Week 5 HW: Protein Design part II
PART A: SOD1 Binder Peptide Design
The sequence for the original protein is:
// sp|P00441|SODC_HUMAN Superoxide dismutase [Cu-Zn] OS=Homo sapiens OX=9606 GN=SOD1 PE=1 SV=2 MATKAVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTS AGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVV HEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ
Mutation occurs at residue 4: Alanine becomes Valine
// 1UXM_1|Chains A, B, C, D, E, F, G, H, I, J, K, L|SUPEROXIDE DISMUTASE [CU-ZN]|HOMO SAPIENS (9606) ATKVVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVS IEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ
Part 1: Generate Binders with PepMLM
Adding Methionine to the mutated sequence, the generated peptides were:
| Nno. Sequence | Binder | Perplexity |
|---|---|---|
| 1 | WRYYVAVVRWKK | 28.003177 |
| 2 | WRYYAAVLEWKE | 16.368043 |
| 3 | WHSYAVVLEWWK | 19.985691 |
| 4 | WLSGPVAVEWKK | 11.674838 |
Comparing to the binding protein, FLYRWLPSRRGG, all of them are quite different. This might be due to the input parameters of the code as well as the fact that the generation is random, which contributes to a higher degree of perplexity.
Part 2: Evaluate Binders with AlphaFold3
The following figures show how the SOD1 protein interacts with each of the four binders:
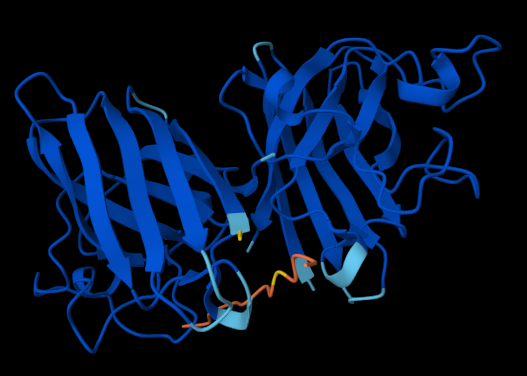
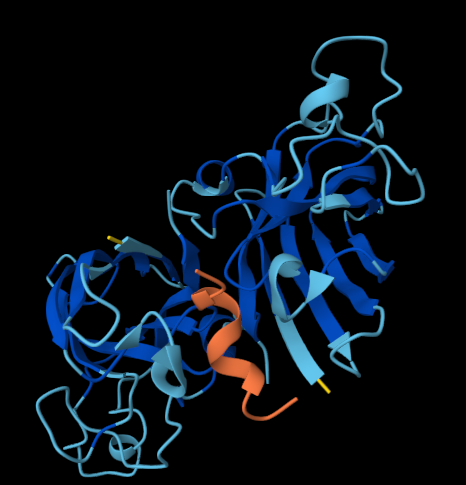
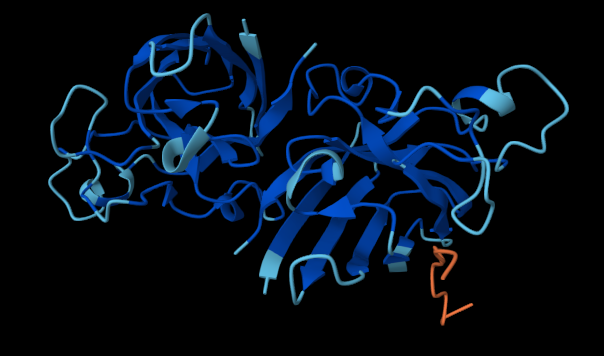
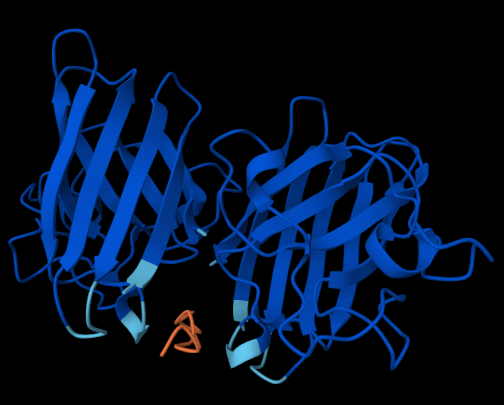
The website also provided the following scores:
| No. Peptide | Sequence | ipTM score | pTM score |
|---|---|---|---|
| 1 | WRYYVAVVRWKK | 0.78 | 0.85 |
| 2 | WRYYAAVLEWKE | 0.64 | 0.77 |
| 3 | WHSYAVVLEWWK | 0.74 | 0.83 |
| 4 | WLSGPVAVEWKK | 0.73 | 0.83 |
Part 3: Evaluate Properties of Generated Peptides in the PeptiVerse
| No. Peptide | Sequence | Solubility | Hemolysis | Binding Affinity | Molecular Weight | Net charge @ pH = 7 | Isoelectric Point | Hydrophobicity (GRAVY) |
|---|---|---|---|---|---|---|---|---|
| 1 | WRYYVAVVRWKK | Soluble | Non-hemolytic (0.066) | Medium (7.377 pKd/pKi) | 1654.0 Da | 3.76 | 10.45 | -0.57 |
| 2 | WRYYAAVLEWKE | Soluble | Non-hemolytic (0.057) | Weak binding (6.837 pKd/pKi) | 1613.8 Da | -0.23 | 6.28 | -0.68 |
| 3 | WHSYAVVLEWWK | Soluble | Non-hemolytic (0.090) | Weak binding (6.133 pKd/pKi) | 1603.8 Da | -0.15 | 6.75 | -0.12 |
| 4 | WLSGPVAVEWKK | Soluble | Non-hemolytic (0.039) | Weak binding (5.301 pKd/pKi) | 1399.6 Da | 0.76 | 8.59 | -0.16 |
Part 4: Generate Optimized Peptides with moPPIt
The generated binders are shown in the following table:
| Binder | Hemolysis | Solubility | Half-Life | Affinity | Motif | Specificity |
|---|---|---|---|---|---|---|
| EAVEGLTAEQIW | 0.95 | 0.5 | 9.92 | 5.91 | 0.34 | 0.61 |
| WIIWVTTTKAQK | 0.94 | 0.5 | 5.66 | 5.77 | 0.77 | 0.67 |
| ITLDEWLKKQCY | 0.88 | 0.67 | 14.46 | 7.16 | 0.78 | 0.57 |
| TDEQKVQLSAYW | 0.84 | 0.67 | 5.64 | 6.41 | 0.66 | 0.53 |
PART C: Final Project: L-Protein Mutants
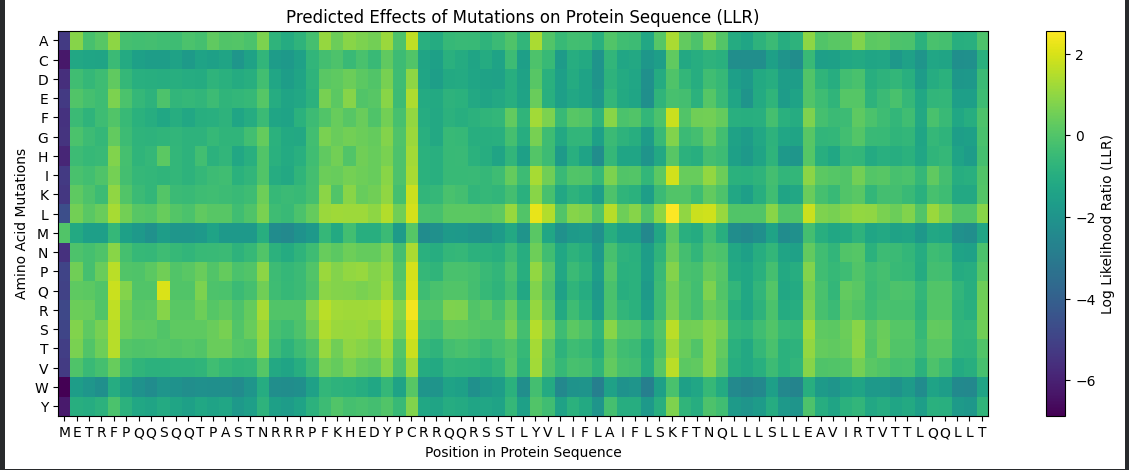
The programs were run and the heatmap along with some mutation candidates were obtained:
| Amino Acid | Position in Protein | Amino Acid in Protein Sequence | Score |
|---|---|---|---|
| L | 50 | K | 2.56 |
| R | 29 | C | 2.39 |
| L | 39 | Y | 2.24 |
| S | 29 | C | 2.04 |
| Q | 9 | S | 2.01 |
| Q | 29 | C | 1.99 |
| P | 29 | C | 1.97 |
| L | 29 | C | 1.96 |
| I | 50 | K | 1.92 |
| L | 53 | N | 1.86 |