Week 3 — Lab Automation
Opentron Python Script Art
Basic Idea
The limited pixel resolution and colors of the petri dish reminded me of the old school bitmap monitors like the IBM PC that I grew up with. Also I wasn’t looking forward to guessing/figuring out a lot of pixel locations by hand, so I took a retro route and wrote some code to provide a terminal like API that let’s you specify a cursor location to write text to using a specific bitmap font and color.
Result
The code to turn the petri dish into a screen was pretty straight-forward to do even by hand. I did bunch of experiments of what I should do with the API and ended up making a logo for my local bio makerspace:
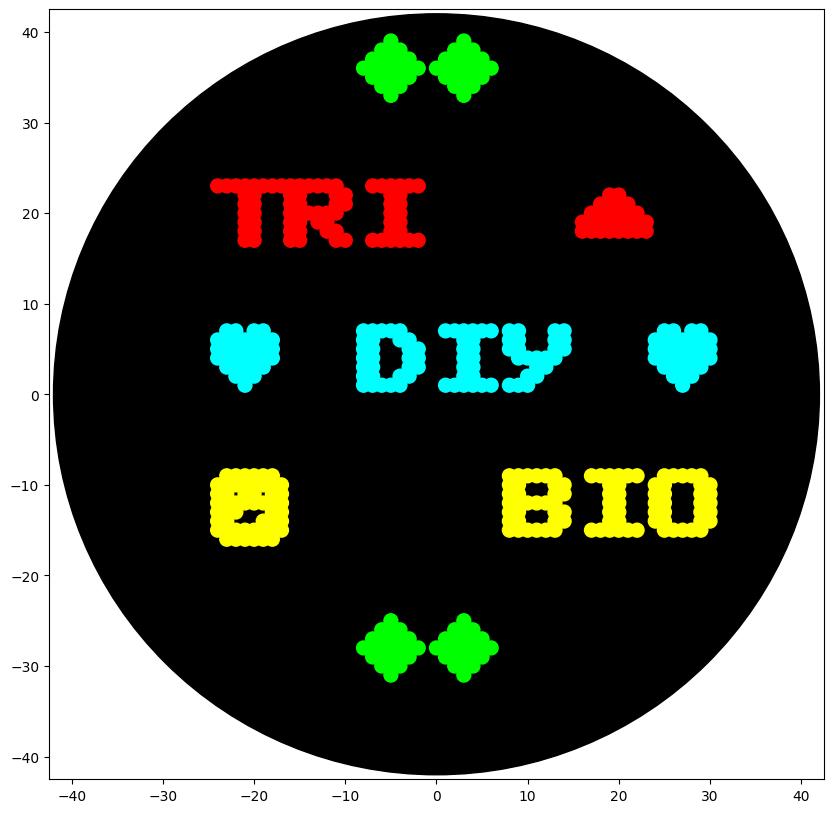

With the screen API drawing this log is only a few lines of python
How
I think AI could have done this really quickly but keeping with the retro theme I did things by hand in my colab notebook. Even by hand this wasn’t too bad and ended up being about 100 lines of python code for the logic.
- Encoded the bytes as dictionaries and binary numbers (so you can sort of see the shape), where highest value digit is upper leftmost digit
- Code supports screens with different bitmap sizes.
- The API uses (row, line) numbers to specify location like an old-fashioned terminal instead of (x,y).
- Since the petri dish is a circle I made (0,0) the center row and line and used negative numbers for rows to left and lines below the center
- The number of rows and lines a petri dish can fit depends on the size of the font and the physical pixel separation which are specified when the Petri Screen is created.
- I experiemented with a bunch of different pixel separations. I ended up using 1mm, but not how this will actually look in real life.
- The python code itself is NOT optimized for speed and uses string operations to manipulate the bytes.
- The use of the robot is optimized because the screen computes all of the pixels before rendering begins, so
- Uses minimal number of tips
- Uses minimal amount of fluid
- The move path is also near optimal because the tip is moved in a left to right scan for each color across the entire screen
There are a lot of sources for bitmap font data all of which use a different format. I went with class IBM 8x8 font data available in this projects header files. I converted this to my format using some simple search and replace and then running this python code outside of the the notebook.
Lab Run
I wasn’t able to make my node’s office hours and didn’t get a reply to if and how we are supposed to run my code, so I wasn’t able to actually run.
Post-Lab Questions
Find and Describe an Opentron Paper
I read this paper Environmental modulators of algae-bacteria interactions at scale. This paper explores the dependence of how autotraphs and hetrotrophs interact as a combined system under various environmental conditions. The space of variables for a two organism system under a variety of environmental conditions is large (~225K), so massively parallel automation is essential. The key technology in this space is to observe nano-liter size well droplets in a microfluidic system using fluorescent bar codes to identify the conditions in each droplet and to also fluroescent intensity to measure protein growth. Given the small scale, number of possibilities, and precision required the solutions used to create the droplets where mixed by an opentron-2 robot (STAR section page e2).
Lab Automation for Final Project
My main current proposed project is building a frugal benchtop bioreactor, which is more along the lines of trying to bring simple automation and sensor control to a wider audience. Given this it isn’t clear how I could use an opentron? Even if I do another project it also isn’t clear that as a remote comitted listener I would have access to either opentron or lab automation in a way that I could do a project?
Final Project Ideas
1. Frugal Benchtop
This was what I outlined in homework #1. Would try to include demo projects with e. coli and/or yeast that have been edited to express visible spectrum colors
2. Non-sterile simple genetic modification
As an extension of sourdough in to create a 2-species system that supports genetic modification of either microbe. Sourdough environment already selects against most competitors.
3. Edit yeast or algae to symbosis with HOB
Not even realy sure how to go about this because it is a massive edit.
Extra Node Specific Questions
I imagine AI could easily answer this question, but I intentially tried to just use the notes that were sent out and my basic understanding of howo things to work to see what I could guess without help.
1. If we were given a random segment of 100,000 bases, how would we determine if it is encoding for eukaryotic or prokaryotic genes? Could we find specific “parts” - promoters, operators, enhancers, silencers?
Assuming it is protein coding section of DNA I think you would have a decent chance of finding enough patterns to make good guesses about the structure and encodiongs
Prokaryote vs Eukaryote
Some things you could look for
- Circular vs linear
- Ribosome Binding Sequences
- Origin of Replications
Specific Parts
If you can identify patterns that you think are potential ori and RBS then you can use those guesses to orient on the strand and look for
- Between Ori and RBS sequence: Common promoters and operators for the domain you think it is
- After the RBS: Look for start codons and stop codons. Do the stop codons seem far enough away from start codon to encode a reasonable size protein.
- If you have multiple potential RBS sites and resonable stop codons, you can than iteratively look for more promoters and operators between stop codon and the next potential RBS.
2. Be able to explain why you chose your sequencing method for HW #2 and what other options you considered. Be able to explain how you would synthesize a particular piece of DNA - is it all in one piece, or are you assembling several synthesized parts together? What editing methods did you consider, and how would you confirm your DNA editing approach worked? Basically, be able to discuss your answer to this question.
Tried to justify in text of last week’s homework
3. What published paper using Opentrons are you analyzing for HW #3? In discussing with your peers, have researchers generally approached this tool for similar uses, or for vastly different fields?
See above.