Week 10 HW: Advanced Imaging & Measurement Technology
Please identify at least one (ideally many) aspect(s) of your project that you will measure. It could be the mass or sequence of a protein, the presence, absence, or quantity of a biomarker, etc.
The main aspect to be measured is the expression and activity of the biosynthetic gene cluster (BGC). This includes:
- Presence and expression of BGC-associated enzymes
- Production of candidate metabolites
- Antibacterial activity against Leptospira
Please describe all of the elements you would like to measure, and furthermore describe how you will perform these measurements.
- BGC enzyme expression: Measured using Western blot to confirm protein presence and approximate expression levels.
- Metabolite production: Measured using LC-MS to detect and quantify candidate compounds produced by the BGC.
- Antibacterial activity: Evaluated through antibiograms to assess inhibition of Leptospira growth.
What are the technologies you will use (e.g., gel electrophoresis, DNA sequencing, mass spectrometry, etc.)? Describe in detail.
- Western blot: To detect specific proteins encoded by the BGC after separation by gel electrophoresis.
- Gel electrophoresis: For protein separation prior to blotting.
- LC-MS (Liquid Chromatography–Mass Spectrometry): Main analytical technique to identify and quantify metabolites based on retention time and mass-to-charge ratio.
- Antibiogram assays: To determine the antibacterial effectiveness of produced compounds.
PART 1
Based on the predicted amino acid sequence of eGFP (see below) and any known modifications, what is the calculated molecular weight? You can use an online calculator like the one at https://web.expasy.org/compute_pi/
MVSKGEELFTG VVPILVELDG DVNGHKFSVS GEGEGDATYG KLTLKFICTT GKLPVPWPTL VTTLTYGVQC FSRYPDHMKQ HDFFKSAMPE GYVQERTIFF KDDGNYKTRA EVKFEGDTLV NRIELKGIDF KEDGNILGHK LEYNYNSHNV YIMADKQKNG IKVNFKIRHN IEDGSVQLAD HYQQNTPIGD GPVLLPDNHY LSTQSALSKD PNEKRDHMVL LEFVTAAGIT LGMDELYKLE HHHHHH
Based on the website provided, the molecular weight is 26.94 kDa, which does not consider the linker and His tag. If we consider them, the new MW will then be 28 kDa. The former value is consistent with other GPFs from other databases such as Q9U6Y4 (26.17 kDa), P42212 (26.89 kDa) and Q9GZ28 (25.91 kDa).
Based on the predicted amino acid sequence of eGFP (see below) and any known modifications, what is the calculated molecular weight? You can use an online calculator like the one at https://web.expasy.org/compute_pi/
The expression is:
$ z = \frac{m/z_{n+1}}{\left(\frac{m}{z_n} - \frac{m}{z_{n+1}}\right)} $Then, considering peaks such as 800.5508 and 824.0635, $z$ is equal to 34.047.
Then, the molecular weight is given by:
$ MW = z \cdot \left(\frac{m}{z_n} - 1\right) $ $ MW = 34.047 \cdot (824.0635 - 1) = 28{,}023.329 \ \text{Da} = 28.02 \ \text{kDa} $With these values, the accuracy will be:
$ \frac{28.02 - 28}{28} = 5.97 \times 10^{-2} $Can you observe the charge state for the zoomed-in peak in the mass spectrum for the intact eGFP? If yes, what is it? If no, why not?
PART 2
Based on learnings in the lab, please explain the difference between native and denatured protein conformations. For example, what happens when a protein unfolds? How is that determined with a mass spectrometer? What changes do you see in the mass spectrum between the native and denatured protein analyses (Figure 2)?
When a protein unfolds, it allows external ions to interact with the structure through ion-molecule interactions such as ion-dipole forces. This increases the amount of charges and, therefore, the number of peaks, which is why it is shown a broader spectrum in the denatured protein than that of the native protein.
Zooming into the native mass spectrum of eGFP from the Waters Xevo G3 QTof MS (see Figure 3), can you discern the charge state of the peak at ~2800? What is the charge state? How can you tell?
TBA
PART 3
How many Lysines (K) and Arginines (R) are in eGFP? Please circle or highlight them in the eGFP sequence given in Waters Part I question 1 above. (Note: adding the sequence to Benchling as an amino acid file and clicking biochemical properties tab will show you a count for each amino acid).
MVS K GEELFTG VVPILVELDG DVNGH K FSVS GEGEGDATYG K LTL K FICTT G K LPVPWPTL VTTLTYGVQC FS R YPDHM K Q HDFF K SAMPE GYVQE R TIFF K DDGNY K T R A EV K FEGDTLV N R IEL K GIDF K EDGNILGH K LEYNYNSHNV YIMAD K Q K NG I K VNF K I R HN IEDGSVQLAD HYQQNTPIGD GPVLLPDNHY LSTQSALS K D PNE K R DHMVL LEFVTAAGIT LGMDELY K LE HHHHHH
Number of Lysines: 20 Number of Arginines: 6
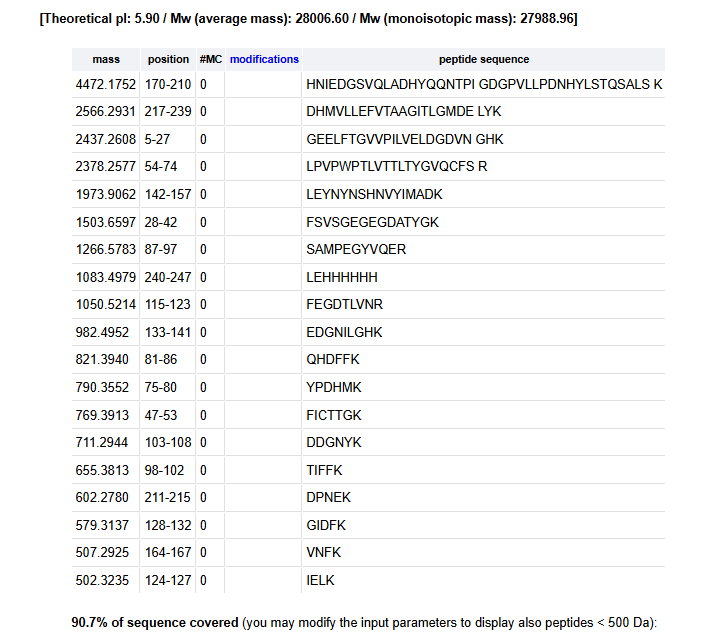
How many peptides will be generated from tryptic digestion of eGFP?
Based on the website provided, there will be 19 fragments