Week 2: DNA Read, Write, and Edit
Week 2: DNA Read, Write, and Edit
Part 1: Benchling & In-silico Gel Art
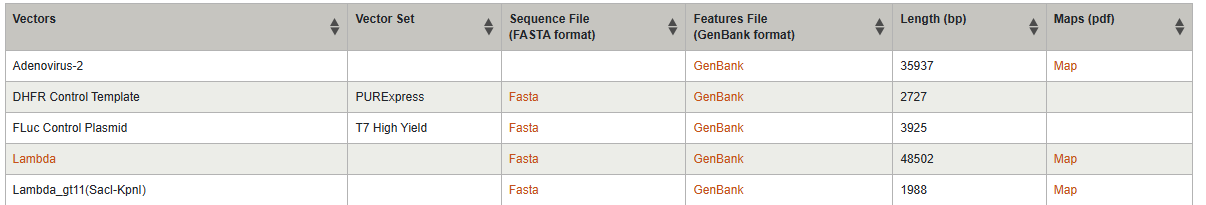
1.1 Import Lambda DNA
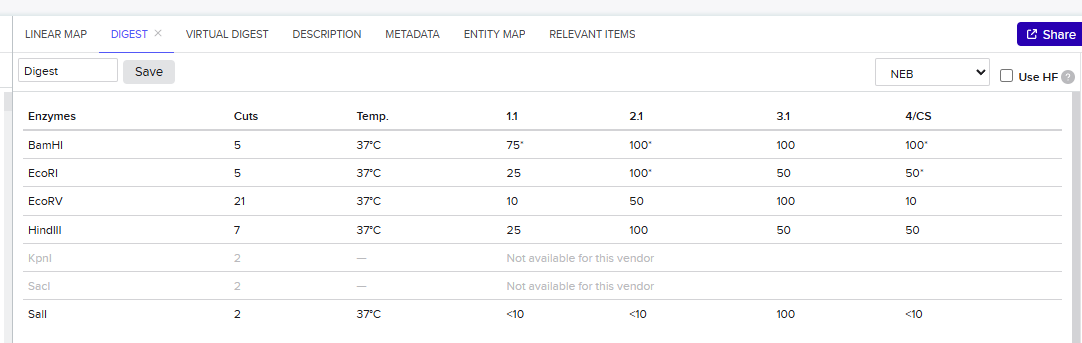
Simulate Restriction Enzyme Digestion
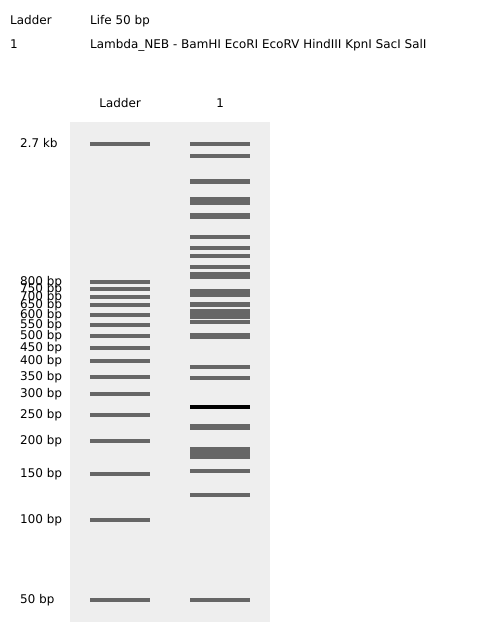
Virtual Gel
Part 2: Gel Art
I have chosen to create a gel art of a person doing a jumping jack through randomization method.
Part 3: DNA Sequence Design
3.1 Protein Selection
I have chosen IL23 as I am interested in autoimmune disease such as psoriasis. This protein is related to inflammation and I am curious to learn more about biologics in general.
- UniProt Link: https://www.uniprot.org/uniprotkb/Q5VWK5/entry#sequences
3.2 Reverse Translation
Reverse translation of sp|Q5VWK5|IL23R_HUMAN Interleukin-23 receptor OS=Homo sapiens OX=9606 GN=IL23R PE=1 SV=3 to a 1887 base sequence of most likely codons.
Reverse translation of sp|Q5VWK5|IL23R_HUMAN Interleukin-23 receptor OS=Homo sapiens OX=9606 GN=IL23R PE=1 SV=3 to a 1887 base sequence of consensus codons.
3.3 Codon Optimization
Original Sequence
- GC Content: 49.34%
- CAI: 0.83
Improved DNA Sequence
- GC Content: 51.56%
- CAI: 0.91
Avoid Cleavage Sites
- BbsI
- BsaI
Why Codon Optimization is Important
Codon optimization is important because there is codon usage bias, which means humans and other organisms like E. coli might prefer different codons for the same amino acid. Expressing human gene like IL23 might be difficult because codons natural to human cells are rare in E. coli. If bacterium has low levels of corresponding tRNAs, then it will be slowed down during translation. There will be low protein yield as a result.
The codon optimization here increased GC content so there will be more mRNA stability. Codon adaptation index has also gone up.
3.4 Protein Expression
Now, we will use this optimized DNA sequence to create IL23 protein. First we clone the codon optimized sequence into expression vector, and we transform a plasmid into E. coli cells. Bacteria will be shocked by heat to start making protein. The cell’s RNA polymerase will read the DNA and makes mRNA copy. Once the transcription is read, it will begin to build protein using tRNAs in the translation process.
Once this is done, there is a chromatography technique which separates protein from everything else in the cell.
Part 4: IL23 Sequence Analysis
Summary
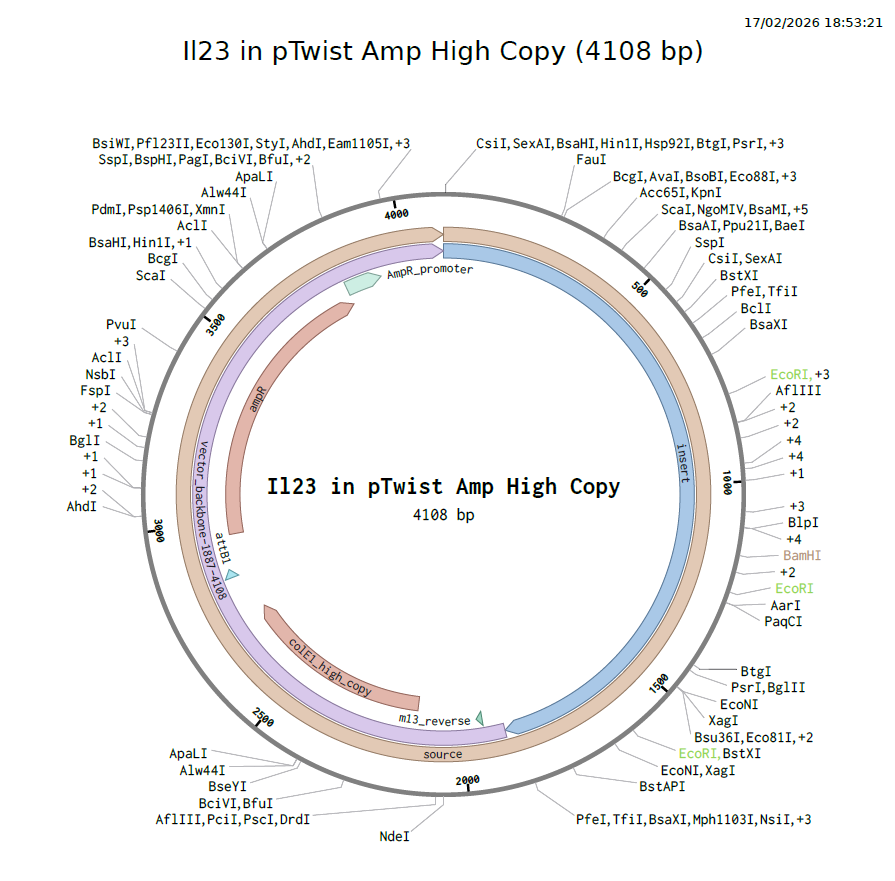

| Property | Value |
|---|---|
| Gene | IL23 |
| Benchling Link | https://benchling.com/s/seq-009SW3mnB5zCD8Vhh1Tp?m=slm-3ISQ8GXHvPtygDx4UjjQ |
| Start Codon (ATG) | Positions 1–3 |
| Coding Sequence | Positions 1 through the end |
| Stop Codon | Missing — needs to be added |
| Promoter, RBS, His-tag, Terminator | All missing — provided by the vector |
Download IL23 Plasmid Map (PDF)
Part 5:
Part 5: DNA Read/Write/Edit
5.1 DNA Read (i) What DNA would you want to sequence (e.g., read) and why? This could be DNA related to human health (e.g. genes related to disease research), environmental monitoring (e.g., sewage waste water, biodiversity analysis), and beyond (e.g. DNA data storage, biobank).
I would like to sequence and read genes that can help facilitate brains-on-chips research, so while human DNA is interesting, I am perhaps more curious about biocompatible materials or bio-glue that can help with assembling living neuronal tissue with physical hardware like microelectrode arrays. This is usually microbial/ environmental DNA where we can look at genetic strands that can be programmed into biocompatible hydrogels.
DNA-based digital data storage technology. Source: Archives in DNA: Workshop Exploring Implications of an Emerging Bio-Digital Technology through Design Fiction - Scientific Figure on ResearchGate. Available from: https://www.researchgate.net/figure/DNA-based-digital-data-storage-technology_fig1_353128454 [accessed 11 Feb 2025] DNA-based digital data storage technology. Source: Archives in DNA: Workshop Exploring Implications of an Emerging Bio-Digital Technology through Design Fiction - Scientific Figure on ResearchGate. Available from: https://www.researchgate.net/figure/DNA-based-digital-data-storage-technology_fig1_353128454 [accessed 11 Feb 2025]
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
If I were to use the Sanger sequencing method, we will need to use ddNTPs to shorten chains and terminate chains.
Also answer the following questions:
Is your method first-, second- or third-generation or other? How so?
First generation uses chain-termination, where Polmerase copies the DNA, but ddNTPs tagged with florescent colors are added, so that it creates fragments and is separated by electrophoresis. Second generation only sequnces short fragments and reading a lot of fragments simutaneously. Third generation pulls single strands through nanopore in a membrane and is is read through currents.
What is your input? How do you prepare your input (e.g. fragmentation, adapter ligation, PCR)? List the essential steps. What are the essential steps of your chosen sequencing technology, how does it decode the bases of your DNA sample (base calling)? What is the output of your chosen sequencing technology?
We need to prepare input but growing bacteriaphage, we will use plasmid purification to extract DNA. Use Benchling to design primer. We will be preparing via chain-termination PCR which will mix DNA sequence with an enzyme Poolymerase, a primer to bind to target DNA, and dNTPs and ddNTPS that are fluorescent to terminate chains.
Electrophoresis will help us separate DNA, RNA, and proteins by electrical charge.
5.2 DNA Write (i) What DNA would you want to synthesize (e.g., write) and why? These could be individual genes, clusters of genes or genetic circuits, whole genomes, and beyond. As described in class thus far, applications could range from therapeutics and drug discovery (e.g., mRNA vaccines and therapies) to novel biomaterials (e.g. structural proteins), to sensors (e.g., genetic circuits for sensing and responding to inflammation, environmental stimuli, etc.), to art (DNA origamis). If possible, include the specific genetic sequence(s) of what you would like to synthesize! You will have the opportunity to actually have Twist synthesize these DNA constructs! :)
Although irrelevant to my final project I’ve always been fascinated by biologics as adillimumab, which is a type of recombinant DNA that instruct living cells to synthesize a therapeutic protein. For the final project probably something that allows biological tissue to be more adhesive to microelectrodes as a part of facilitating electrical communication. Also interested in bioprinting microfluidics.
See some famous examples of DNA design
DNA origami by Paul W. K. Rothemund, California Institute of Technology, 2004. 100 nanometers in diameter.
(ii) What technology or technologies would you use to perform this DNA synthesis and why?
Benchling, which is a platform that can help copy and paste DNA sequence, import DNA and protein sequences, perform in silico restriction digestion, and to design gel layouts. We will cut with restrictions enzyme,copy through polymerase chain reactions, and perform DNA cloning to synthesize in silico.
Also answer the following questions:
What are the essential steps of your chosen sequencing methods? What are the limitations of your sequencing method (if any) in terms of speed, accuracy, scalability?
5.3 DNA Edit (i) What DNA would you want to edit and why? In class, George shared a variety of ways to edit the genes and genomes of humans and other organisms. Such DNA editing technologies have profound implications for human health, development, and even human longevity and human augmentation. DNA editing is also already commonly leveraged for flora and fauna, for example in nature conservation efforts, (animal/plant restoration, de-extinction), or in agriculture (e.g. plant breeding, nitrogen fixation). What kinds of edits might you want to make to DNA (e.g., human genomes and beyond) and why?
If working on neural tissues, I am curious to edit neuroplasticity-related genes so that I can consider how plasticity can be modified or reinforced. I would like to facilitate electrical and chemical stimulation to make it easier for reinforcement learning experiments.
Colossal Biosciences Inc., a biotechnology company using genetic engineering to de-extinct various historic animals such as the woolly mammoth, dodo, and dire wolf.
(ii) What technology or technologies would you use to perform these DNA edits and why? Also answer the following questions:
How does your technology of choice edit DNA? What are the essential steps? What preparation do you need to do (e.g. design steps) and what is the input (e.g. DNA template, enzymes, plasmids, primers, guides, cells) for the editing? What are the limitations of your editing methods (if any) in terms of efficiency or precision?
Electrophoresis will help us separate DNA, RNA, and proteins by electrical charge.
I would like to first perform PCR and digest, and then conduct assembly by converting GFP into RFP.