with RNA synthesis. It helps decompose RNA into ribonuclease (RNase)
- GMP
- Guanosine monophosphate (GMP) is key for RNA synthesis and regulates cellular signaling. Helps polymerize RNA
- UMP
- Uridine monophosate (UMP) is a pyrimidine compound. It also helps polymerize RNA.
- Guanine
- Nucleic acid base that pairs with cystosine in double-stranded DNA. It’s used to build RNA during the transcription process.
- Ribose
- Translation Mix (Amino Acids)
- 17 Amino Acid Mix
- The mix provides a group of compatible amino acids for translation by ribosomes. These are the materials ribosomes work with for translation into proteins
- Tyrosine
- Assists with protein synthesis. Also asists with phosphate group post-translational modification (PTM).
- Cysteine
- An amino acid used by ribosomes to build a protein chains during the translation process. Helps with protein folding and stability.
- 17 Amino Acid Mix
- Additives
- Nicotinamide
- Helps manage the process of cellular nutrients converting themselves to ATP and vice versa. Think this means it also might help with reaction energetic stability
- Nicotinamide
- Backfill
- Nuclease Free Water
- Ensures appropriate reaction concentration. It also ensures no extraneous enzymes destroy reaction byproducts
- Nuclease Free Water
- Describe the main differences between the 1-hour optimized PEP-NTP master mix and the 20-hour NMP-Ribose-Glucose master mix shown in the Google Slide above.
- Believe the main difference between the 1-hour optimized PEP-NTP master mix and the 20-hour NMP-Ribose-Glucose master mix is that the 1-hour optimized PEP-NTP master mix optimizes for the fast production of flourescent proteins, while the 20-hour NMP-Ribose-Glucose master mix optimizes for flourescent proteins across a longer timespan. While this might seem obvious based on the slide content, my understanding is that the PEP-NTP master mix is more energy intensive (i.e., it essentially consumes more energy faster to create flourescent protein output) while the NMP-Ribose-Glucose master mix is comparatively less energy intensive (i.e., it essentially consumes either less energy slower across its 20 hour-reaction timespan to create flourescent protein output or it consumes the same amount of energy across its 20 hour-reaction timespan in a less energy intensive fashion). So, in essence, I think the main difference between these reactions comes down to their respective energy constumption levels
Supporting prompts for this section listed below:
| Supporting Prompt | Model |
|---|---|
| Under the ‘Salts/Buffer’ subsection under the 1st question in Part B, I think glutamates help with the creation of a given chemical (in this case potassium or magnesium). Not sure how/why these salts/buffers are relevant in a cell-free protein synthesis reaction. Any insights you might have into the roles of the glutamates here, as well as all the various types of potassium would be useful. Do NOT hallucinate/make things up when replying to this prompt | Gemini |
| When we say something is a buffer for a chemical reaction, what exactly do we mean when we say that? Do NOT hallucinate/make things up when replying to this prompt and keep the answer relatively succinct and plainspoken (4-5 sentences max) | Gemini |
| Looking at the ‘Energy/Nucleotide’ subsection under the 1st question in Part B, and given a passing understanding of genomics, I understand that guanine (G) pairs with Cytosine (C). In the context of this subsection, does this mean that a cell-free Guanine mix translates or outputs Cysteine in some may? Is that a relationship between these two things? If so, what’s the relationship? Do NOT hallucinate/make things up when replying to this prompt and keep the answer relatively succinct and plainspoken (5-6 sentences max) | Gemini |
| Looking at the ‘Backfill’ subsection under the 1st question in Part B, my understanding of nuclease free water’s function in a cell-free protein synthesis reaction is to basically provide a clean backdrop for the reaction to occur, or to determine what’s what post-reaction. Are either of those high-level explanations correct or sensible? Do NOT hallucinate/make things up when replying to this prompt and keep the answer relatively succinct and plainspoken (4-5 sentences max) | Gemini |
| Doing a sanity check: ribosomes turn amino acids into proteins as part of the translation process, correct? Do NOT hallucinate/make things up when replying to this prompt and keep the answer relatively succinct and plainspoken (2-3 sentences max) | Gemini |
Part C: Planning the Global Experiment | Cell-Free Master Mix Design
Given the 6 fluorescent proteins we used for our collaborative painting, identify and explain at least one biophysical or functional property of each protein that affects expression or readout in cell-free systems. (Hint: options include maturation time, acid sensitivity, folding, oxygen dependence, etc) (1-2 sentences each)
- sfGFP
- It has a relatively quick maturation time (13.6 min.). This means researchers can find out whether or not the cell-free reaction occurred successfully rather quickly if they were solely measuring this protein’s flourescence
- mRFP1
- It has a comparably longer maturation time (60 min.). This means researchers might need to wait a bit to determine whether or not the cell-free reaction occurred successfully if they were solely measuring this protein’s flourescence
- mKO2
- Acid sensitivitiy levels are 5.5 pKa. This means that if pH drops from the typical 7.5 pH of a common cell-free reaction, this will cause the flourescence to not show
- mTurquoise2
- It’s acid sensitivitiy levels are 5.5 pKa and its maturation time is also relatively quick (33.5 min.). This means its realtively resistant to drops in pH (i.e., a flourescence readout will still occur) and a researcher can discern whether or not a successful reaction occurred relatively quickly if they were solely measuring this protein’s flourescence
- mScarlet_1
- It’s acid sensitivitiy levels are 5.3 pKa and its maturation time is comparatively long (174 min.). This means the protein’s relatively senstive to pH drops from the common mean, and it will also take several hours for a researcher to discern whether or not a successful reaction occurred if they were solely measuring this protein’s flourescence
- Electra2
- It’s the second brightest of all the proteins in this list. It has a 61.48 brightness readout (the brightest protein is mScarlet_1 with a 70.0 brightness readout)
- sfGFP
Create a hypothesis for how adjusting one or more reagents in the cell-free mastermix could improve a specific biophysical or functional property you identified above, in order to maximize fluorescence over a 36-hour incubation. Clearly state the protein, the reagent(s), and the expected effect.
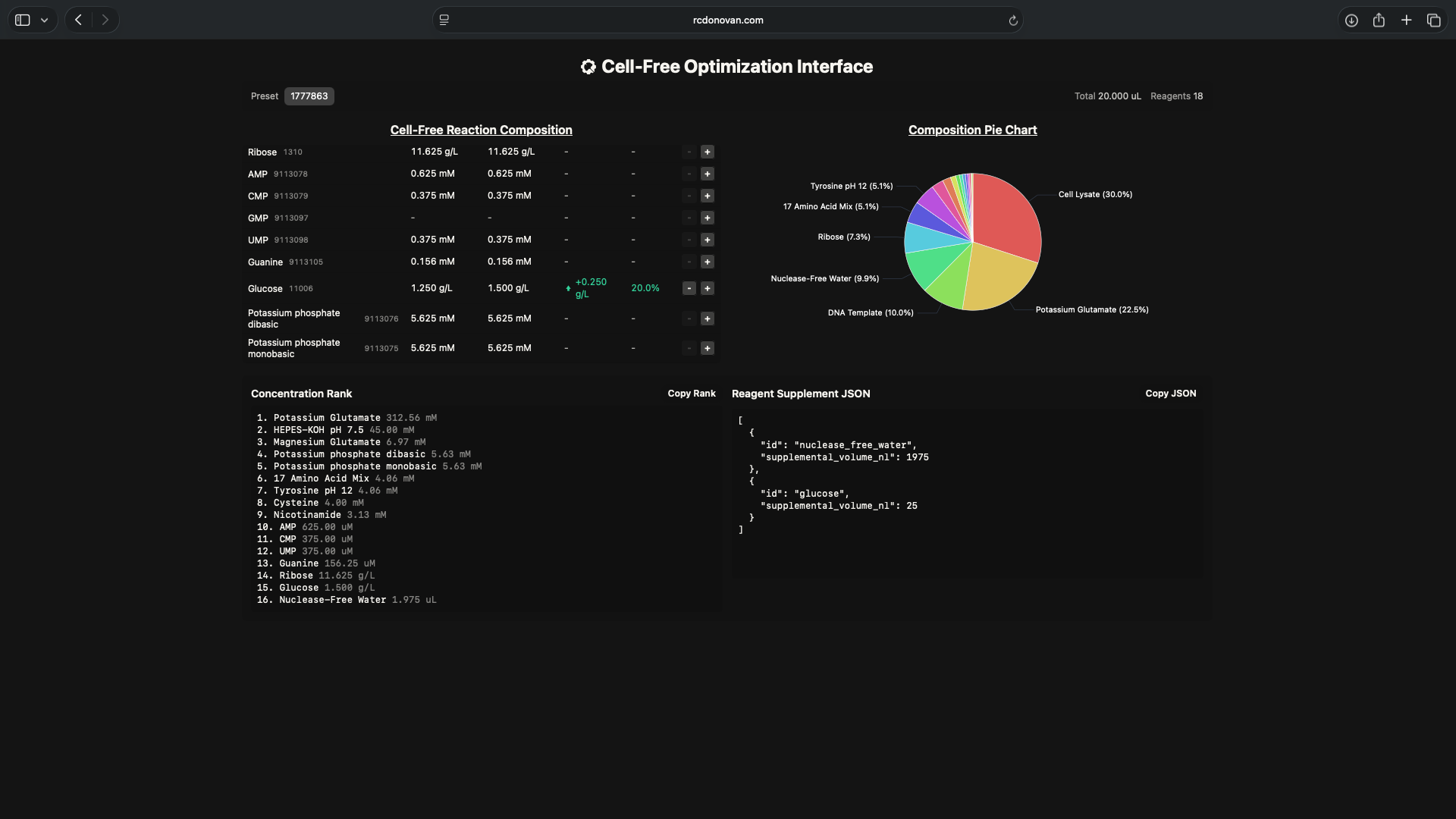
- I hypothesize that if I increase ribose and/or glucose reagent concentrations in the cell-free mastermix, it will increase sfGFP brightness over a 36-hour incubation period relative to its nominal brightness rate (54.15)
The second phase of this lab will be to define the precise reagent concentrations for your cell-free experiment. You will be assigned artwork wells with specific fluorescent proteins and receive an email with instructions this week (by April 24). You can begin composing master mix compositions here 1
- Test reagent master mix compositions based on the hypothesis above shown in the screenshots below
The final phase of this lab will be analyzing the fluorescence data we collect to determine whether we can draw any conclusions about favorable reagent compositions for our fluorescent proteins. This will be due a week after the data is returned (date TBD!). The reaction composition for each well will be as follows:
- 6 μL of Lysate
- 10 μL of 2X Optimized Master Mix from above
- 2 μL of assigned fluorescent protein DNA template
- 2 μL of your custom reagent supplements
Supporting prompts for this section listed below:
| Supporting Prompt | Model |
|---|---|
| In the ‘Attributes’ table in this tab, what does the ‘Maturation (min.)’ value mean? Why does it matter in practical terms? Do NOT hallucinate/make things up when replying to this prompt and keep the answer relatively succinct and plainspoken (3-4 sentences max) | Gemini |
| On this tab, when we say mK02 has moderate acid sensitivity, what does that actually mean? What acids is it sensitive to? Any type of acid? Why does acid sensitivity matter in practical terms? Do NOT hallucinate/make things up when replying to this prompt and keep the answer relatively succinct and plainspoken (5-6 sentences max) | Gemini |
| So based on mK02’s moderate acid sensitivity readout (5.5) would we say that that readout is suboptimal/undesirable relative to normal cell-free reaction pH? Do NOT hallucinate/make things up when replying to this prompt and keep the answer relatively succinct and plainspoken (2-3 sentences max) | Gemini |
| Looking over the subsections in Part B, does every single subcategory consist of reagents, or do only some subcategories consist of reagents? Do Do NOT hallucinate/make things up when replying to this prompt and keep the answer relatively succinct and plainspoken (3-4 sentences max) | Gemini |
| This might be a dumb question. If I give a cell-free reaction more power in the form of higher concentrations of some of the ATP/cellular energy-associated reagents under the ‘Energy/Nucleotide System’ subsection, could I expect a decreased maturation time for fluorescent protein readout/indication of fluorescence? Do Do NOT hallucinate/make things up when replying to this prompt and keep the answer relatively succinct and plainspoken (5-6 sentences max) | Gemini |
| In the response to the last prompt, the energy reagents listed were AMP, CMP, GMP, or UMP. What if ribose or glucose levels were increased? Could I expect a decreased maturation time for fluorescent protein readout/indication of fluorescence for sfGPF? Do Do NOT hallucinate/make things up when replying to this prompt and keep the answer relatively succinct and plainspoken (5-6 sentences max) | Gemini |
| Understood. What about the impact of increased energy reagents on sfGFP brightness? If I added increased concentrations of some of the ATP/cellular energy-associated reagents under the ‘Energy/Nucleotide System’ subsection, such as Ribose or Glucose, in principle could I expect to get brighter/more fluorescent sfGFP as an output? Do Do NOT hallucinate/make things up when replying to this prompt and keep the answer relatively succinct and plainspoken (5-6 sentences max) | Gemini |