Week 6 — Genetic Circuits Part I: Assembly Technologies
DNA Assembly
- What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
- Endonucleases to eat away at the ends of all the DNA strands to expose single stranded ends
- DNA Ligase to take matching ends and stitch them together
- What are some factors that determine primer annealing temperature during PCR?
- One factor is the proportion of G and C vs A and T because G and C have 3 hydrogren points they require more energy/higher temperature to break than the 2 hydrogen bonds linking A and T
- 3 dimensional structure like hairpins can also have an impact on the temper
- There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
- Restriction enzymes cut the DNA into a linear piece and most of them leave sticky ends which can be joined together by DNA ligase directly after they are cut which is simple because you can cut and join at the same time. The downside is that enzymes have specific recognition sites that you must use and engineer around. These sites are not that long and are often palindromic, which increases the chance of cuts where you don’t want them. On the other hand, the very fact that they recognition sites are limited makes standard like BioBrick and MoClo possible.
- PCR uses custom primers and to walk the DNA in two different directions during the amplification cycles.
- The primers are longer than most recognition sites and can be almost anything. This gives you a lot of flexiblity to custom match your DNA without having to look at specific sites.
- How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
- You need to make sure that your primer sequences are unique
- You need to make sure your primer sequences line up so that the pieces get glued together in the order you want.
- How does the plasmid DNA enter the E. coli cells during transformation?
- Describe another assembly method in detail (such as Golden Gate Assembly)
- Is a restriction enzyme method
- Uses Type IIS enzymes which cuts to one side of the recognition site, in parituclar it doesn’t destroy the recognition site when it cuts.
- Because the recognition site stays intact you can make sure that the piece you want to glue to doesn’t have the recognition site. This means that if your DNA glues back to the piece it was just cut from in stead of the piece you want it to glue to it will again be a target for the Type IIS enzyme that is being used.
Asimov Kernel Modeling
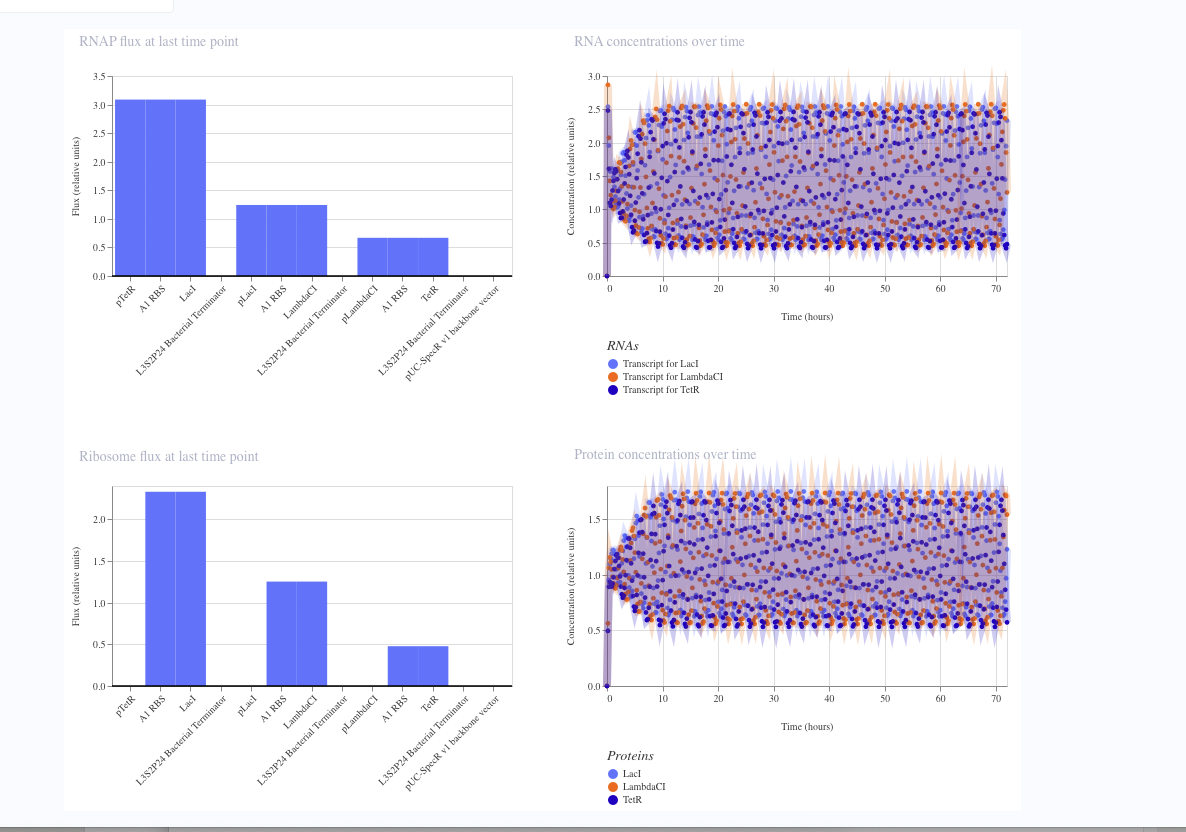
I simulated a represillator in Aismov and got following results: