Week 3: Lab Automation
Python Script for Opentrons Artwork
Question 1
Generate an artistic design using the GUI at opentrons-art.rcdonovan.com.
I first generated a design using an image input of a voxelated ragdoll. The pixels should help simplify the image so that it can be plotted in the dish similarly.


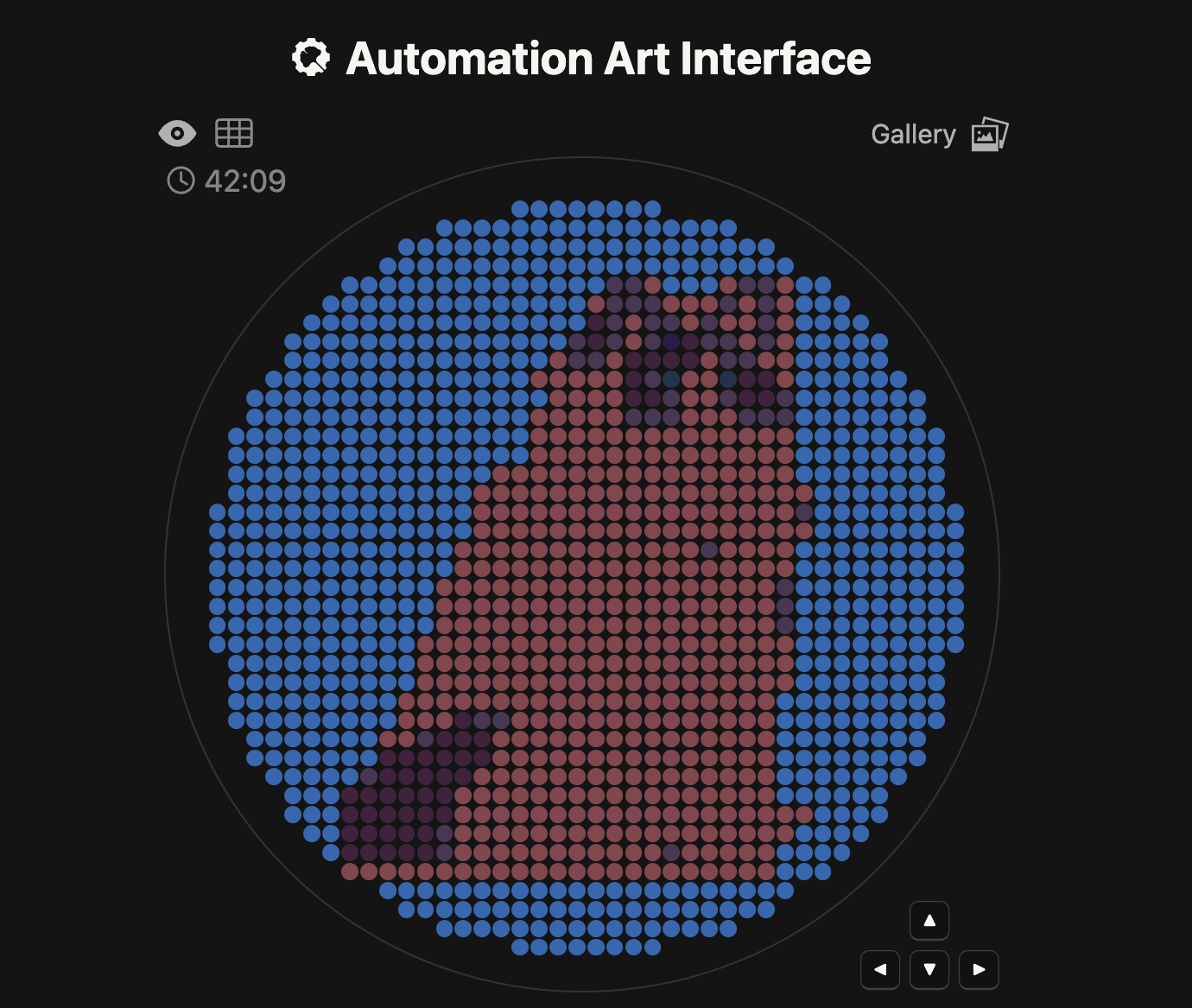

Because of the lack of contrast and limitations in the range of colors, the image looked different than expected.
Meanwhile at LifeFabs, we only had access to the colors Pink, Blue, and Purple. So I ended up simplifying the number of fluorescent proteins used to three and generated the coordinates appropriately.
Question 2
Using the coordinates from the GUI, follow the instructions in the HTGAA26 Opentrons Colab to write your own Python script which draws your design using the Opentrons.
These were the coordinates generated from the GUI using three fluorescent proteins:
Using the Opentrons Colab document, I successfully integrated the point data into the code:
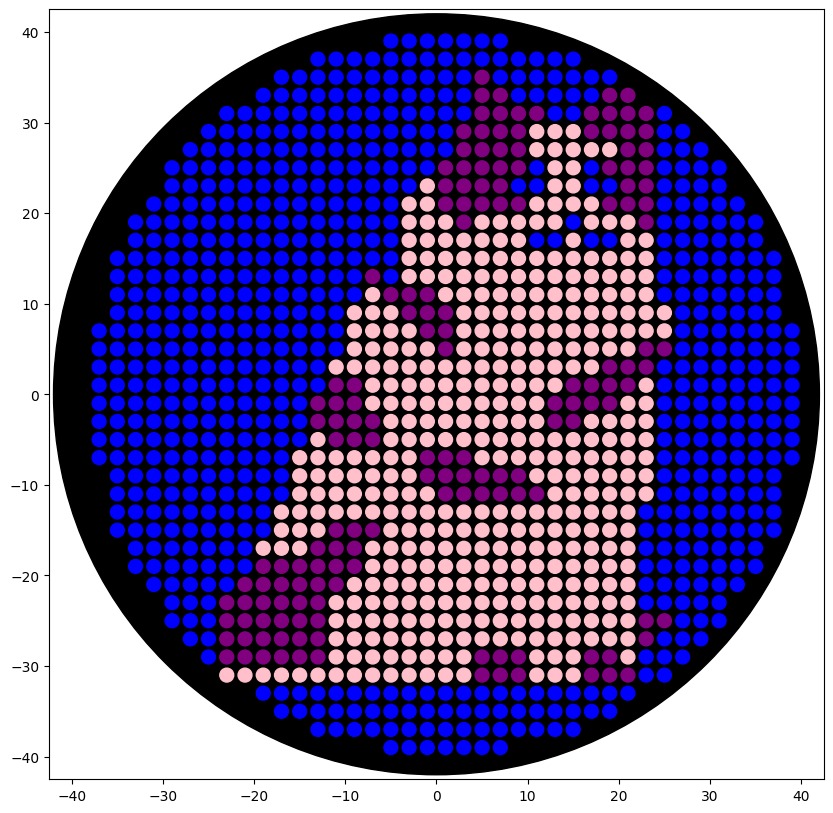

This is the result of the final preview on the colab document, using the three colors available.
Post-Lab Questions
Question 1
Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
Answer:
‘Fluidic Programmable Gravi-maze Array for High Throughput Multiorgan Drug Testing’ by Wong et al. proposes OrganRX, which is a multi-organ-on-a-chip system that is compatible with automated liquid dispensing robots such as Opentrons, OT2.The programmable part of the microfluidic architecture uses robotic liquid handlers and automated plate readers, which can help researchers program how much media reaches each organ compartment.
There is also a programmable tilting recirculation mechanism that drives flow between the corner wells of the chip, allowing for directional flow.
The developers developed a Bluetooth-enabled iOS app that allows for remote control of the recirculation system, allowing users to select from multiple shear flow rates, set programmable waiting times between tilt-direction changes, and conduct system reset.
Question 2
Write a description about what you intend to do with automation tools for your final project. You may include example pseudocode, Python scripts, 3D printed holders, a plan for how to use Ginkgo Nebula, and more.
As my research focuses on brains-on-chips and facilitating closed-loop interactions between living substrates and software systems, I’m curious to develop something similar to the OrganRX platform that utilizes Opentrons for chemical I/O with synthetic bioengineered intelligence. The direction is to look into facilitating biochemical feedback loops and designing custom plates for Opentrons via 3D printing.
WIP JSON code for custom labware:
Final Project Ideas — DUE BY START OF FEB 24 LECTURE
As explained in this week’s recitation, add 1-3 slides with 3 ideas you have for an Individual Final Project in the appropriate slide deck for MIT/Harvard/Wellesley students or for Committed Listeners. Be sure to put your name on your slide(s); for CLs, also put your city and country on your slide(s) and be sure you’re putting your slide(s) in your Node’s section of the deck.
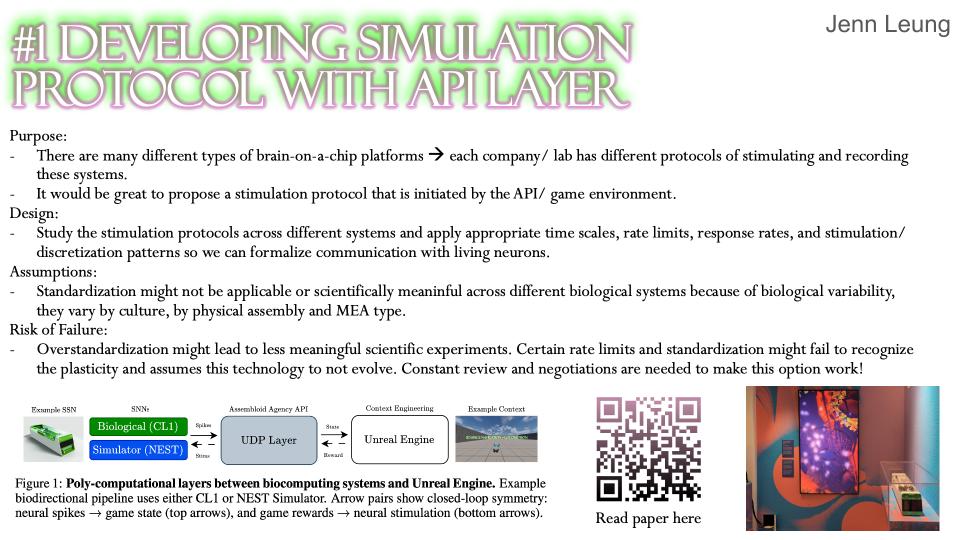
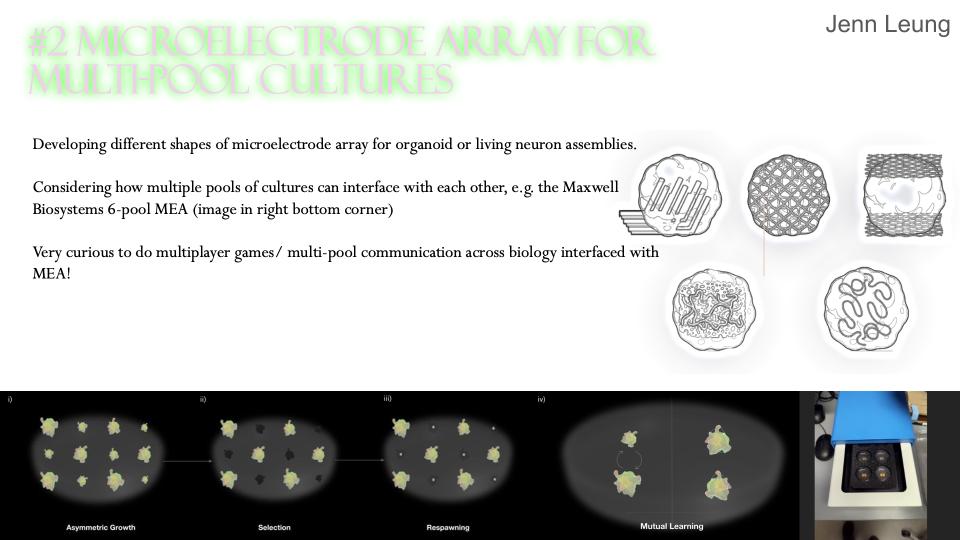
Assembloid Agency is a bio-digital interface platform designed to facilitate closed-loop biochemical communication between synthetic neural substrates and automated software systems. I will be integrating the Opentrons OT-2 with Multi-Electrode Array to create chemical I/O bridge between neural substrates and software systems.
I’m looking into using DREADDs to allow software-controlled chemical I/O as well as designing custom 3D-printed labware, housing the biological assembly while providing microfluidic channels for automated media exchange, chemical reinforcement signals, and waste management. The aim is to conduct real-time closed-loop chemical communication with the substrate.







Reading & Resources
- Opentrons API Documentation: https://docs.opentrons.com/python-api/
- Opentrons Artwork GUI Website: http://opentrons-art.rcdonovan.com/
- Opentrons Artwork Colab: HTGAA26 Opentrons Colab
- Automation Equipment: HTGAA 2026 Recitation: Lab Automation, Opentrons Art, Intro to Cloud Laboratories