Jason Ross — HTGAA Spring 2026


About me
Passionate about space-based biotechnology applications and strengthening biosecurity. Excited to grow my technical acumen through this course!
Contact info
Please feel free to reach out via the HTGAA Discourse forum at 2026a-jason-ross!
Homework
- Week X HW: Test
- Week 1 HW: Principles and Practices
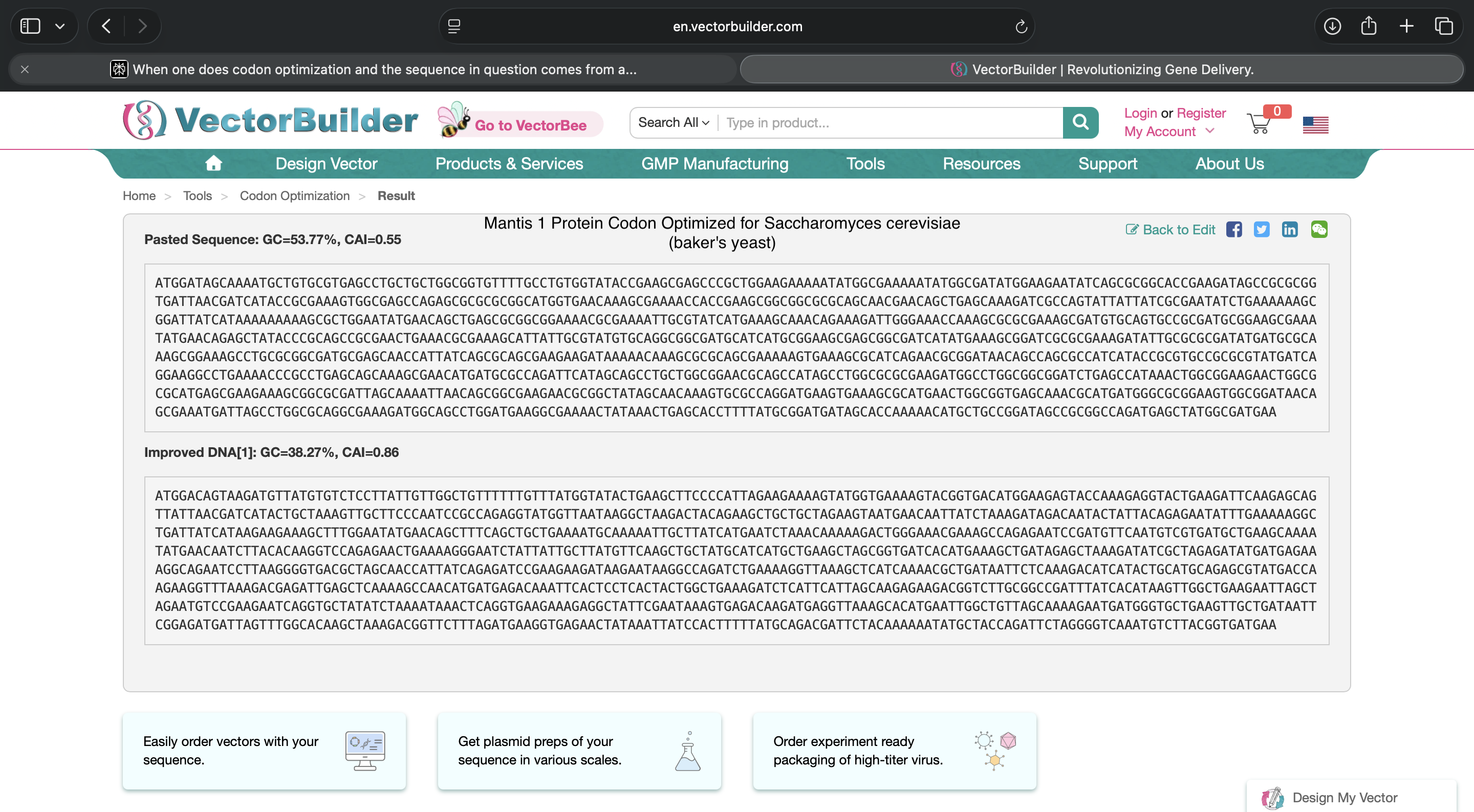
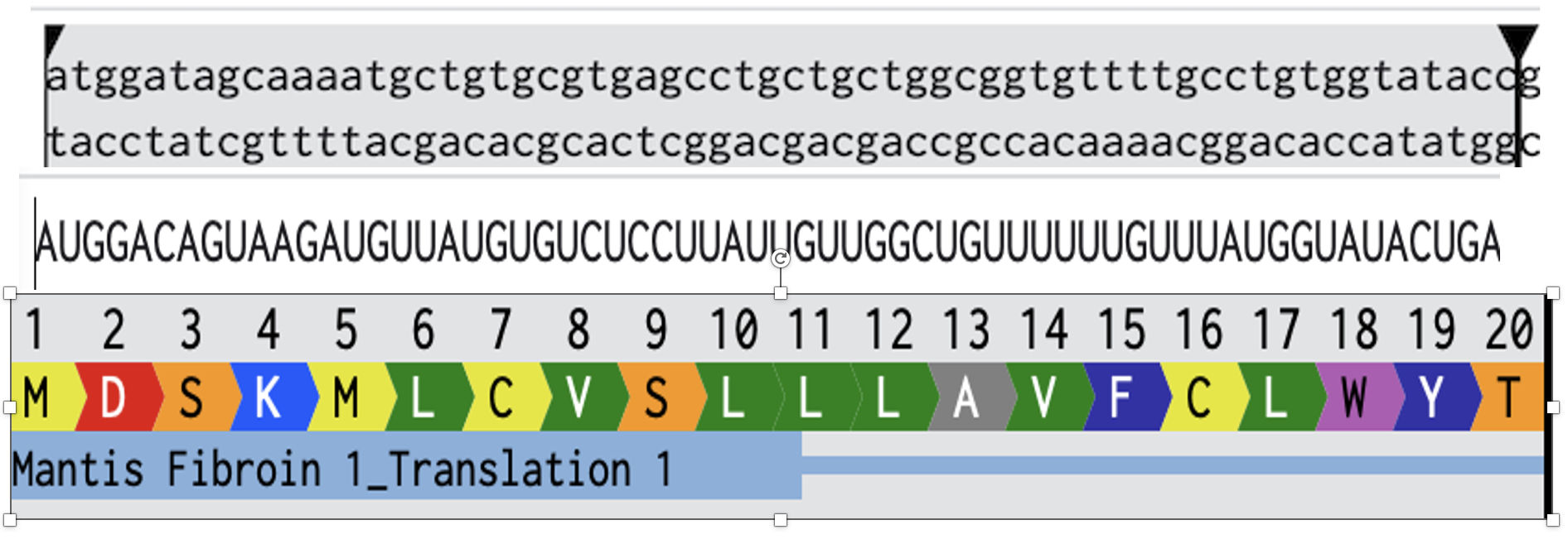
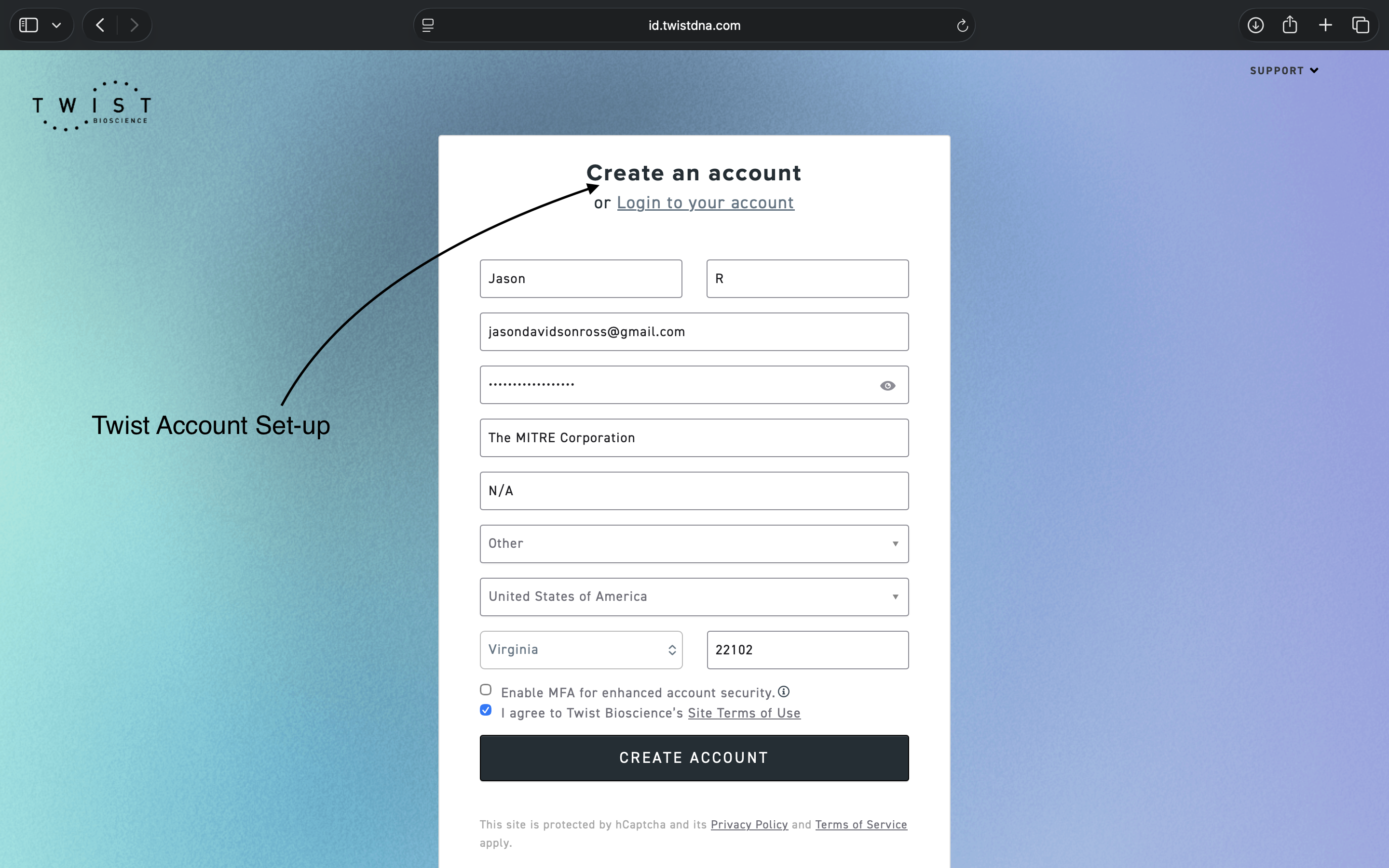
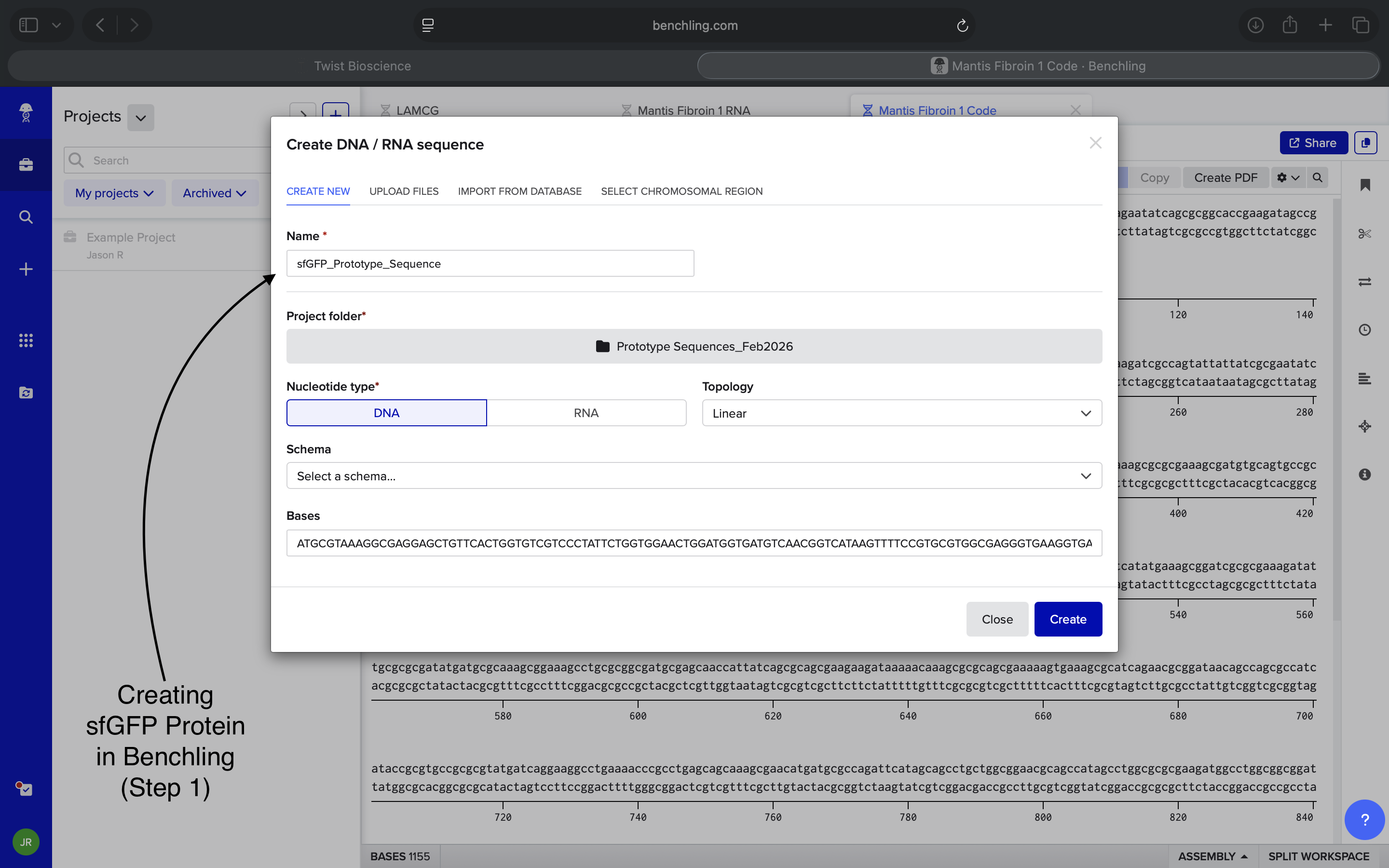
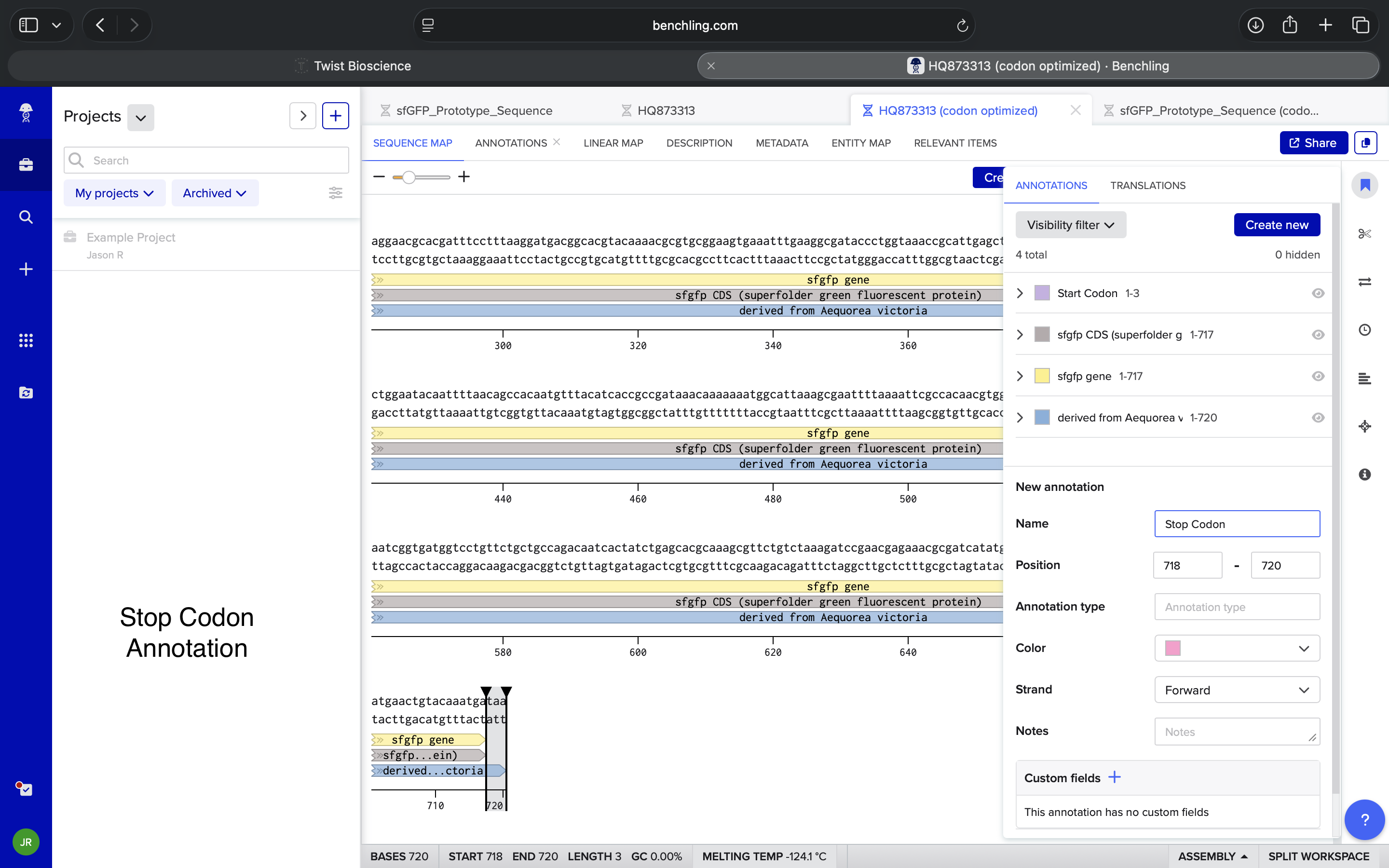
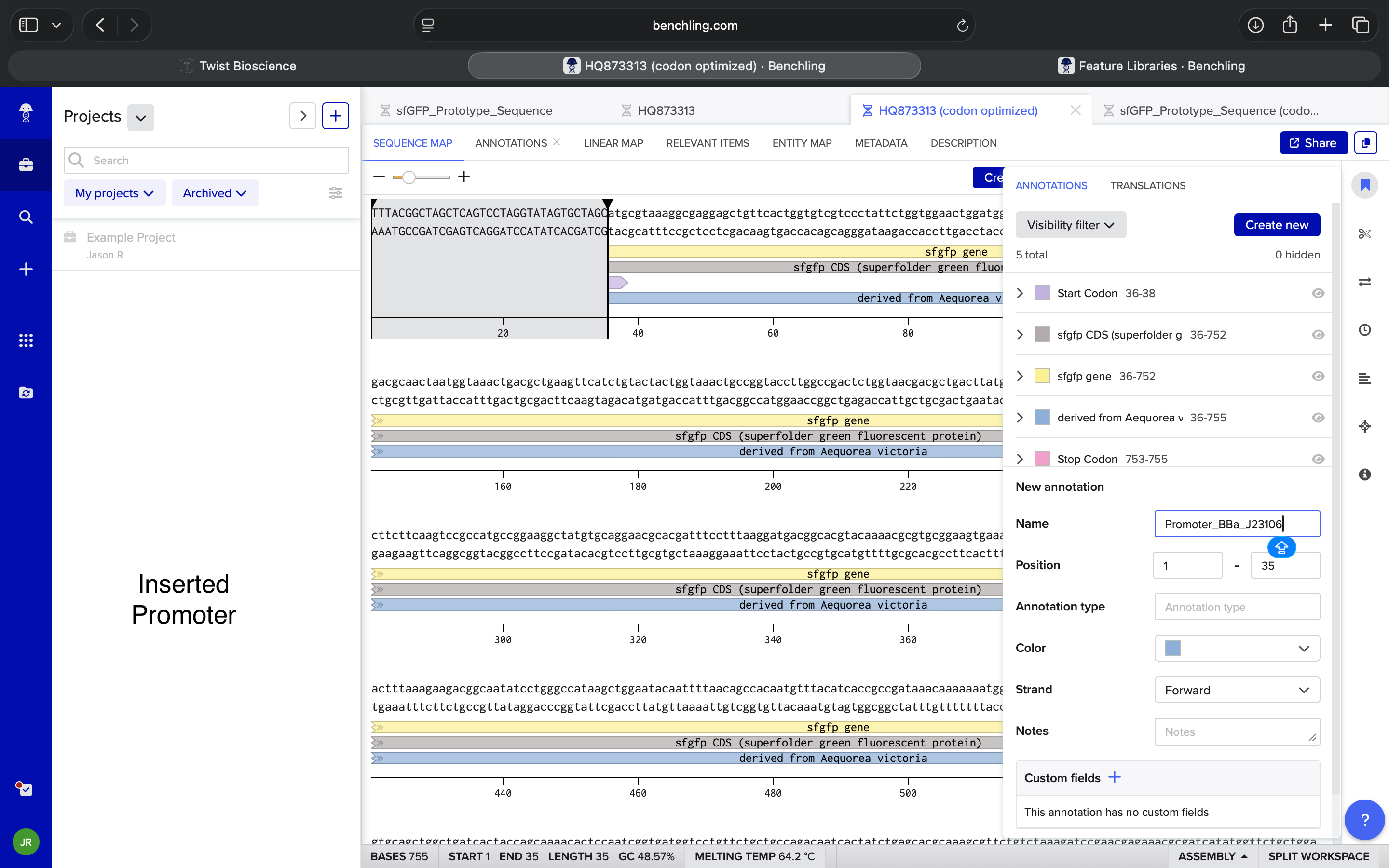
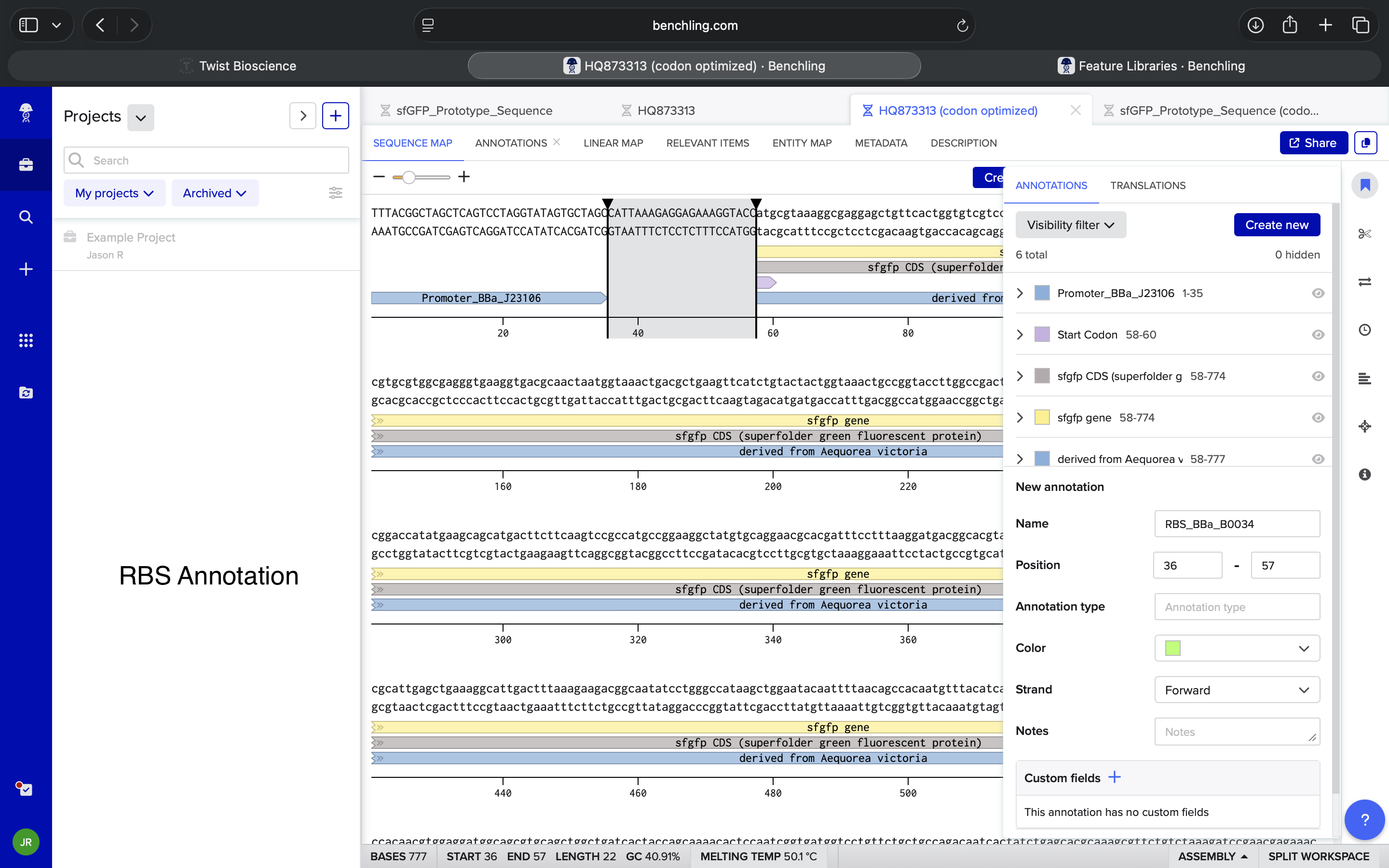
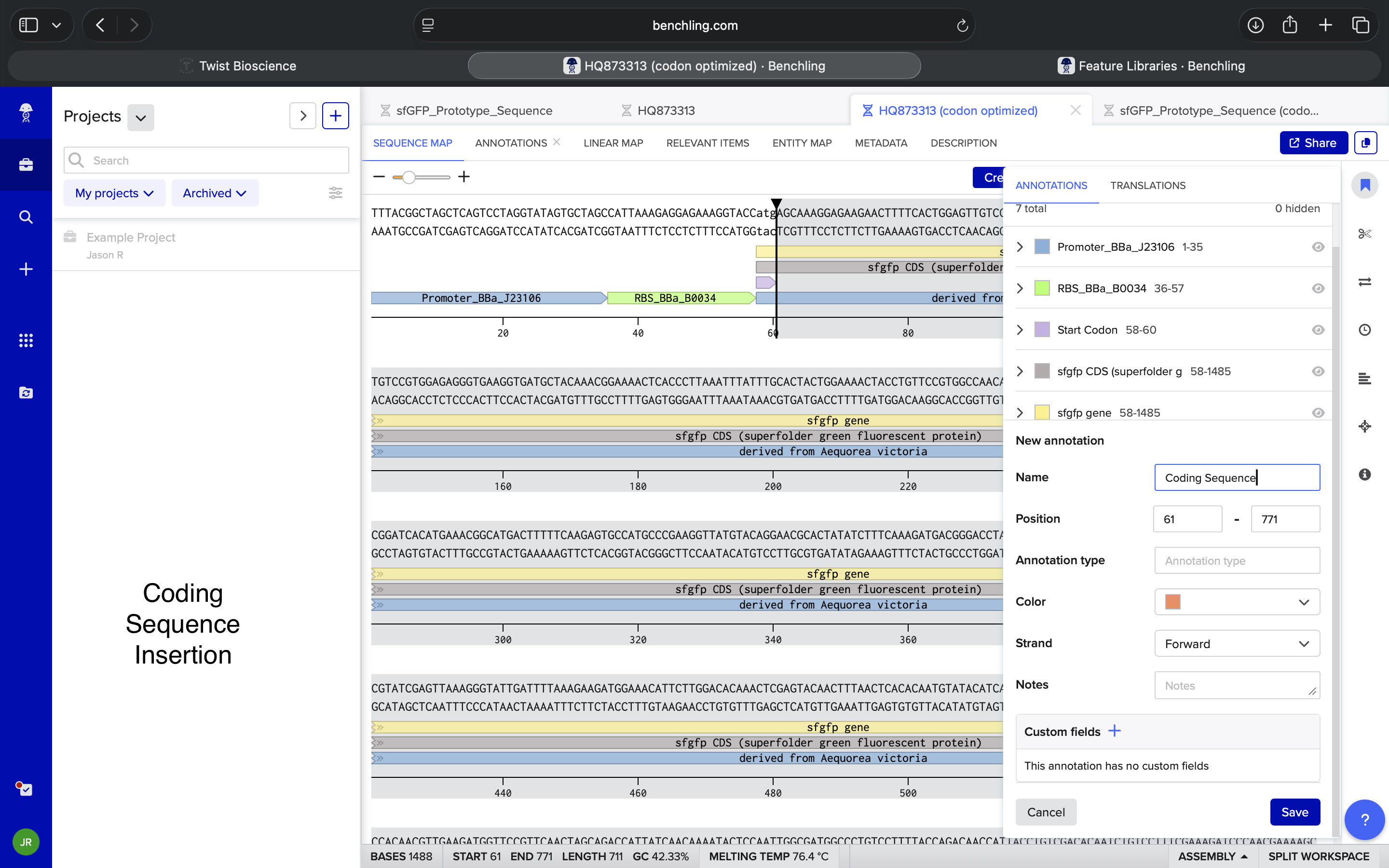
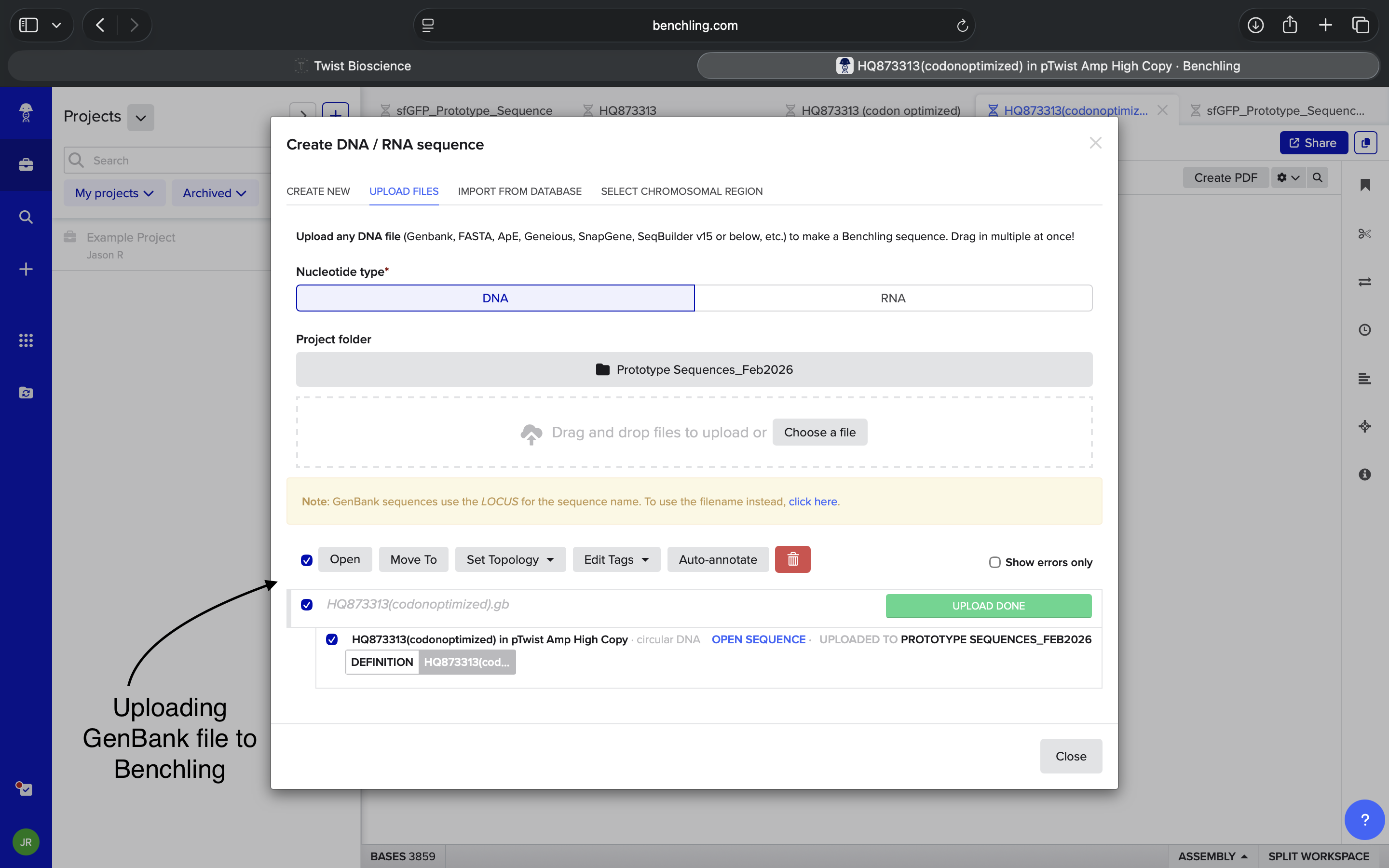
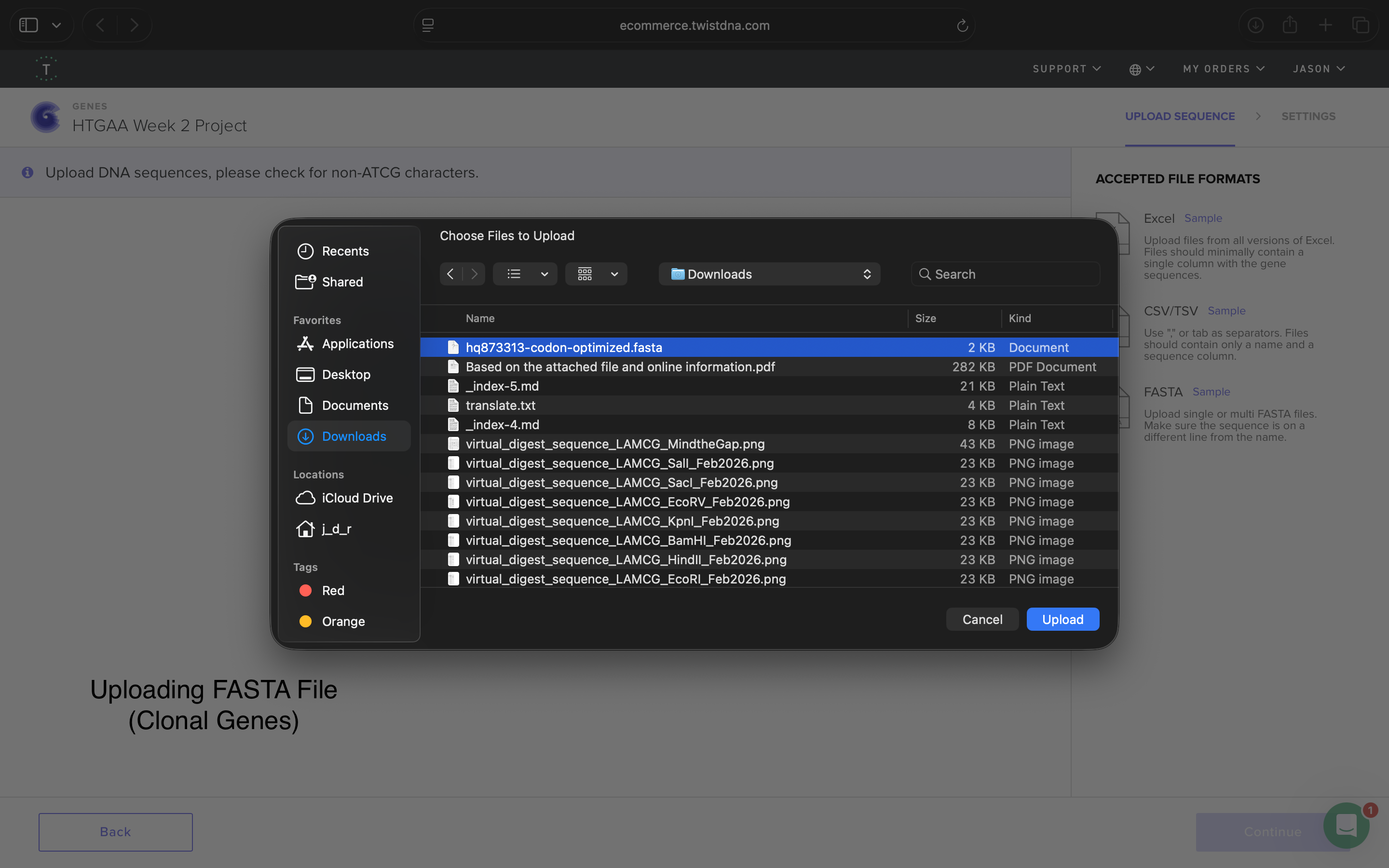
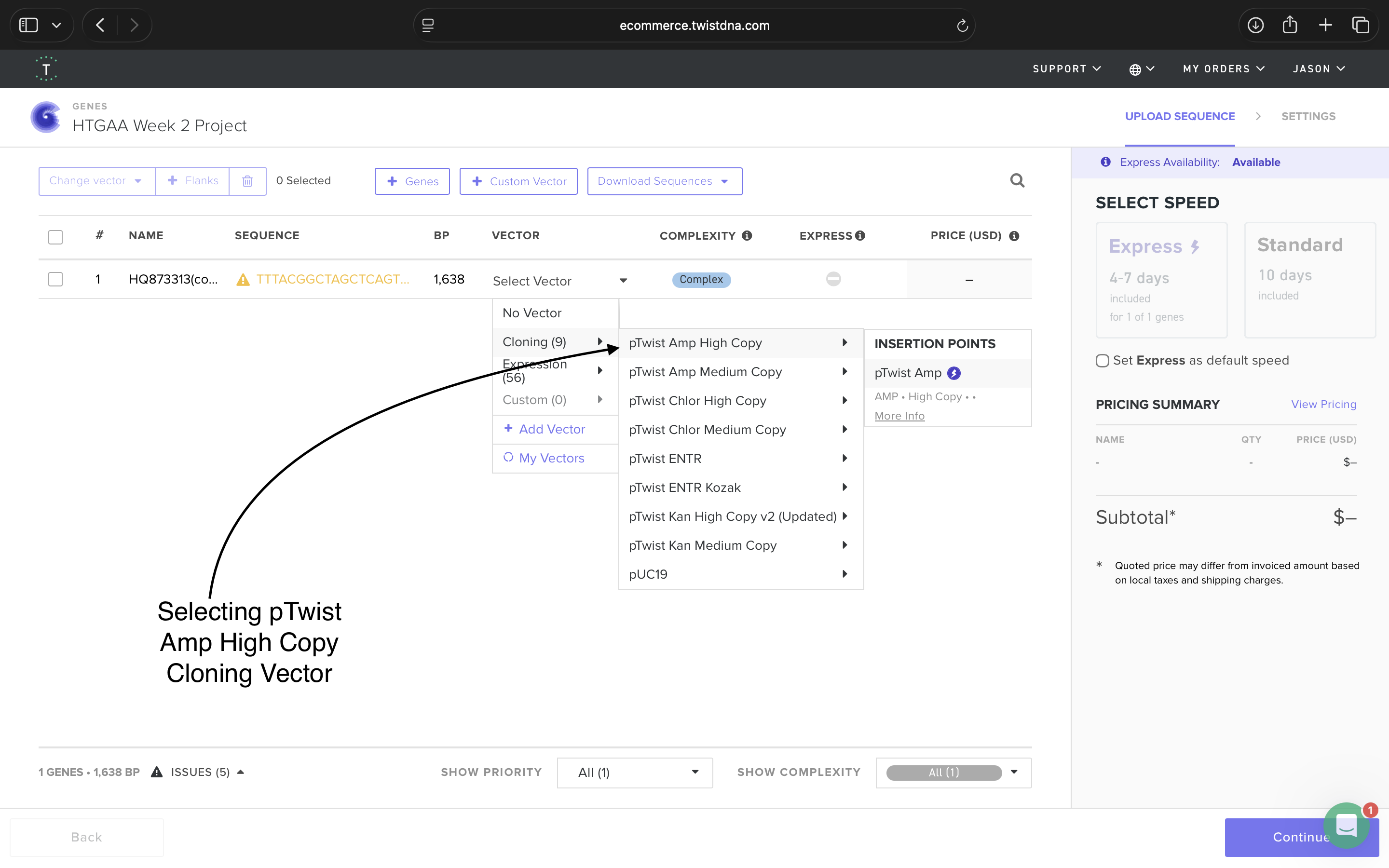
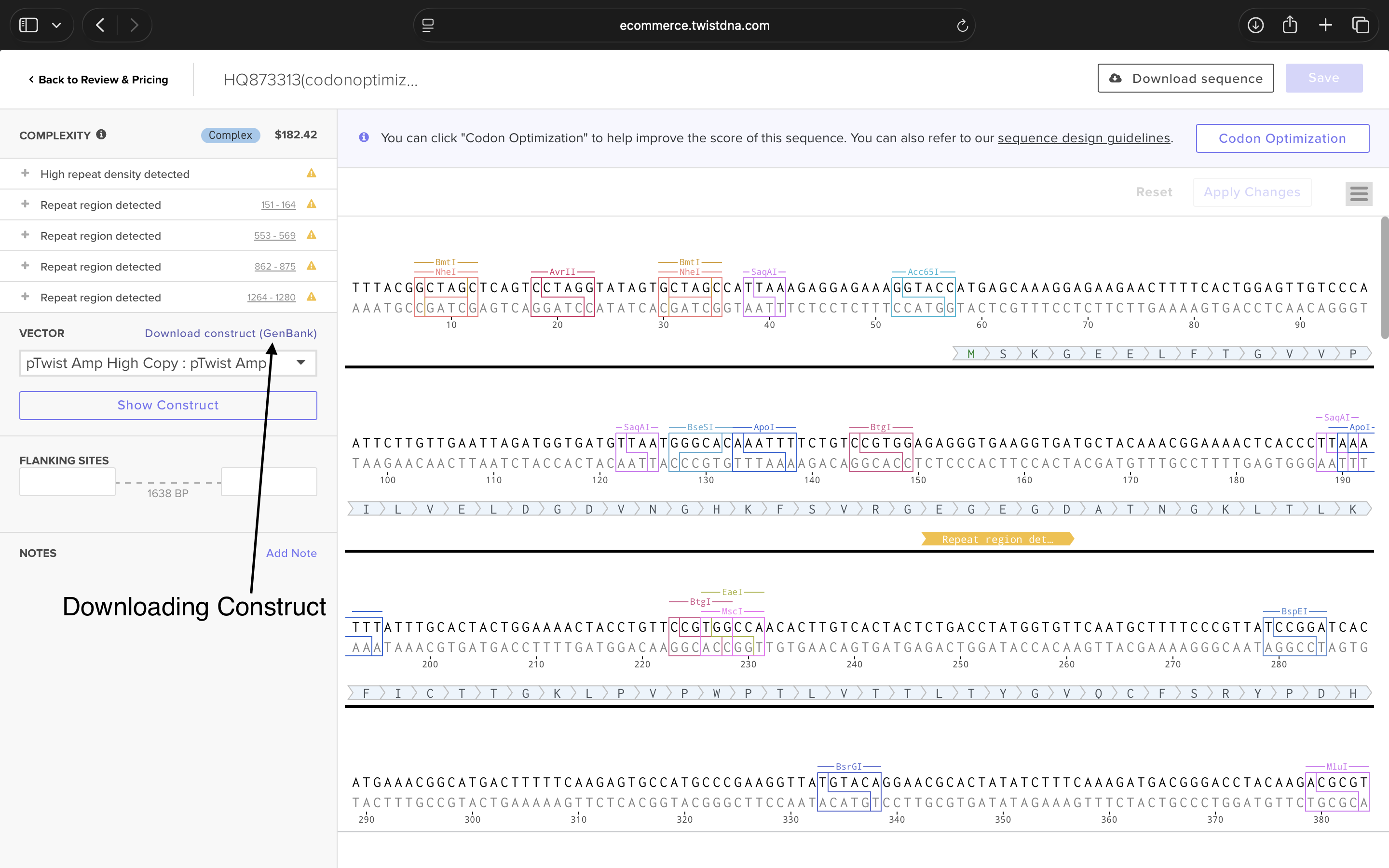
- Week 2 HW: DNA Read, Write, and Edit
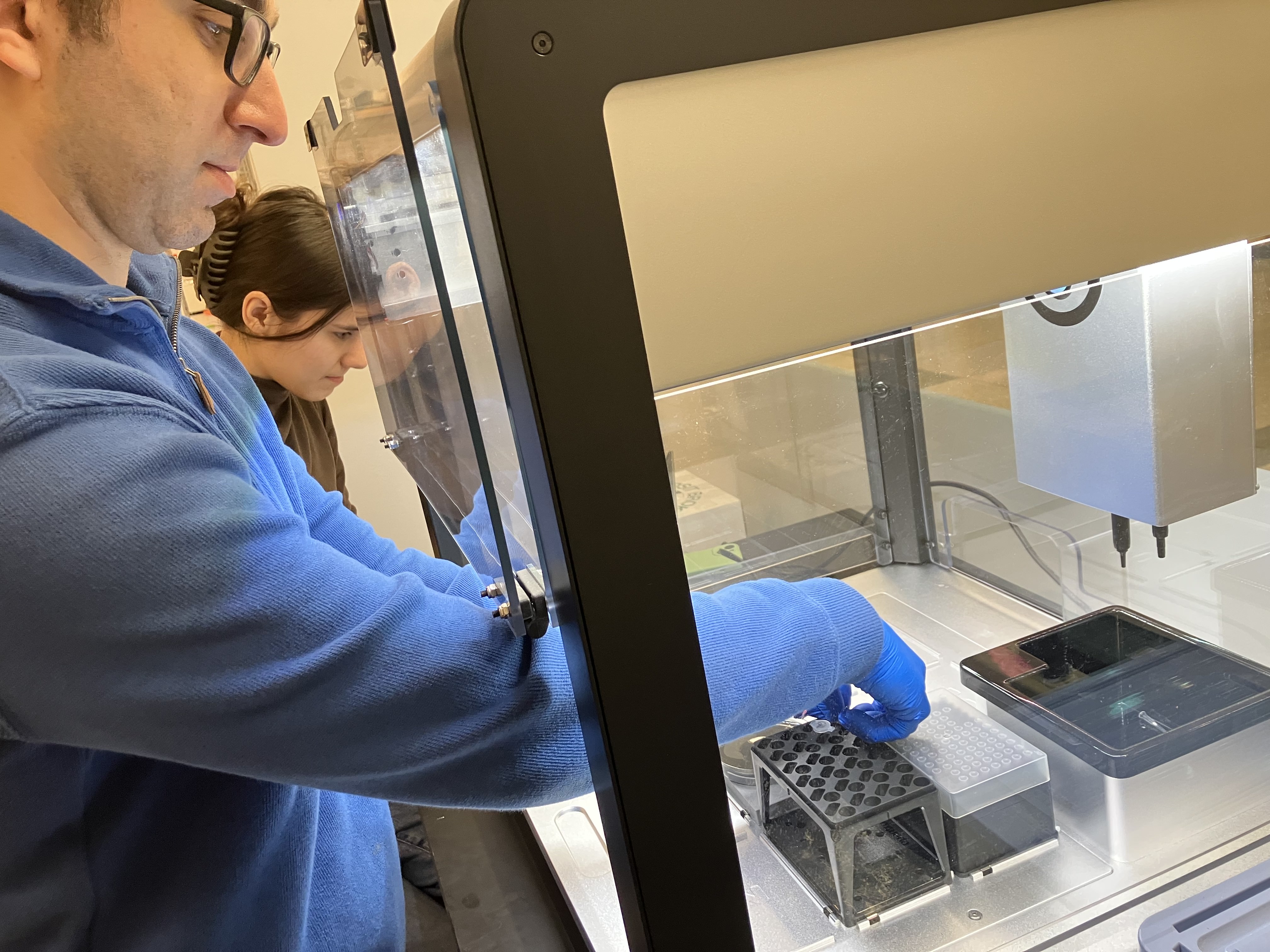
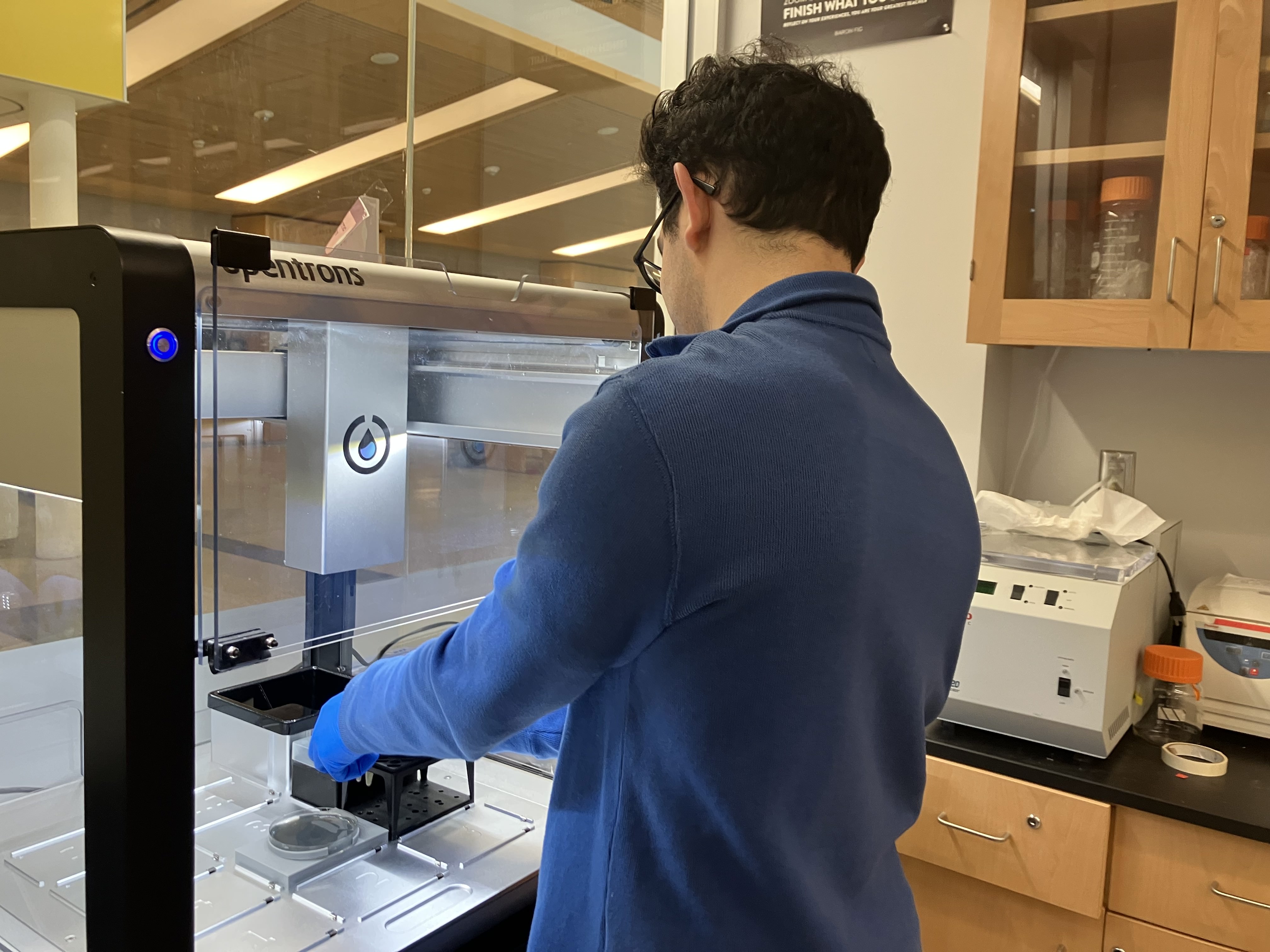
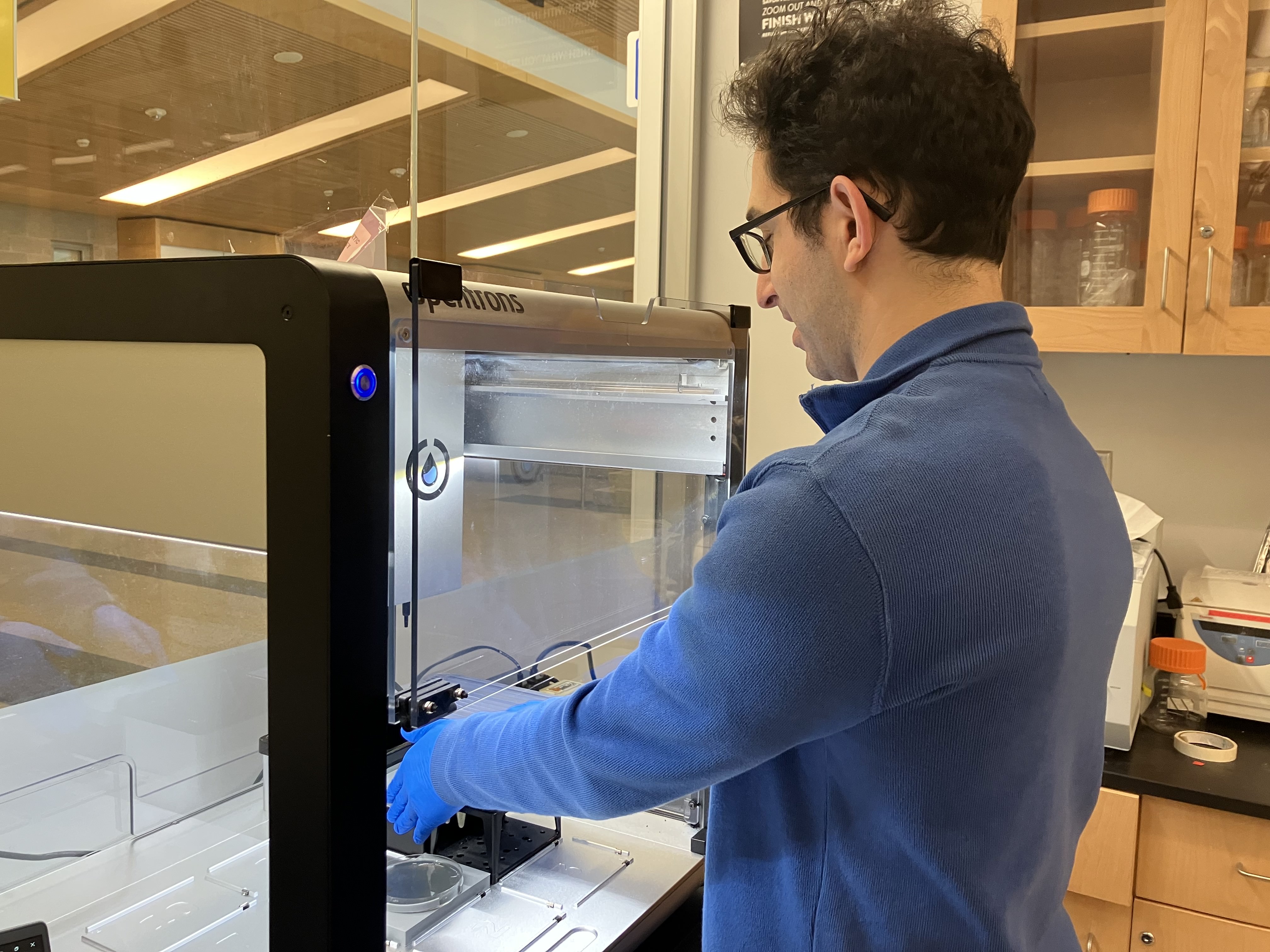
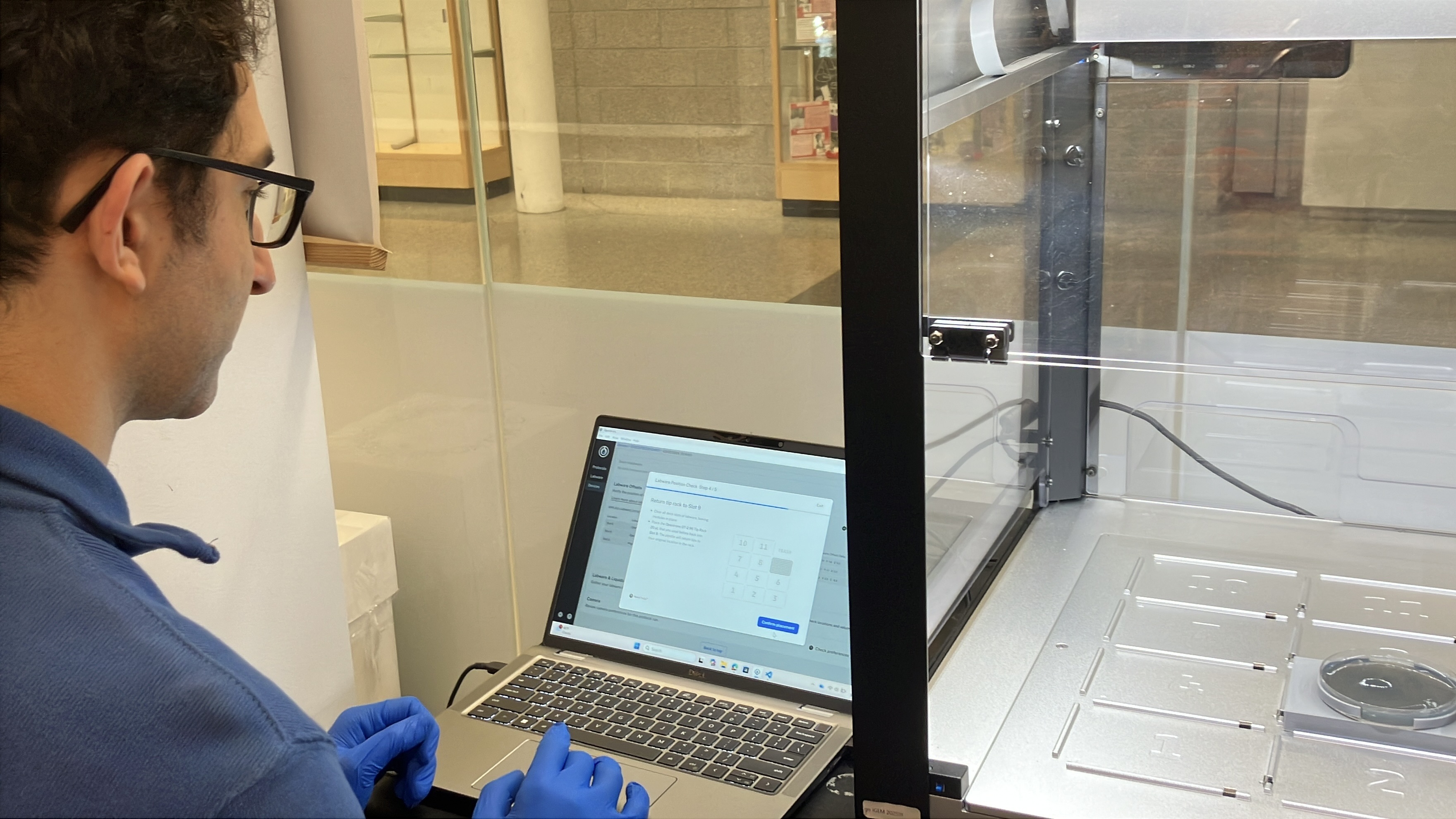
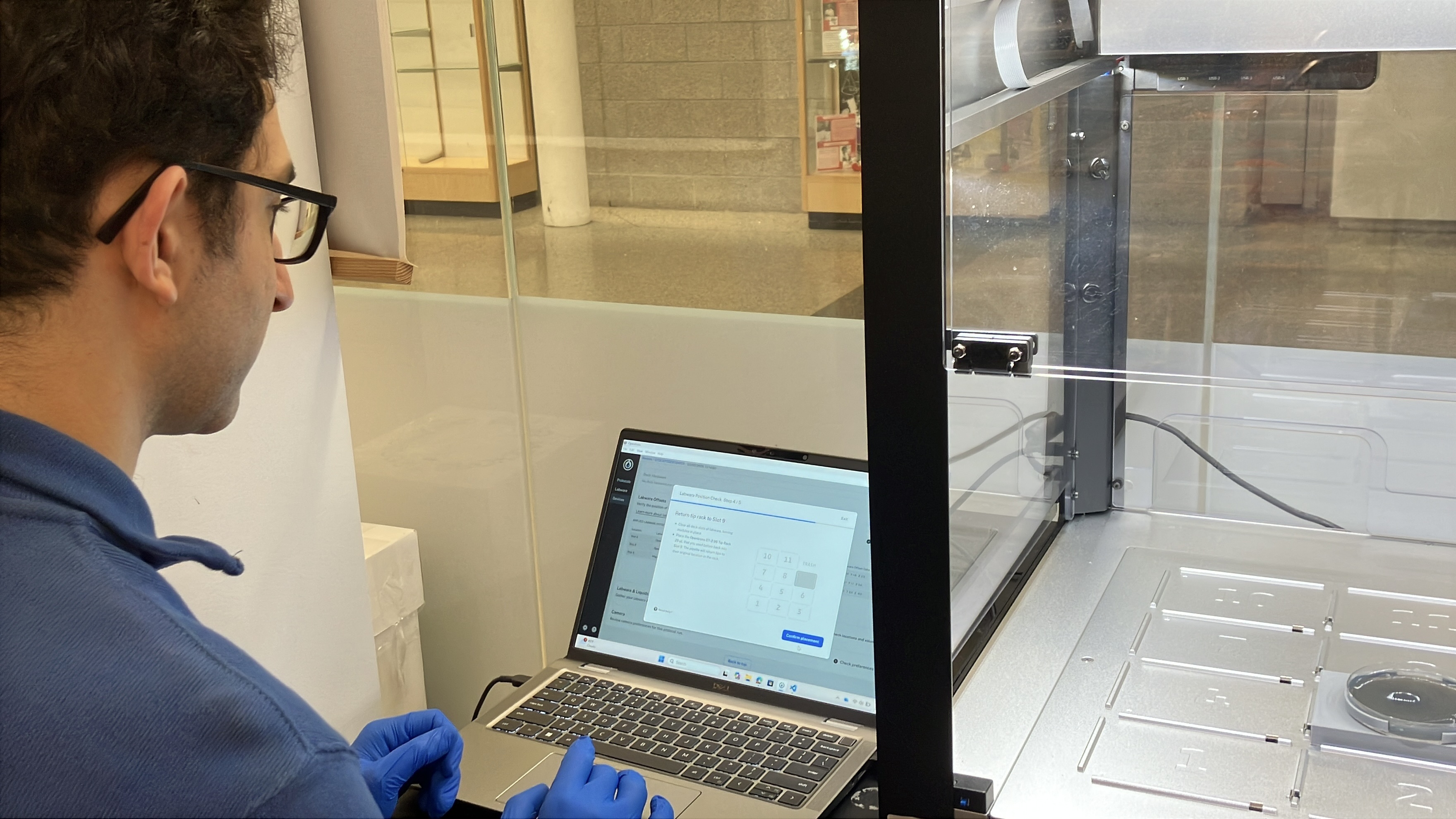
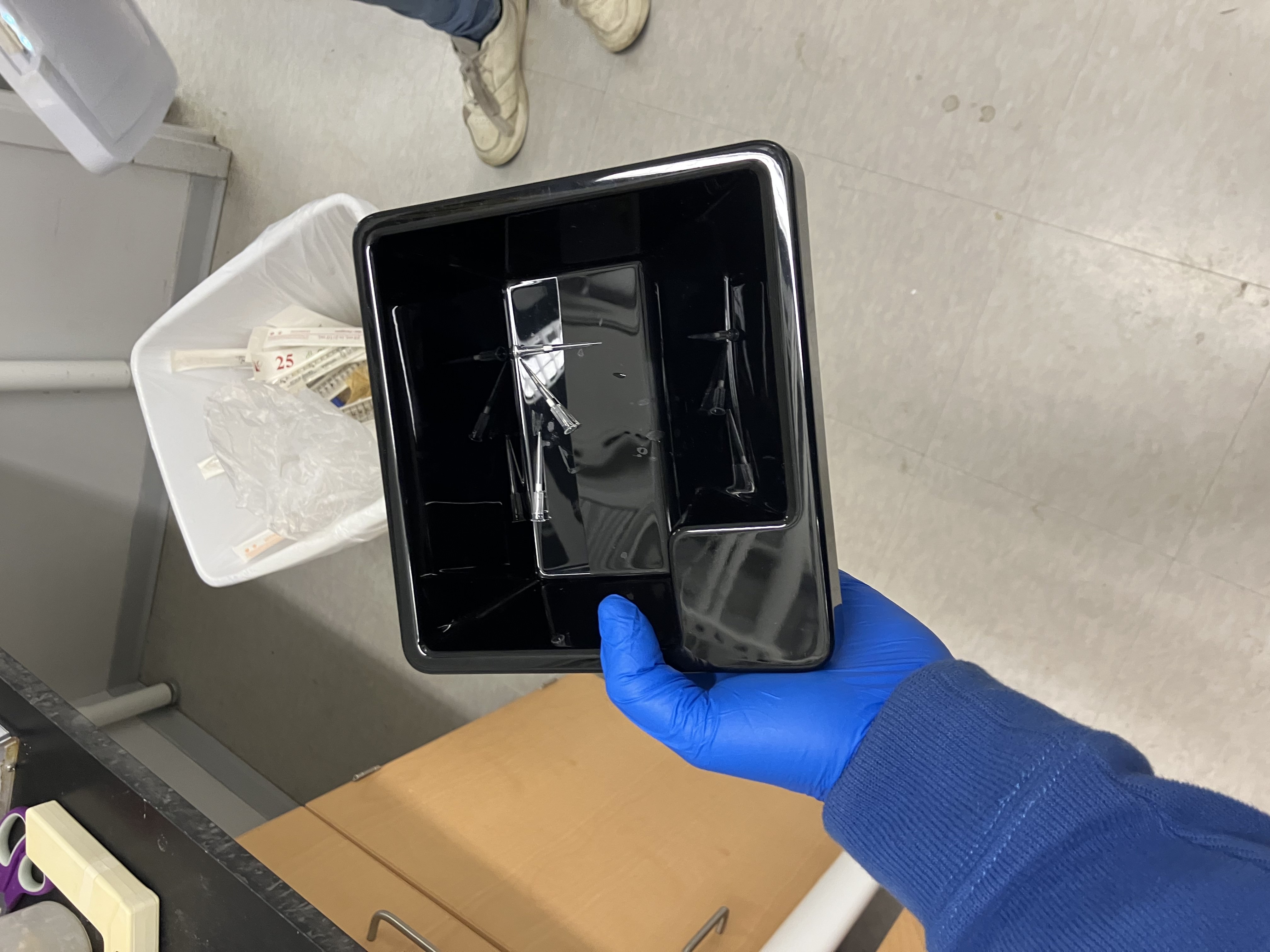
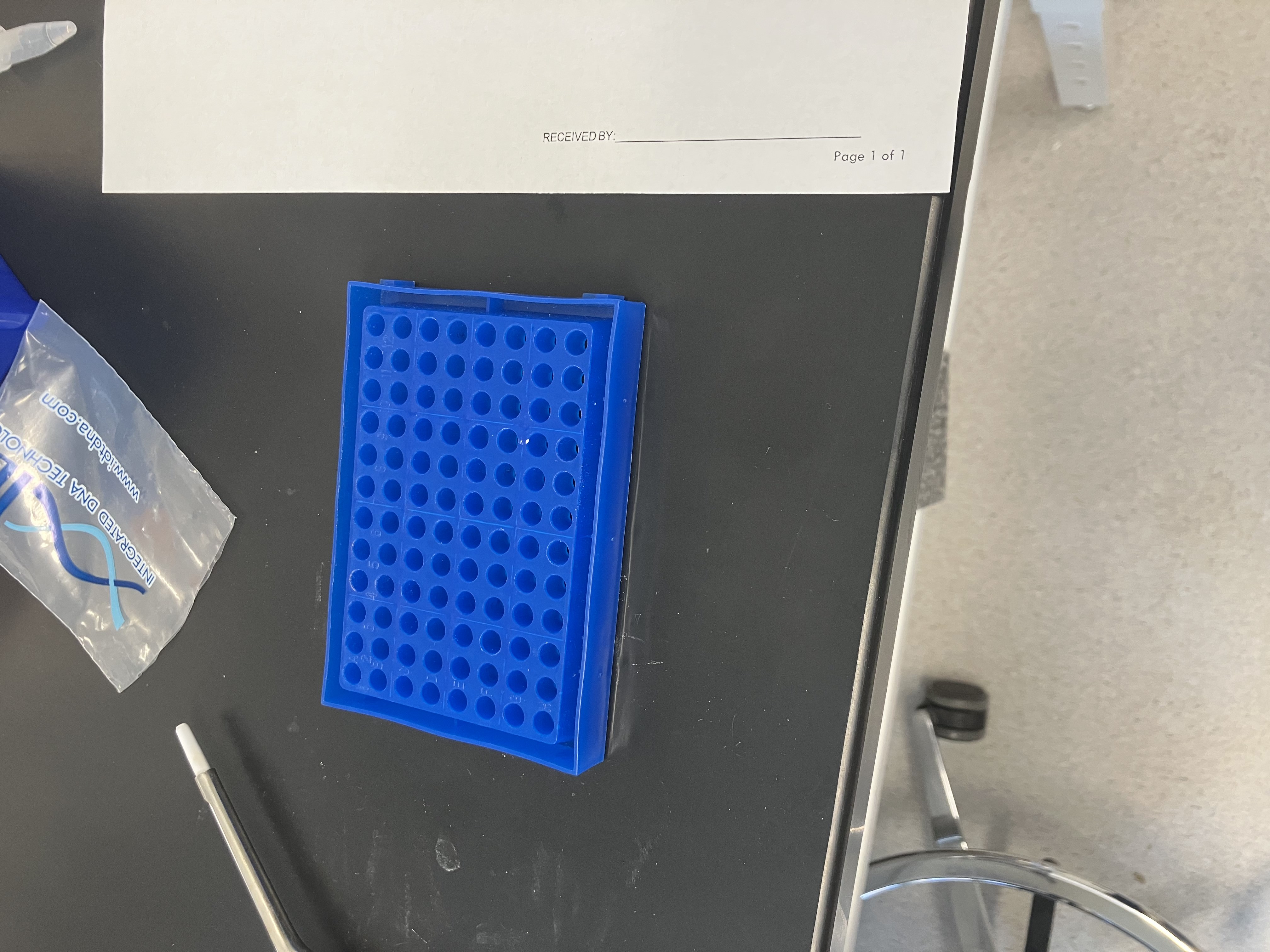
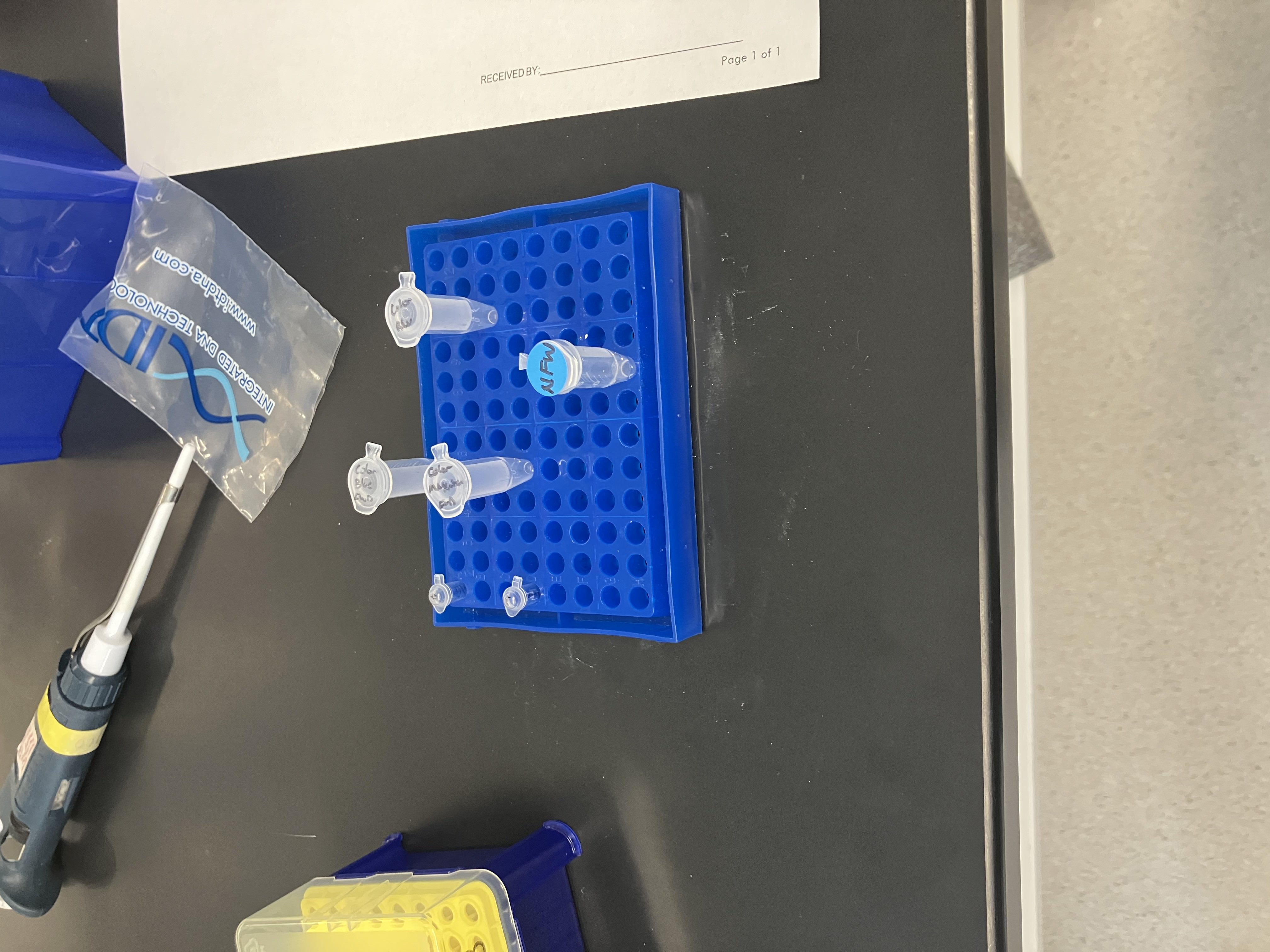
- Week 3 HW: Lab Automation
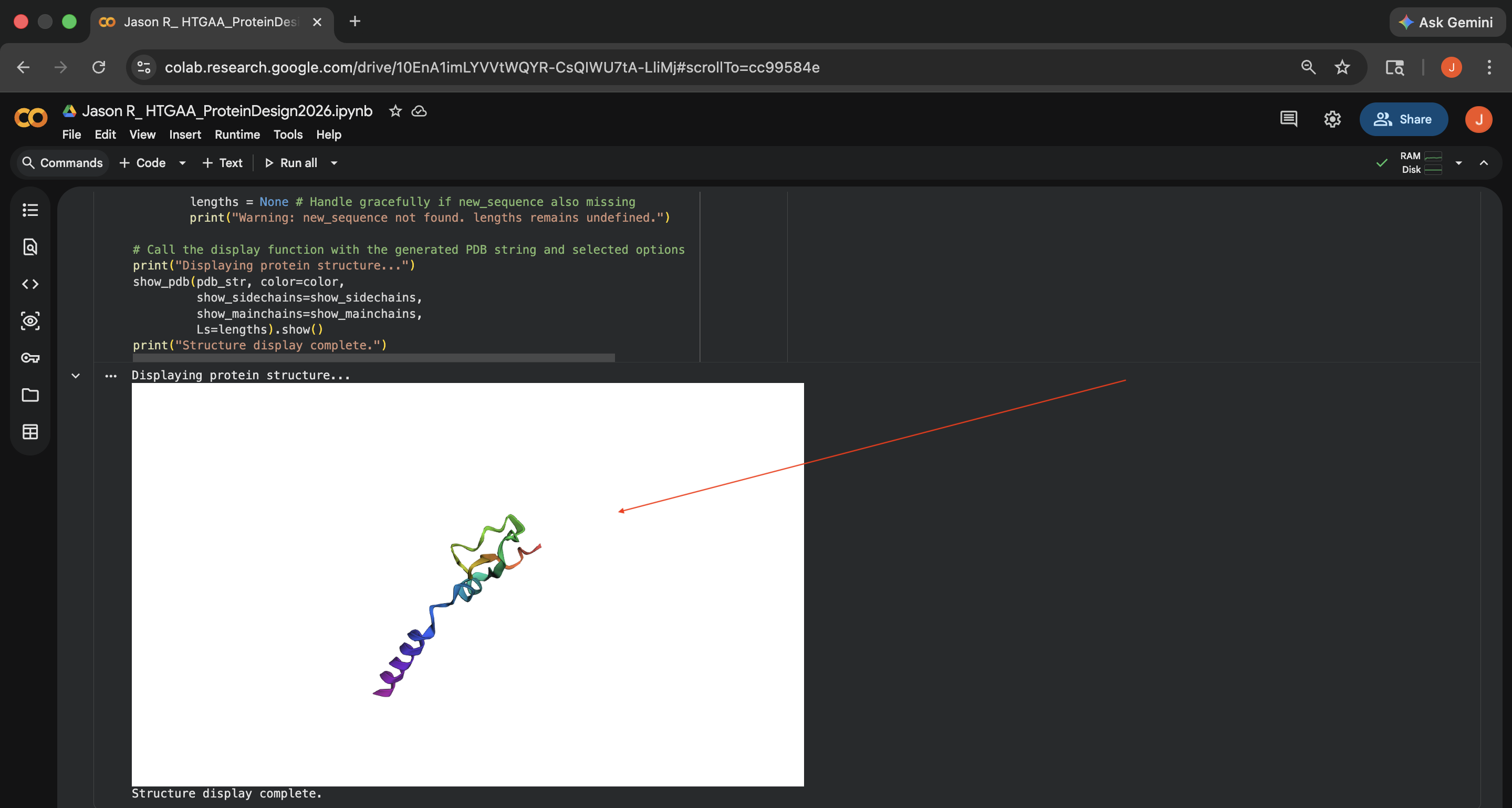
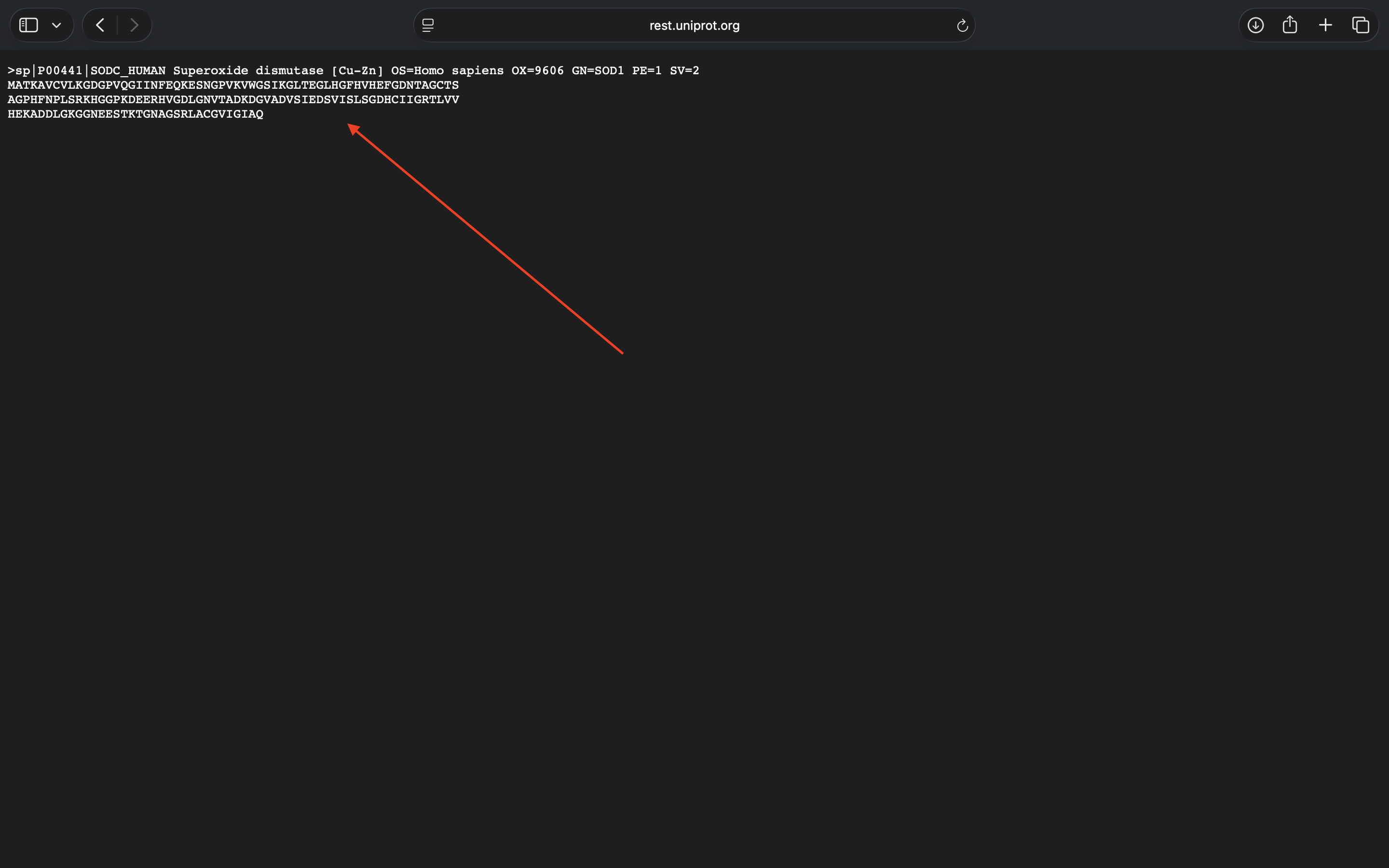
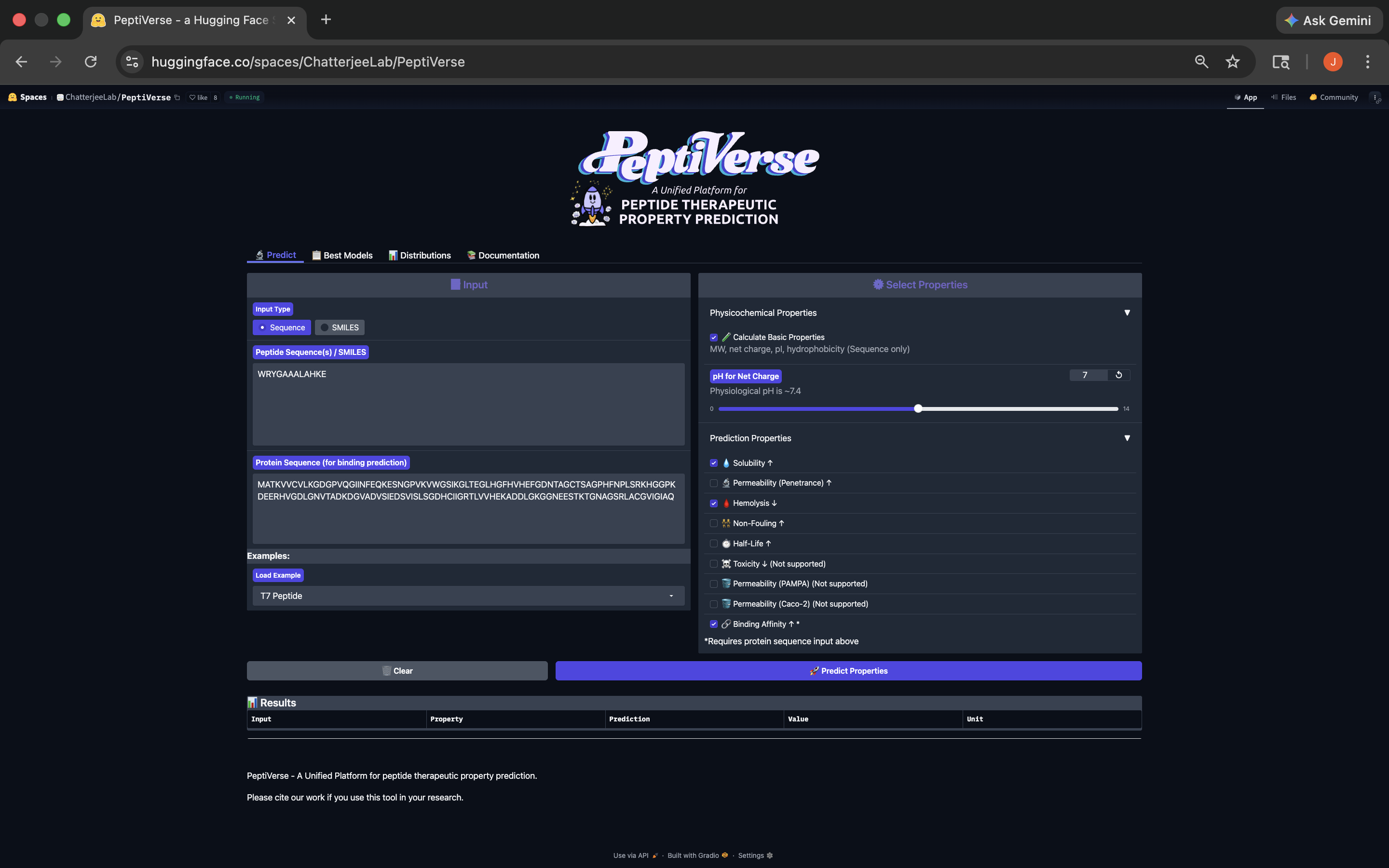
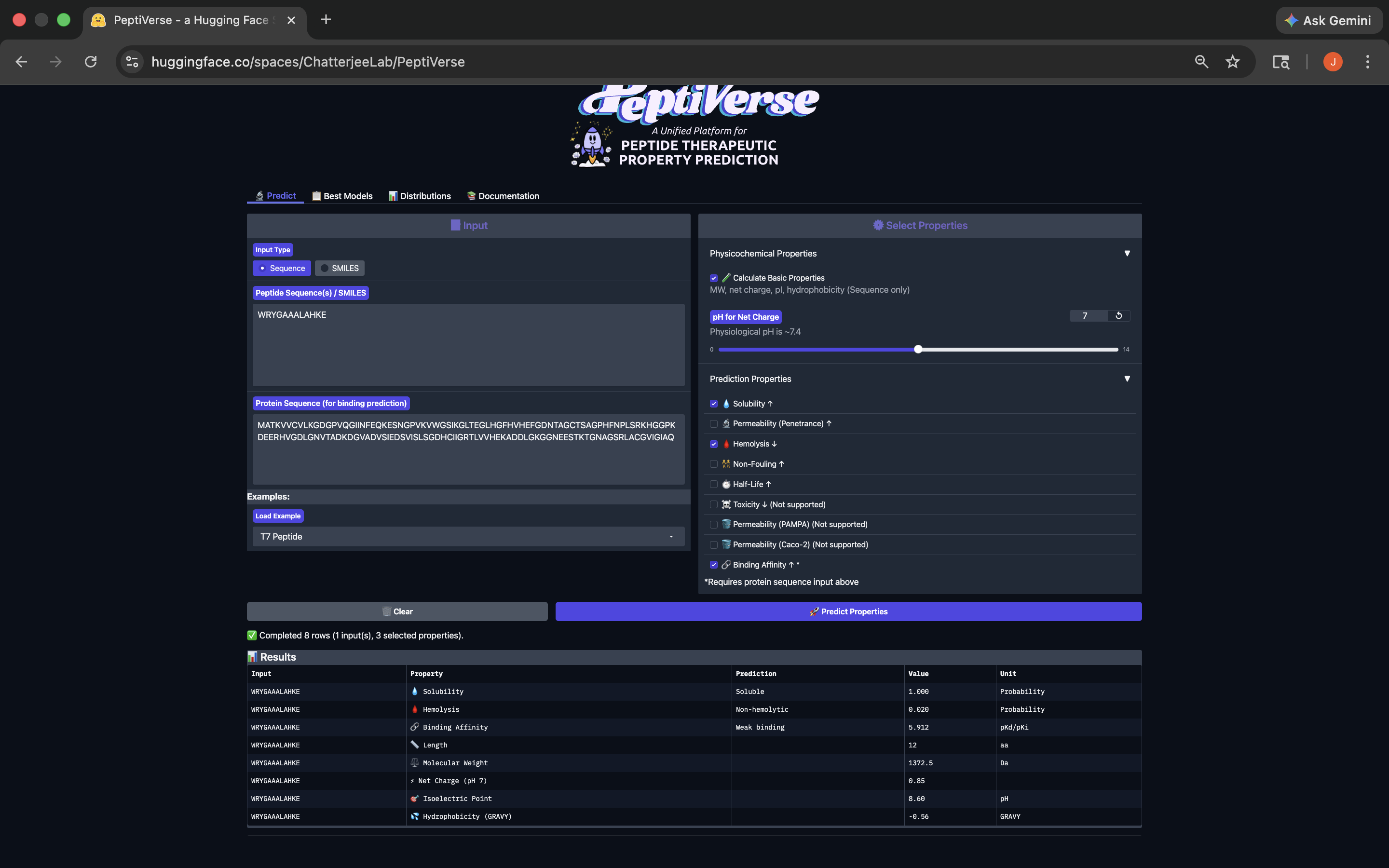
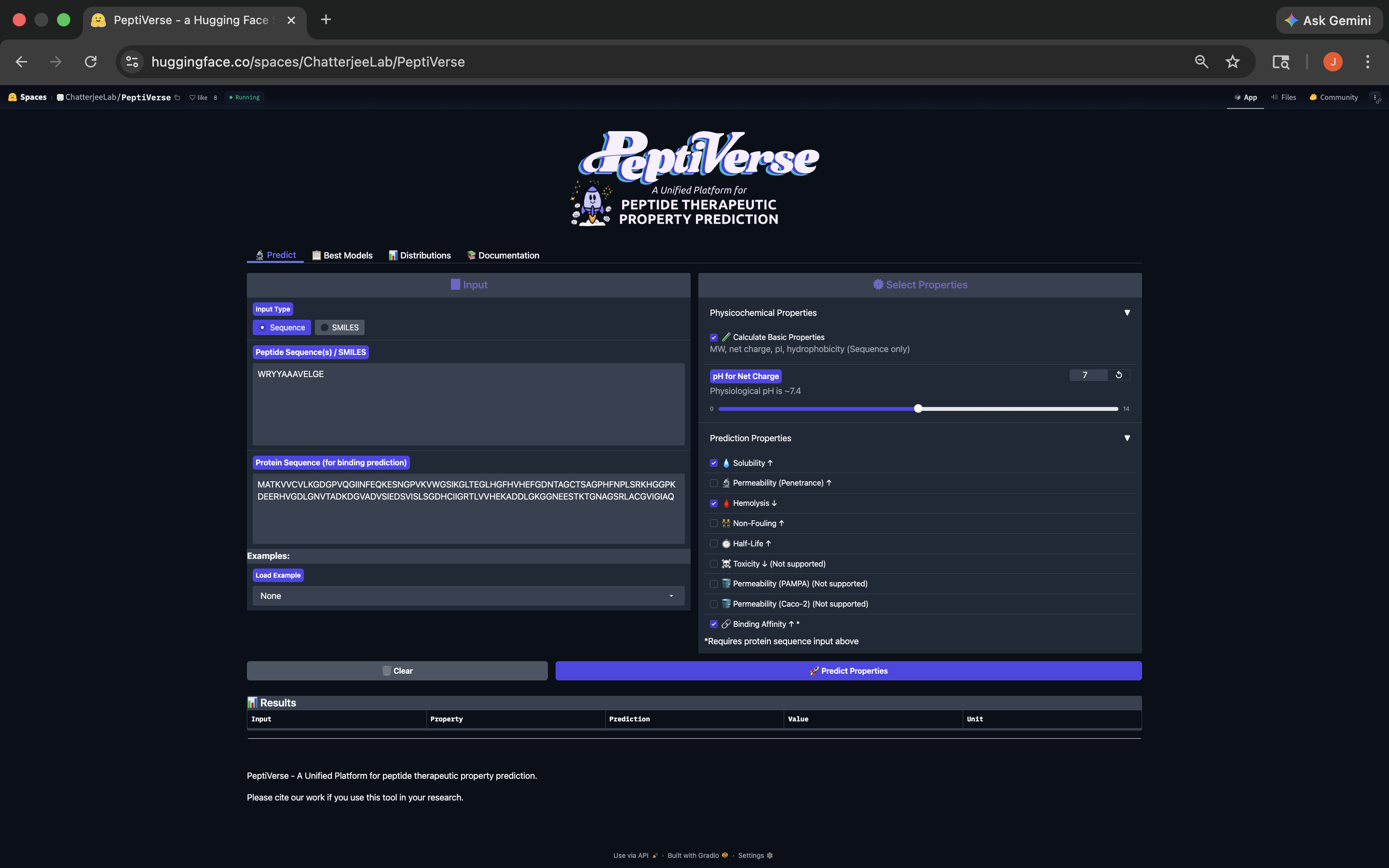
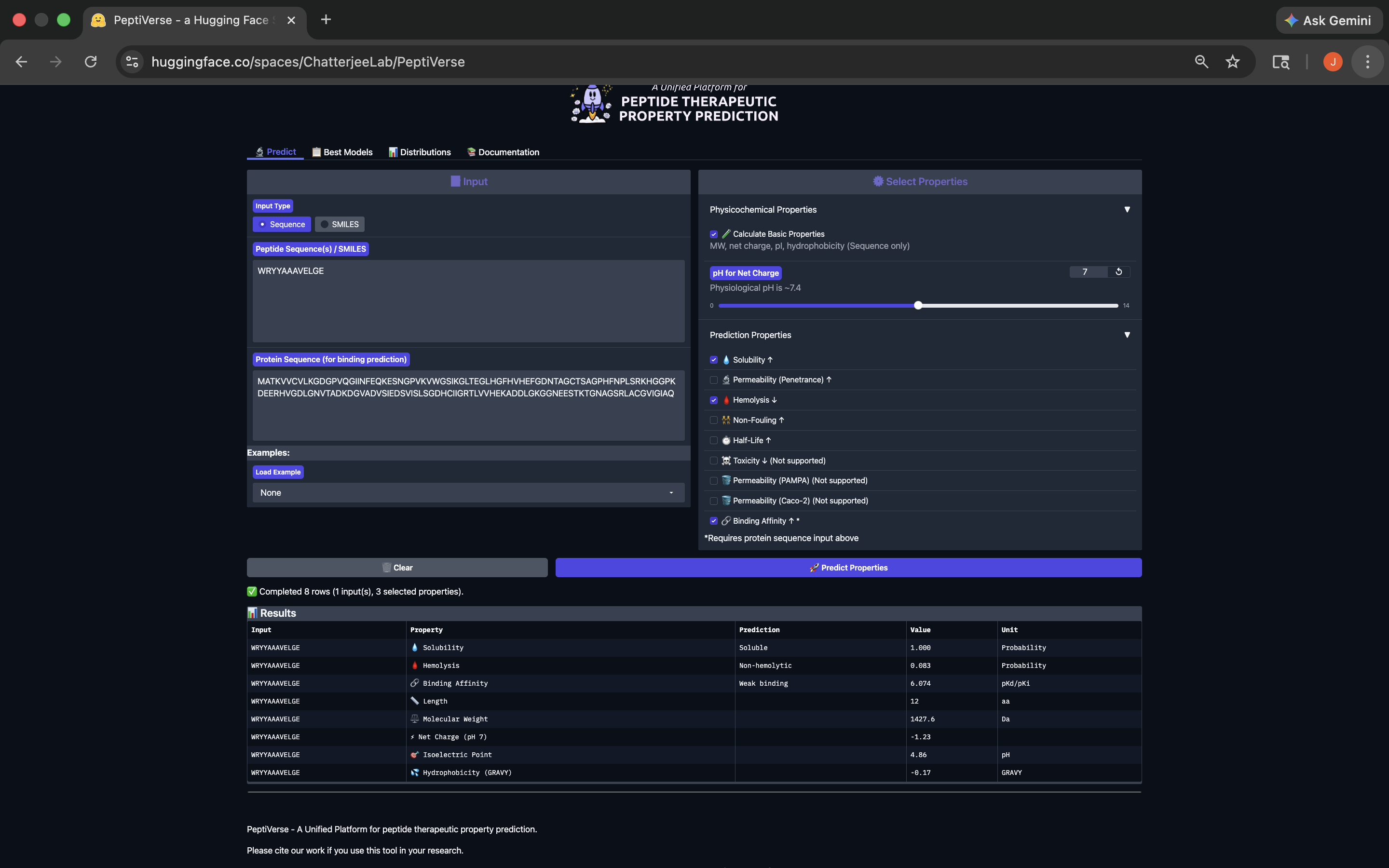
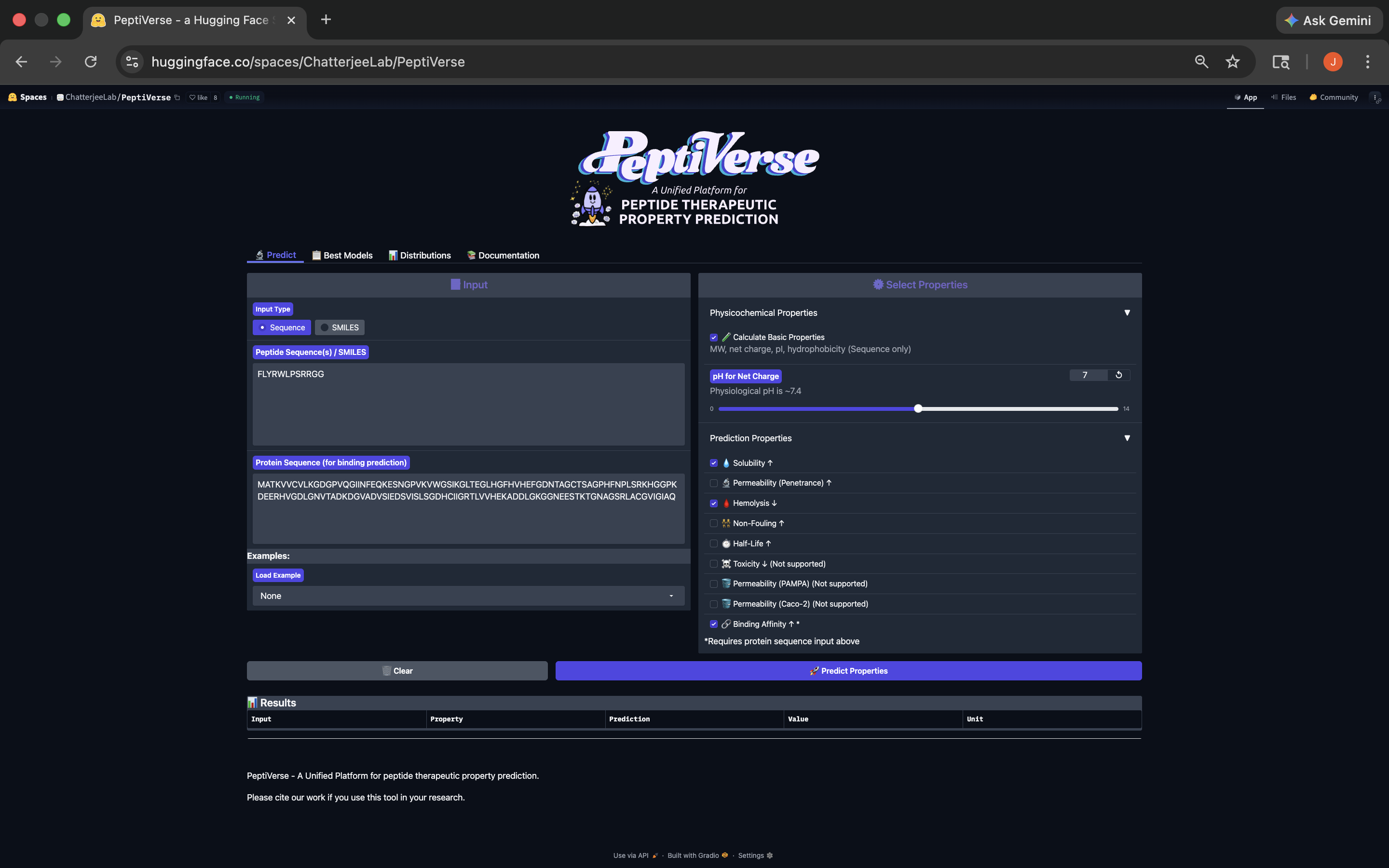
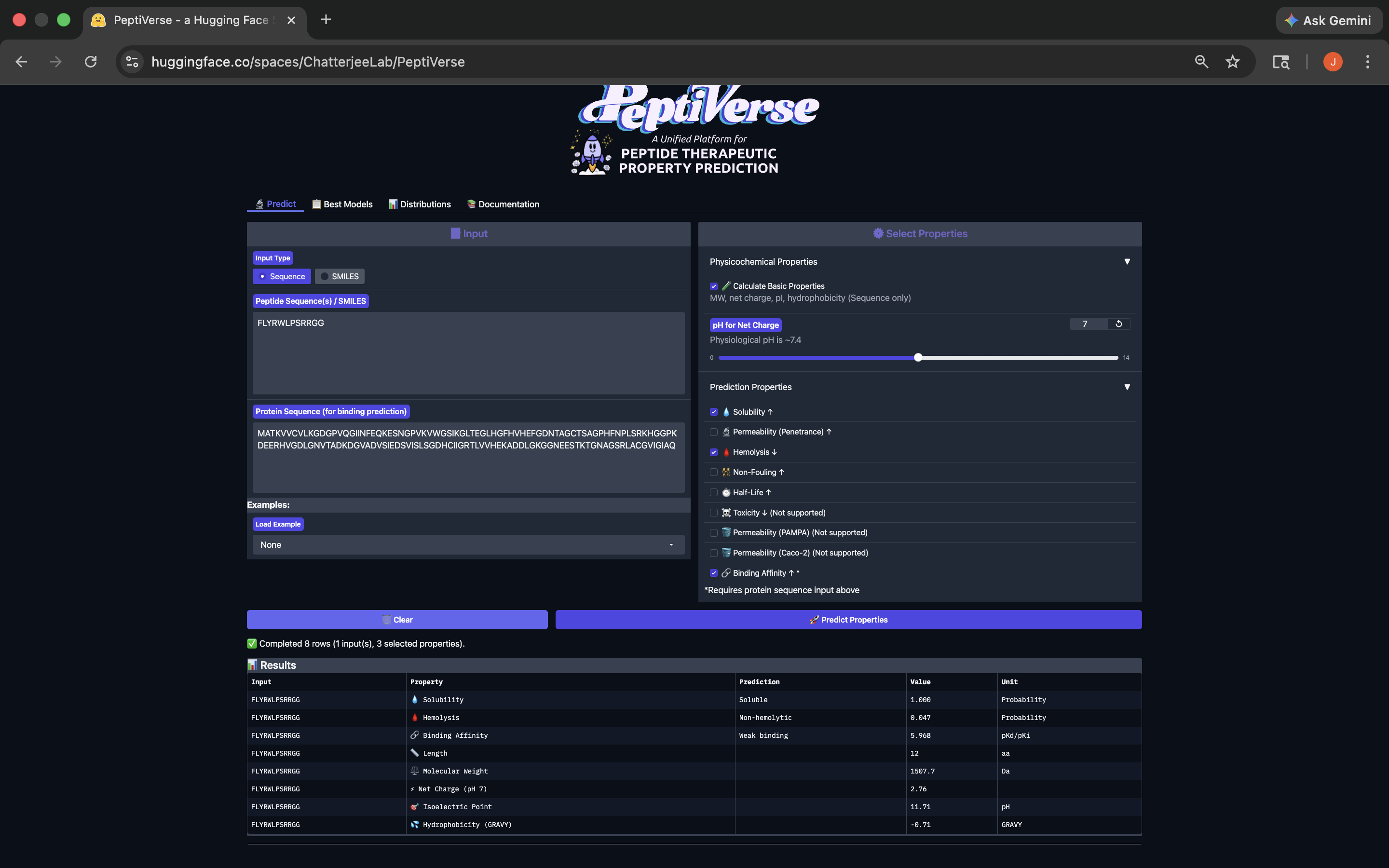
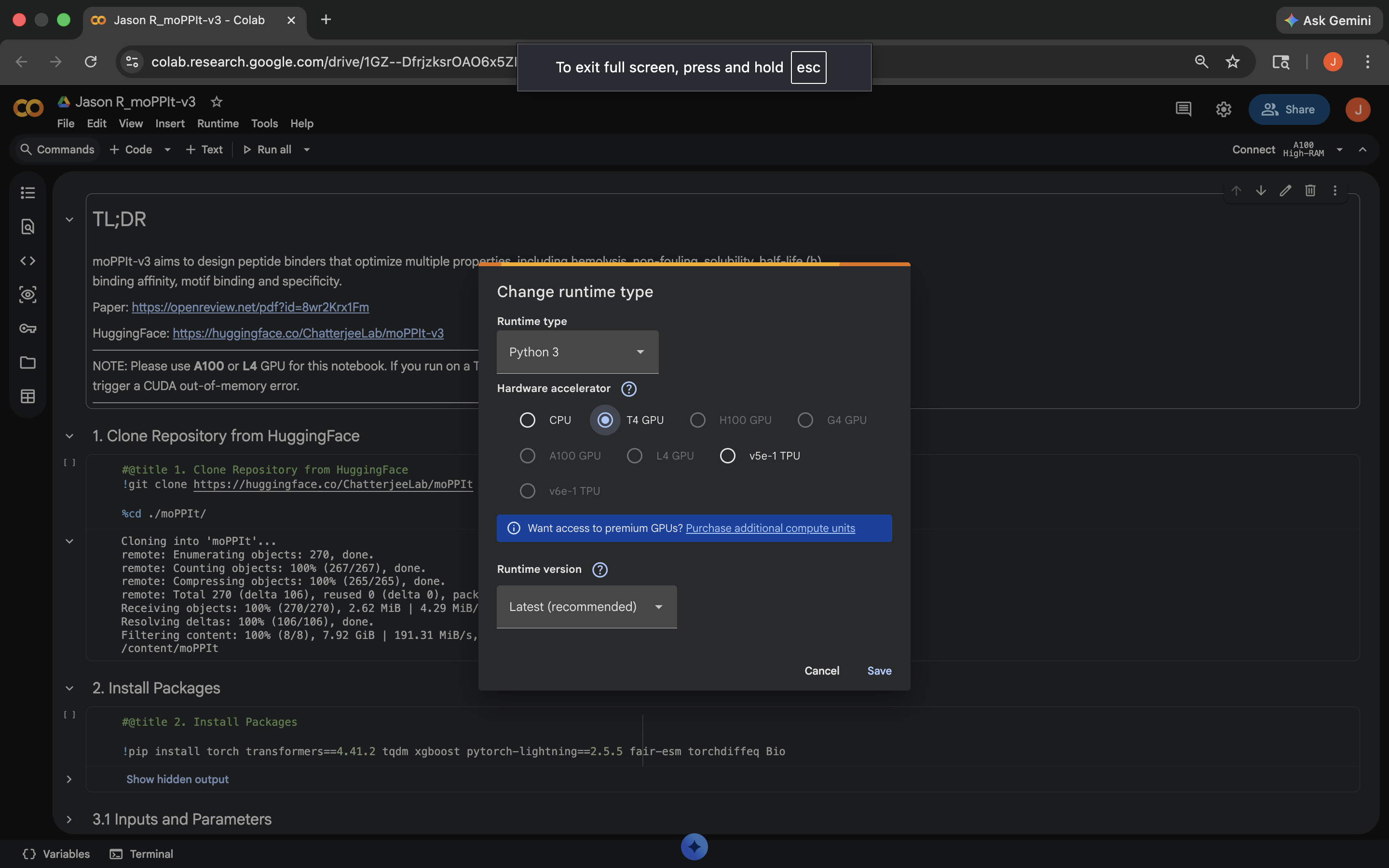
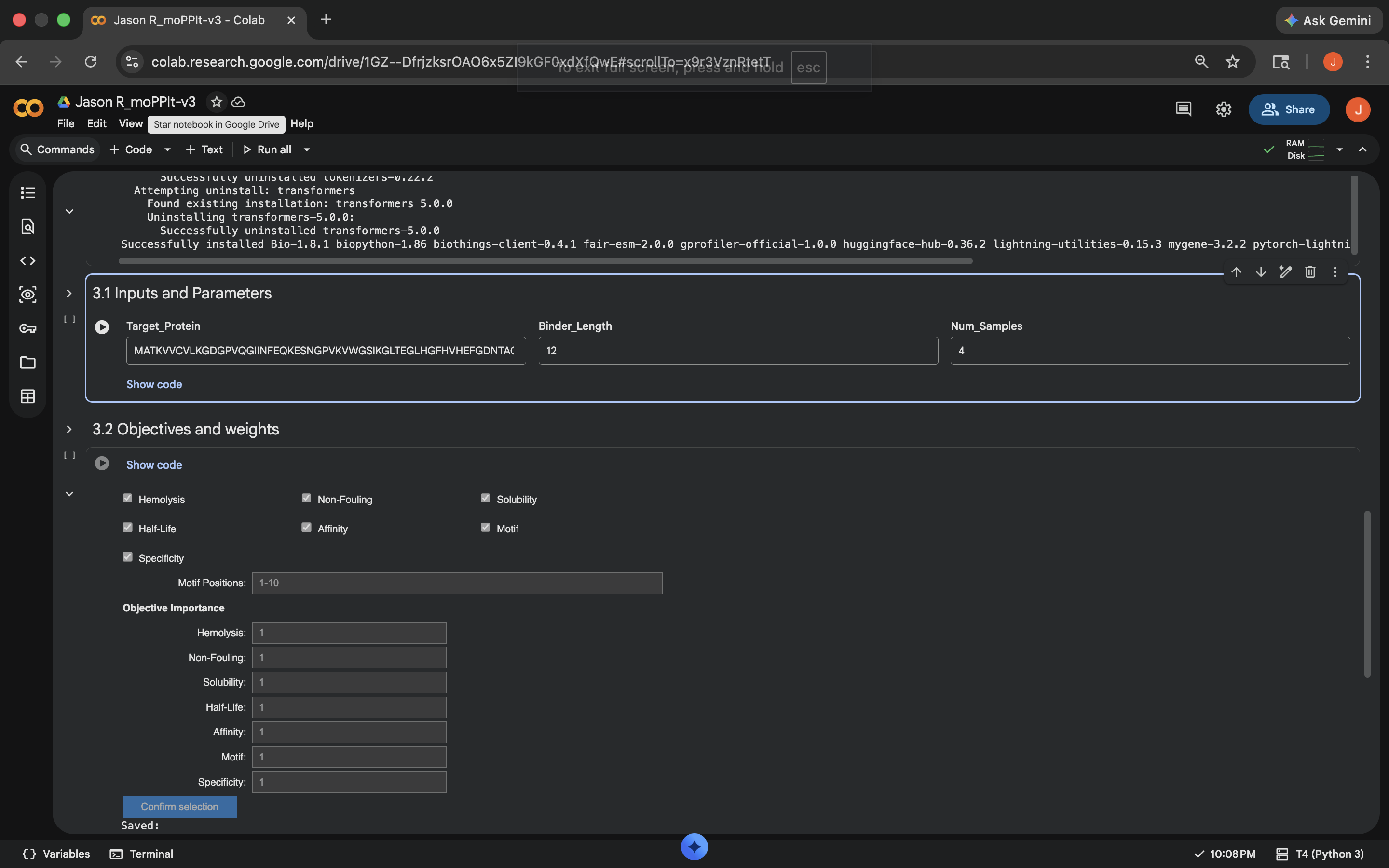
- Week 4 HW: Protein Design Part 1
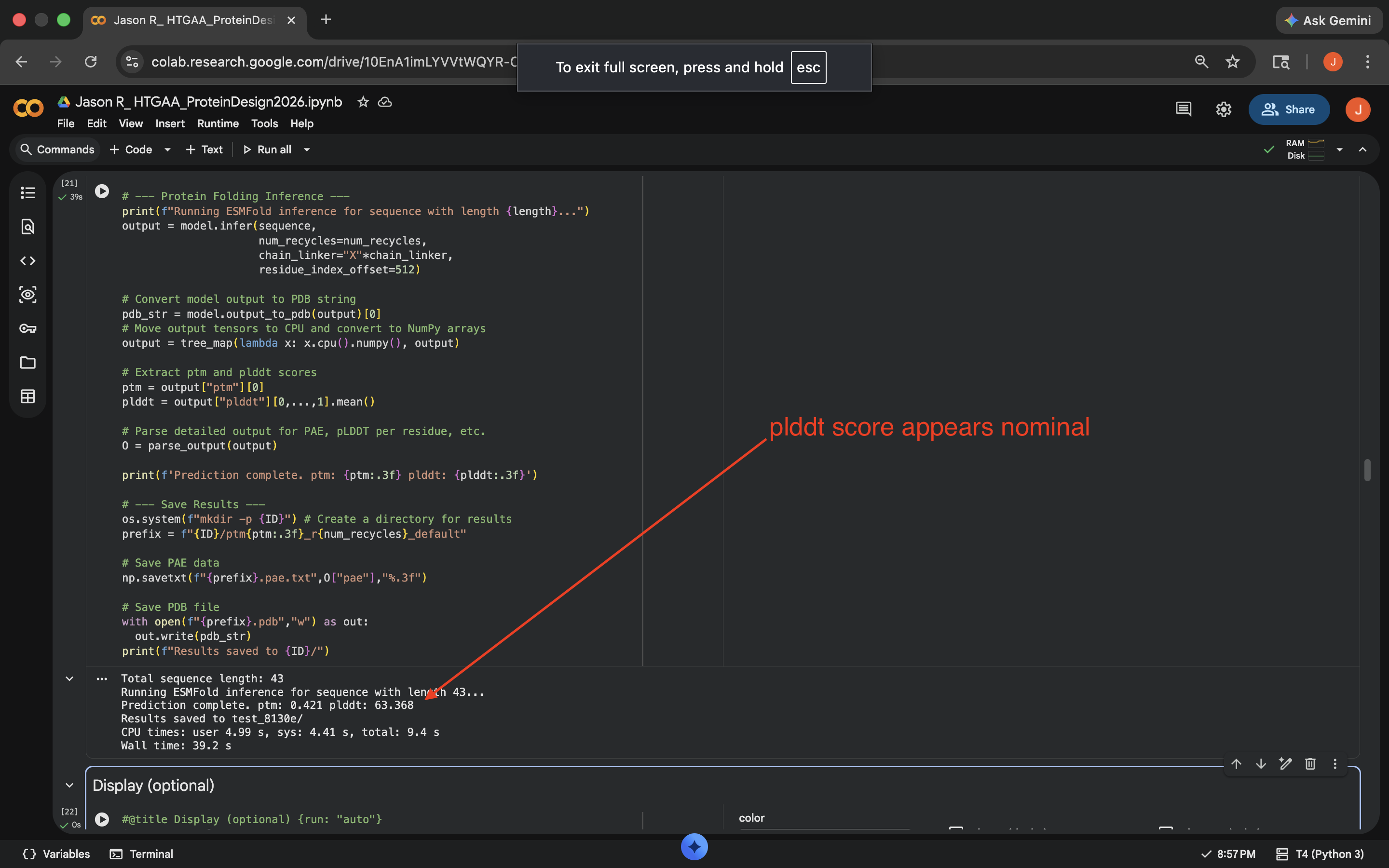
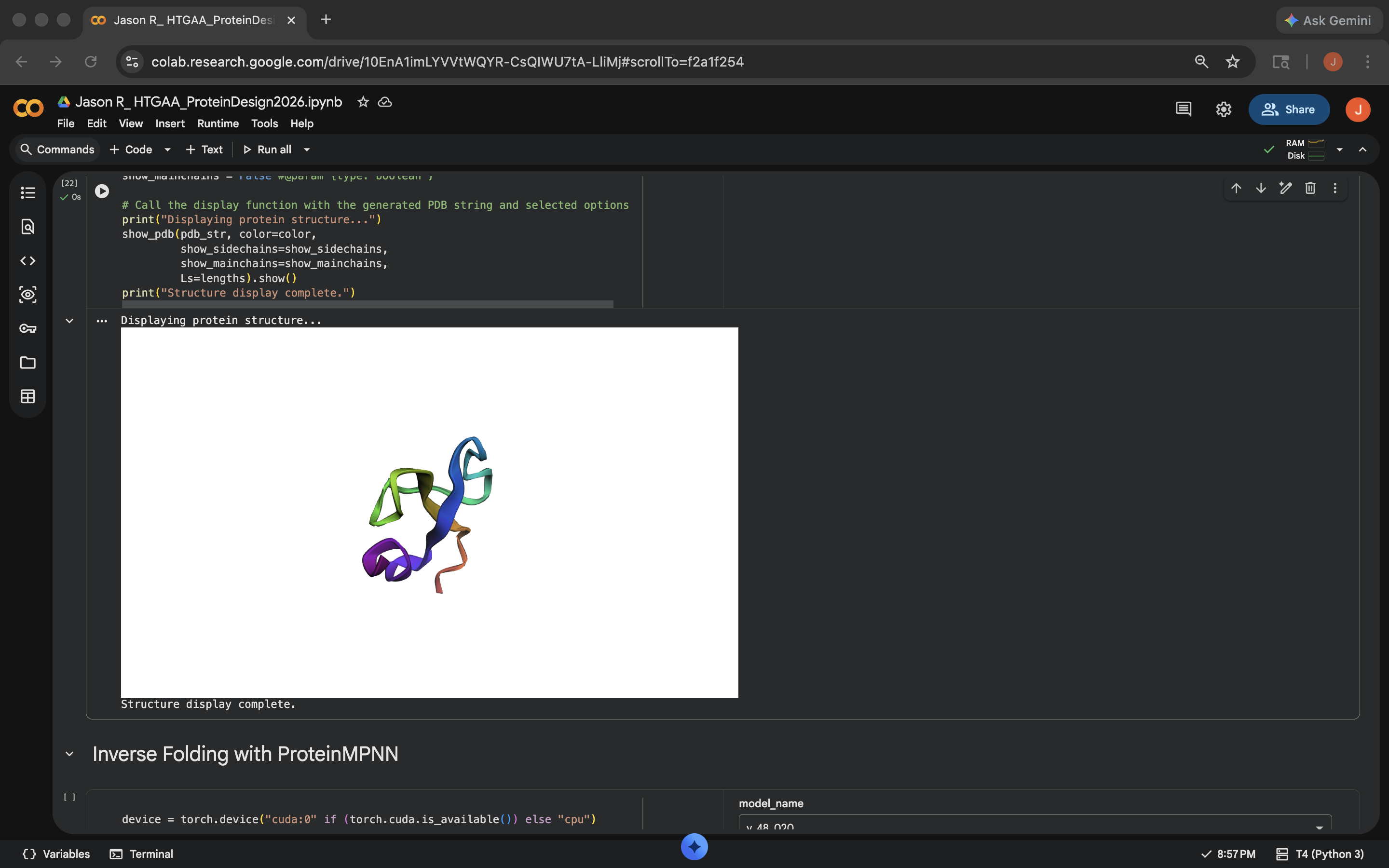
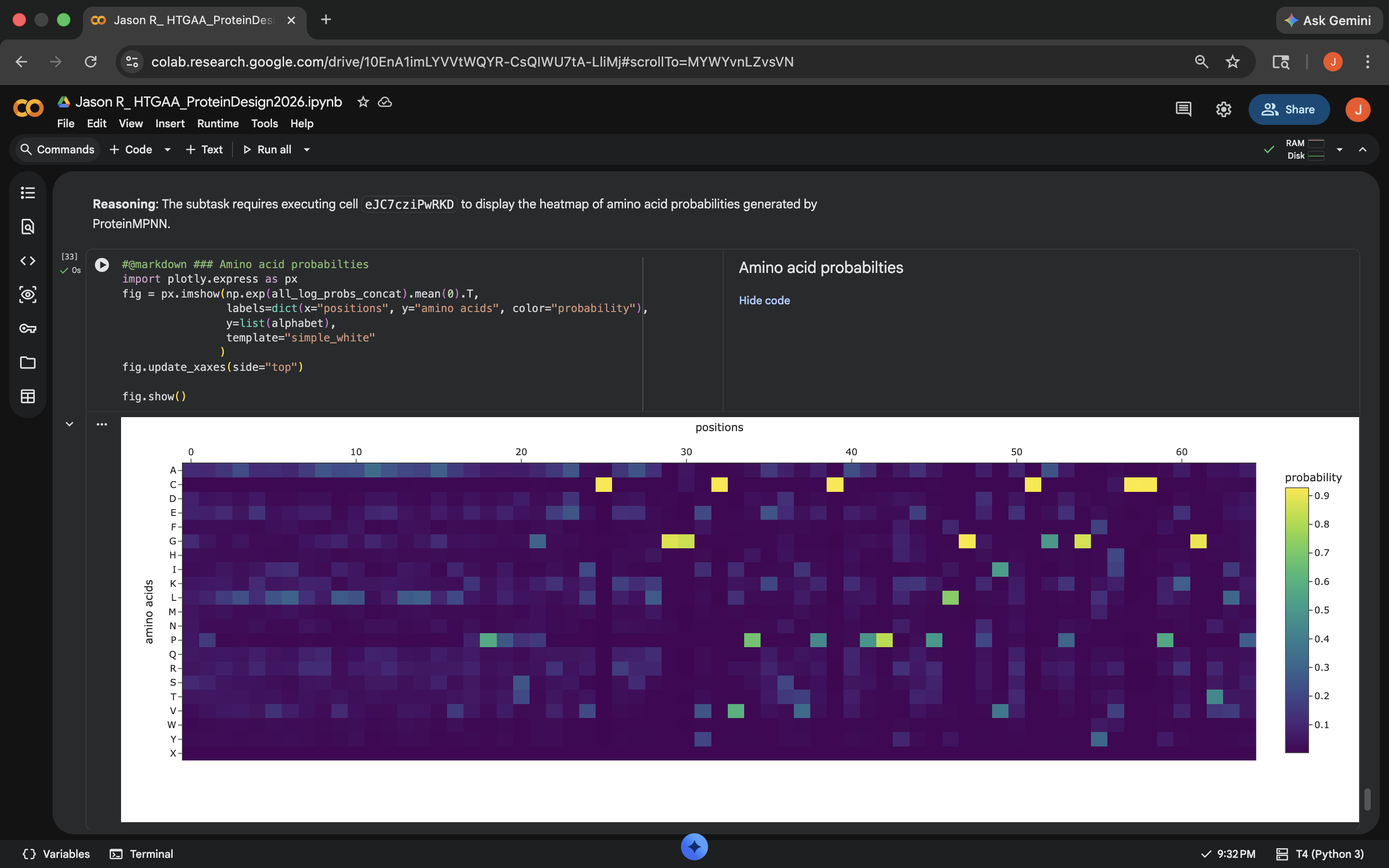
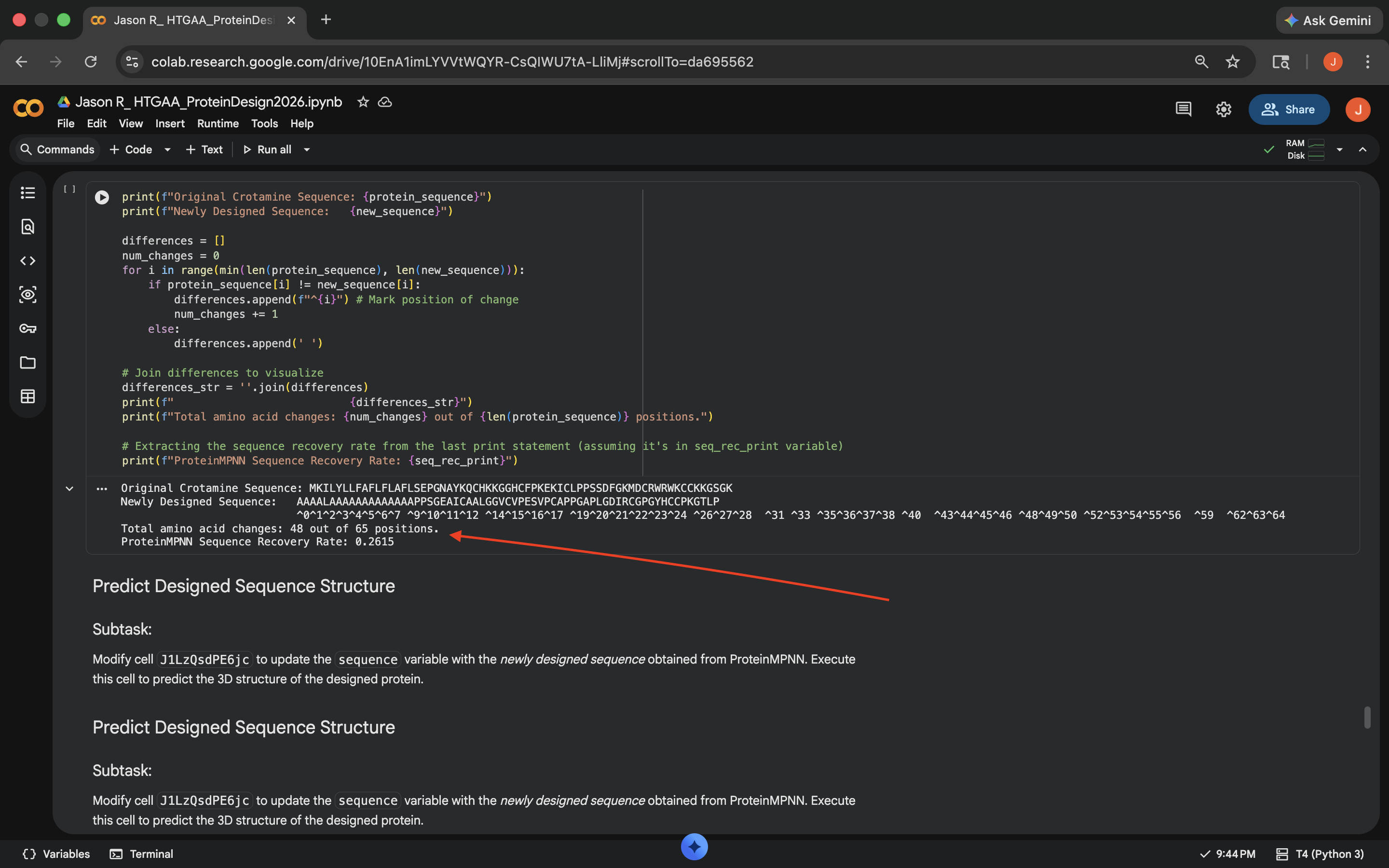
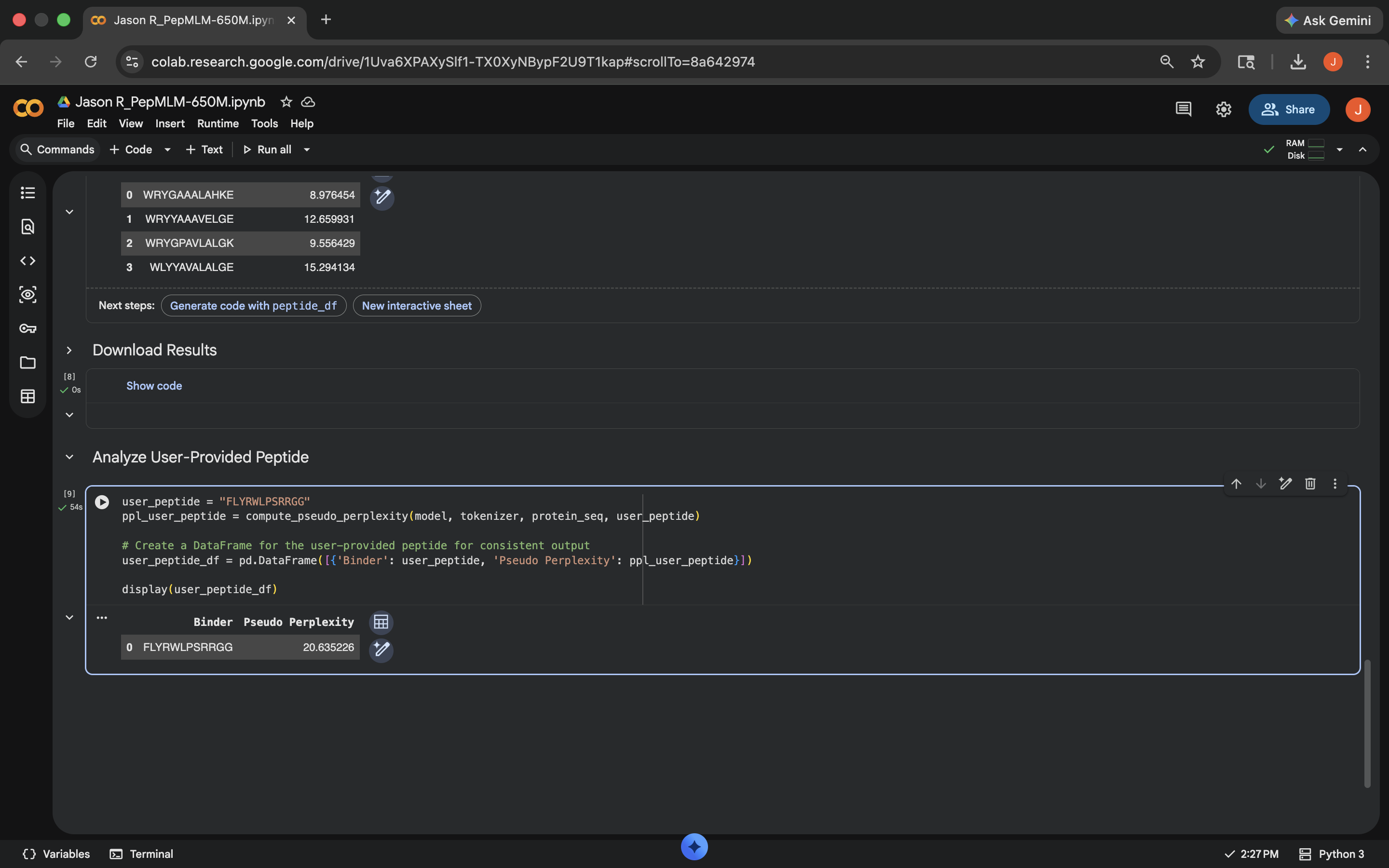
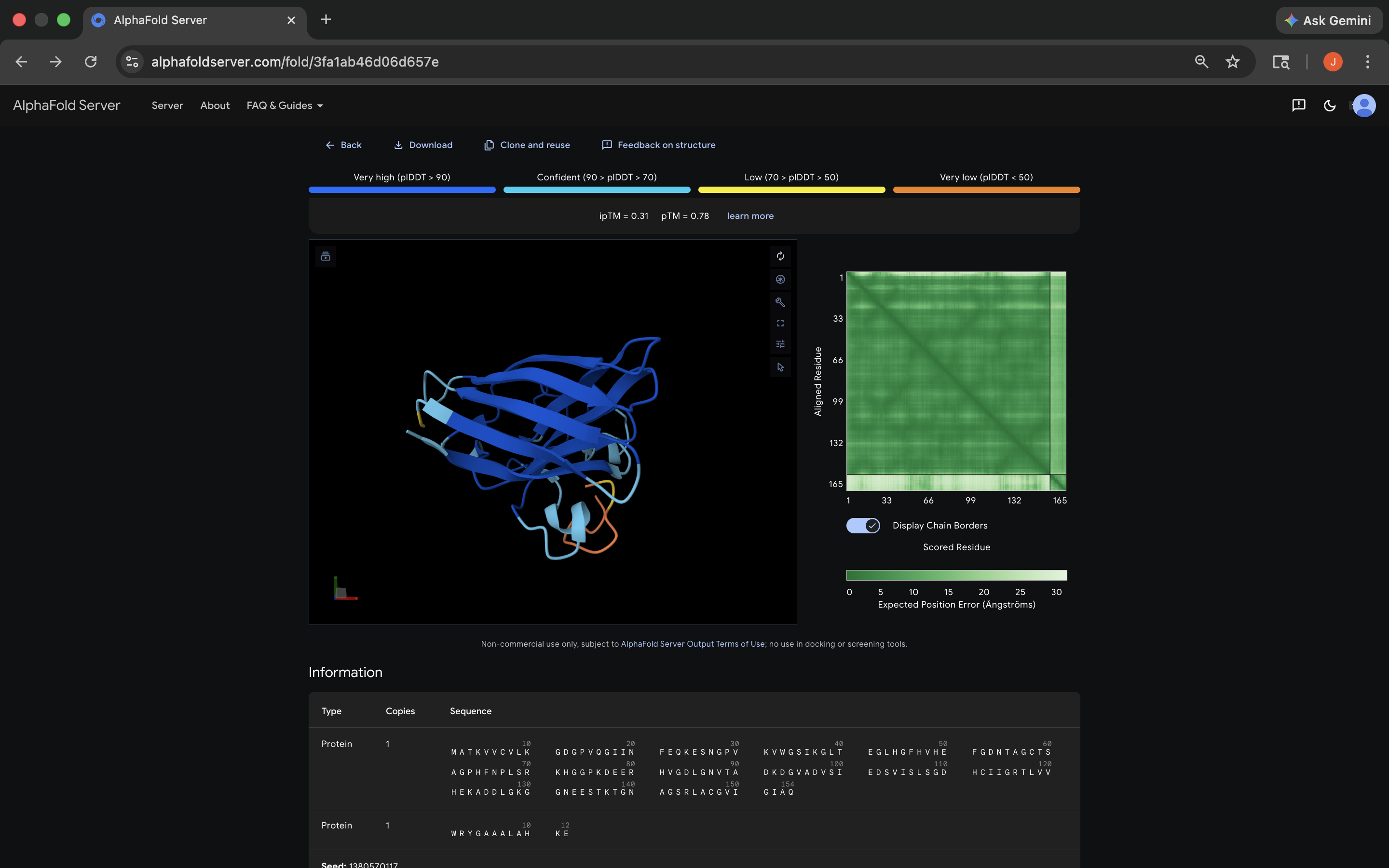
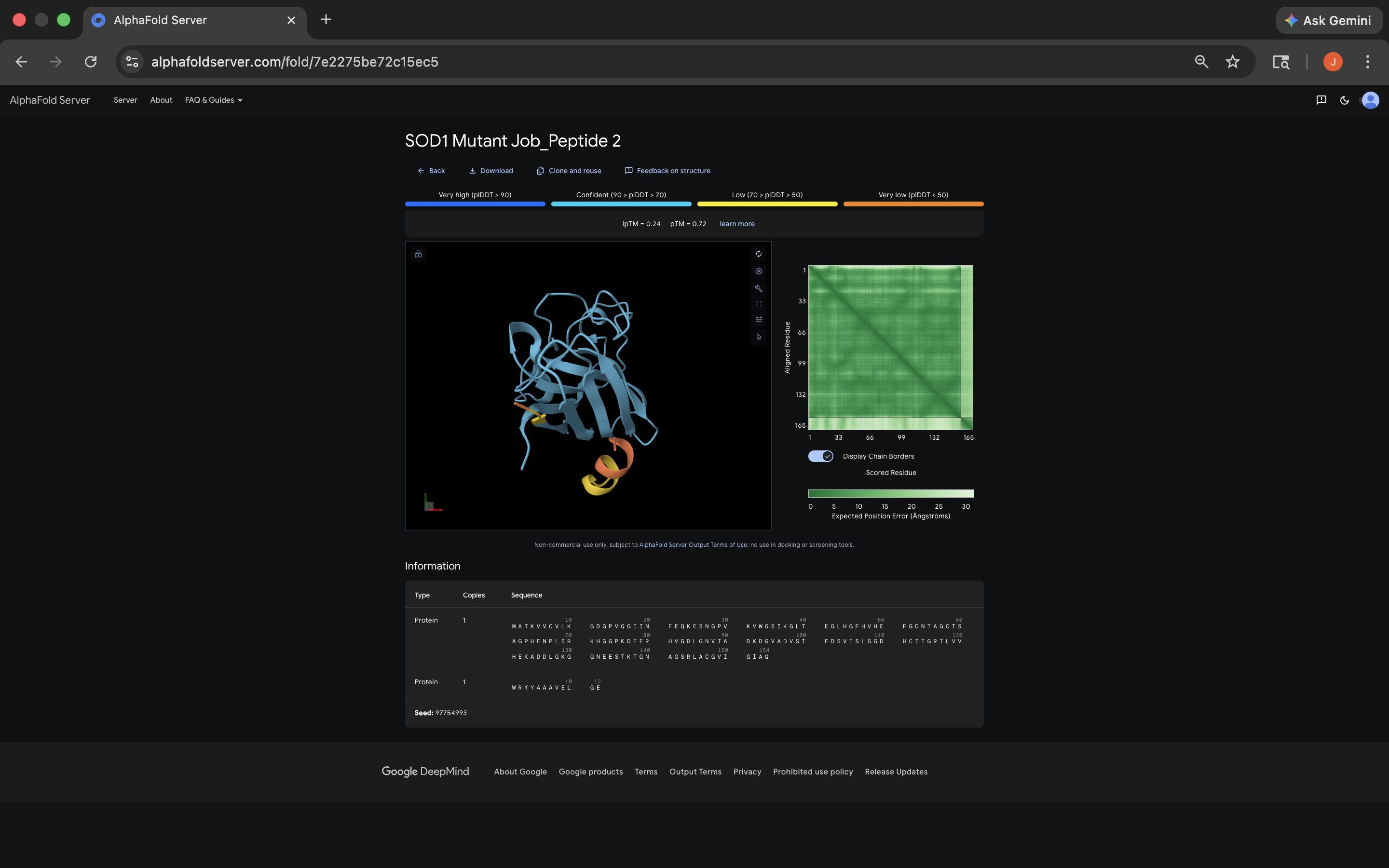
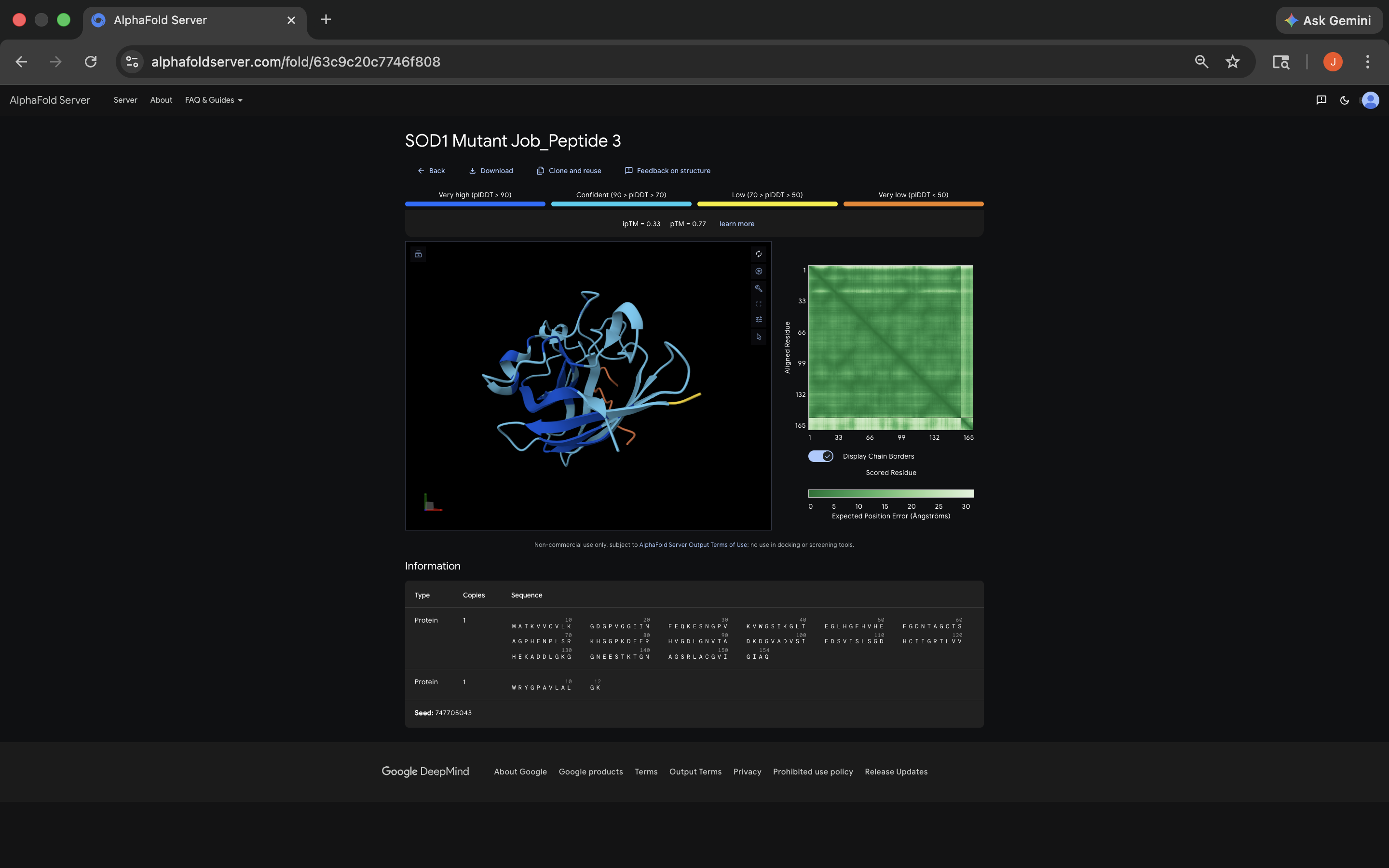
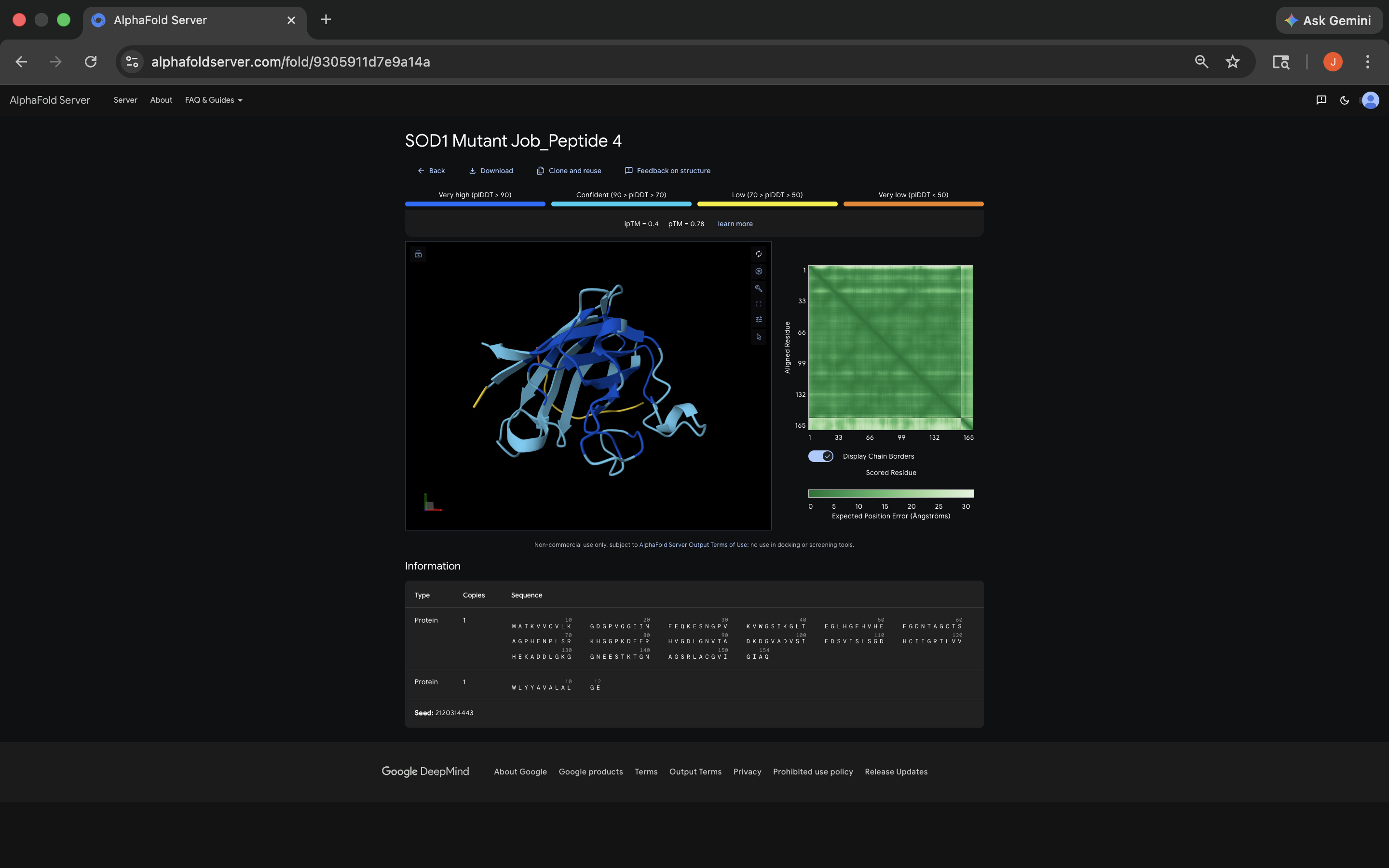
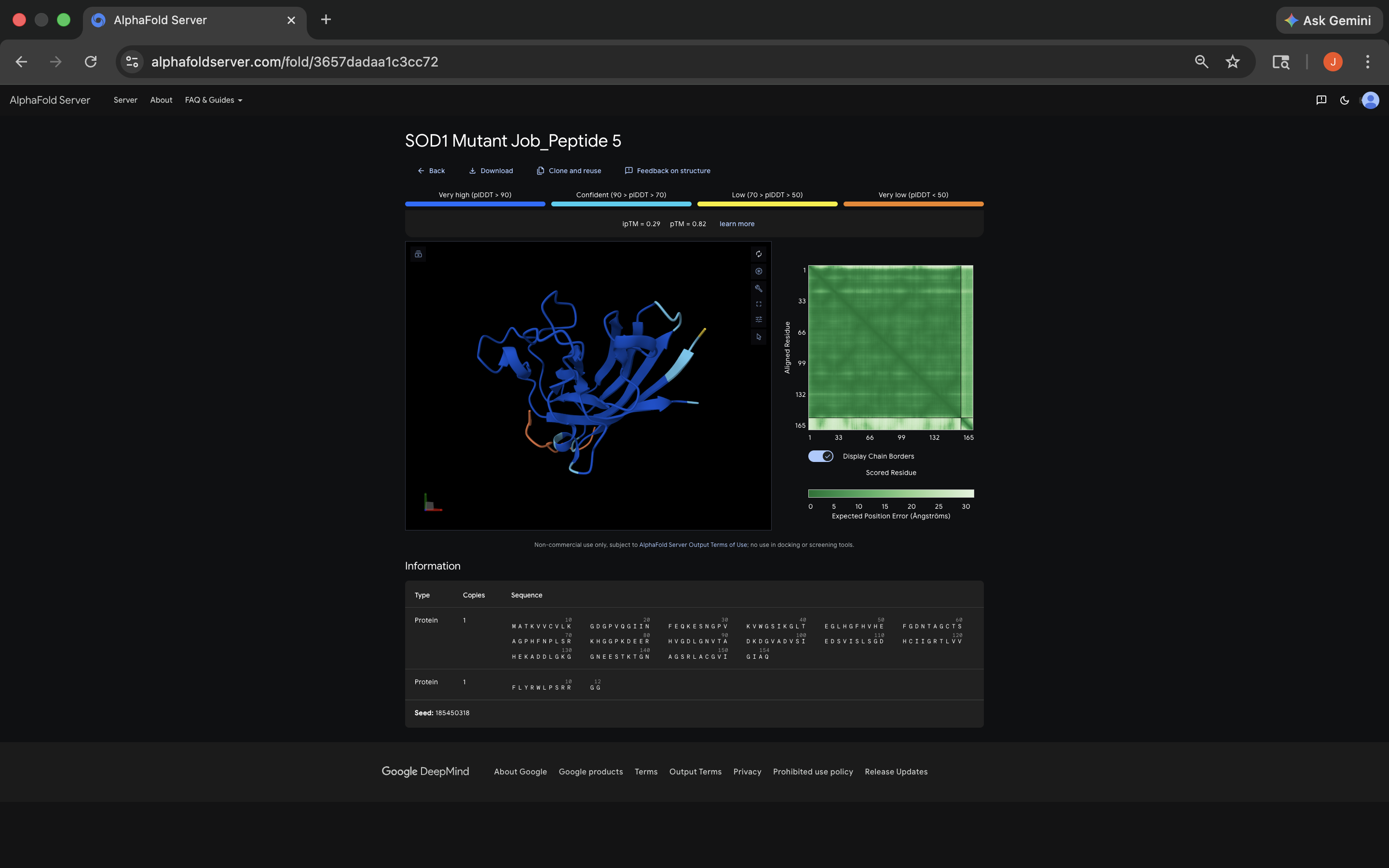
- Week 5 HW: Protein Design Part 2
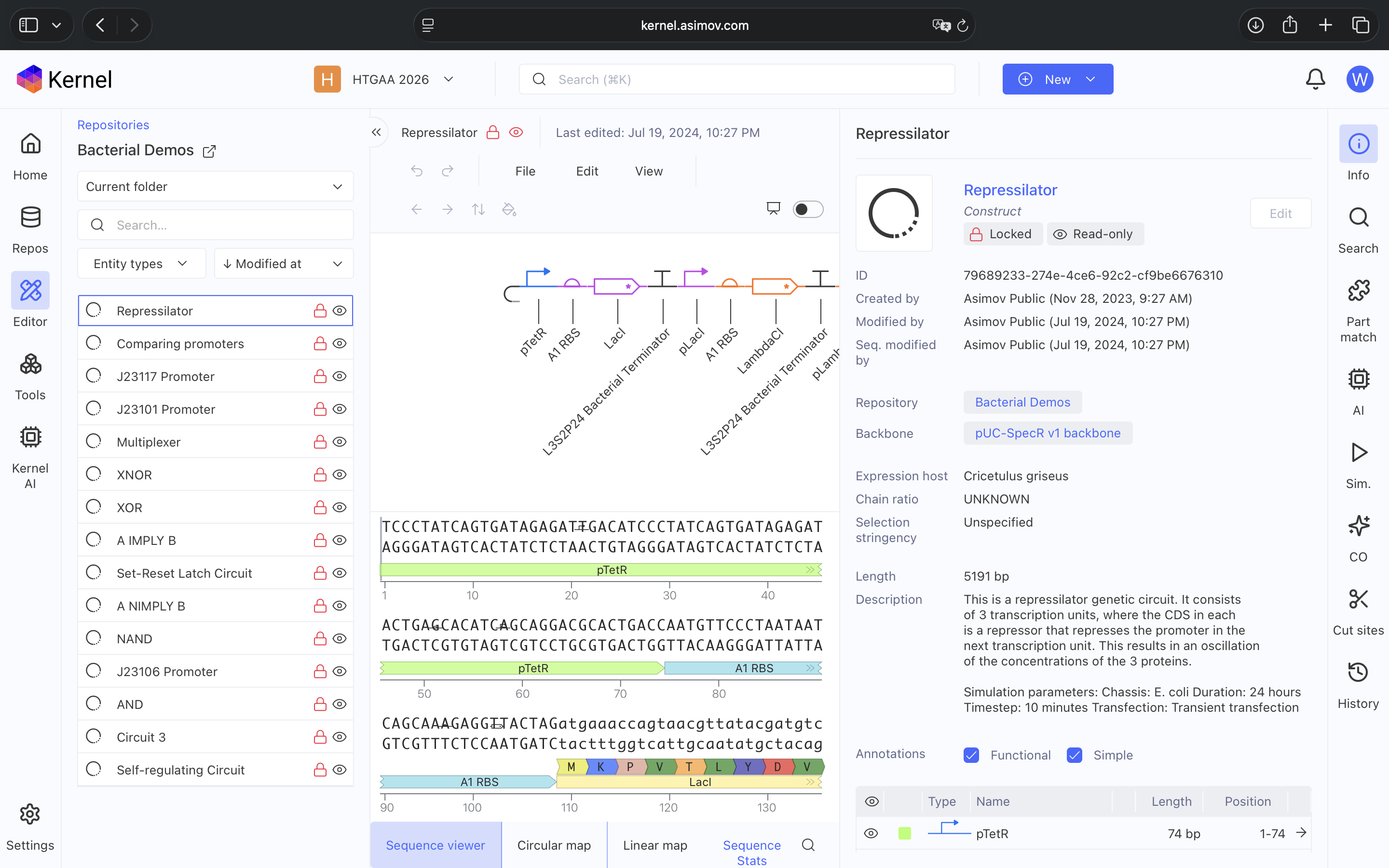
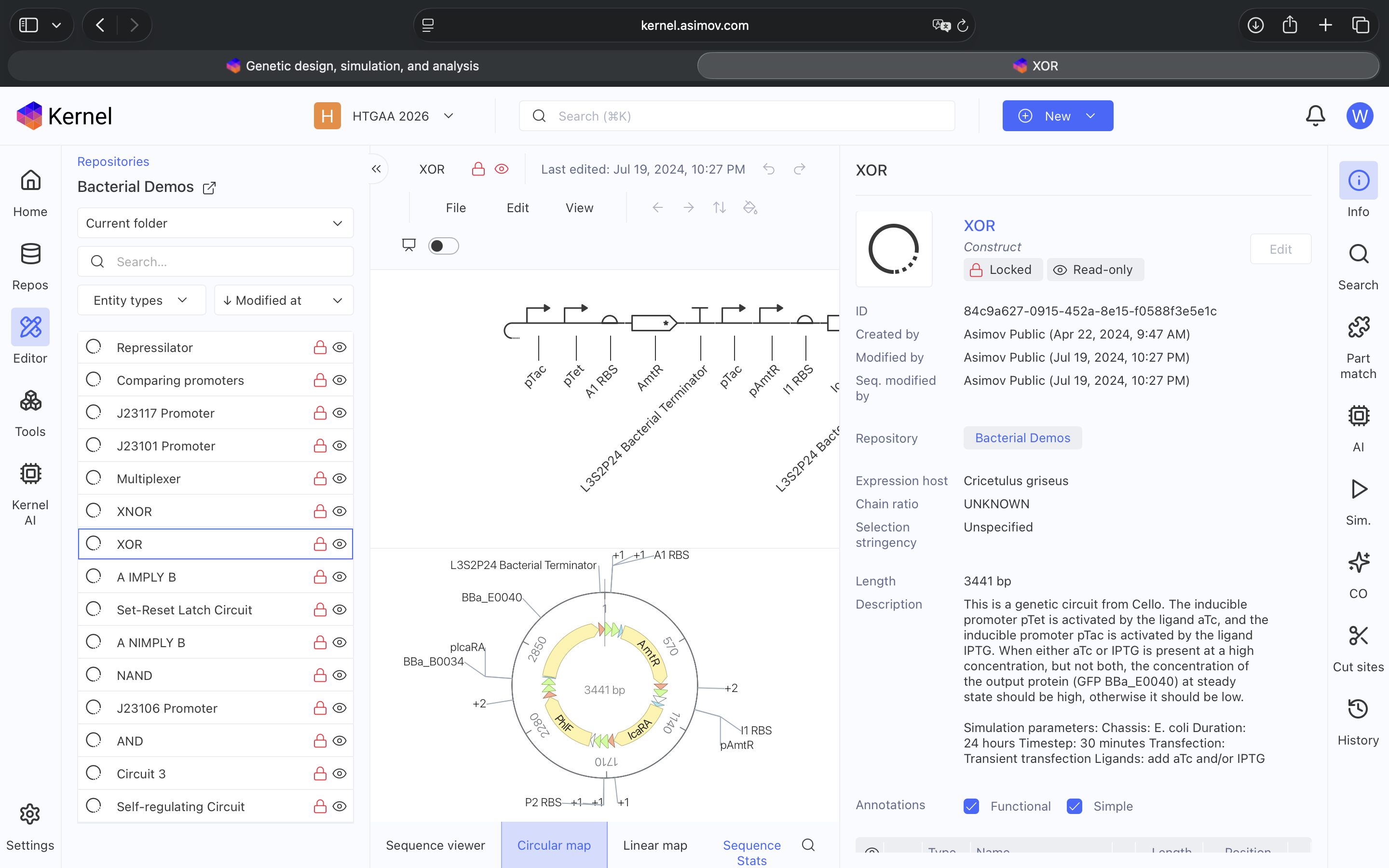
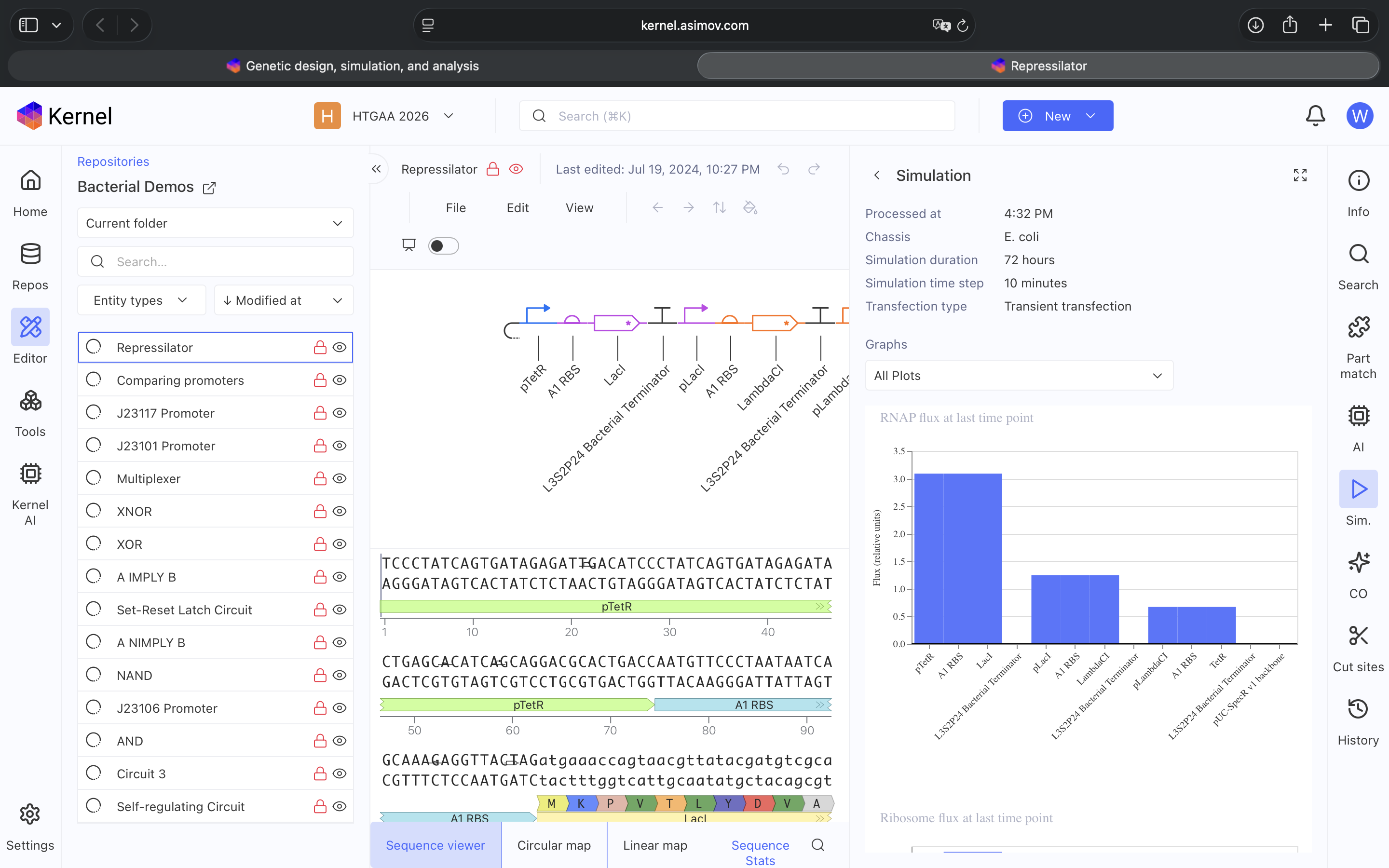
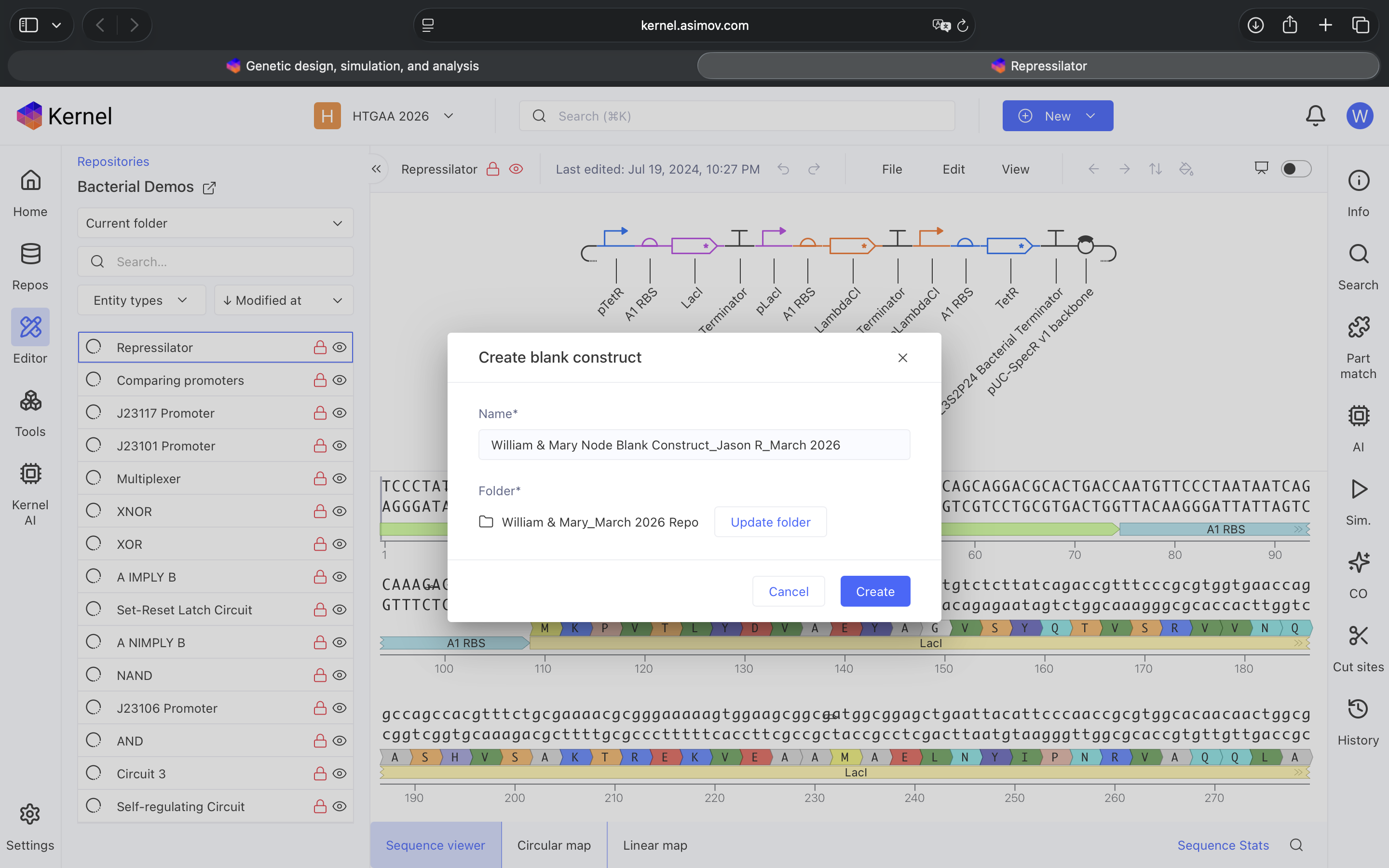
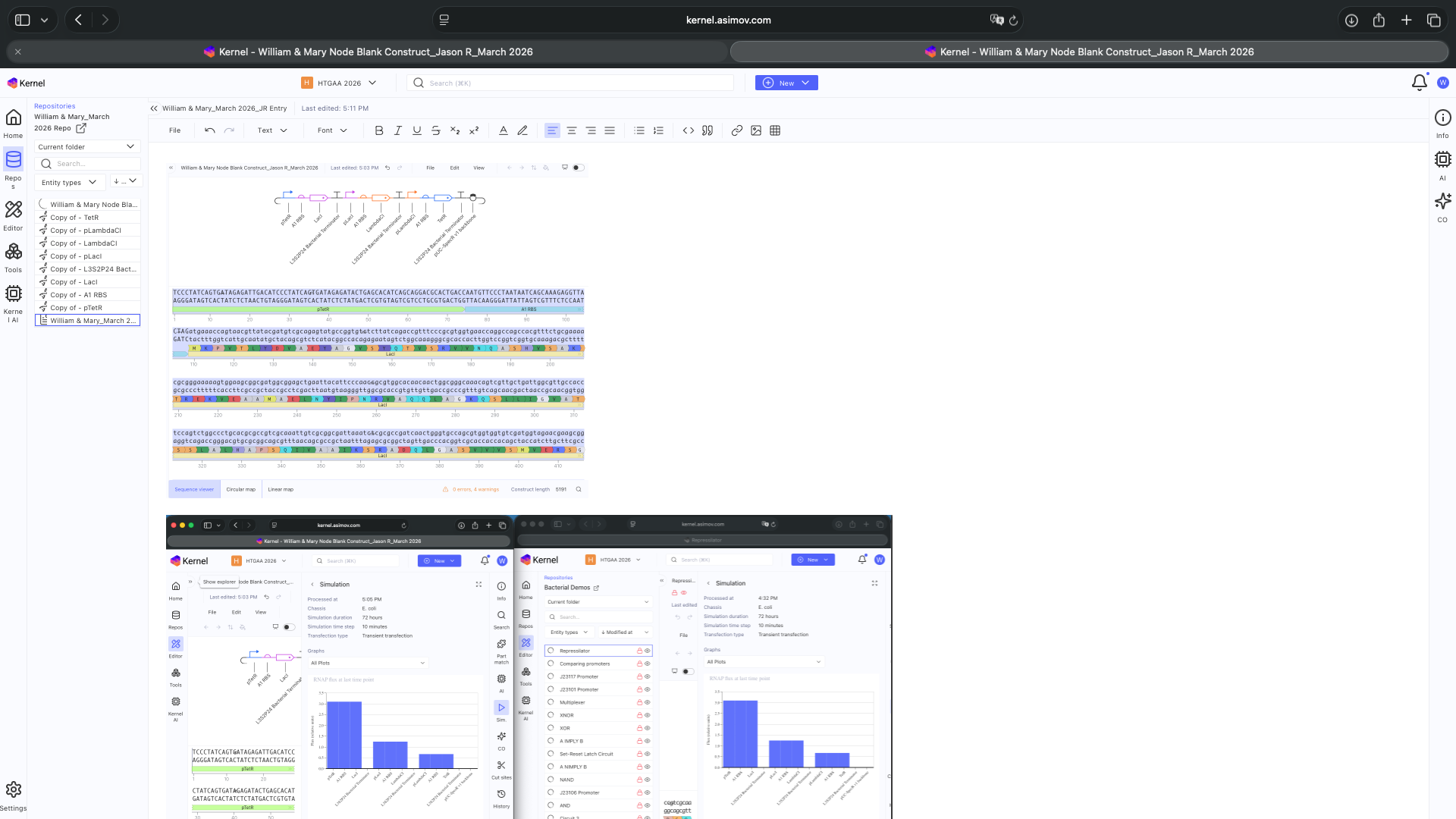
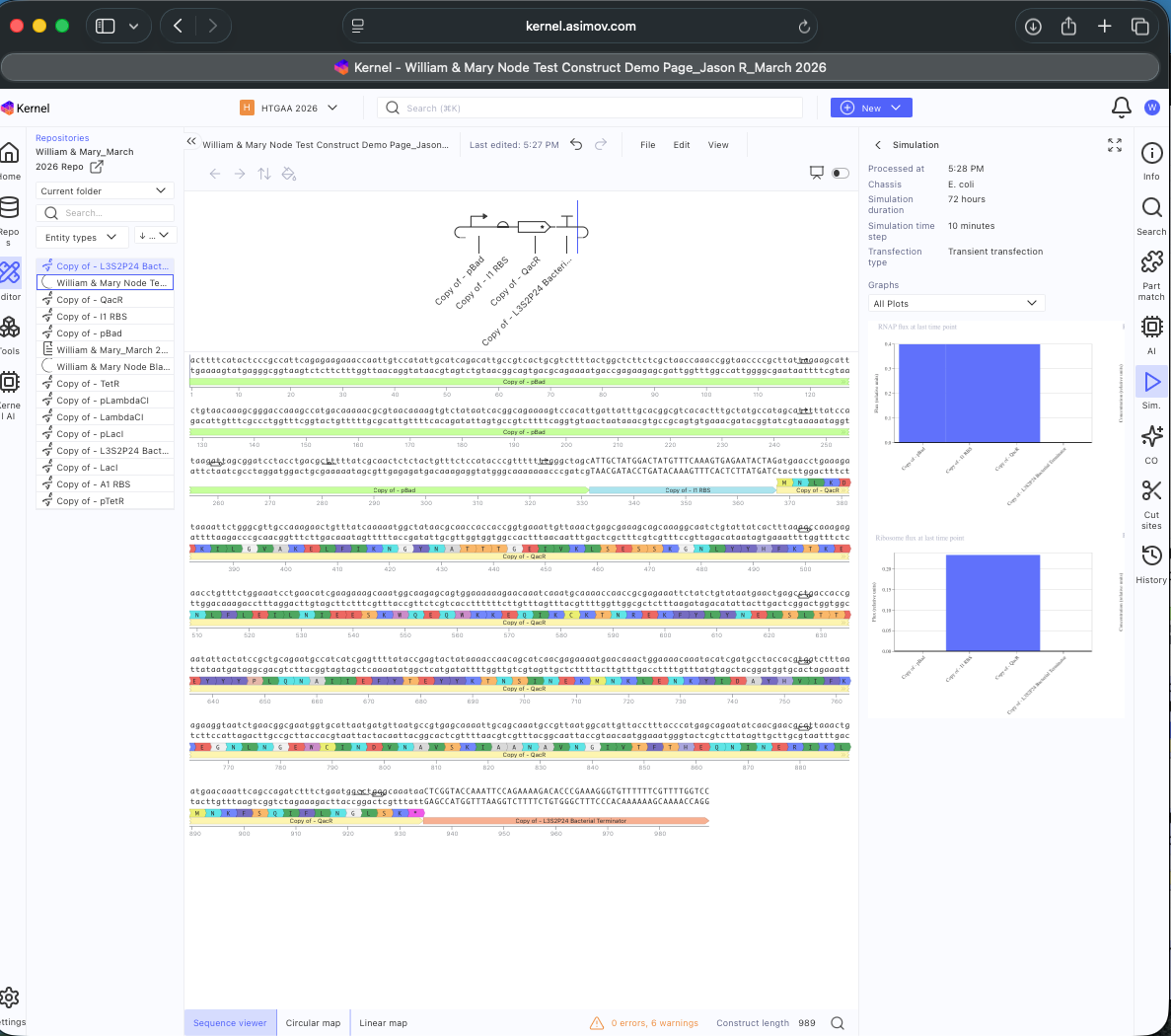
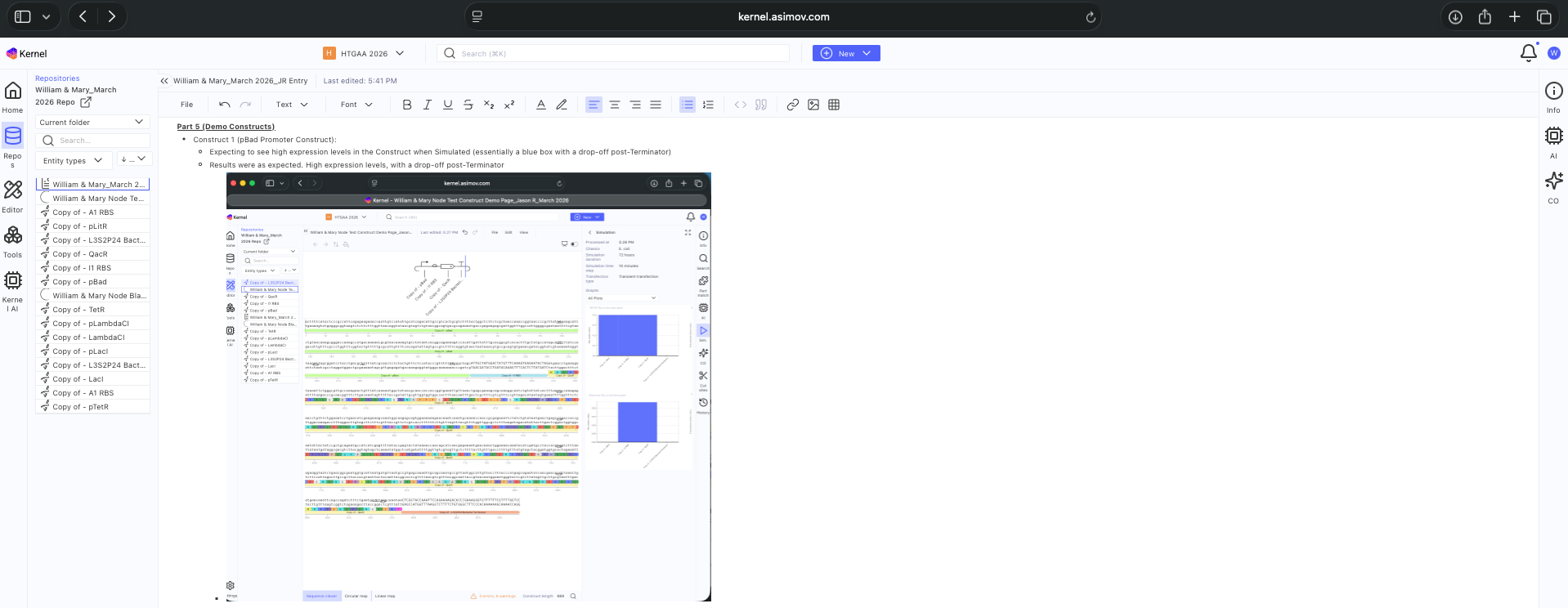
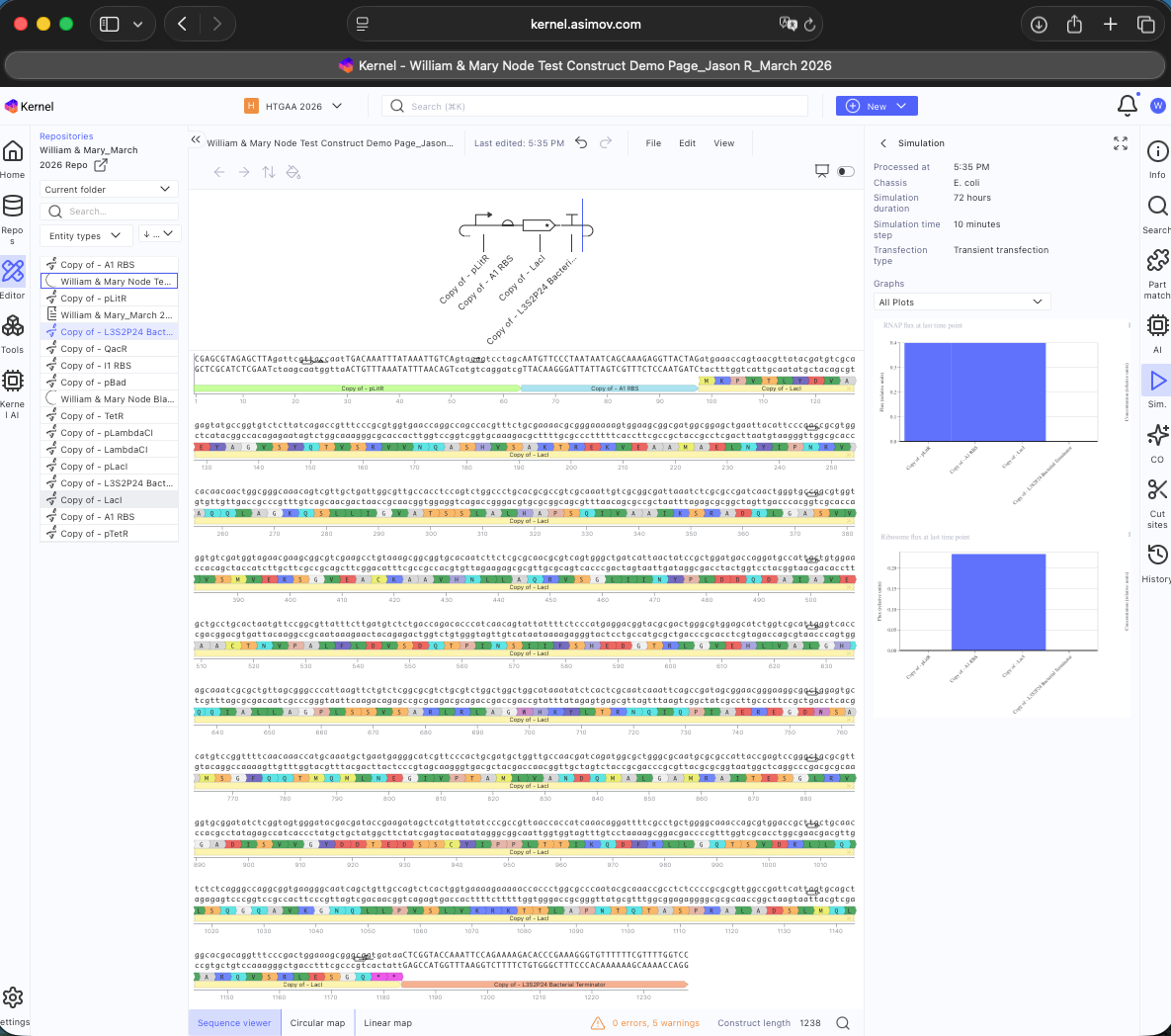
- Week 6 HW: Genetic Circuits Part 1
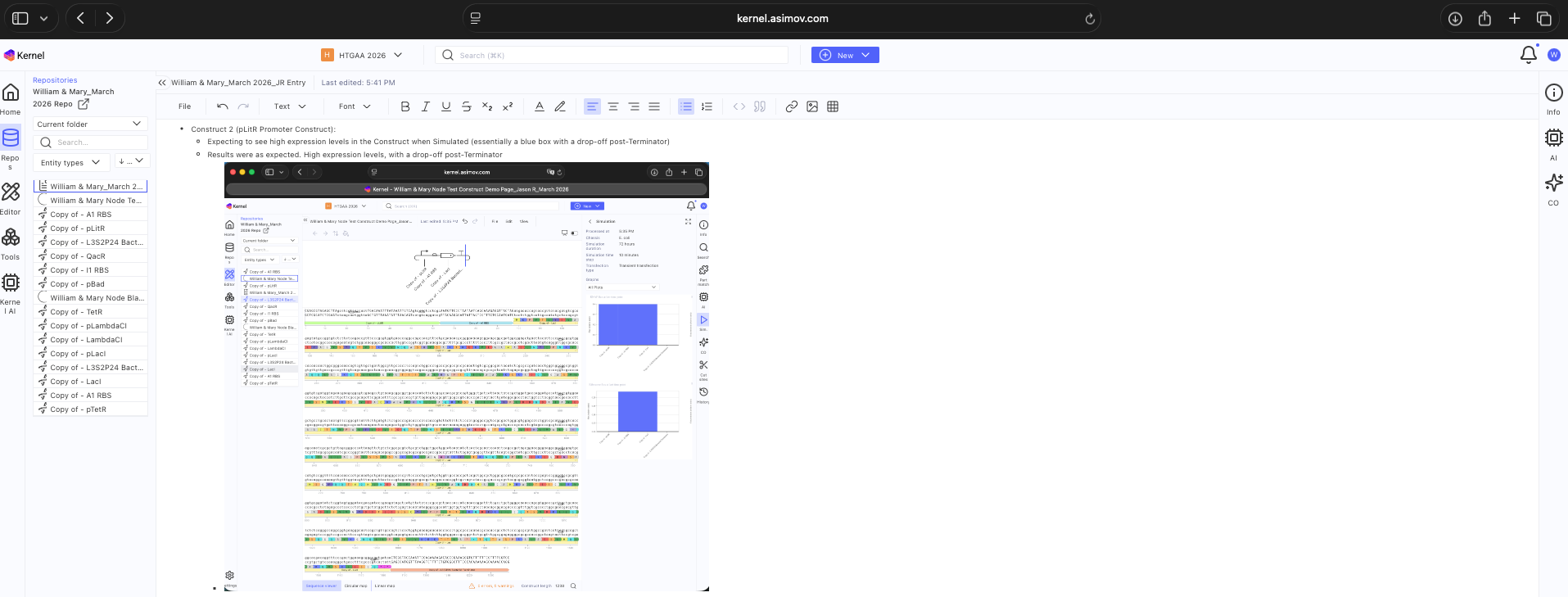
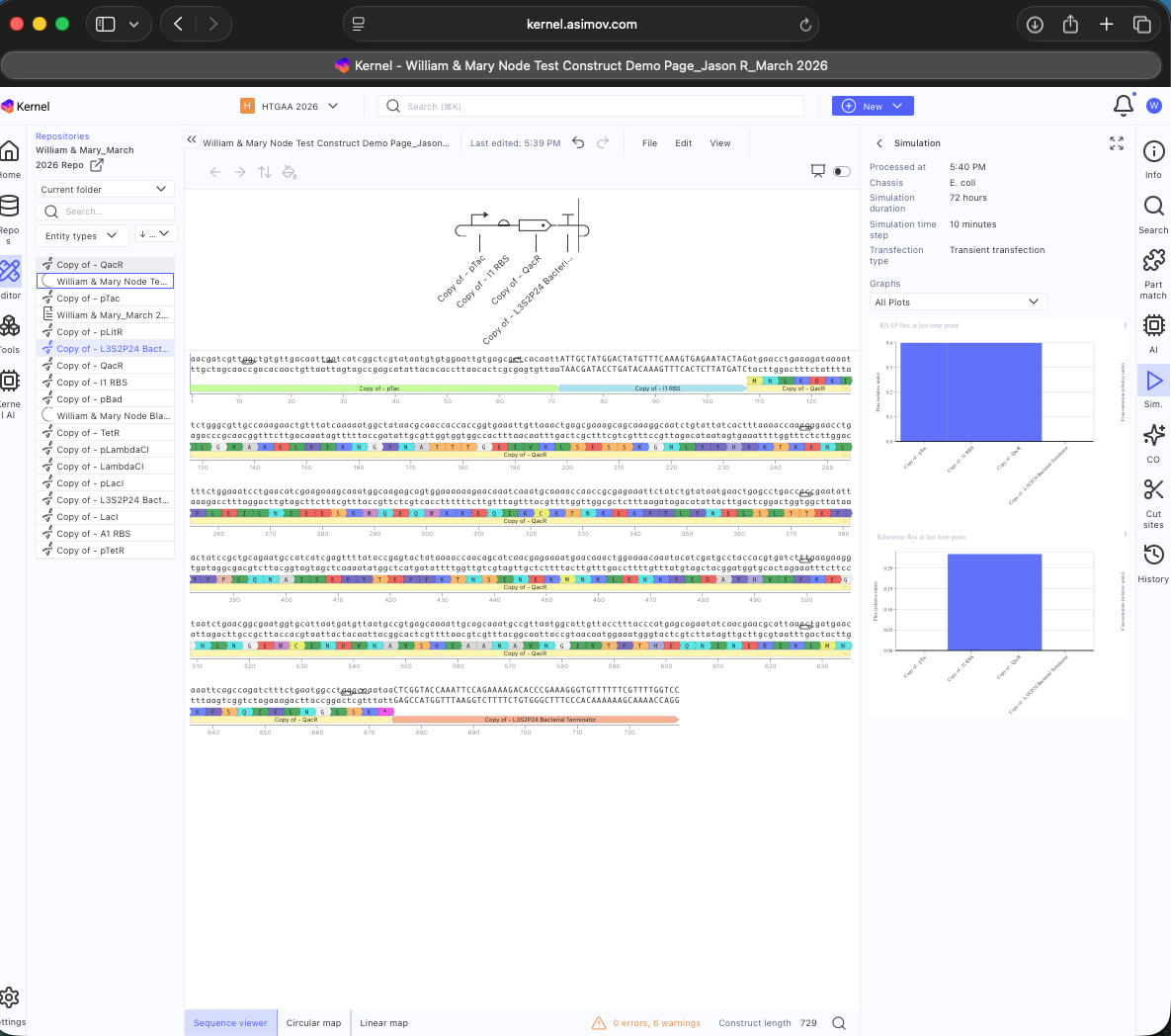
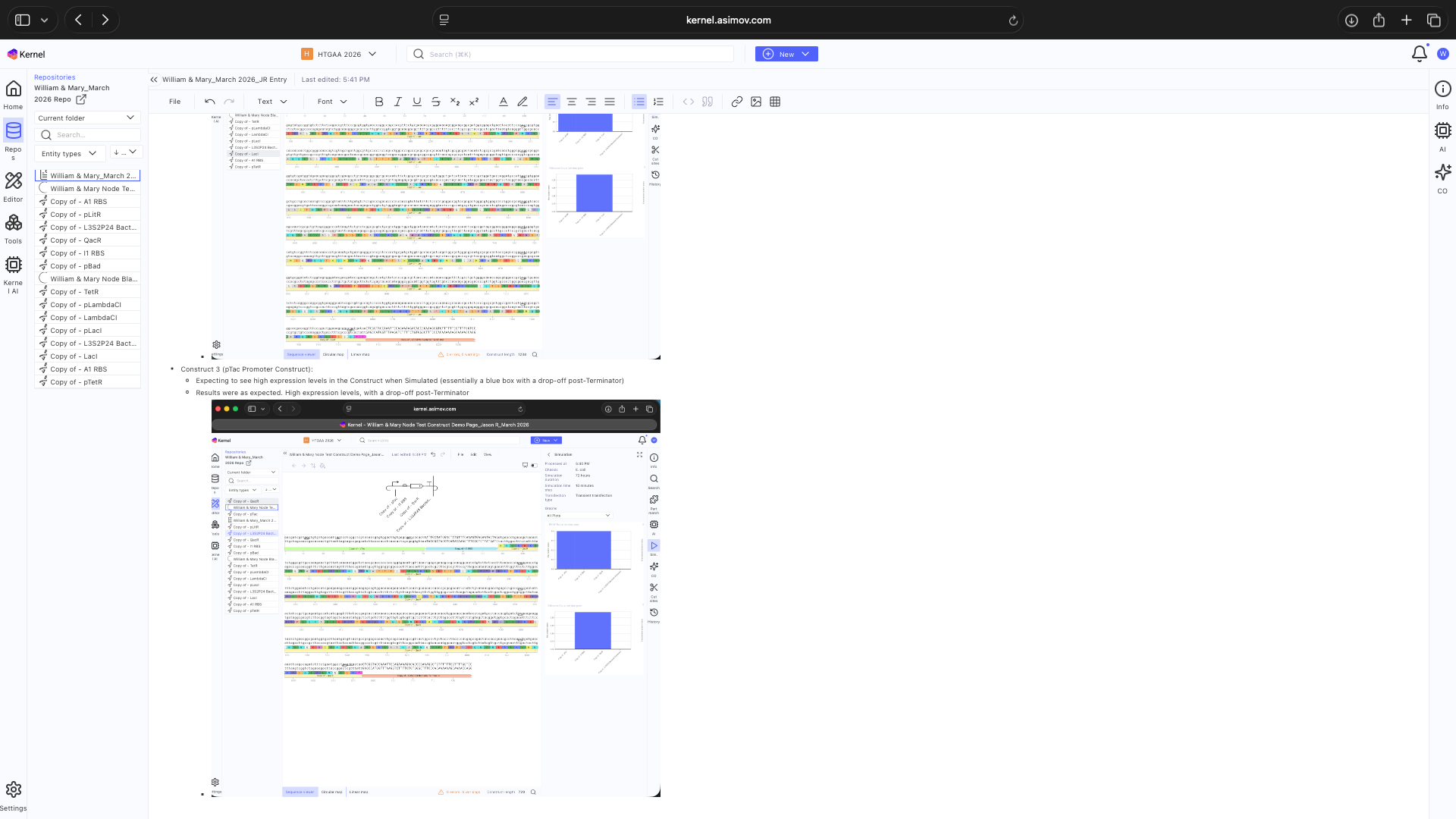
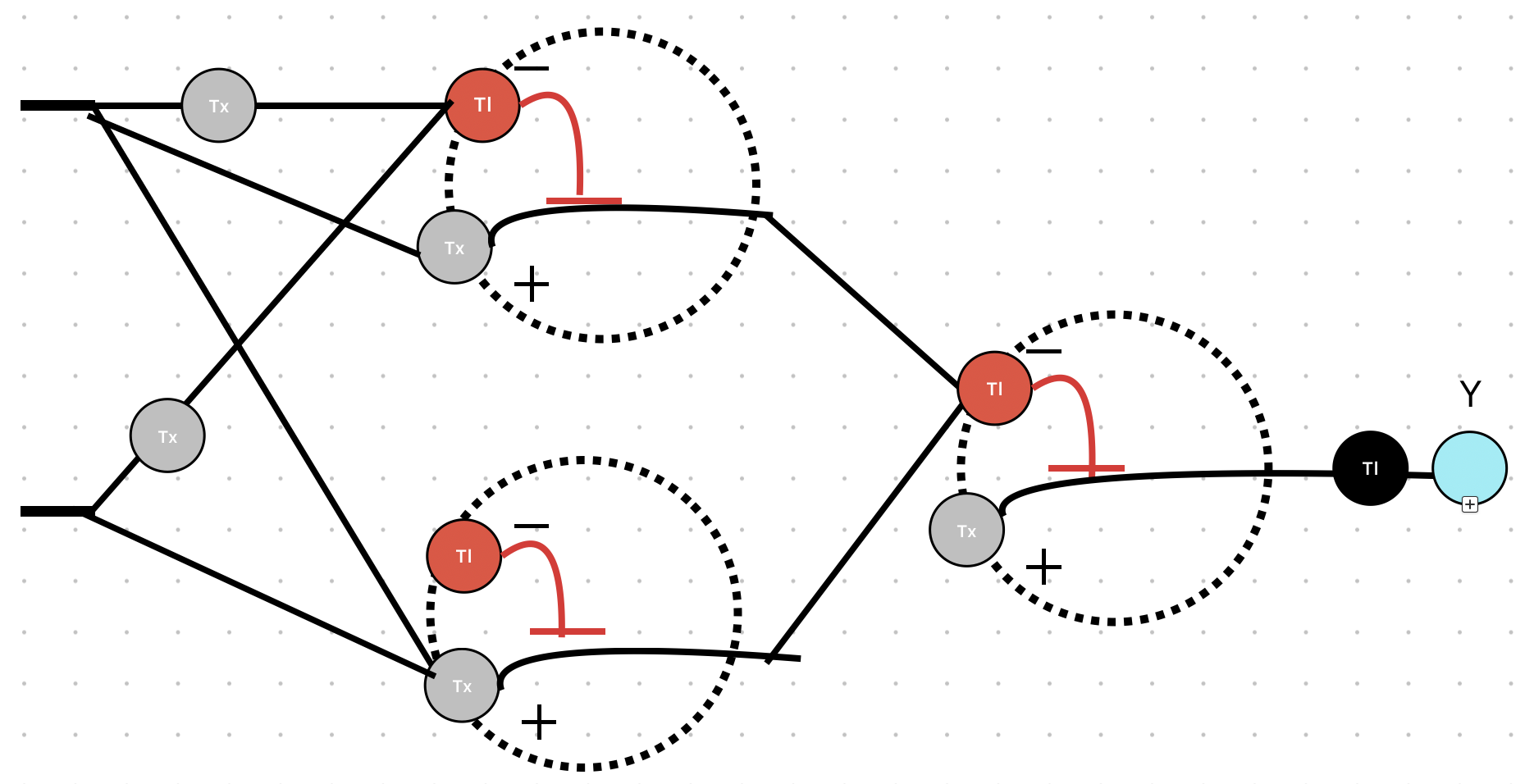
- Week 7 HW: Genetic Circuits Part 2
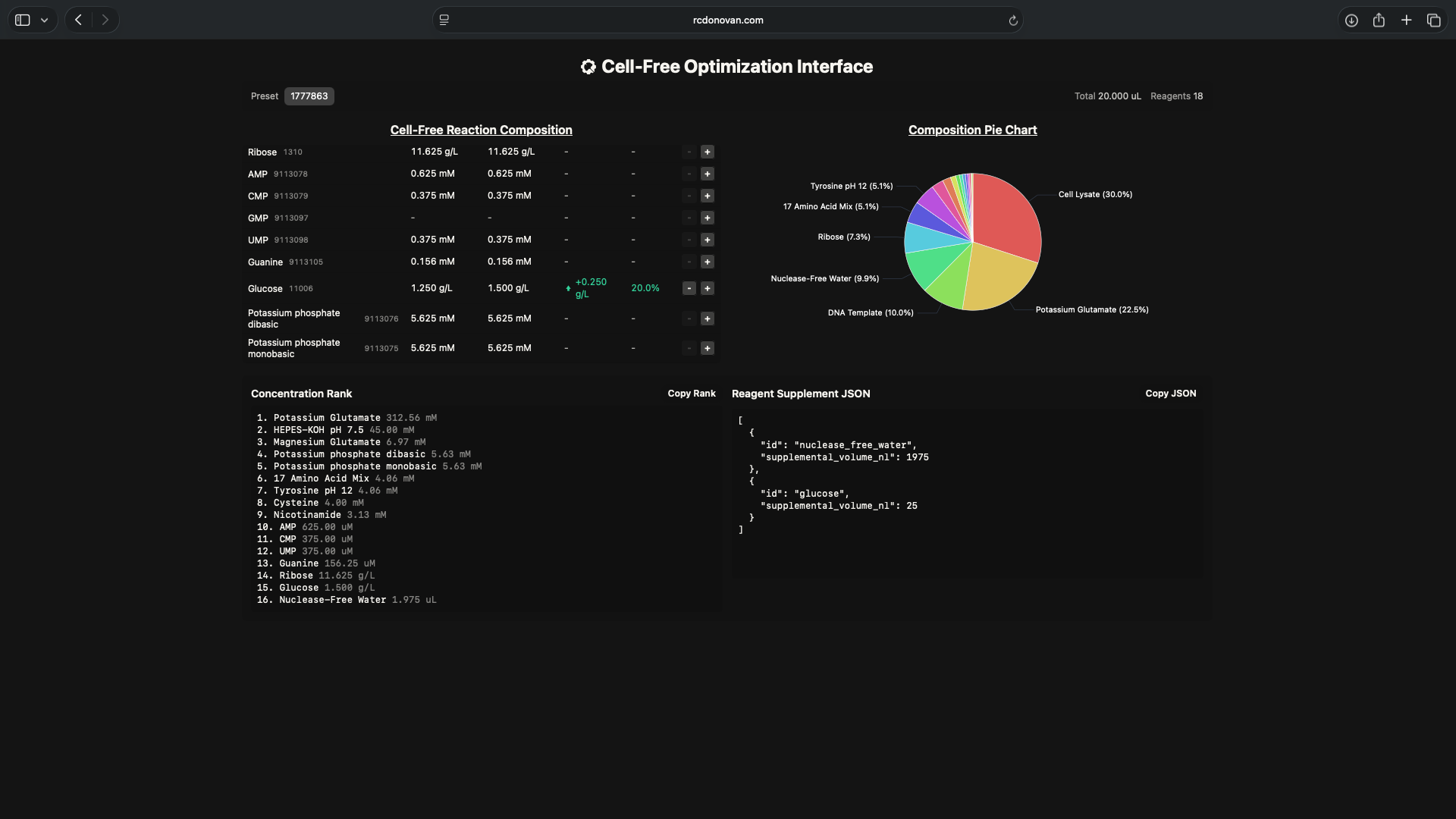
- Week 9 HW: Cell-Free Systems
- Week 10 HW: Advanced Imaging & Measurement Technology
- Week 11 HW: Bioproduction and Cloud Labs
- Week 12 HW: Bioproduction & Cloud Labs Part 2
- Week 13 HW: Scaling Health Innovation
Labs
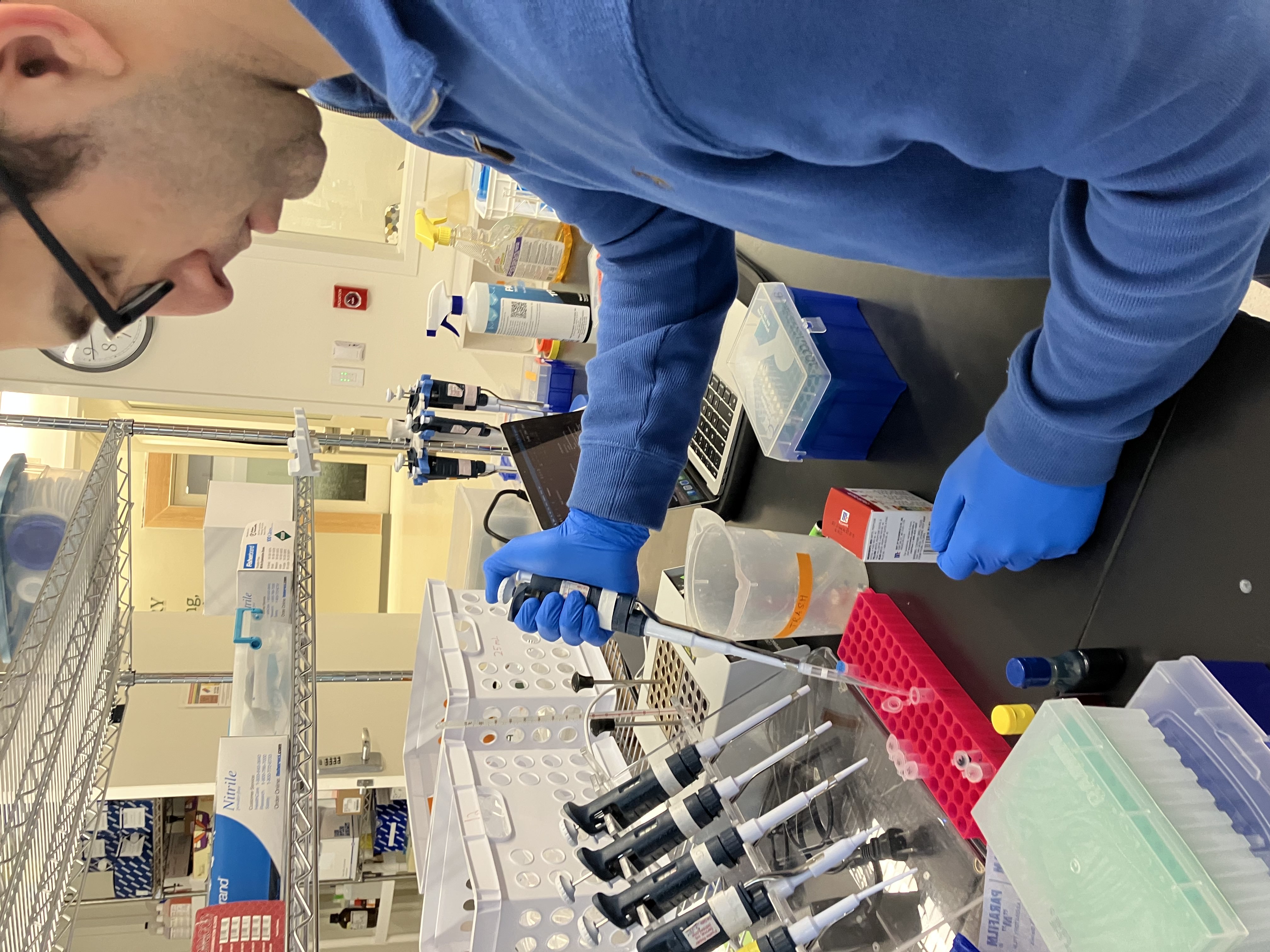
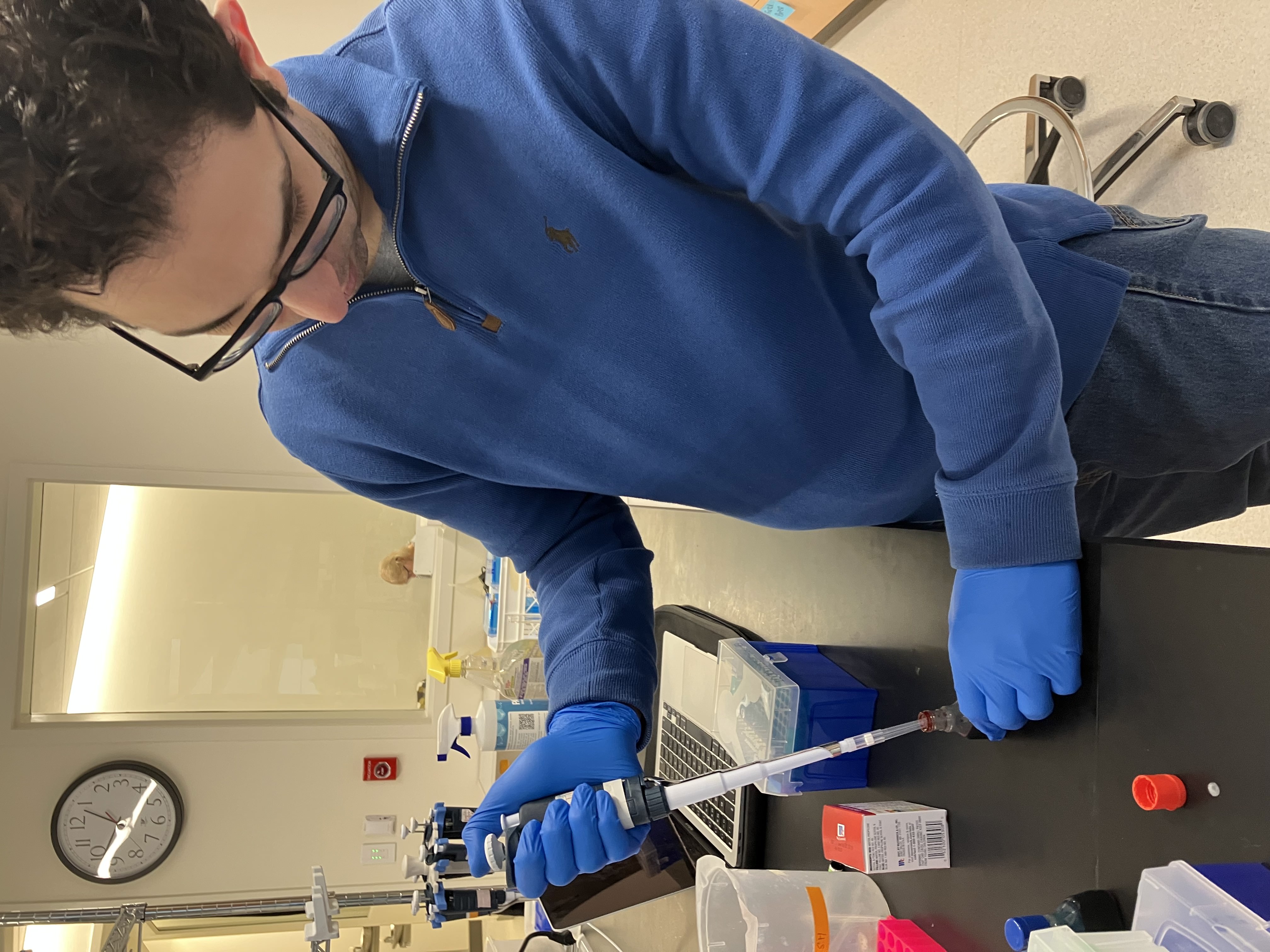
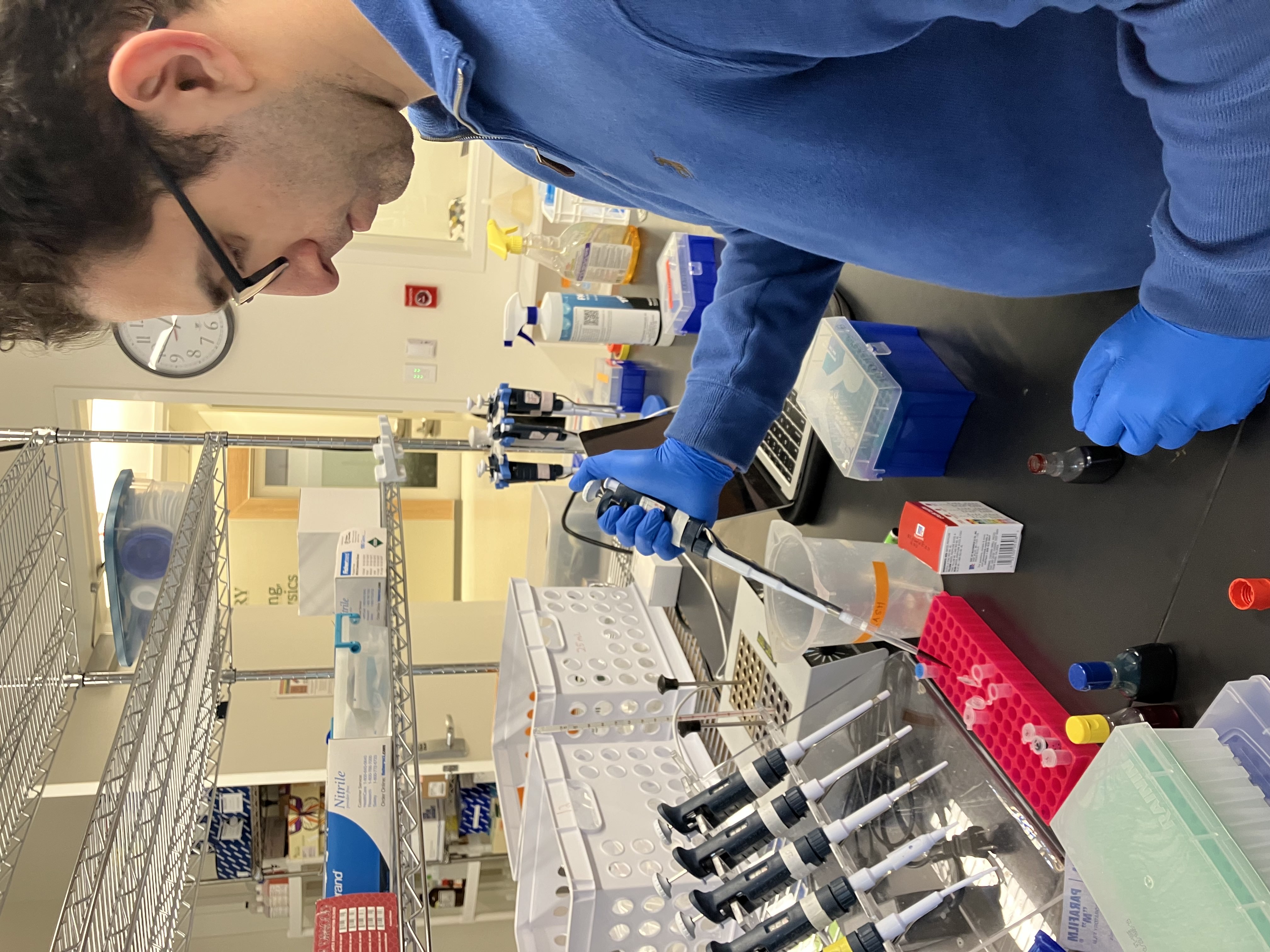
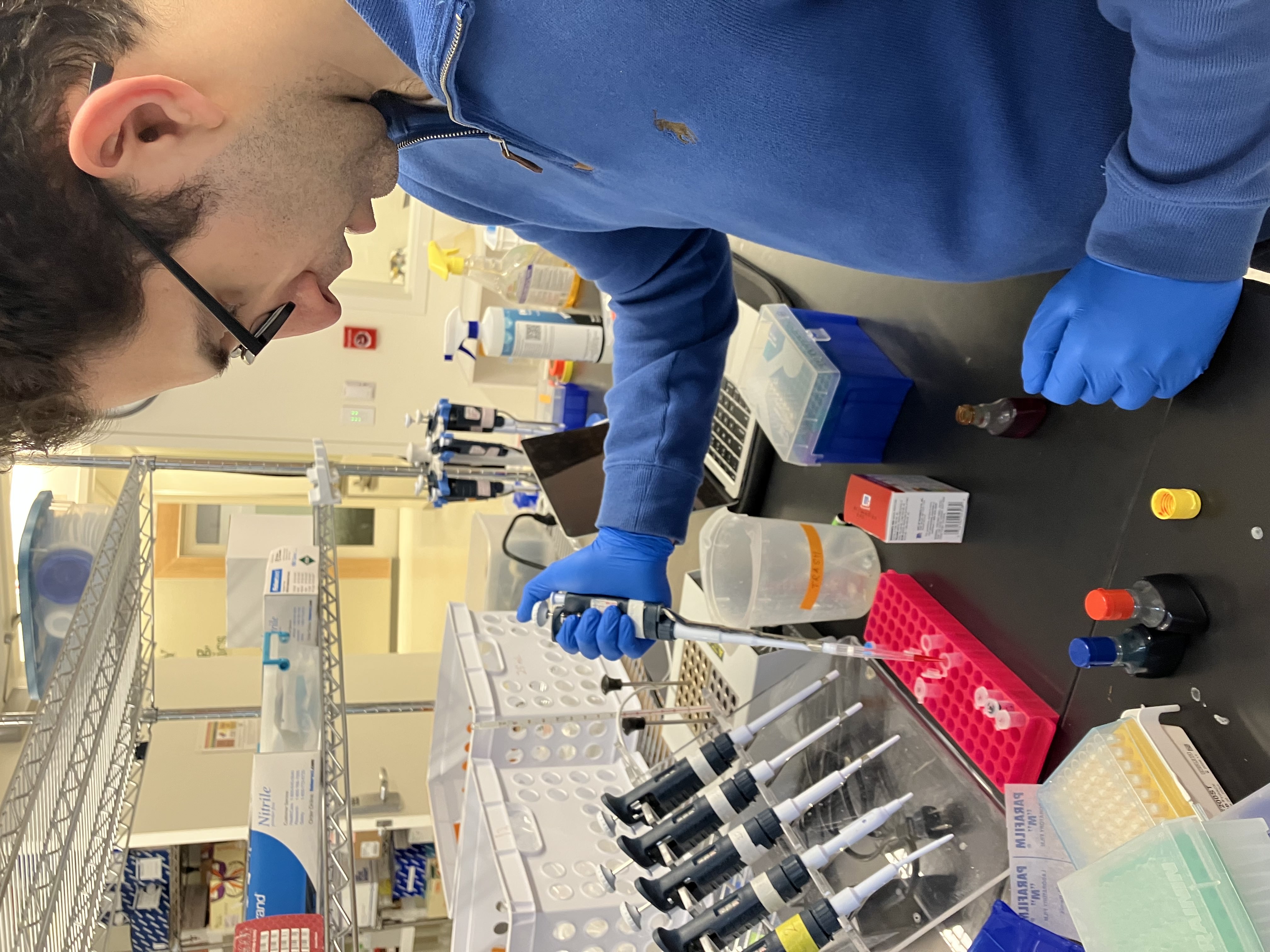
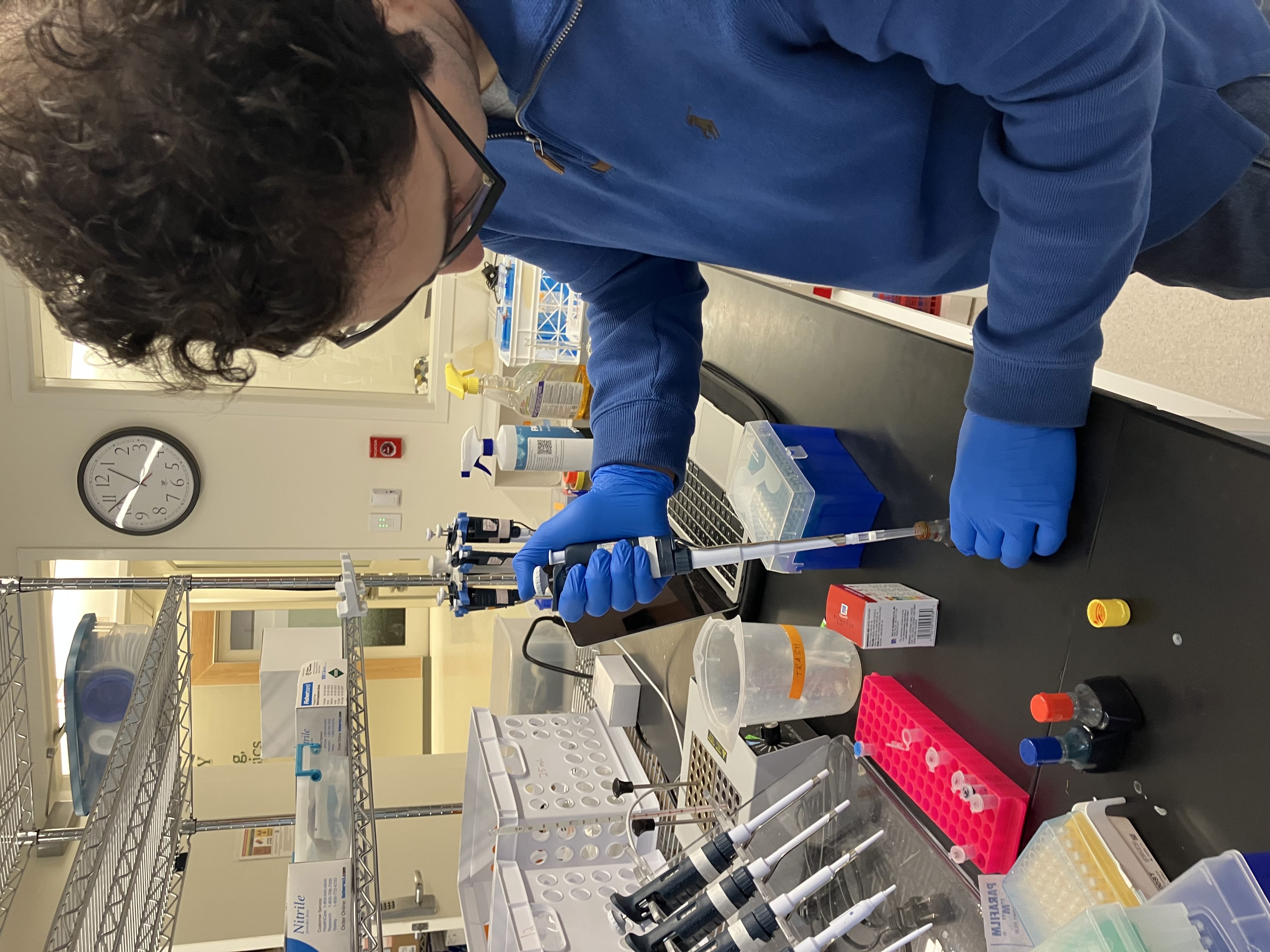
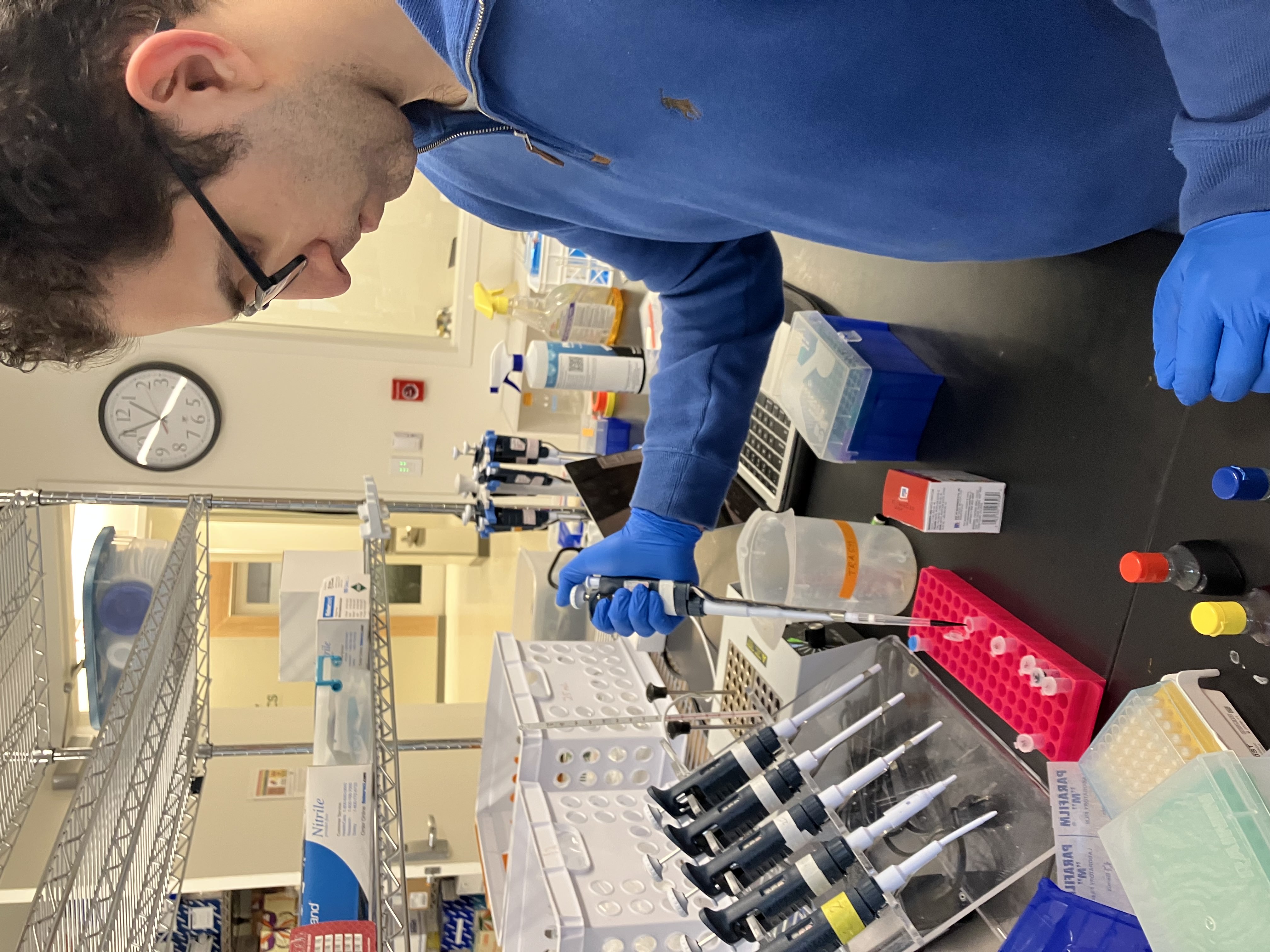
- Week 1 Lab: Introduction to Pipetting and Dilutions
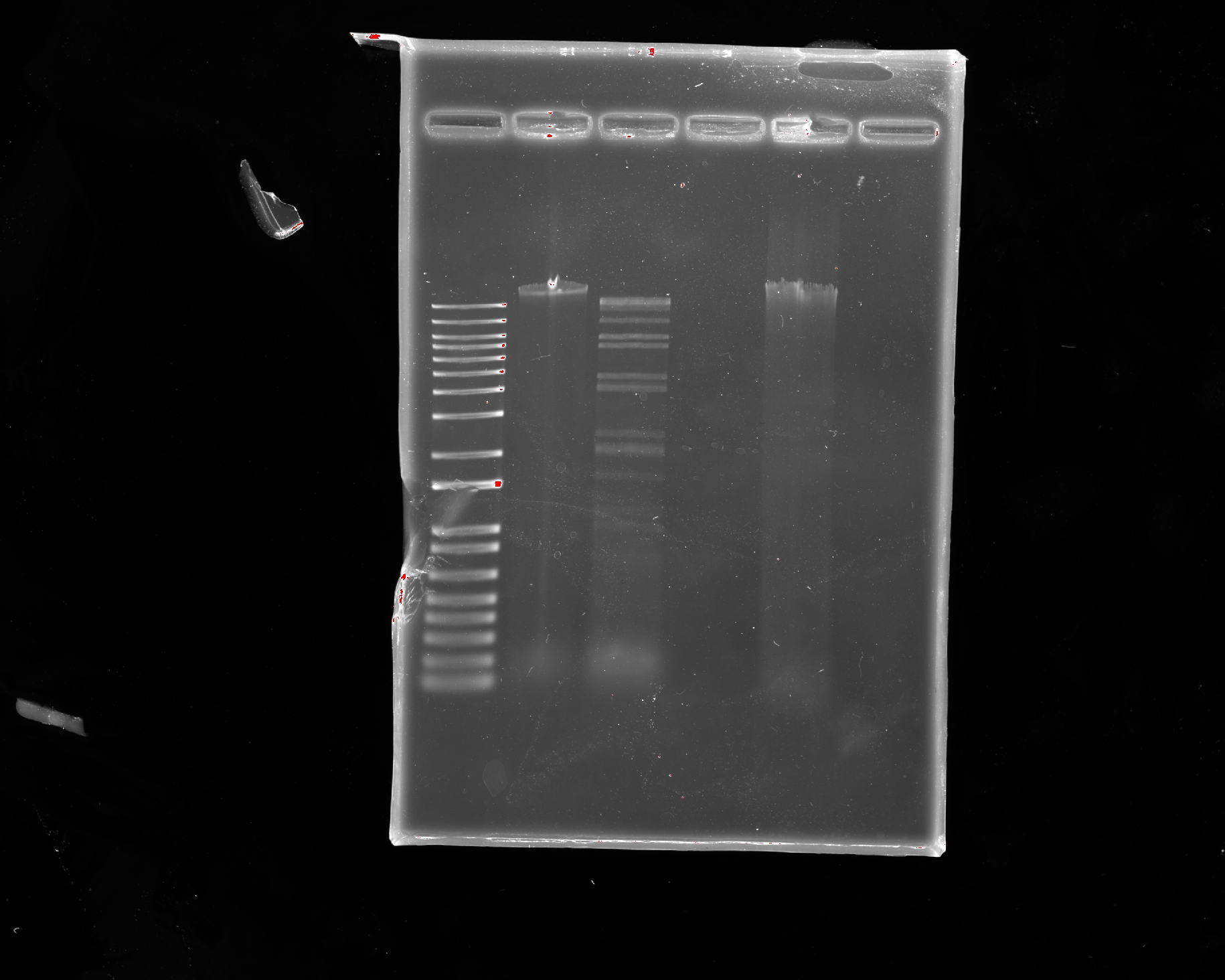
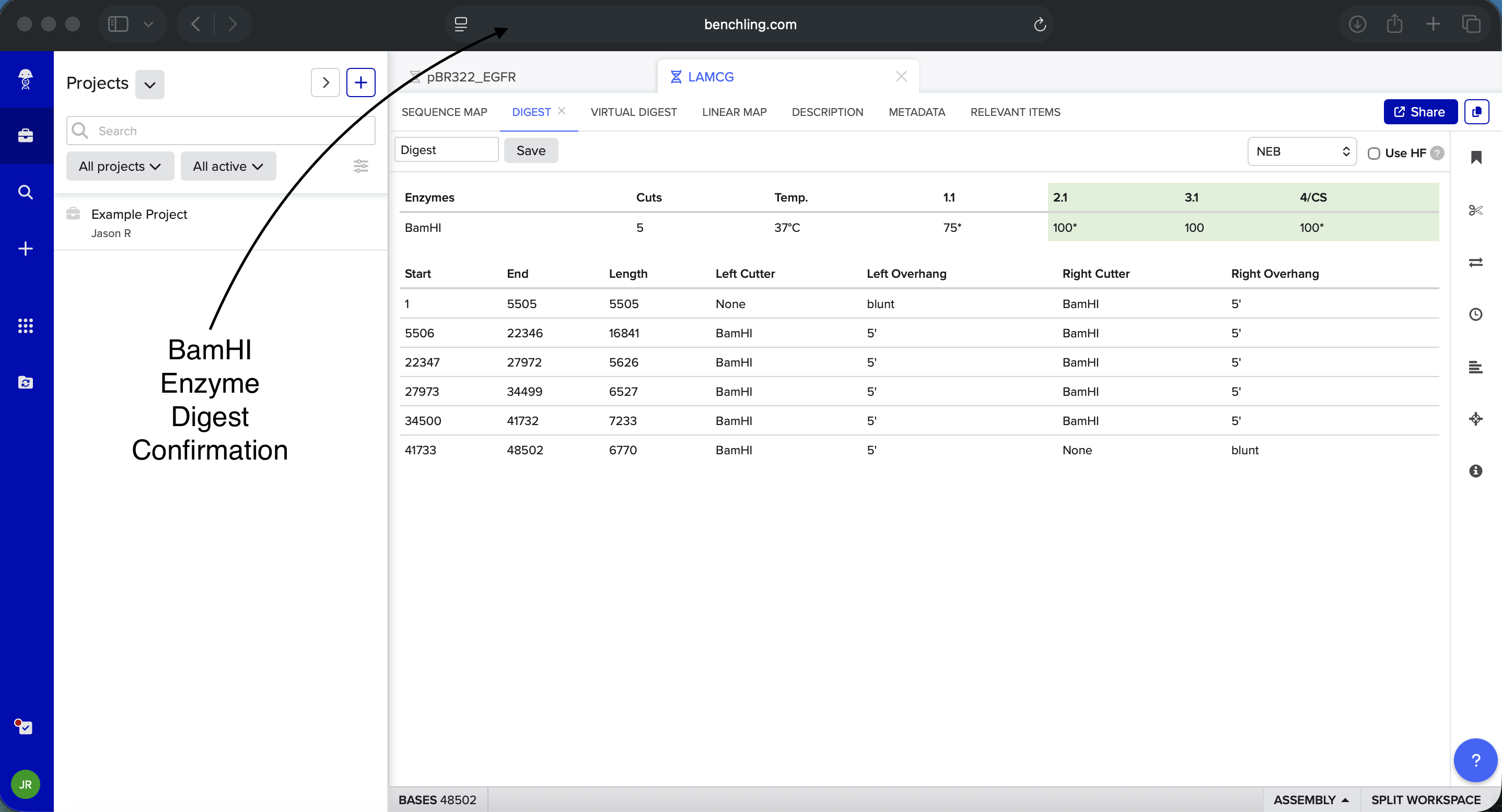
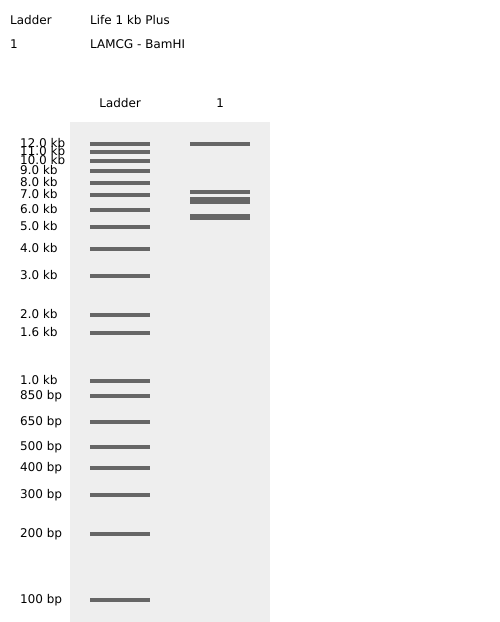
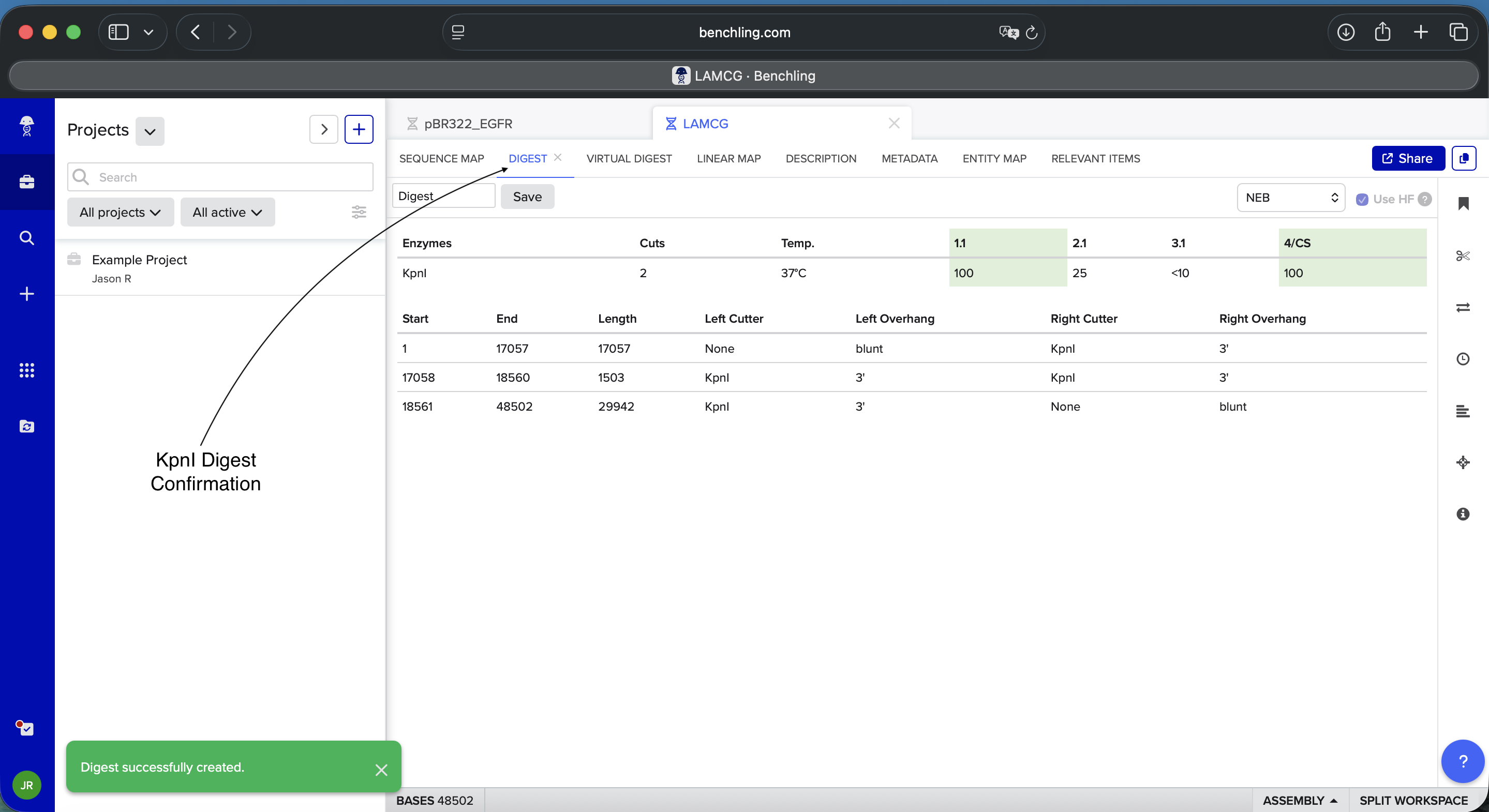
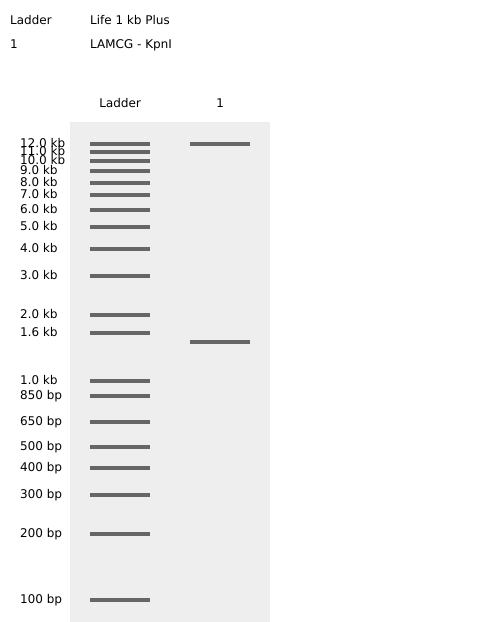
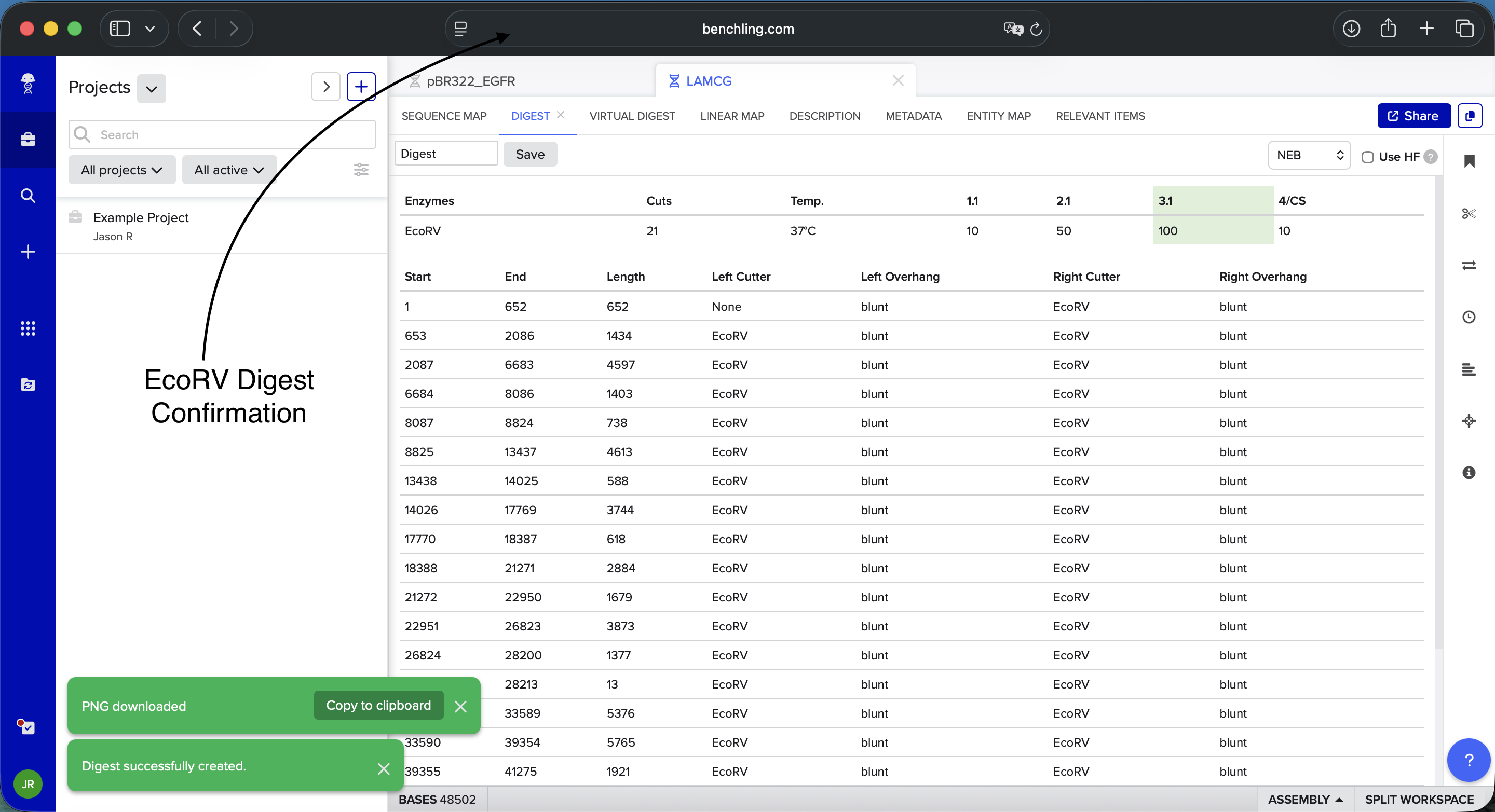
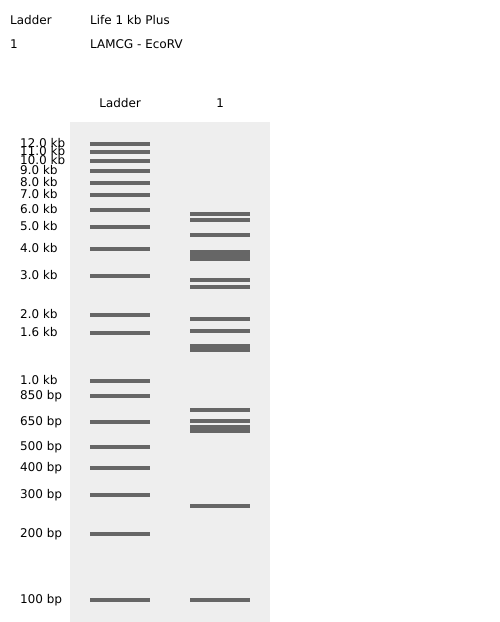
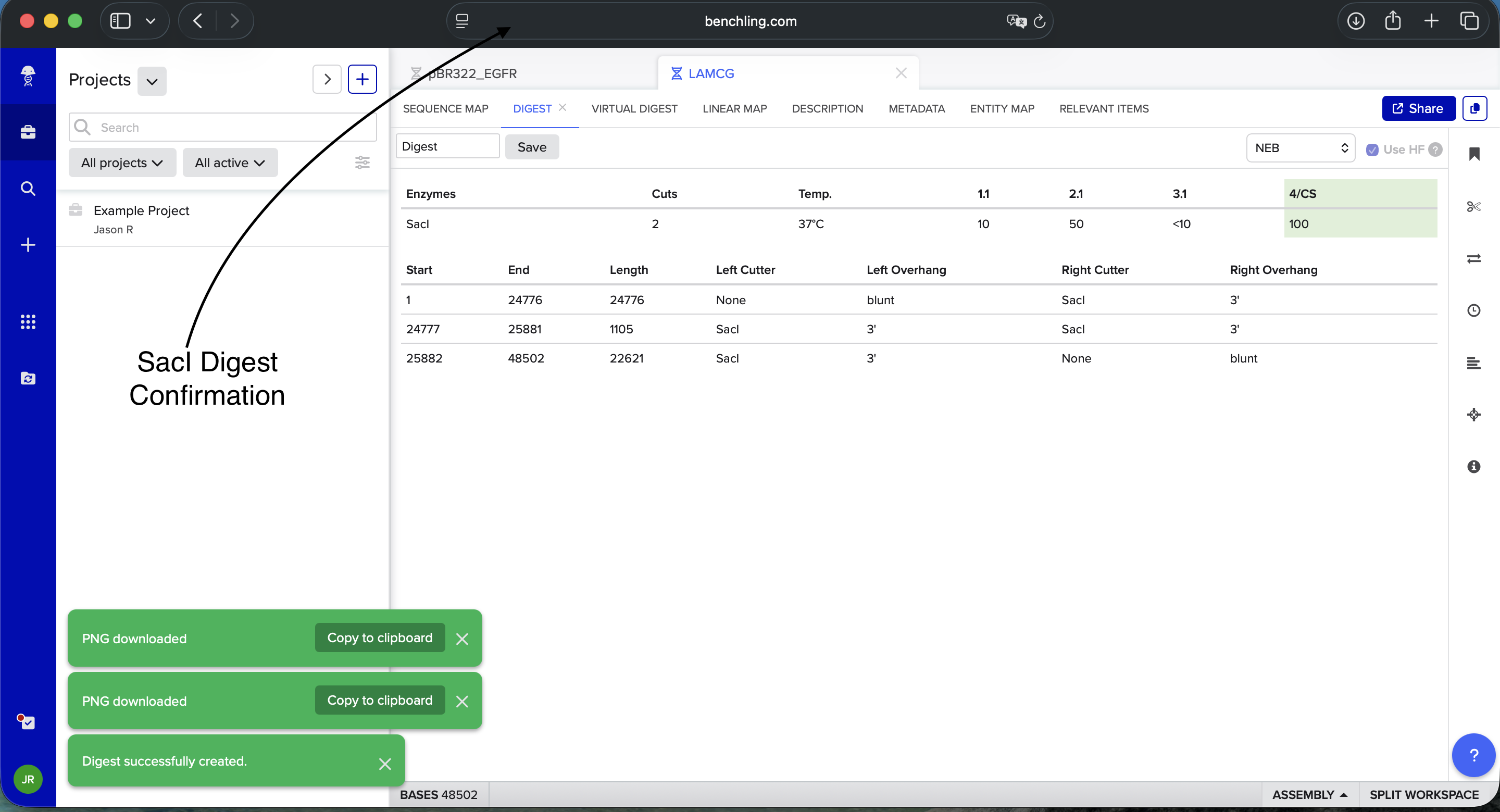
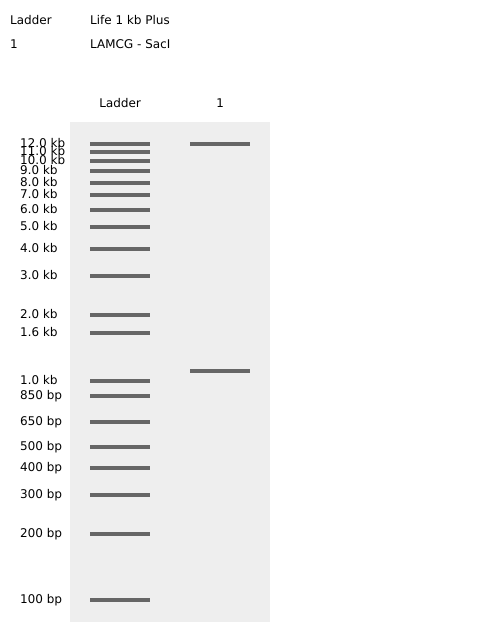
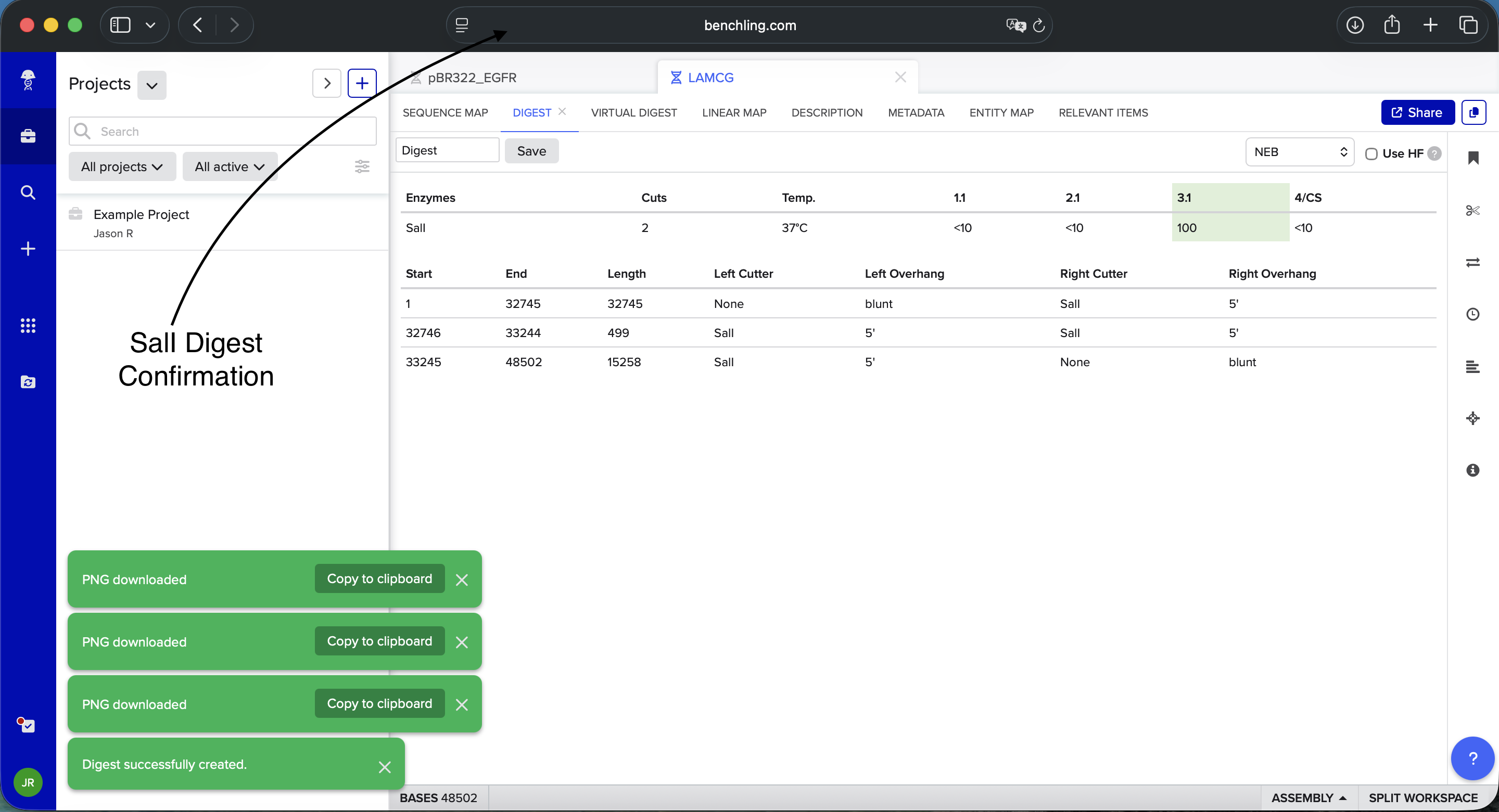
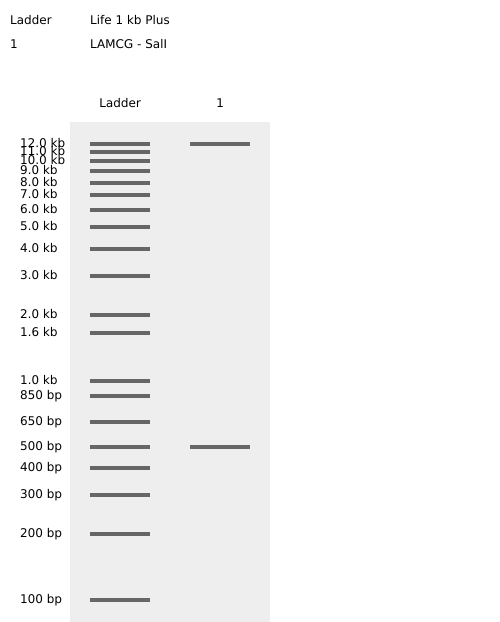
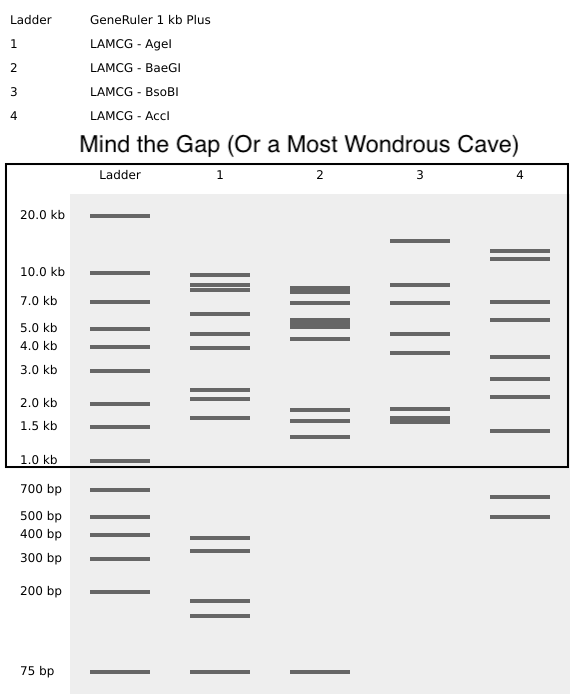
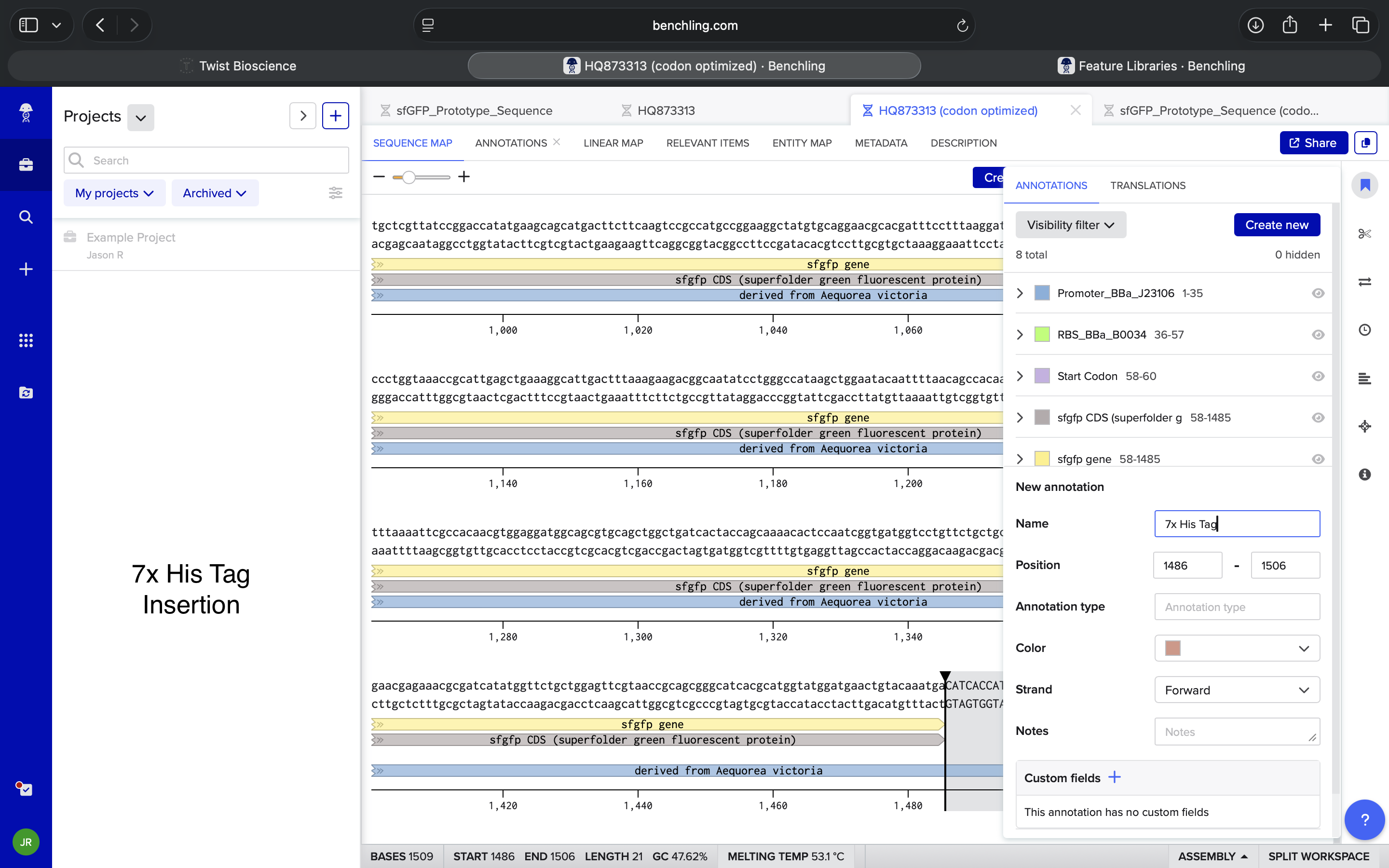
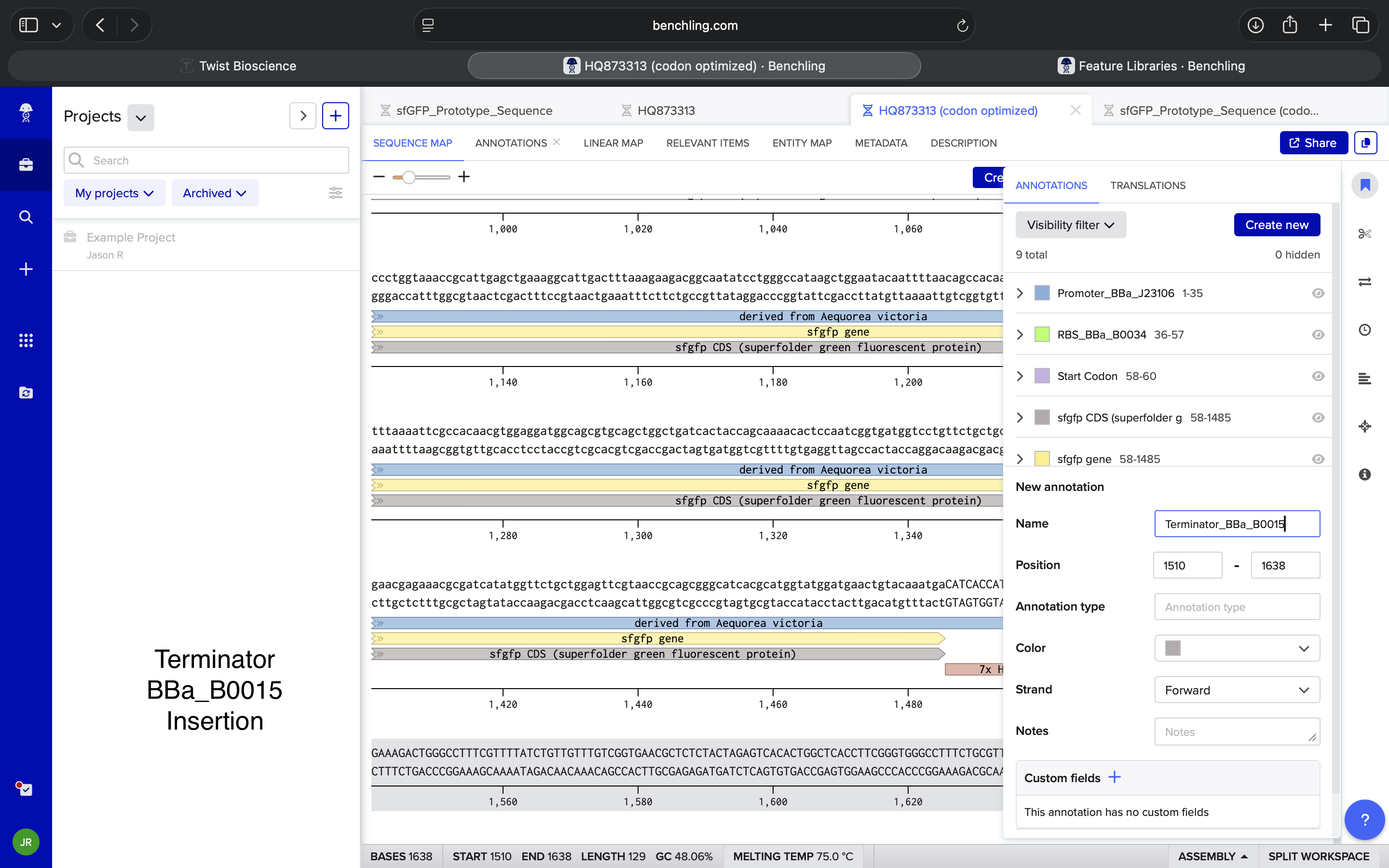
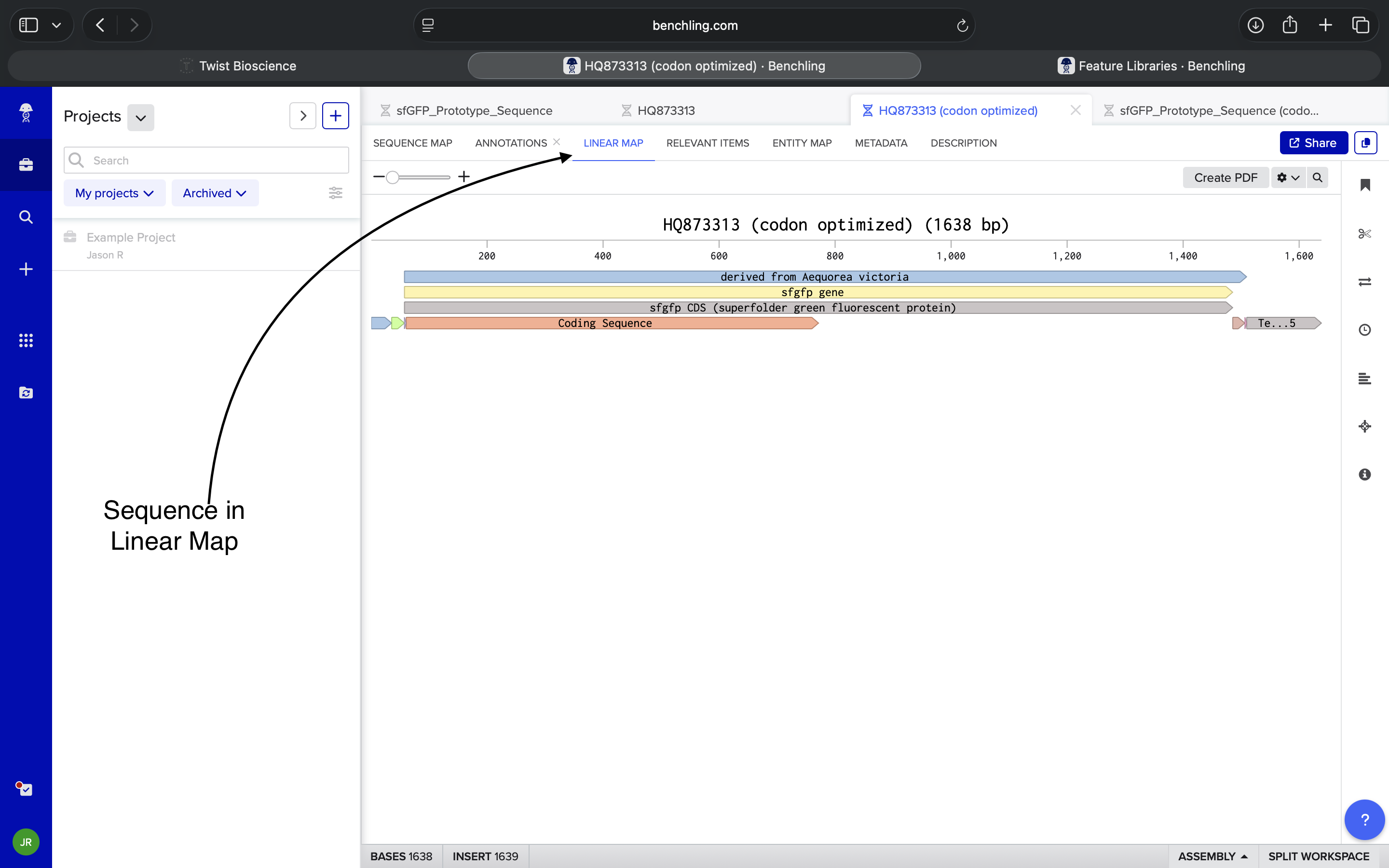
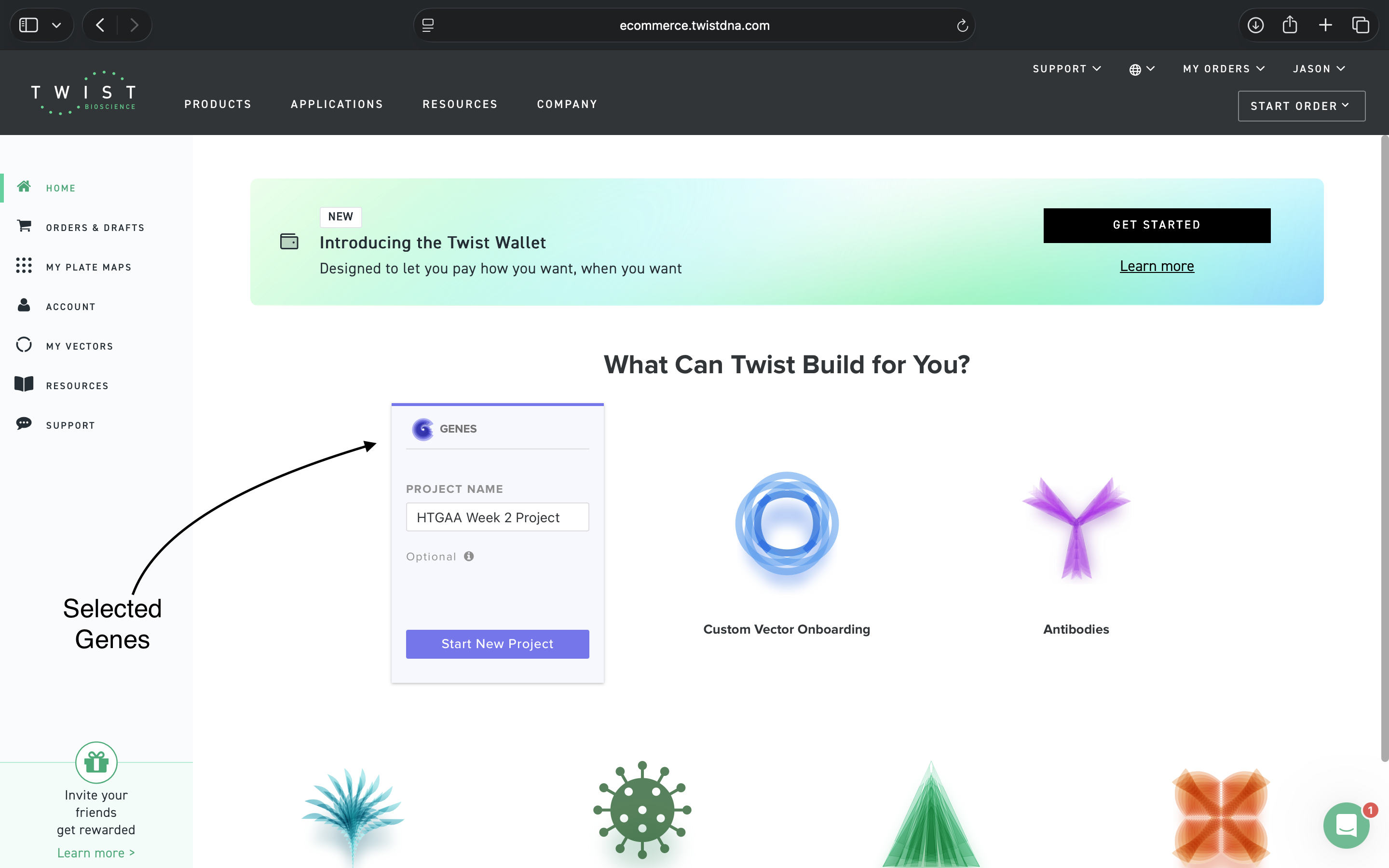
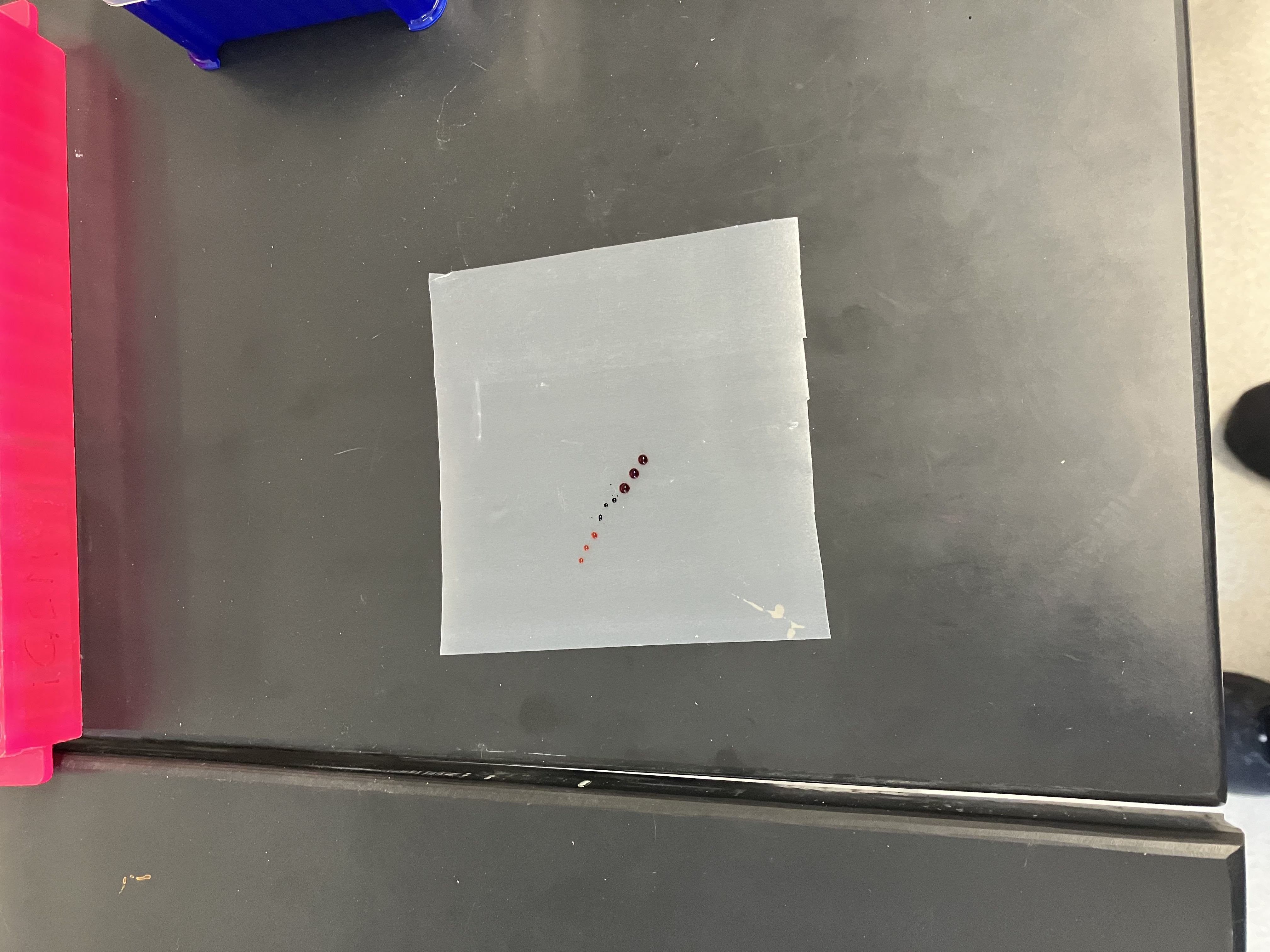
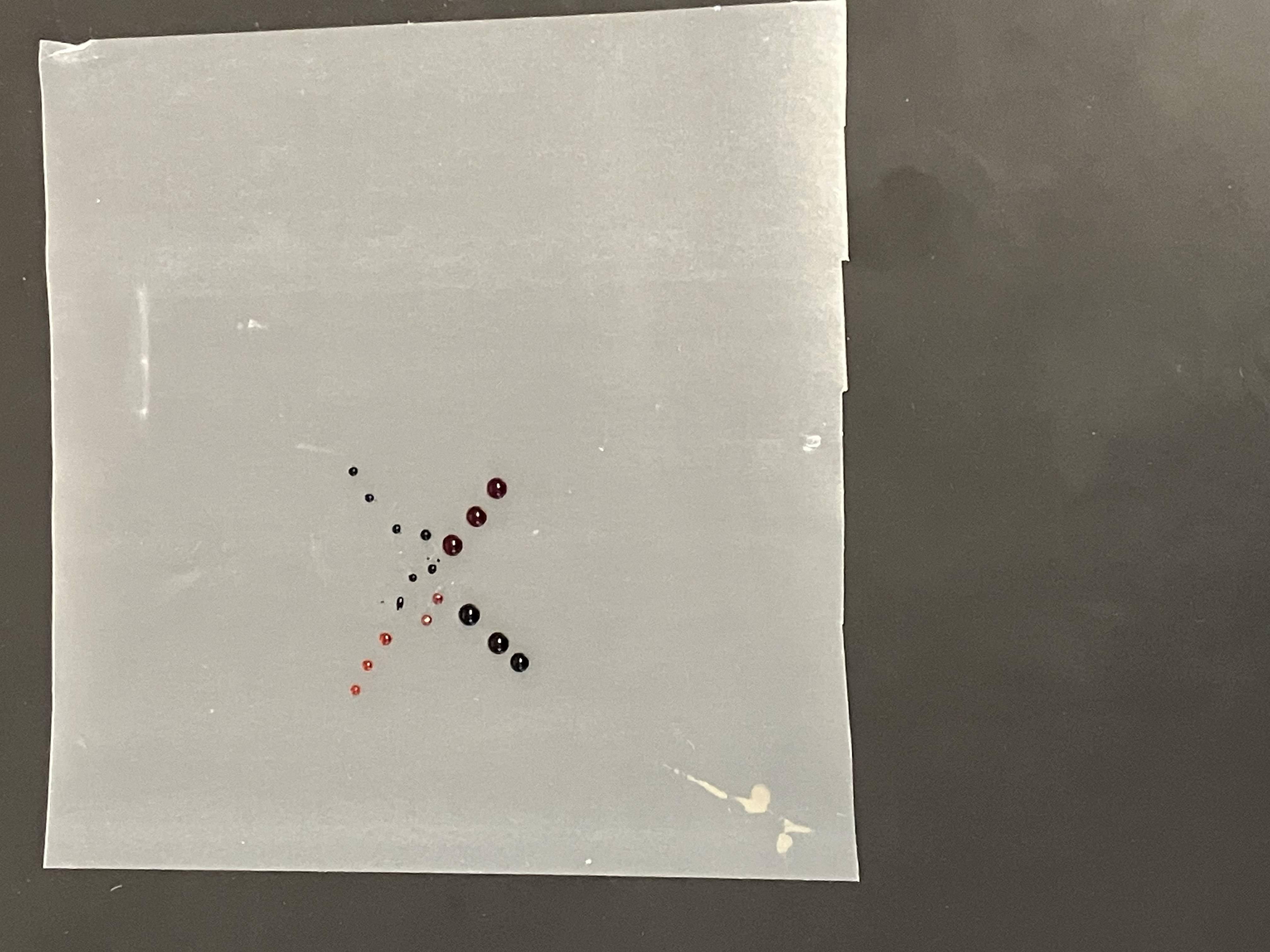
- Week 2: DNA Gel Art
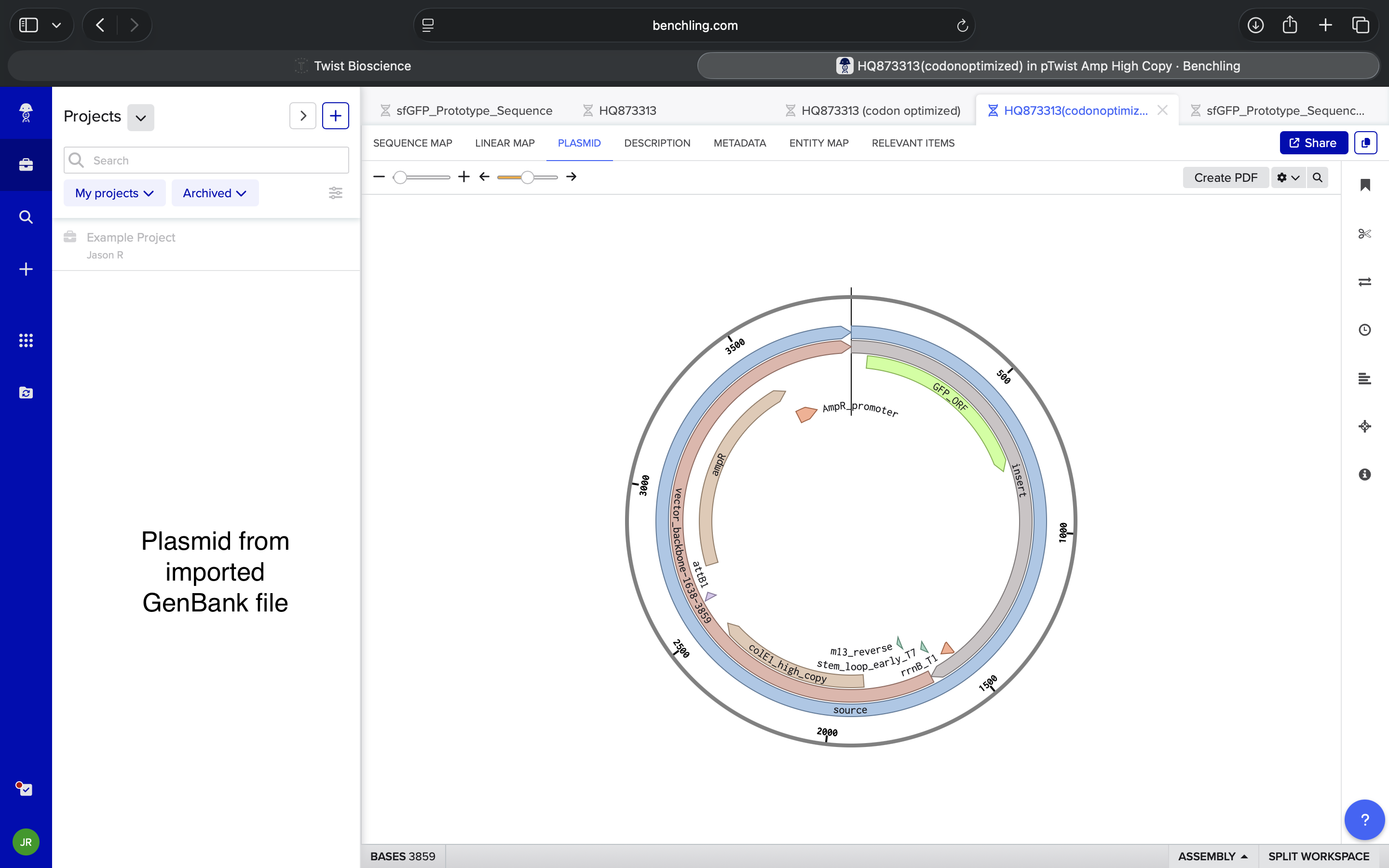
- Week 3 Lab: Opentrons Art
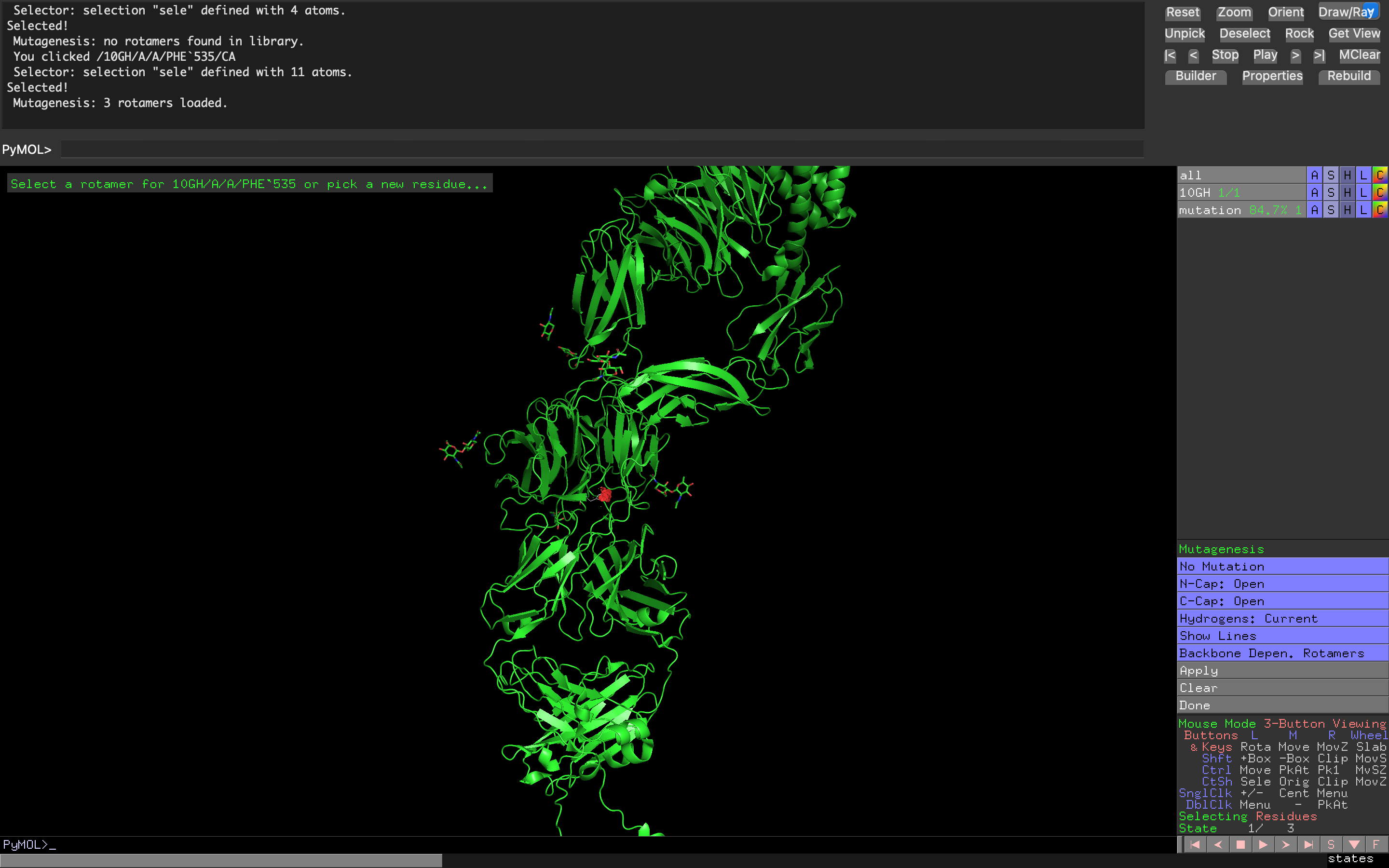
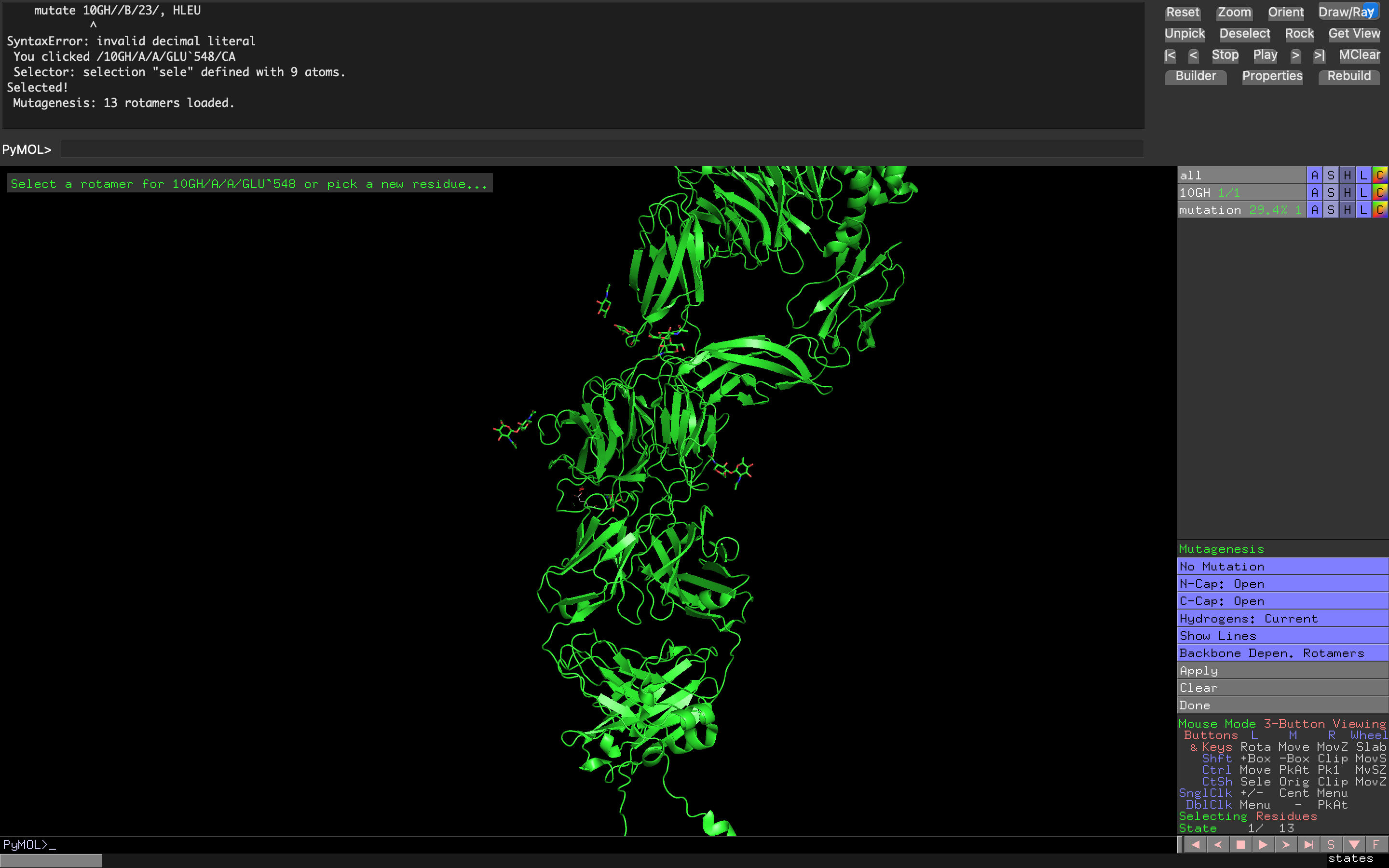
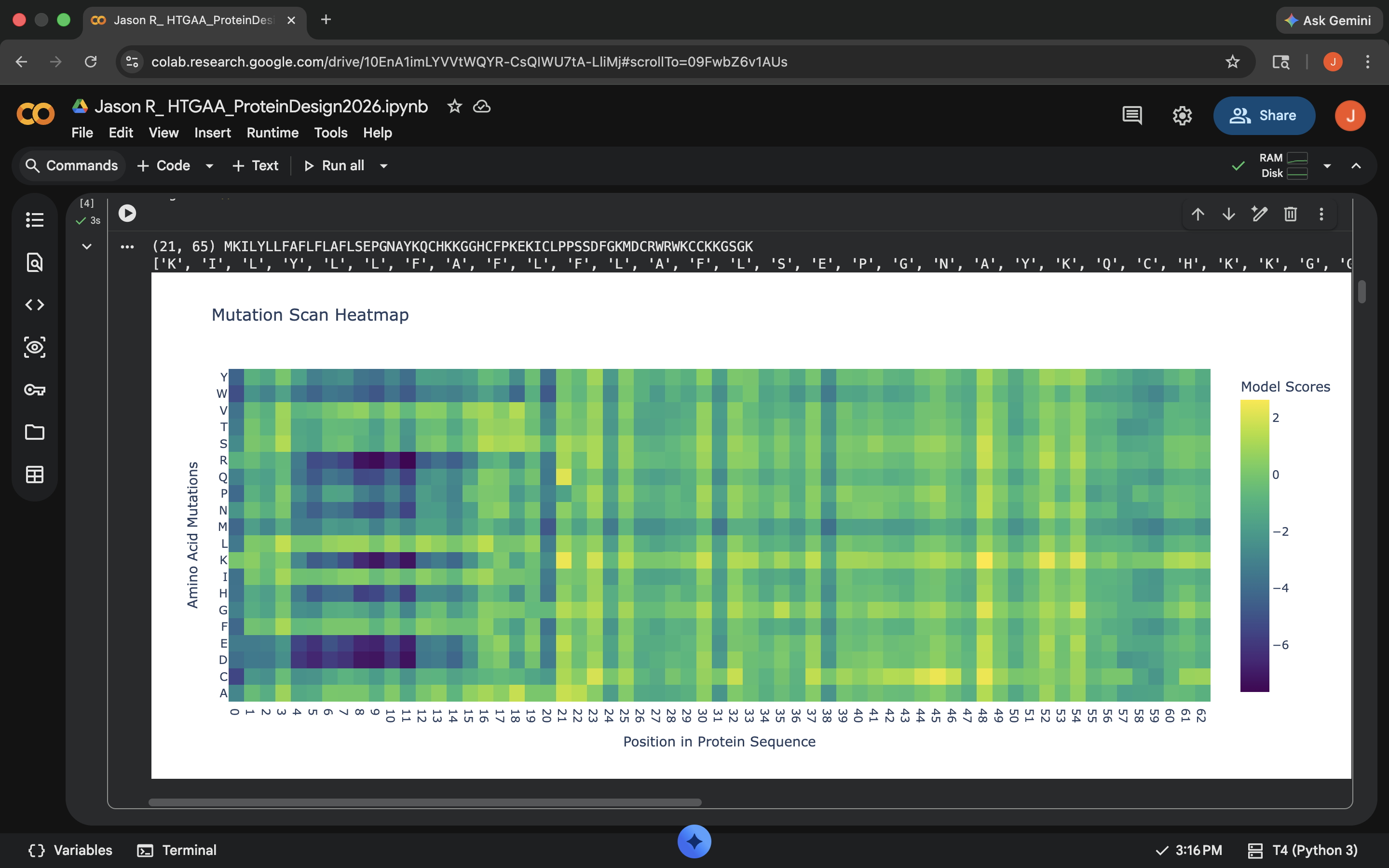
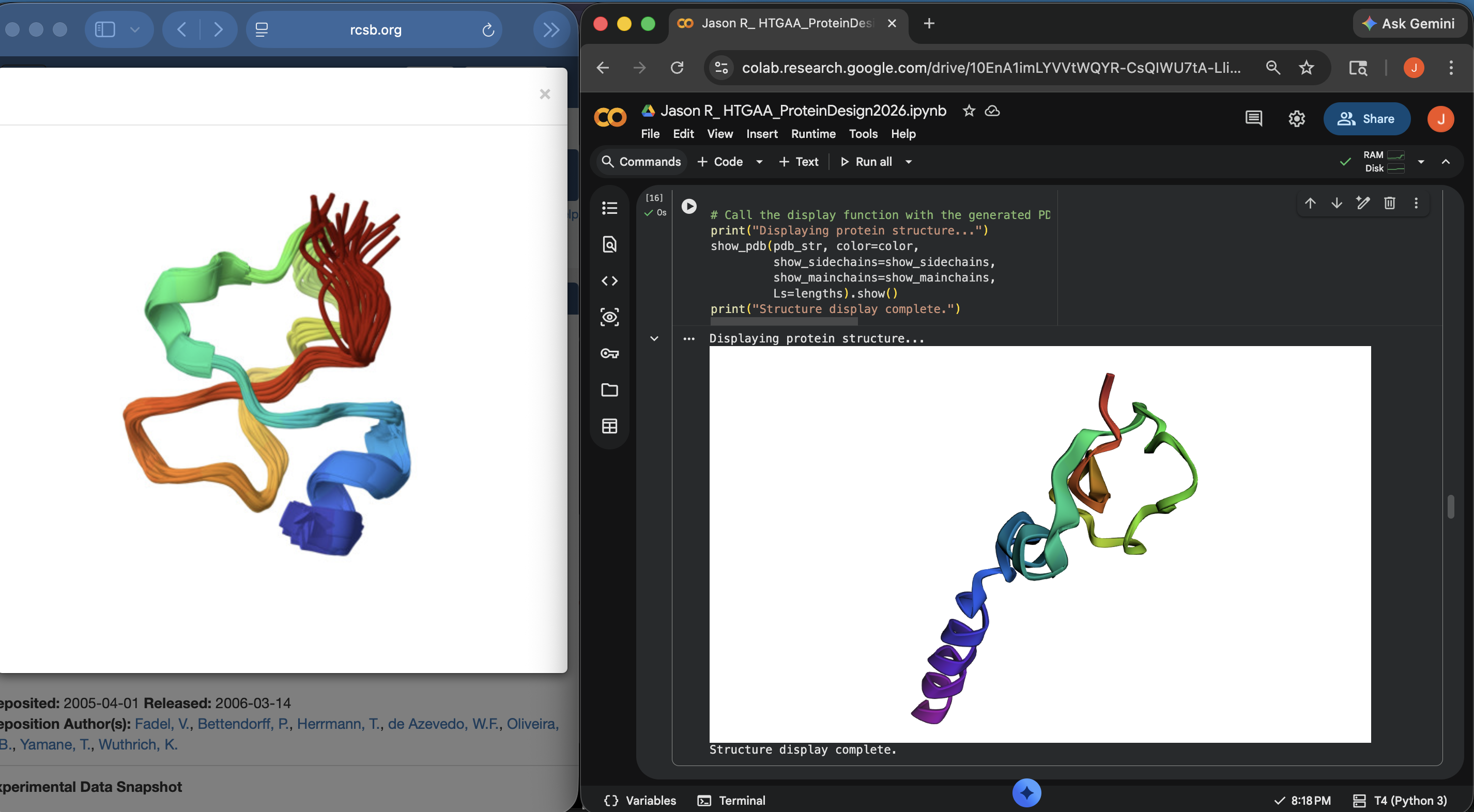
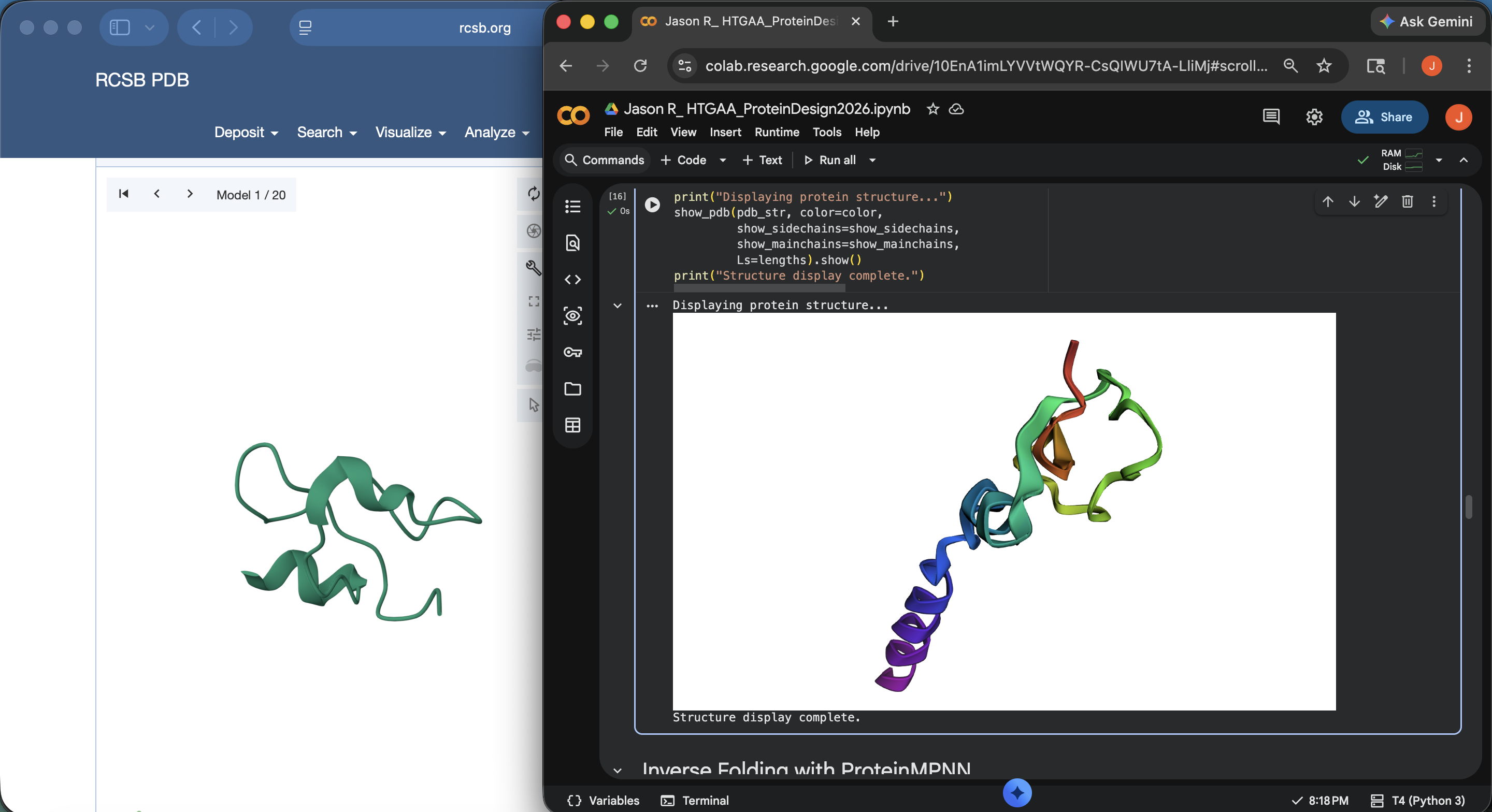
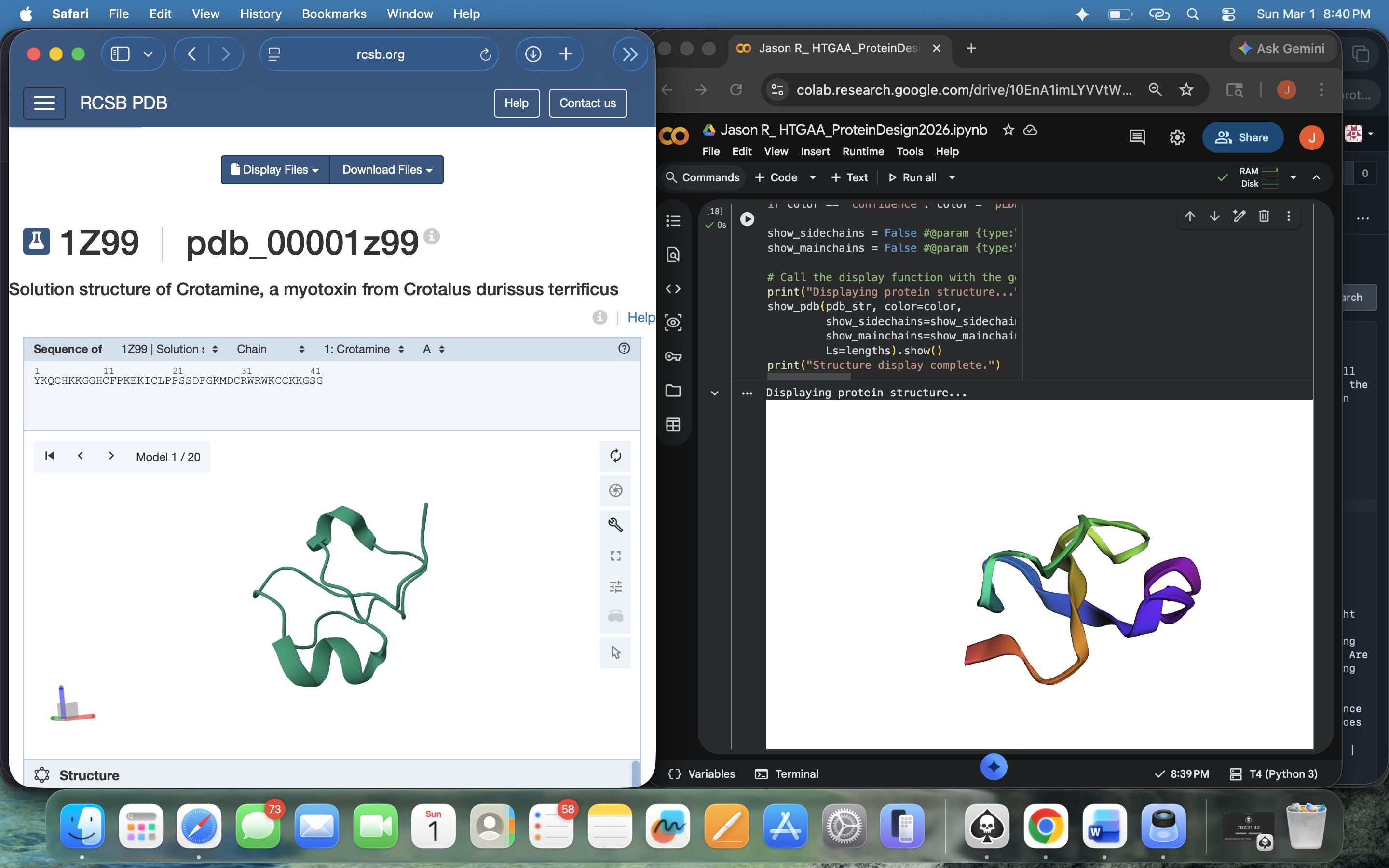
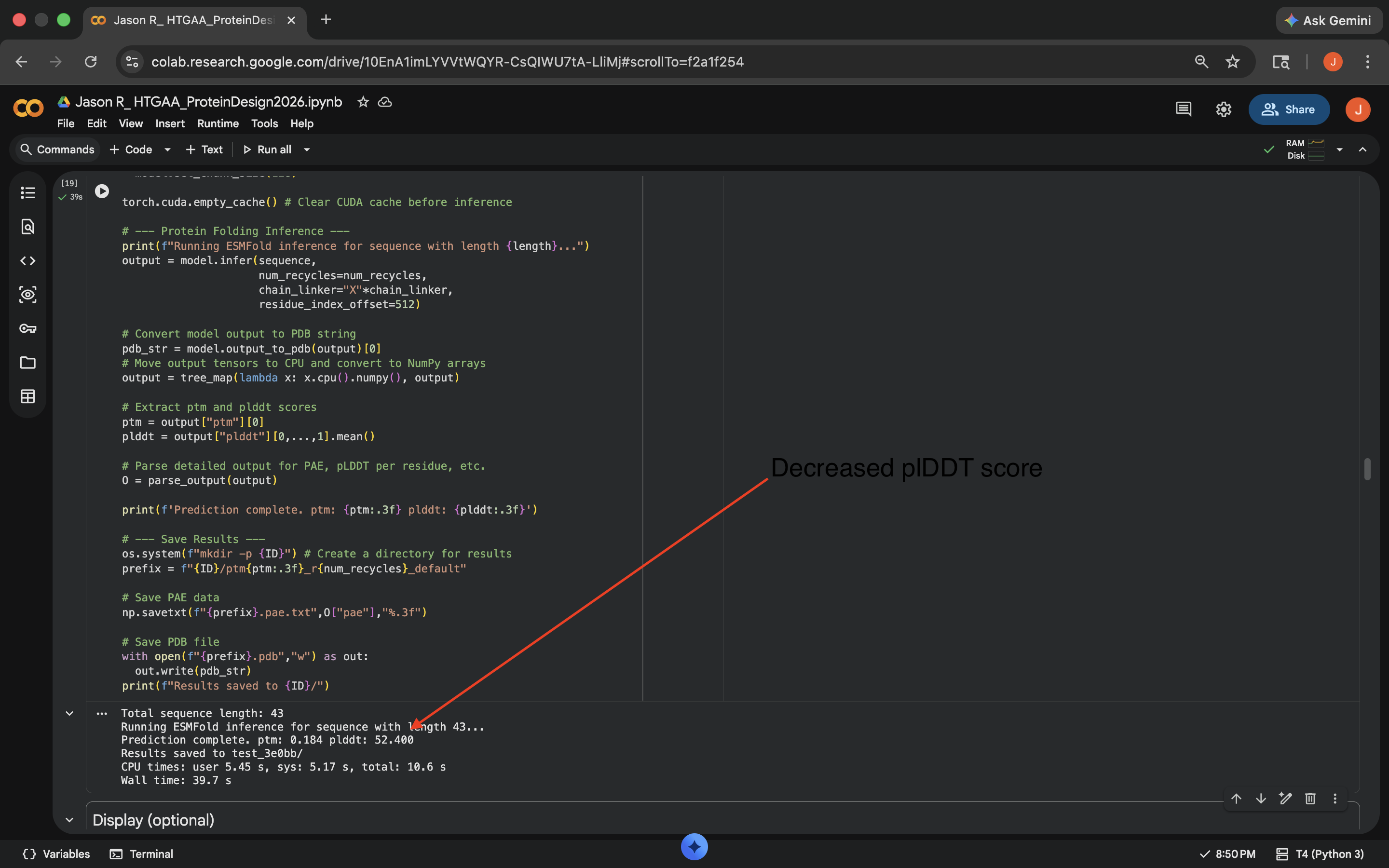
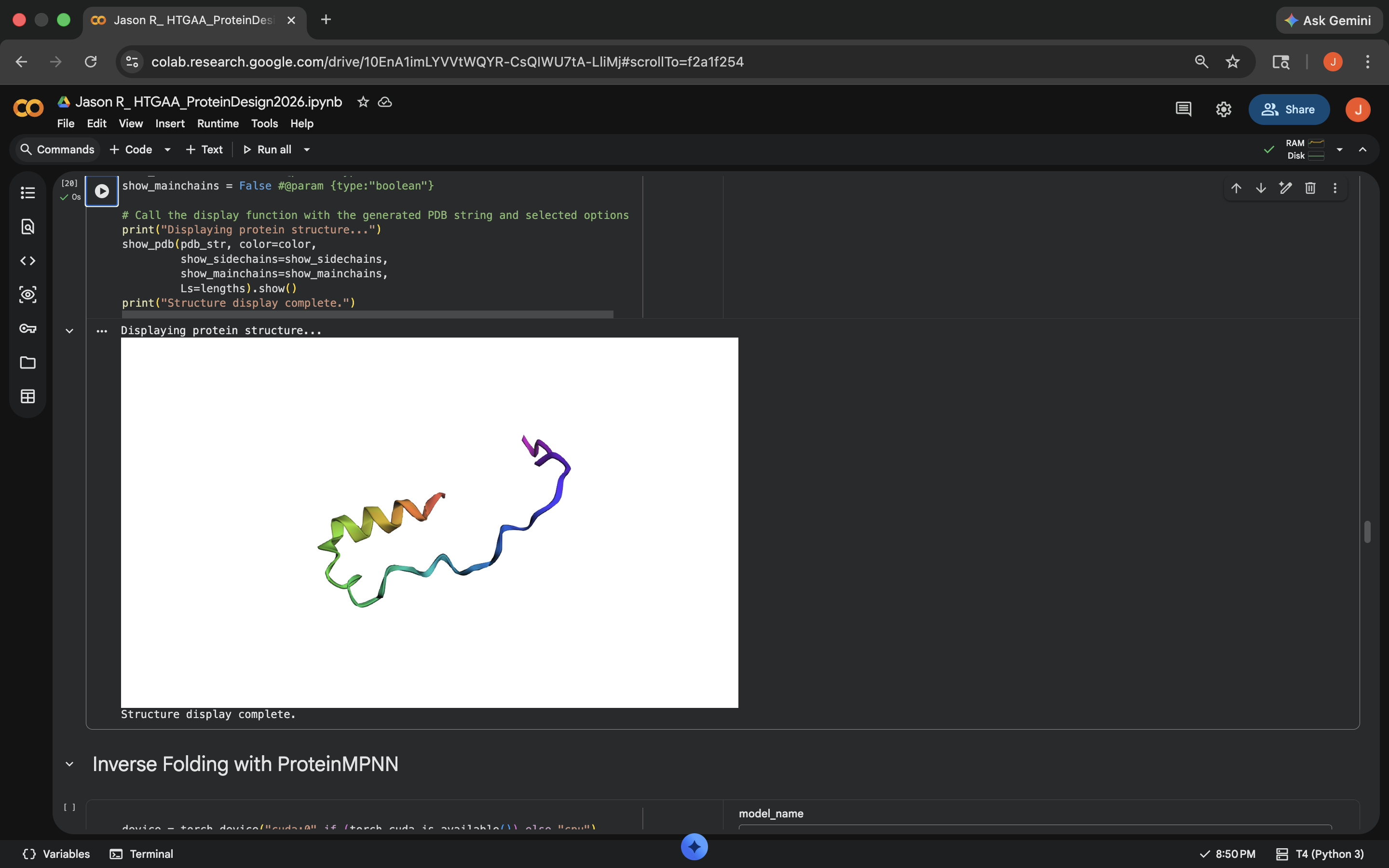
- Week 4 Lab: Protein Design Part I
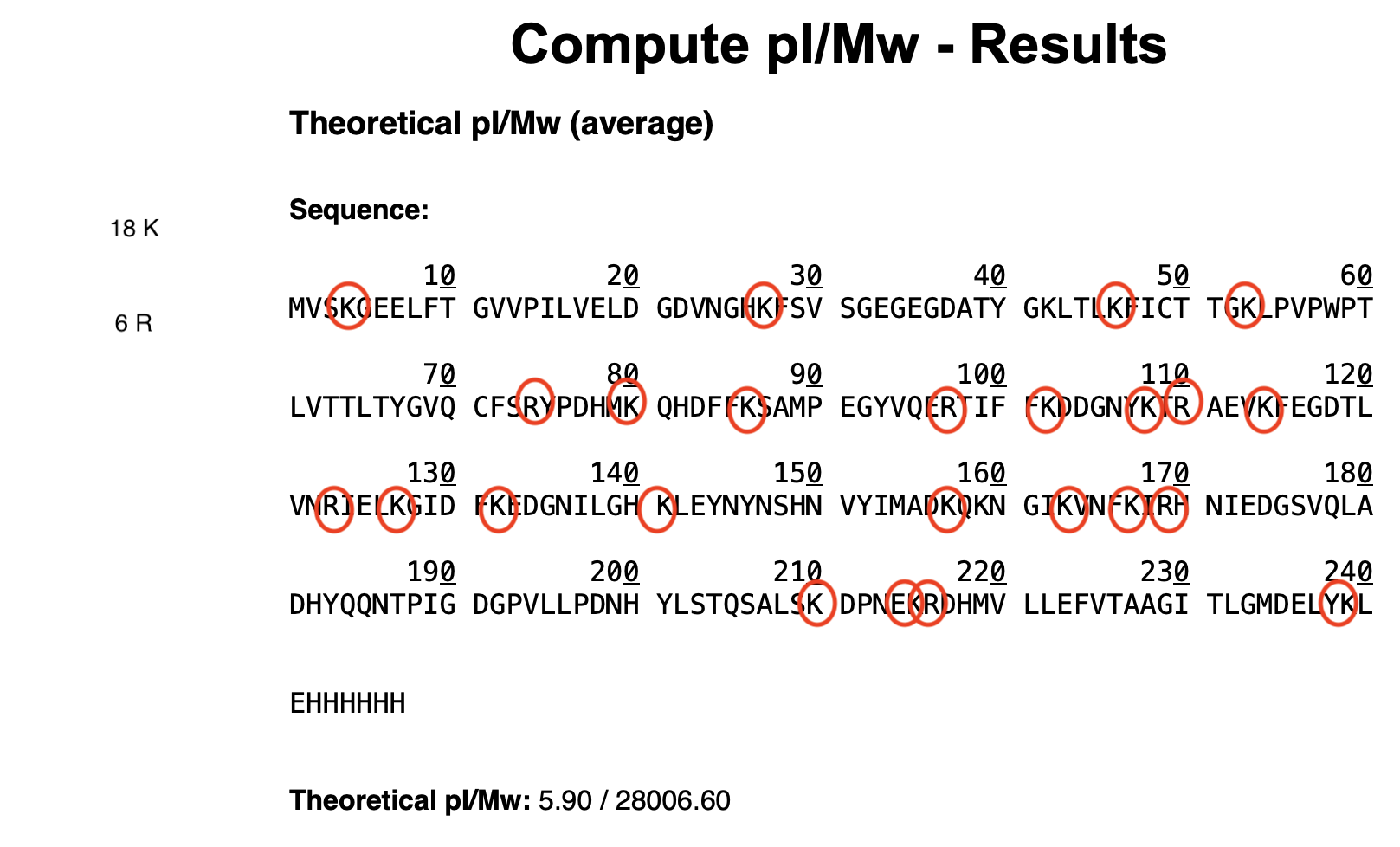
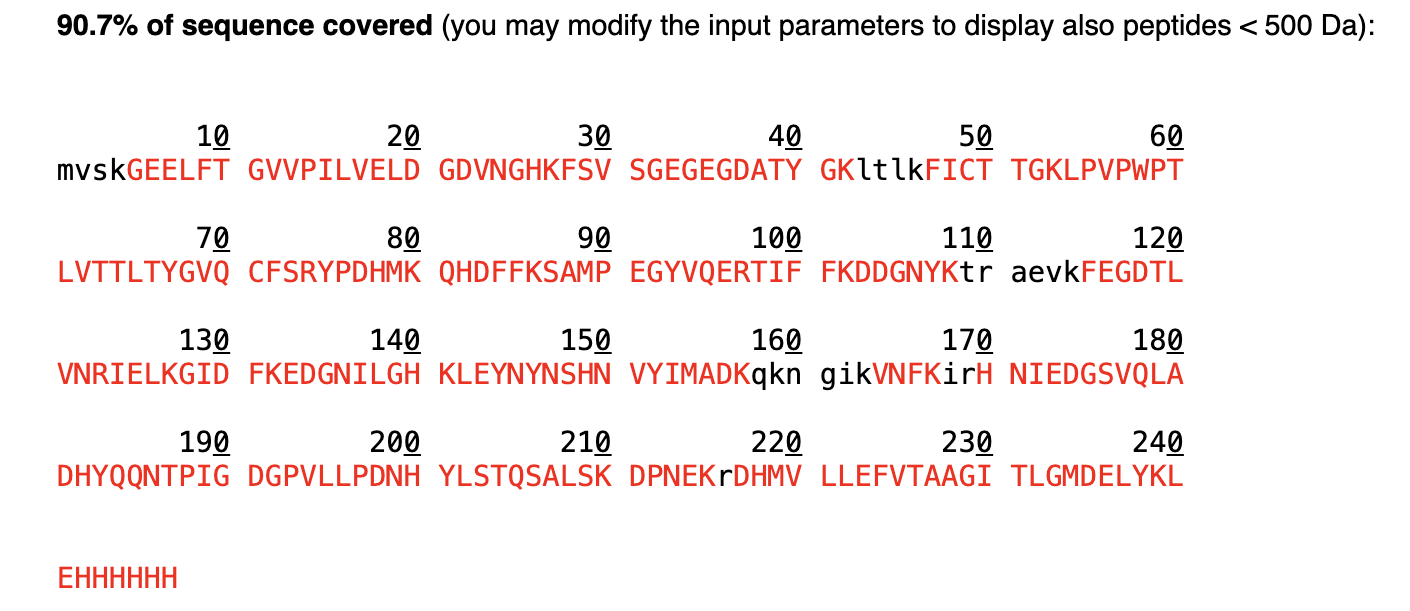
- Week 5 Lab: Protein Design Part II
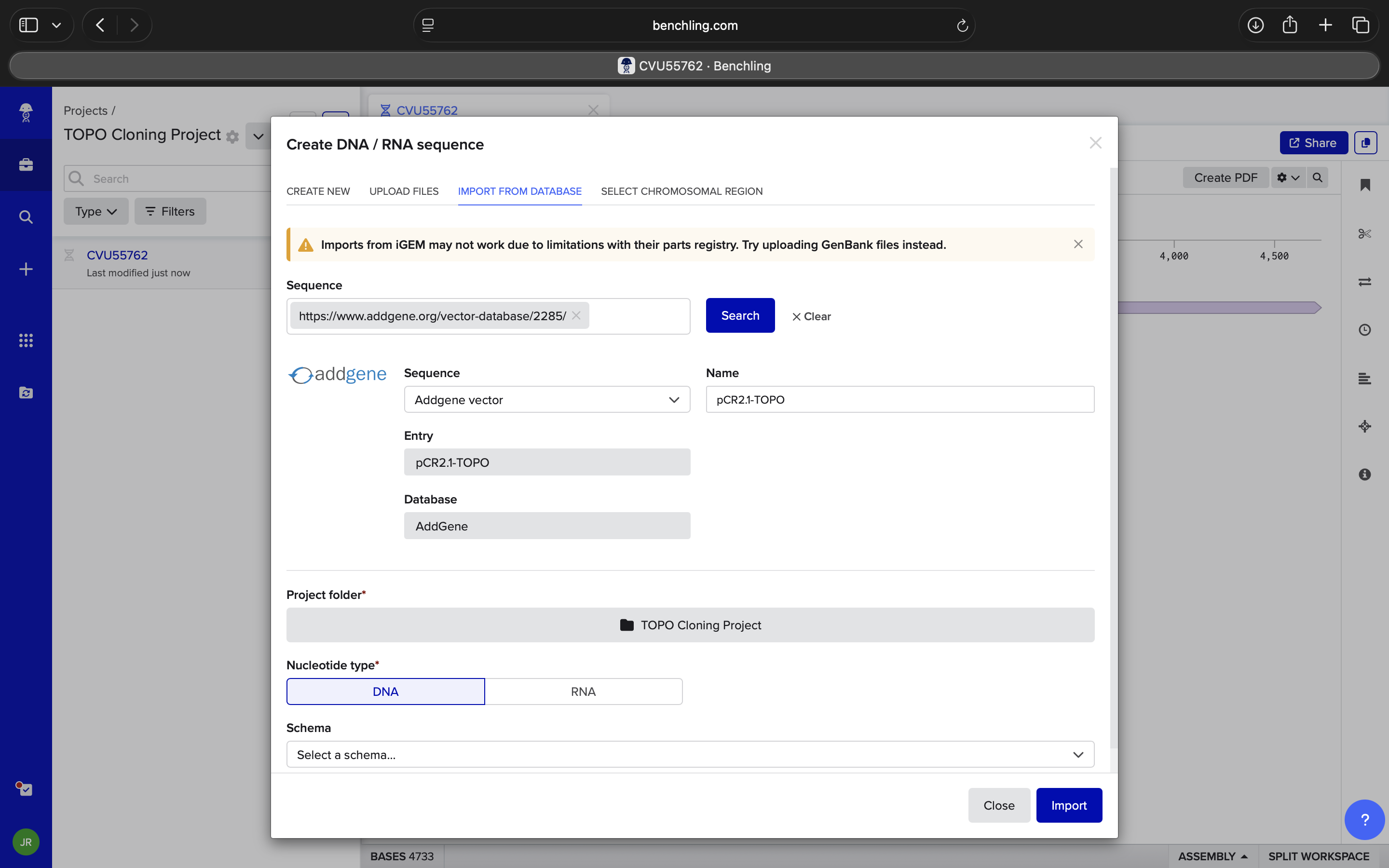
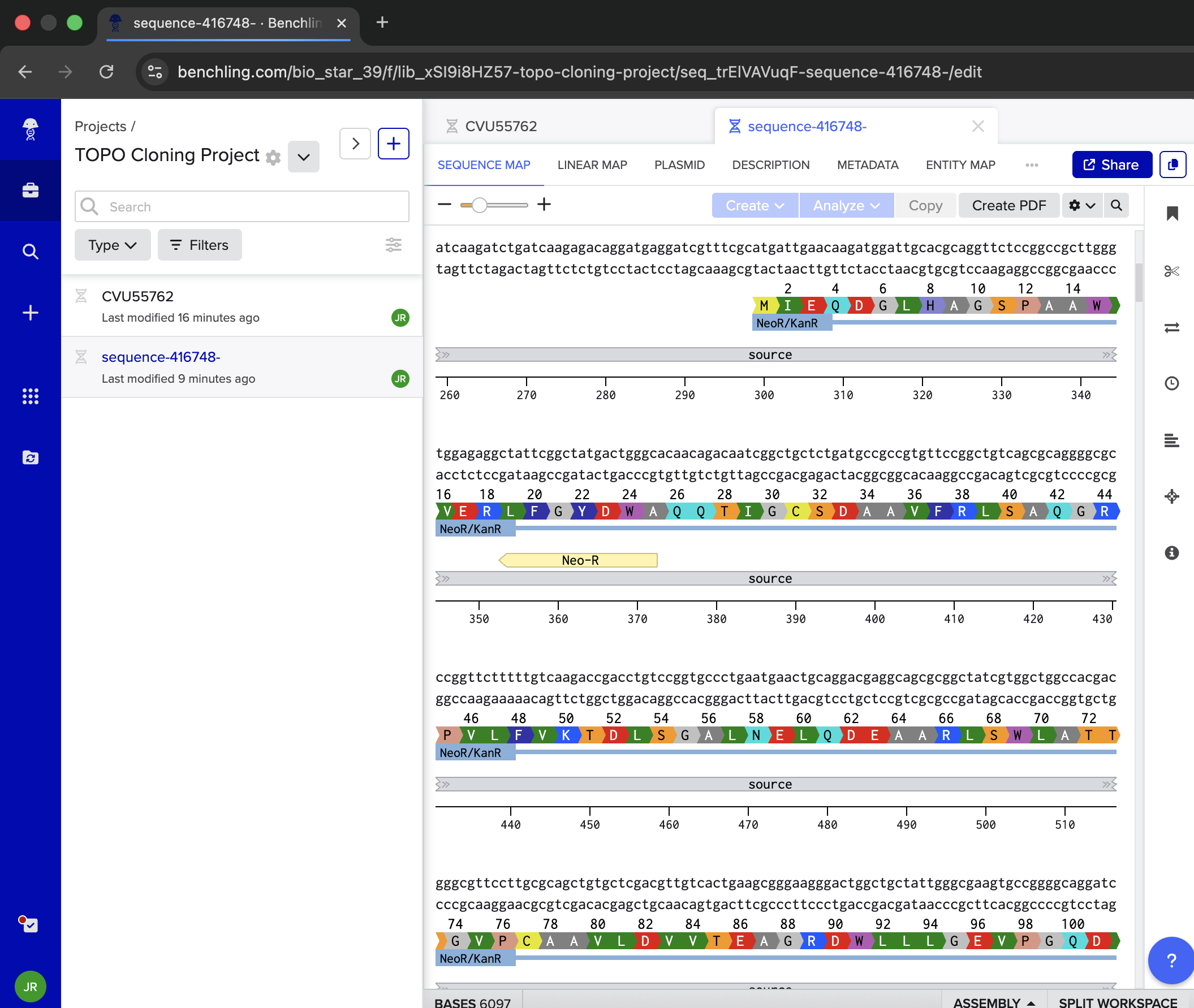
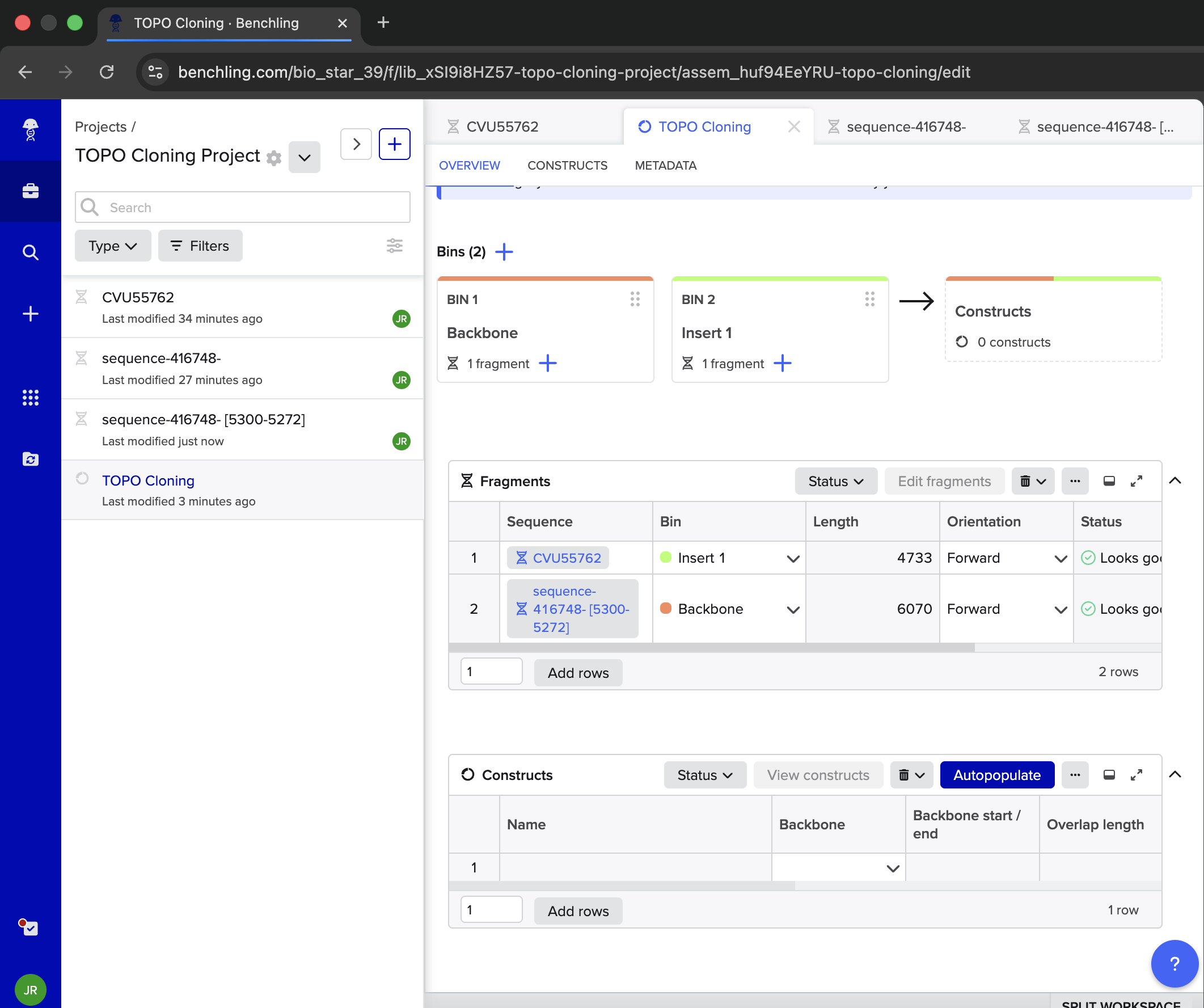
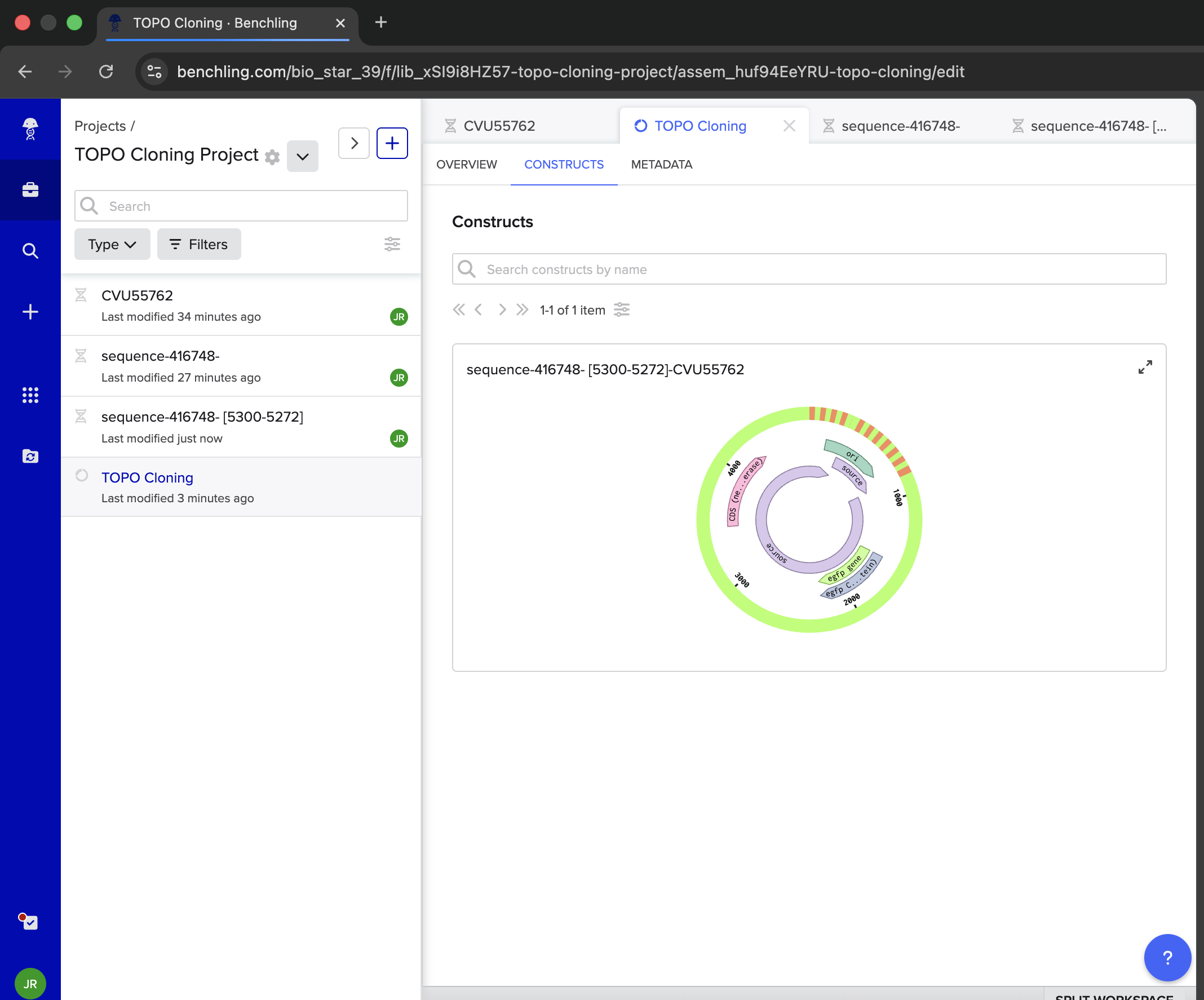
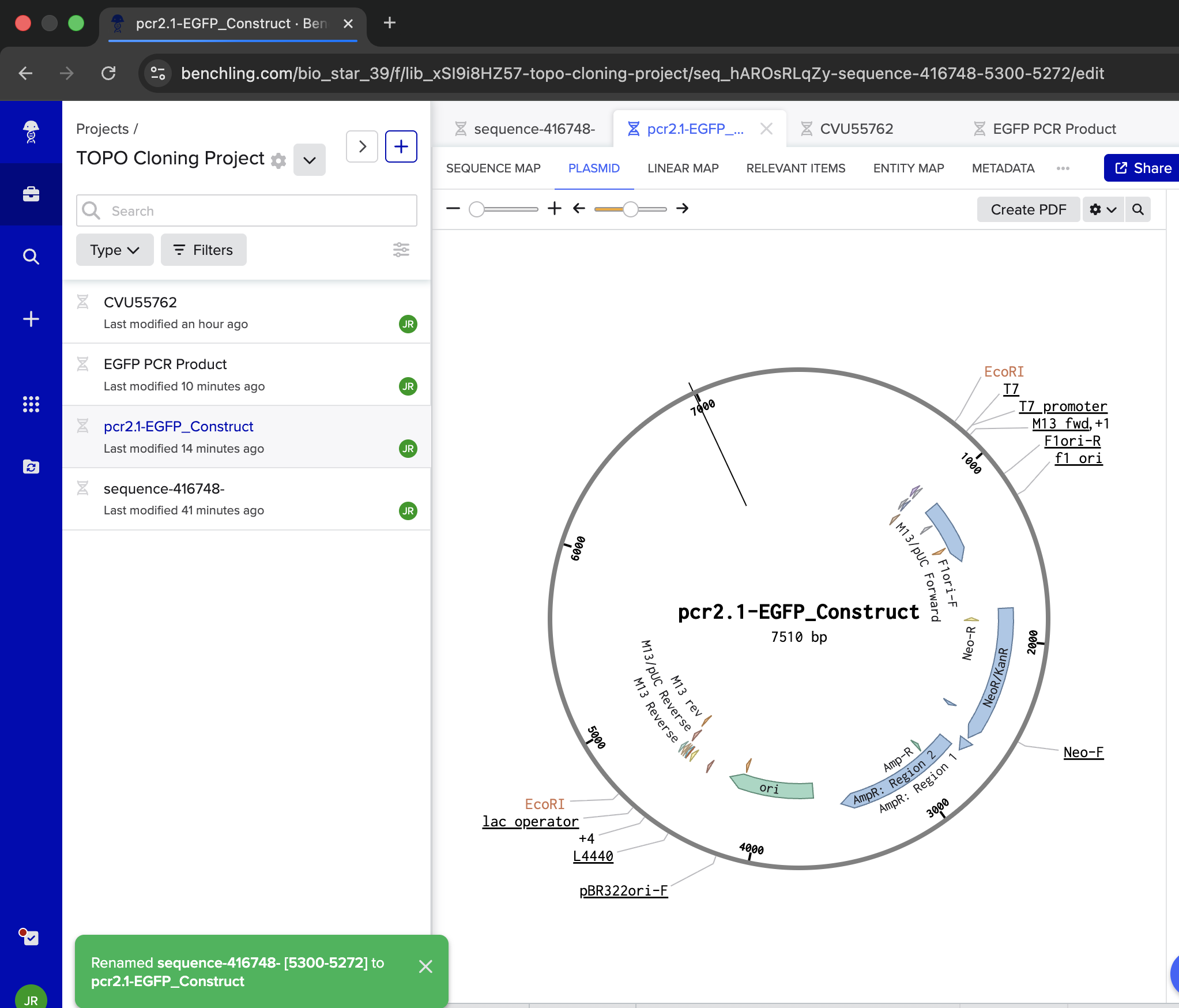
- Week 6 Lab: Gibson Assembly
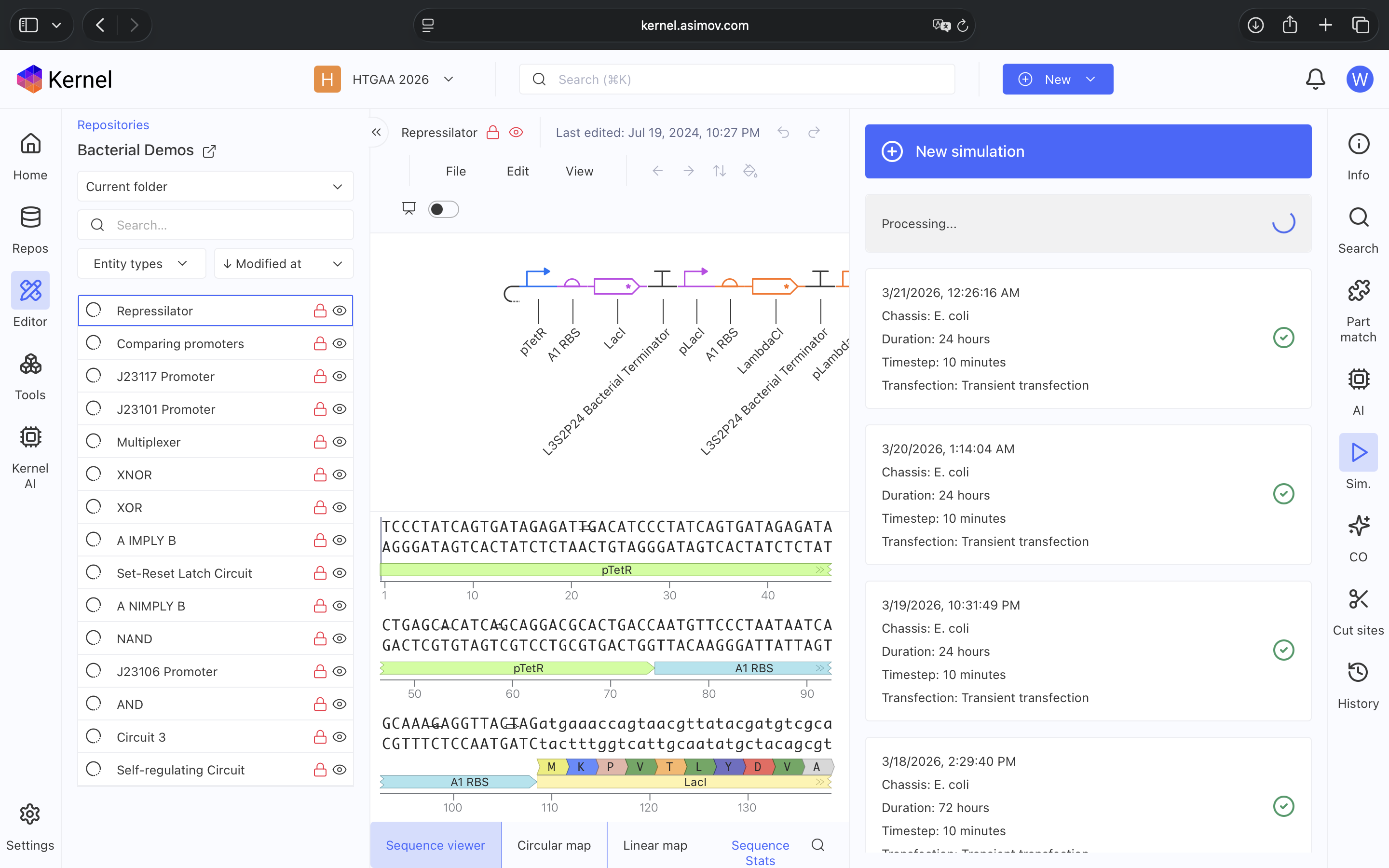
- Week 7 Lab: Neuromorphic Circuits
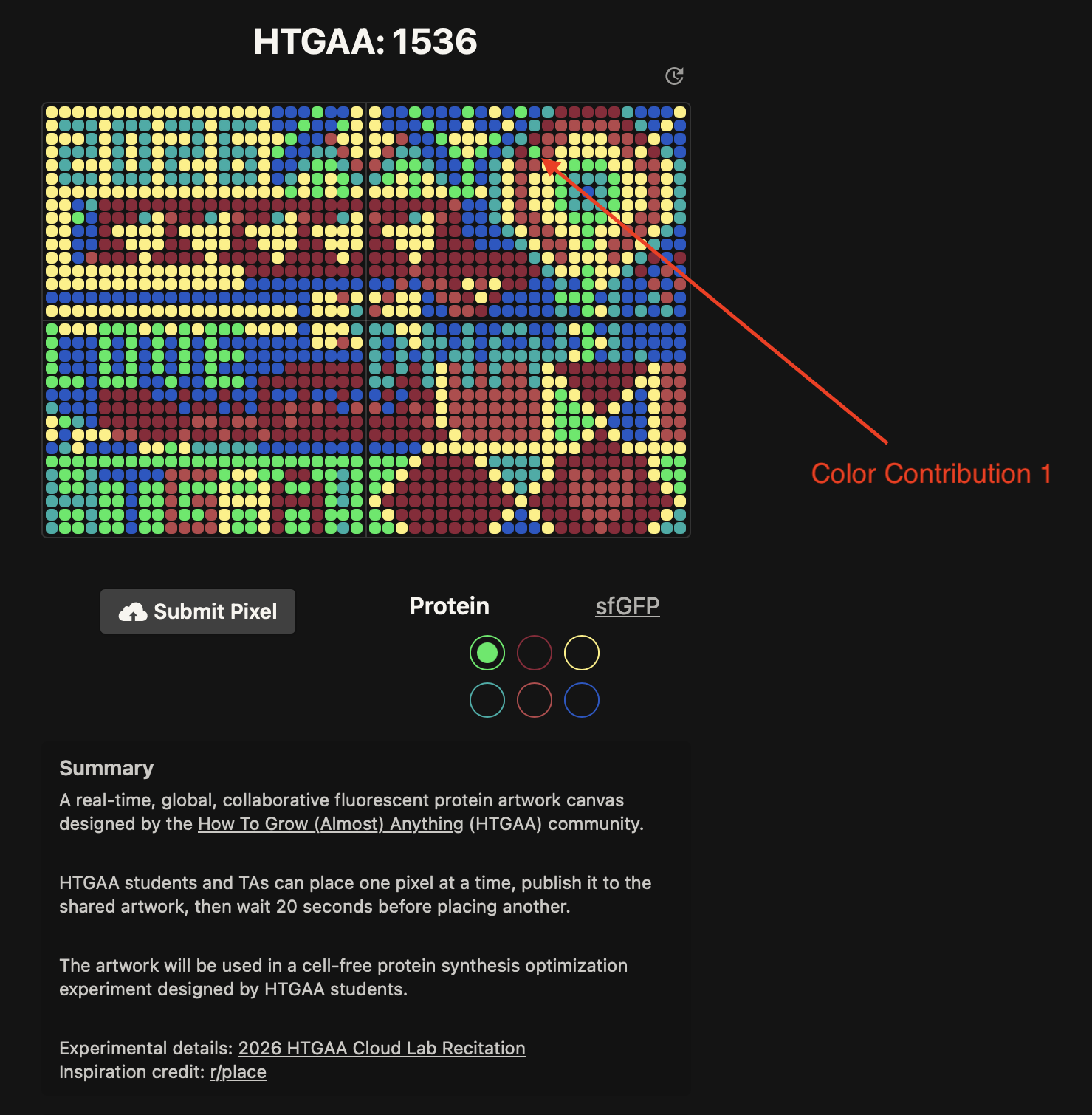
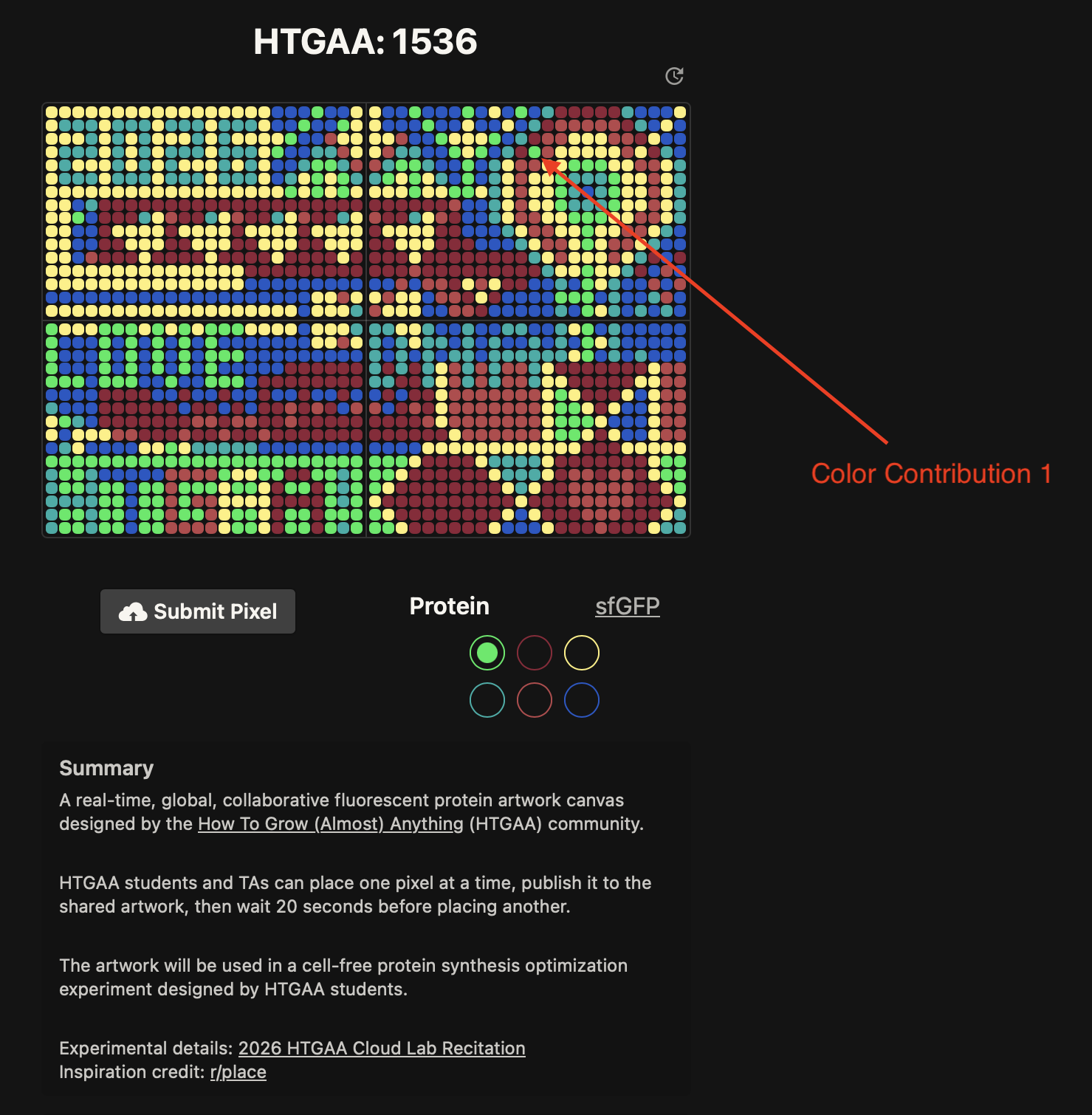
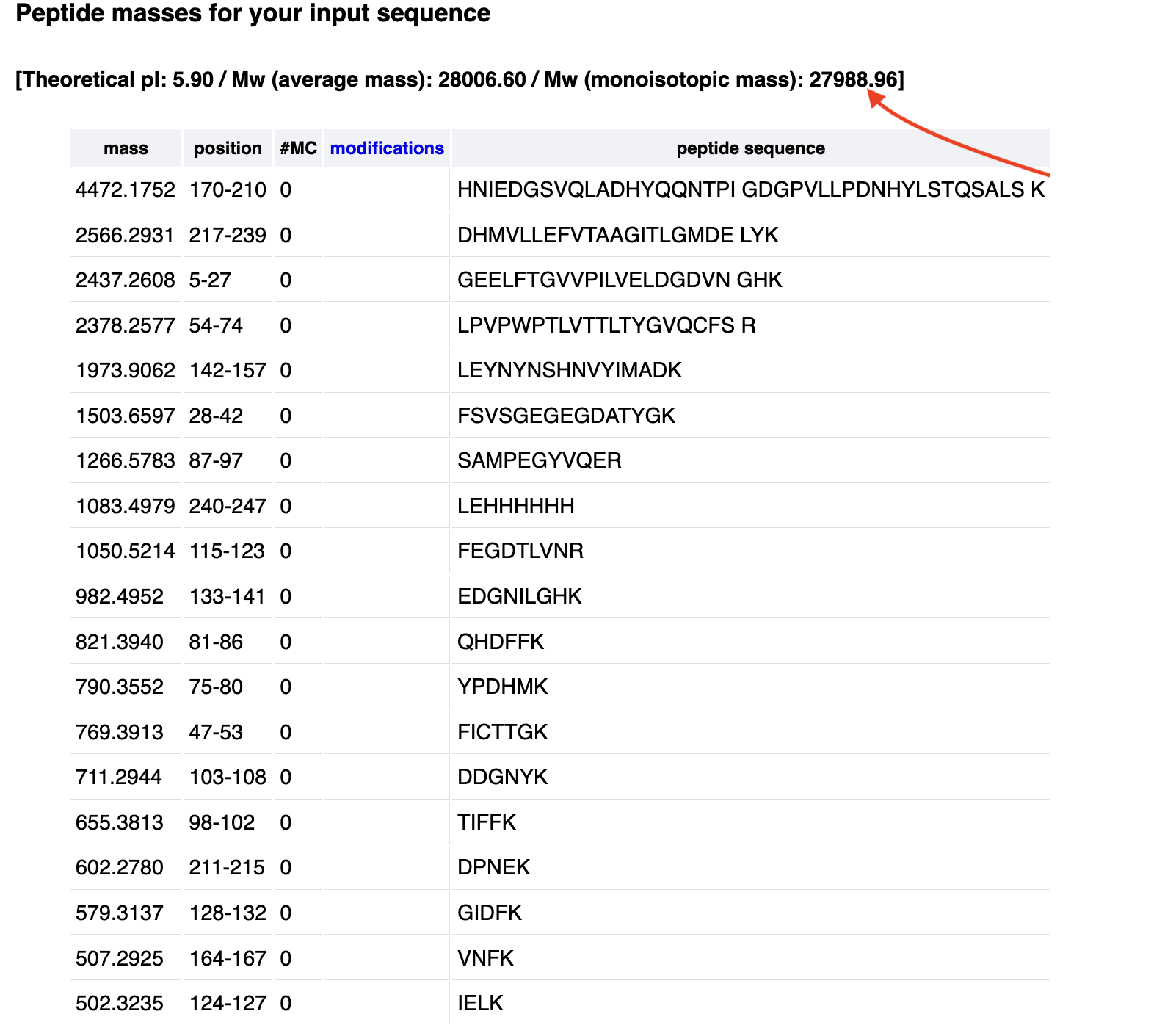
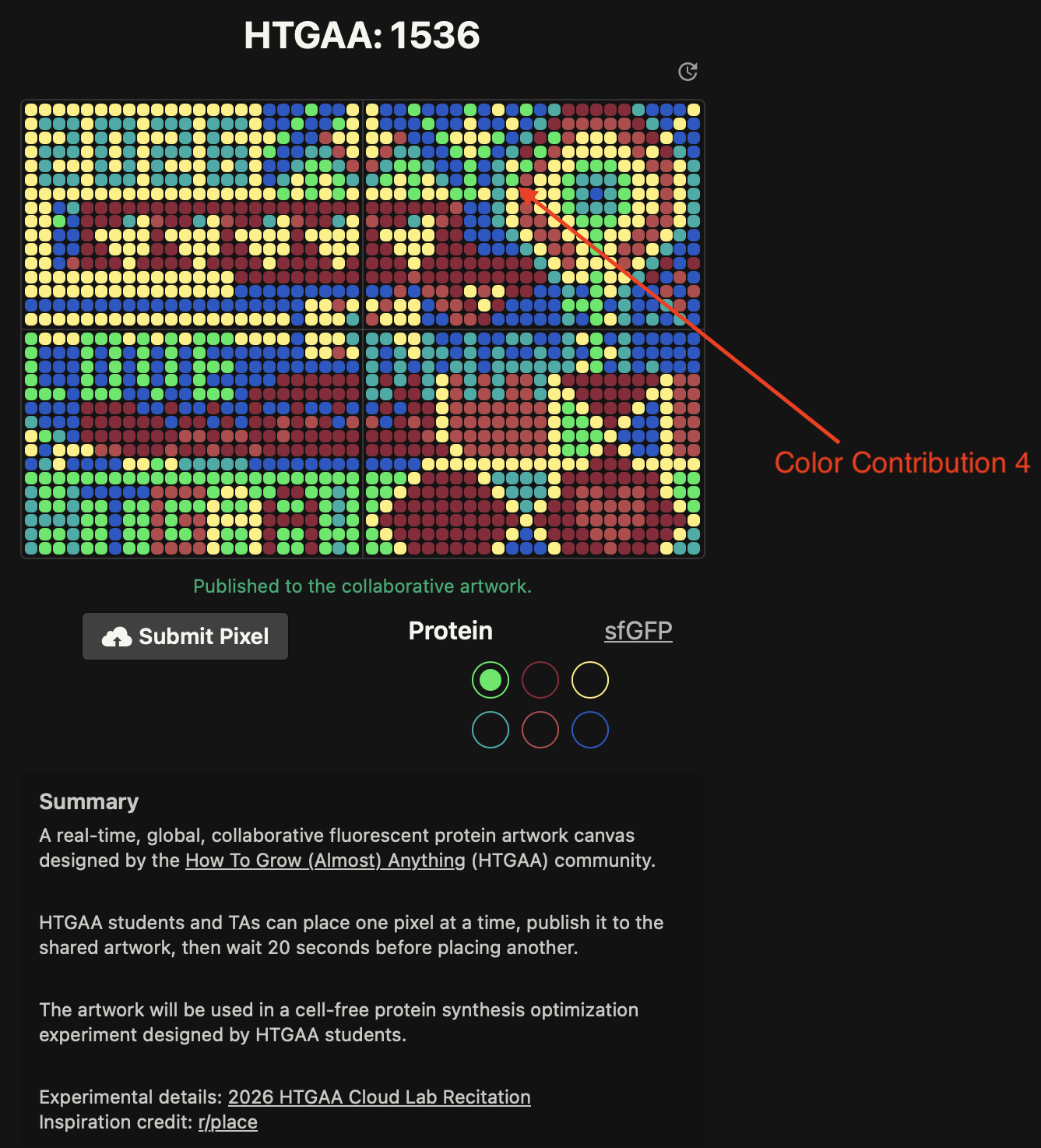
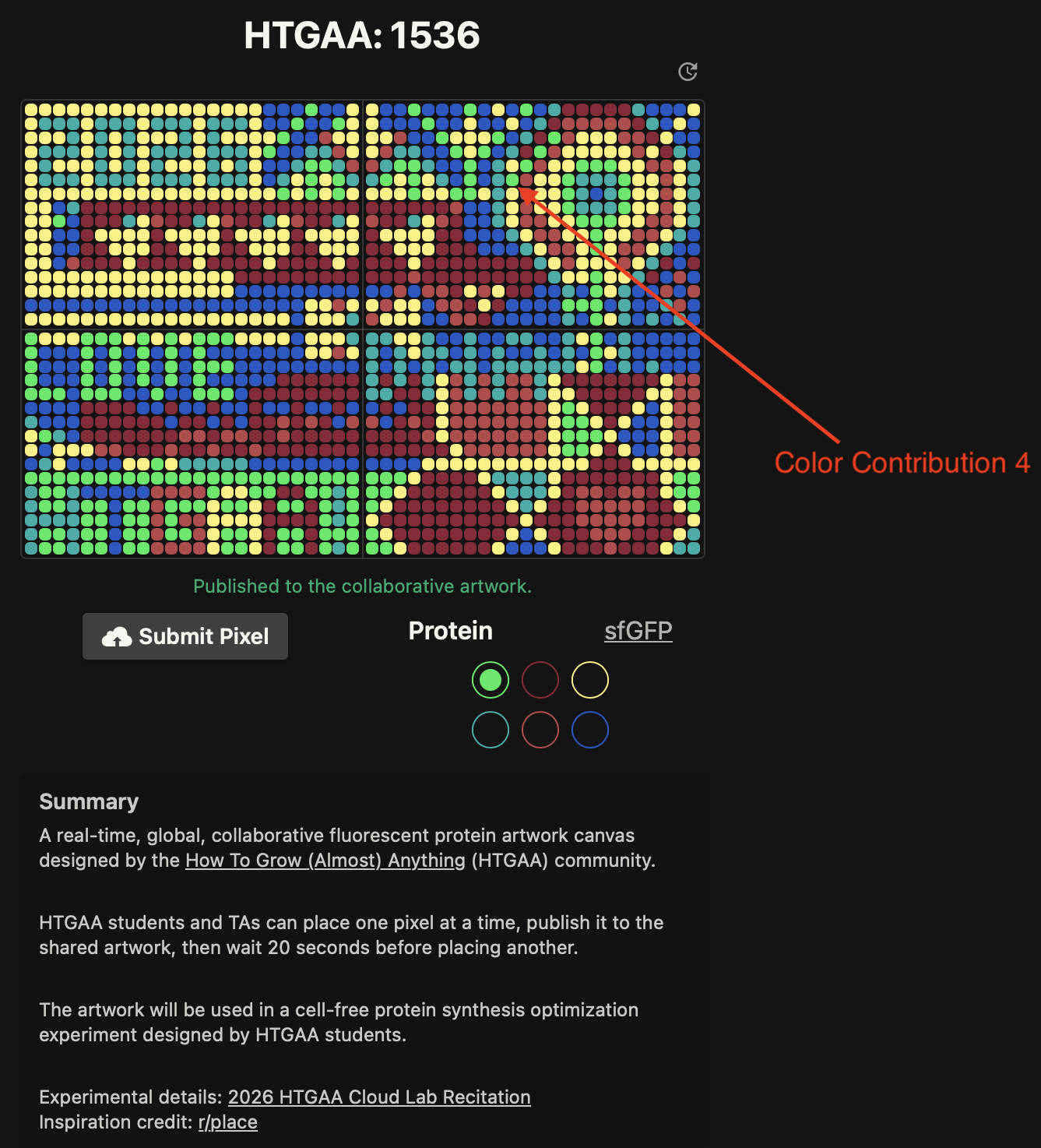
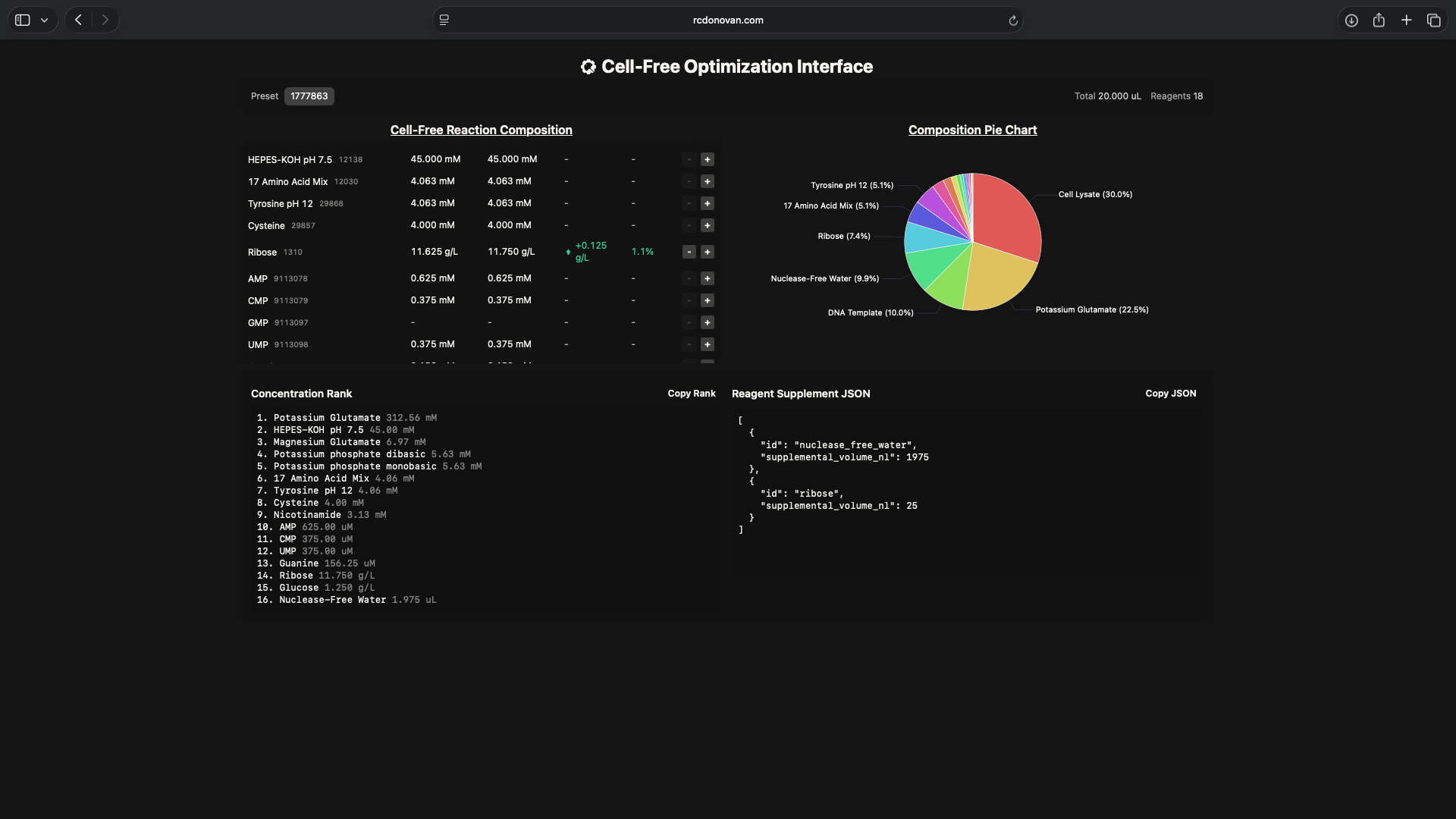
- Week 9 Lab: Cell-Free Systems


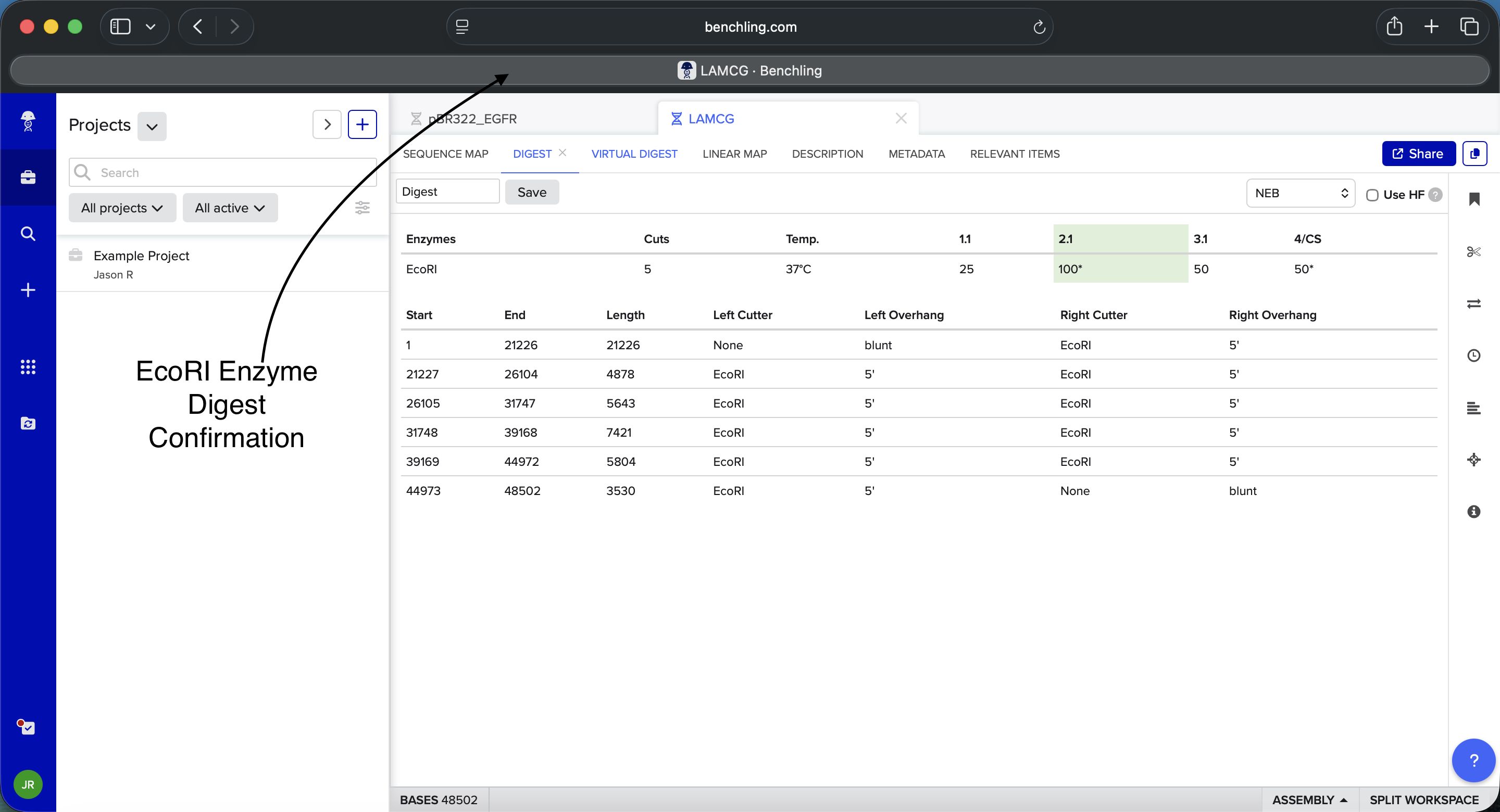
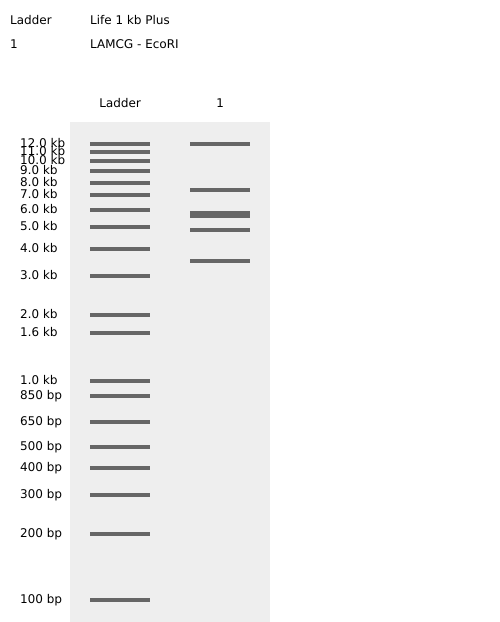
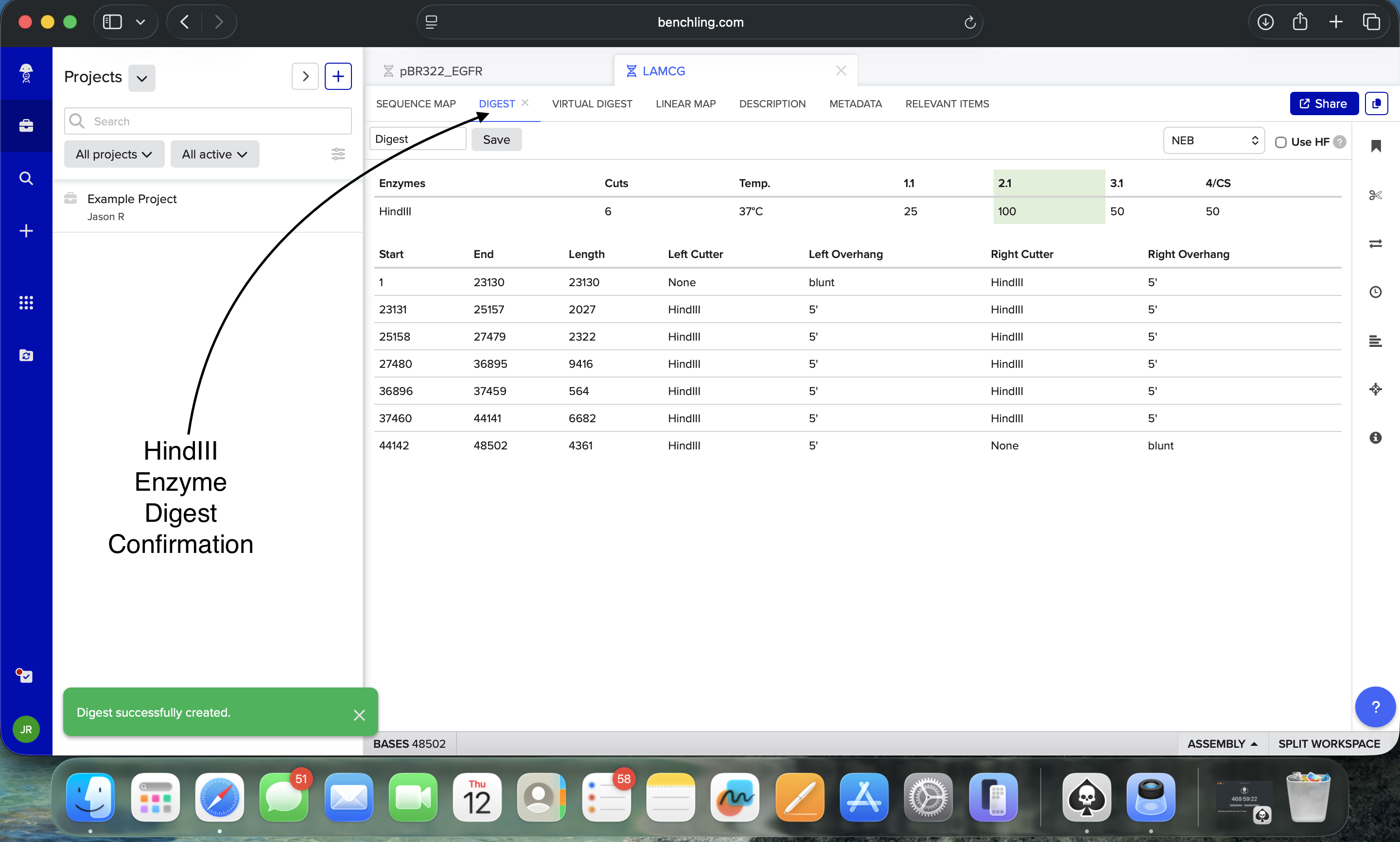
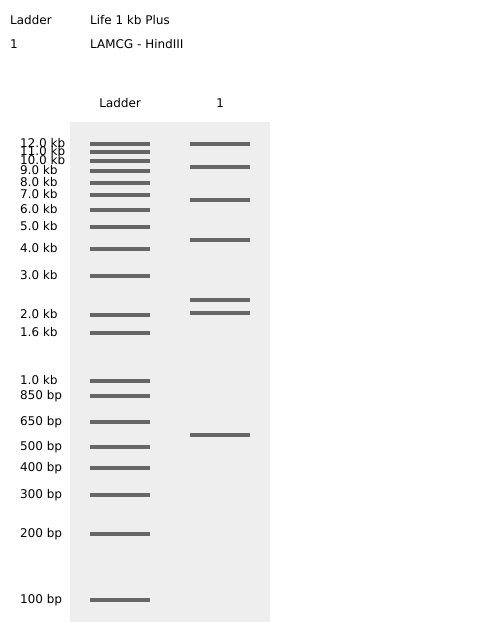


.png)
.png)
.png)
.png)
.png)
.png)
.png)

.png)
.png)
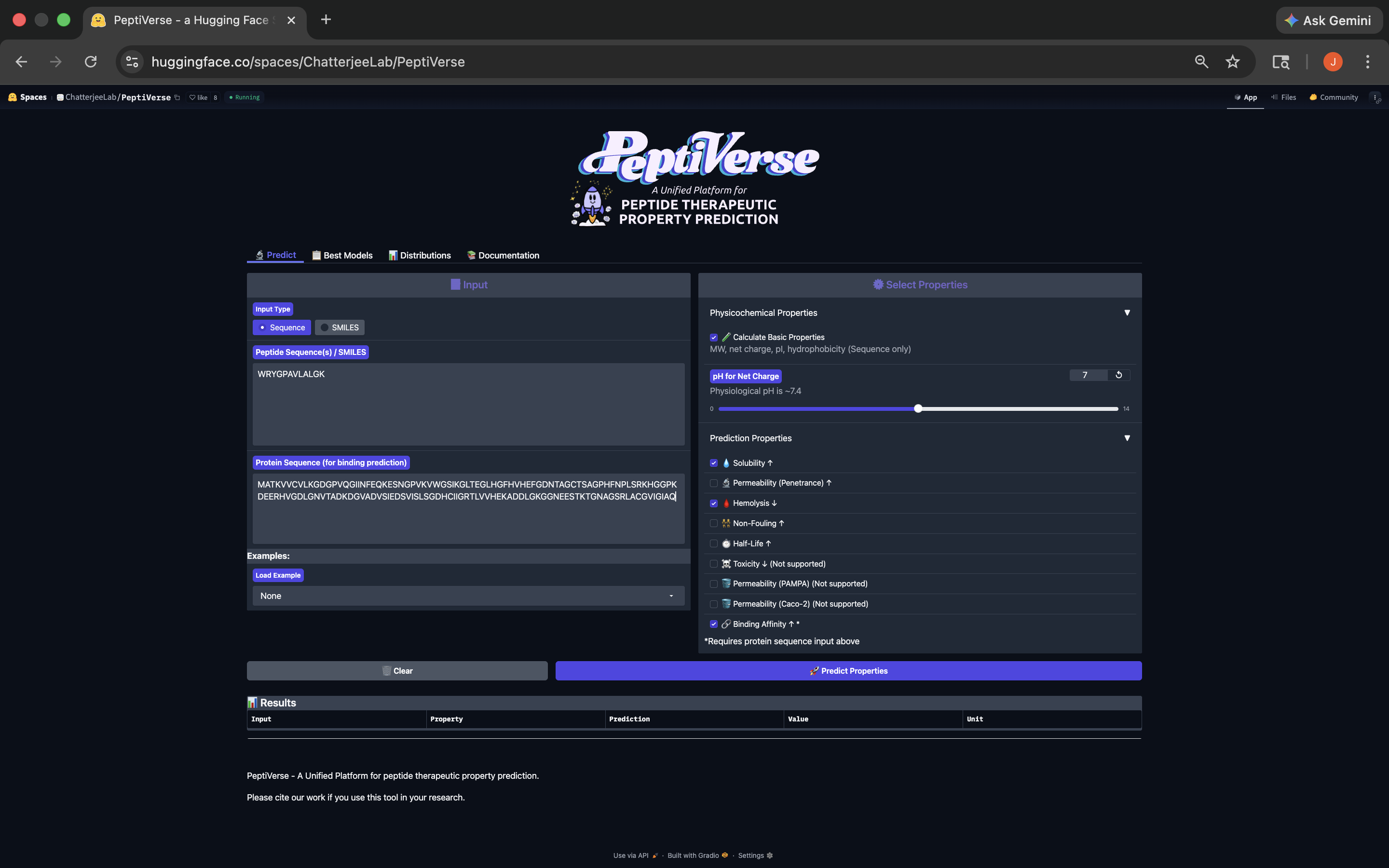
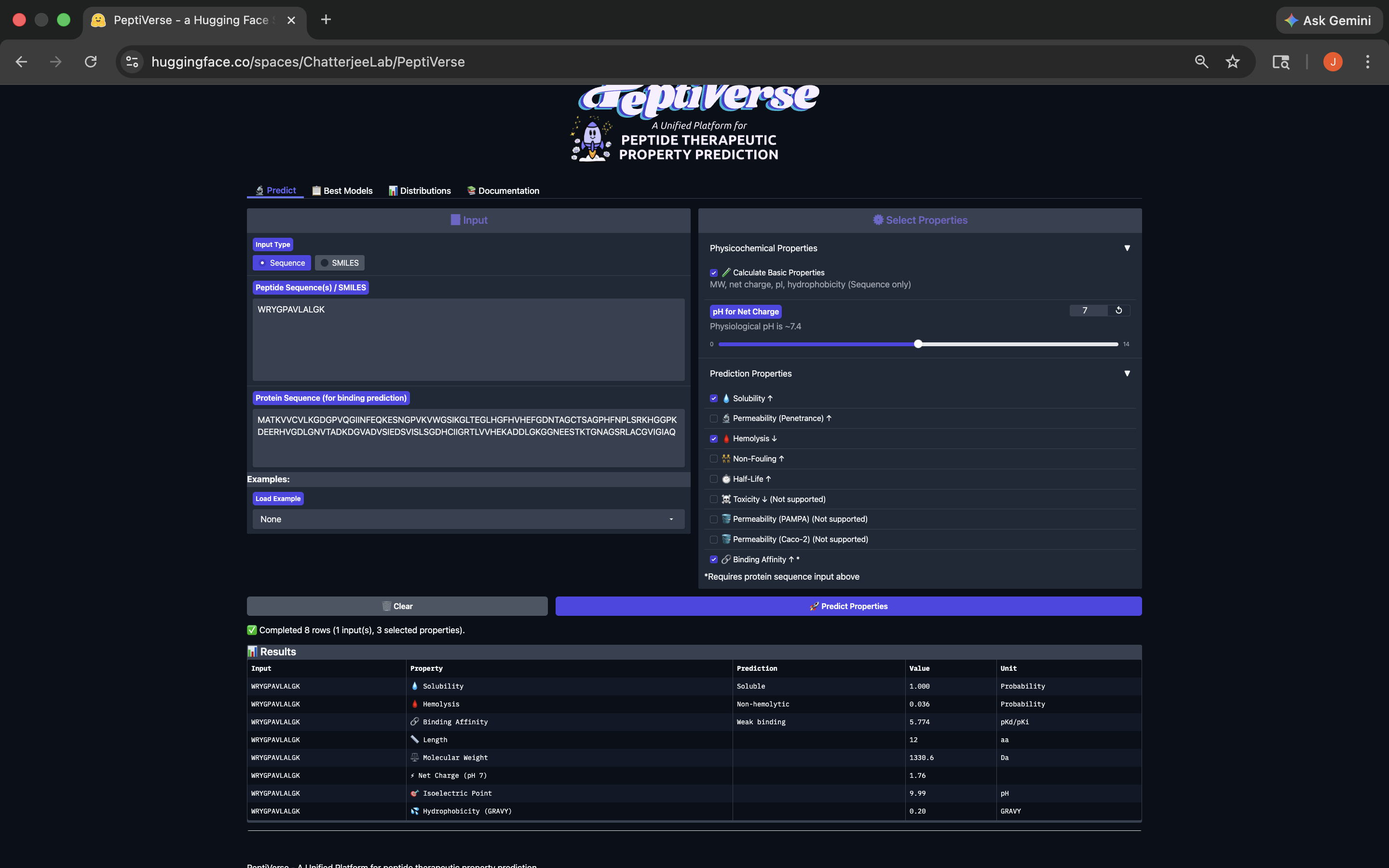
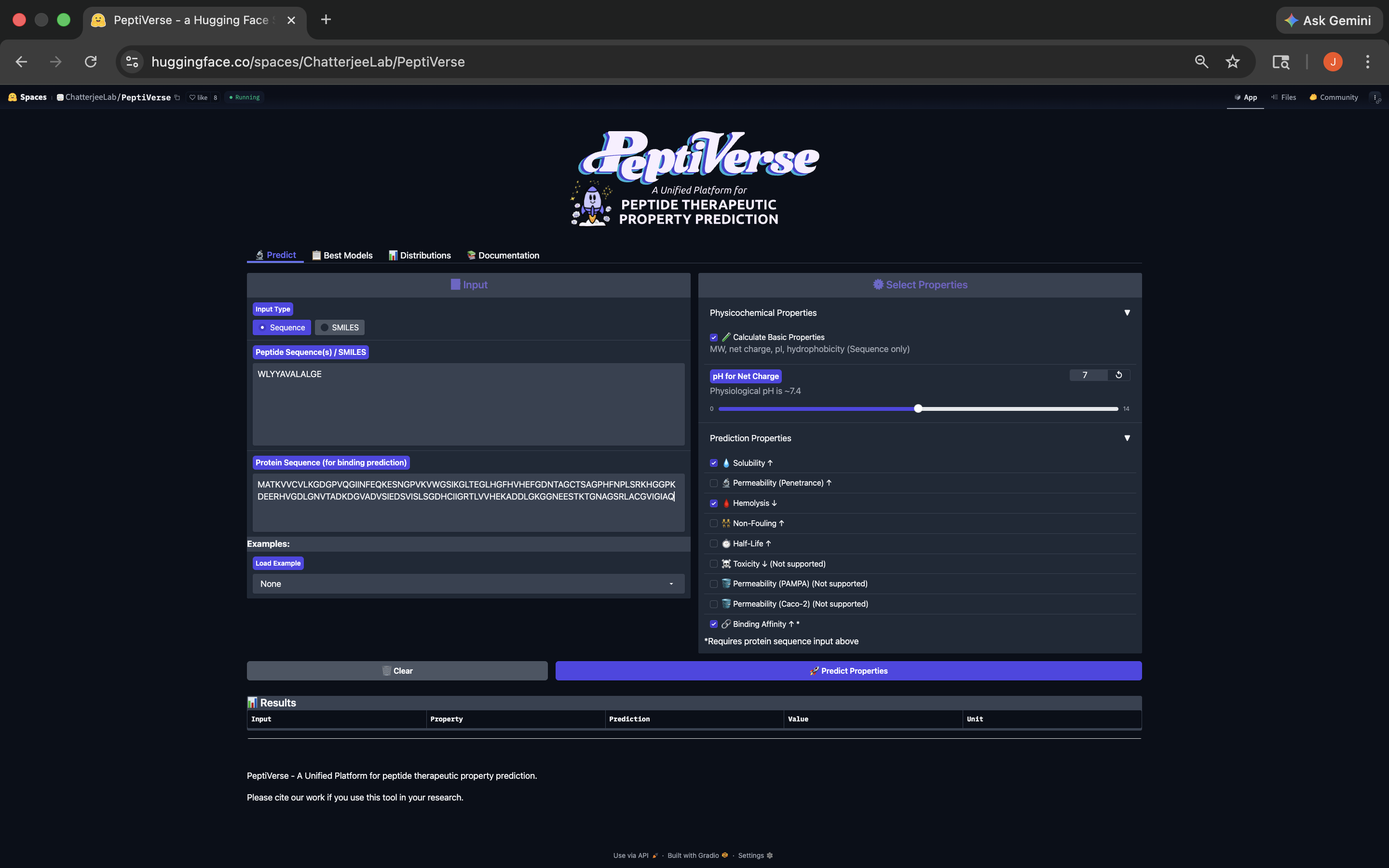
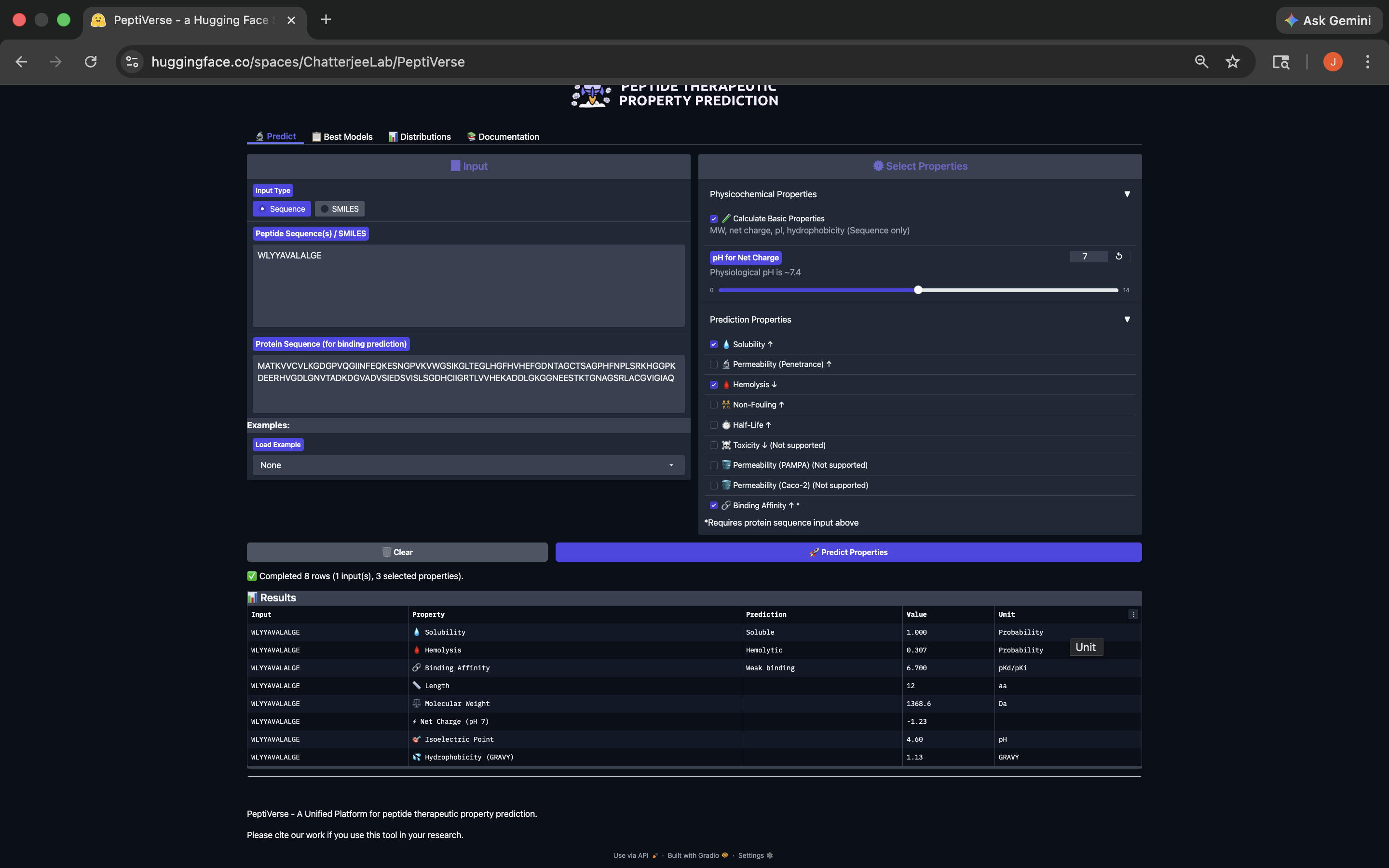
.png)














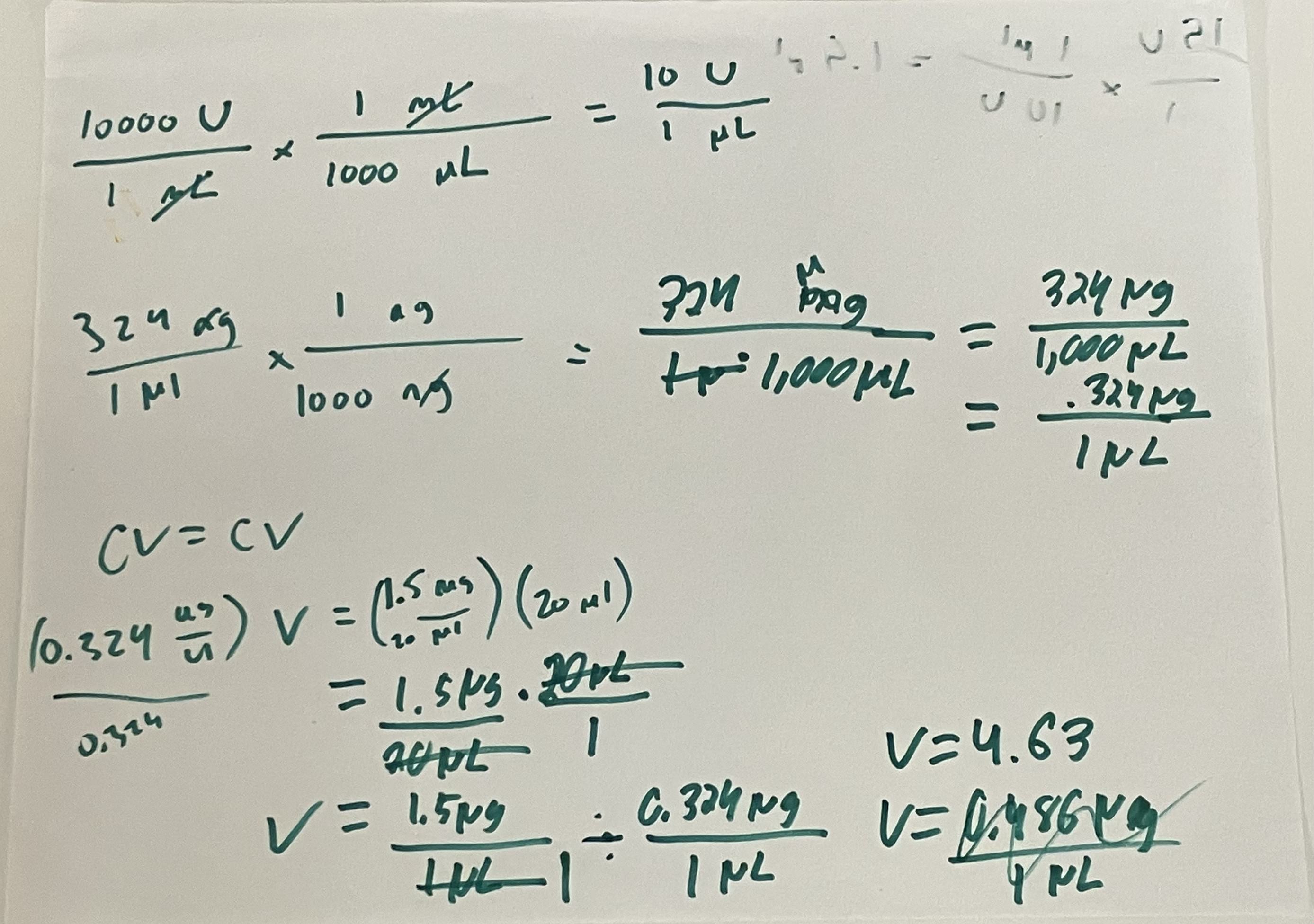










![virtual_digest_sequence_Mycobacterium-phage-Kampy-complete-sequence-51378-bp-including-10-base-3-overhang-CGGCCGGTAA-Cluster-A4_[HiddenHello]](/2026a/jason-ross/labs/week-02-lab-dna-gel-art/virtual_digest_sequence_Mycobacterium-phage-Kampy-complete-sequence-51378-bp-including-10-base-3-overhang-CGGCCGGTAA-Cluster-A4_[HiddenHello].png)