Week 7: Genetic Circuits Part II
Week 7: Genetic Circuits Part II
Assignment Part 1: Intracellular Artificial Neural Networks (IANNs)
- What advantages do IANNs have over traditional genetic circuits, whose input/output behaviors are Boolean functions?
An artificual neuron is a weighted summation through an activation function that produces outputs, eventually they form networks to become ANN. Intracellular artificial networks still have weighted summation and a non-linear activation function, but we can consider implementing gene circuits as these activation functions. The main difference is that IANNs will have two inputs that can do addition and subtraction. On the one hand, a promoter that through transcription makes a gene, and through translation we create proteins, we can perform addition on this. To subtract, we can treat input x1 as an endoribonuclease CasE that will bind and cleaves the RNA on the sequence and produce output. x1 is negative weight and x2 is positve weight, where the function is max(x2-x1,0). This is also referred to as Sequestration. Sequestration involves using an endorribonucleus to transcribe into mRNA to produce non-linearity (applying single turnover enzyme to remove it out of circulation).
- Describe a useful application for an IANN; include a detailed description of input/output behavior, as well as any limitations an IANN might face to achieve your goal.
I think an interesting use of IANNs would be in culturing and programming organoids - Weiss Lab also focuses a lot on programmable patterning to trigger cell changes, and I can see this leading to very useful applications in supporting organoid intelligence. I think using endoribonuclease in waste management in microfluidics might be useful.
Microfluidics supporting organoid growth may end up accumulating extracellular vesicles, might shed RNA and dead cells, and these RNA-protein aggregates that can clog channels. We can use RNase H to clear RNA or use Cas13 endoribonuclease to cleave transcripts or CasE to degrade fragments so they can pass through filters, then maybe more layers of flushing and binding for removal? Limitations - not sure how they will exist the microfluidics- more resesarch needed.
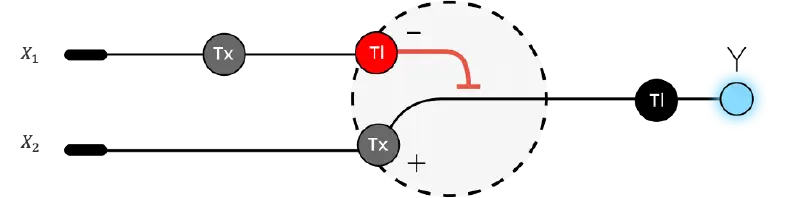
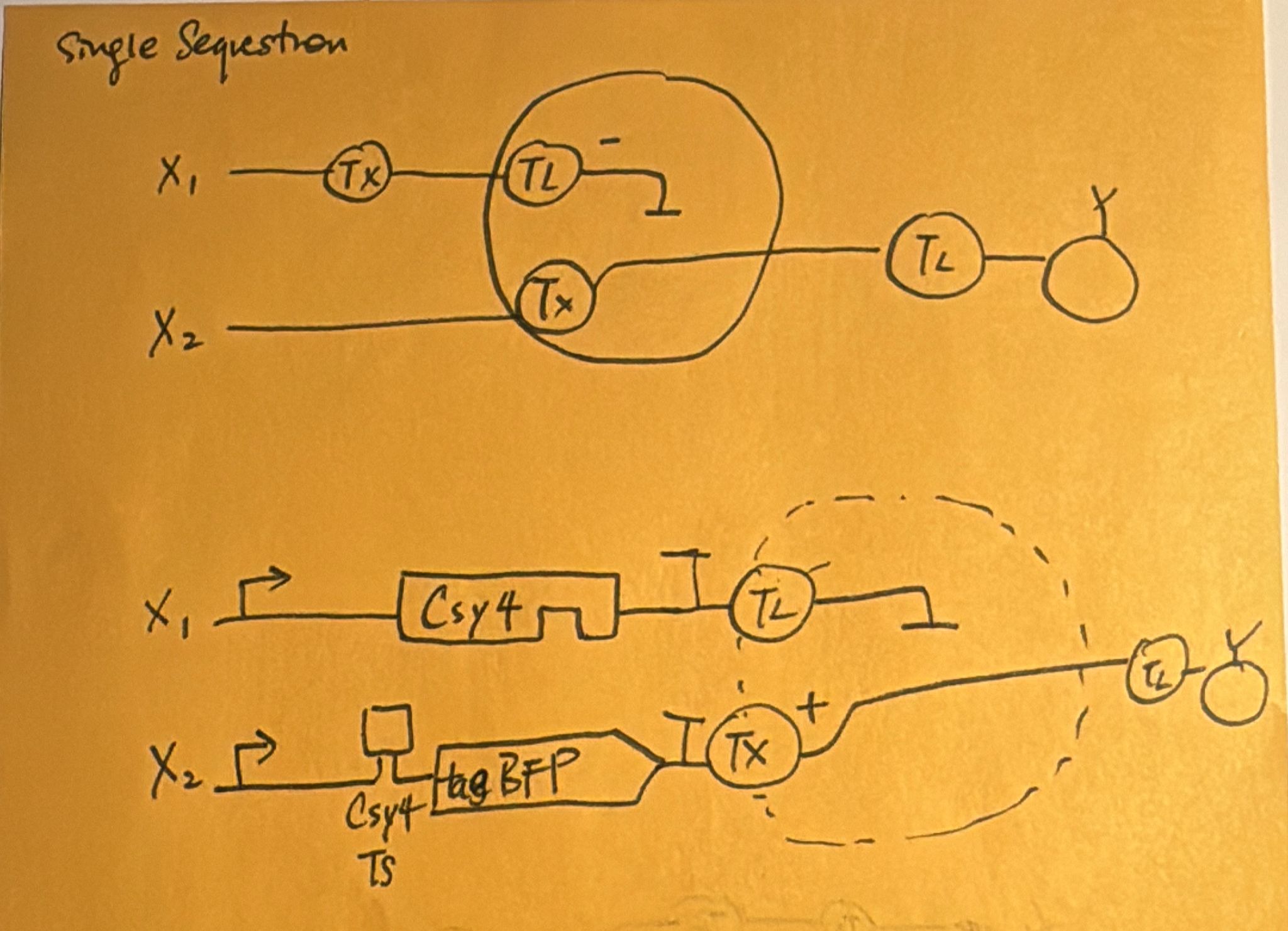
- Below is a diagram depicting an intracellular single-layer perceptron where the X1 input is DNA encoding for the Csy4 endoribonuclease and the X2 input is DNA encoding for a fluorescent protein output whose mRNA is regulated by Csy4. Tx: transcription; Tl: translation.
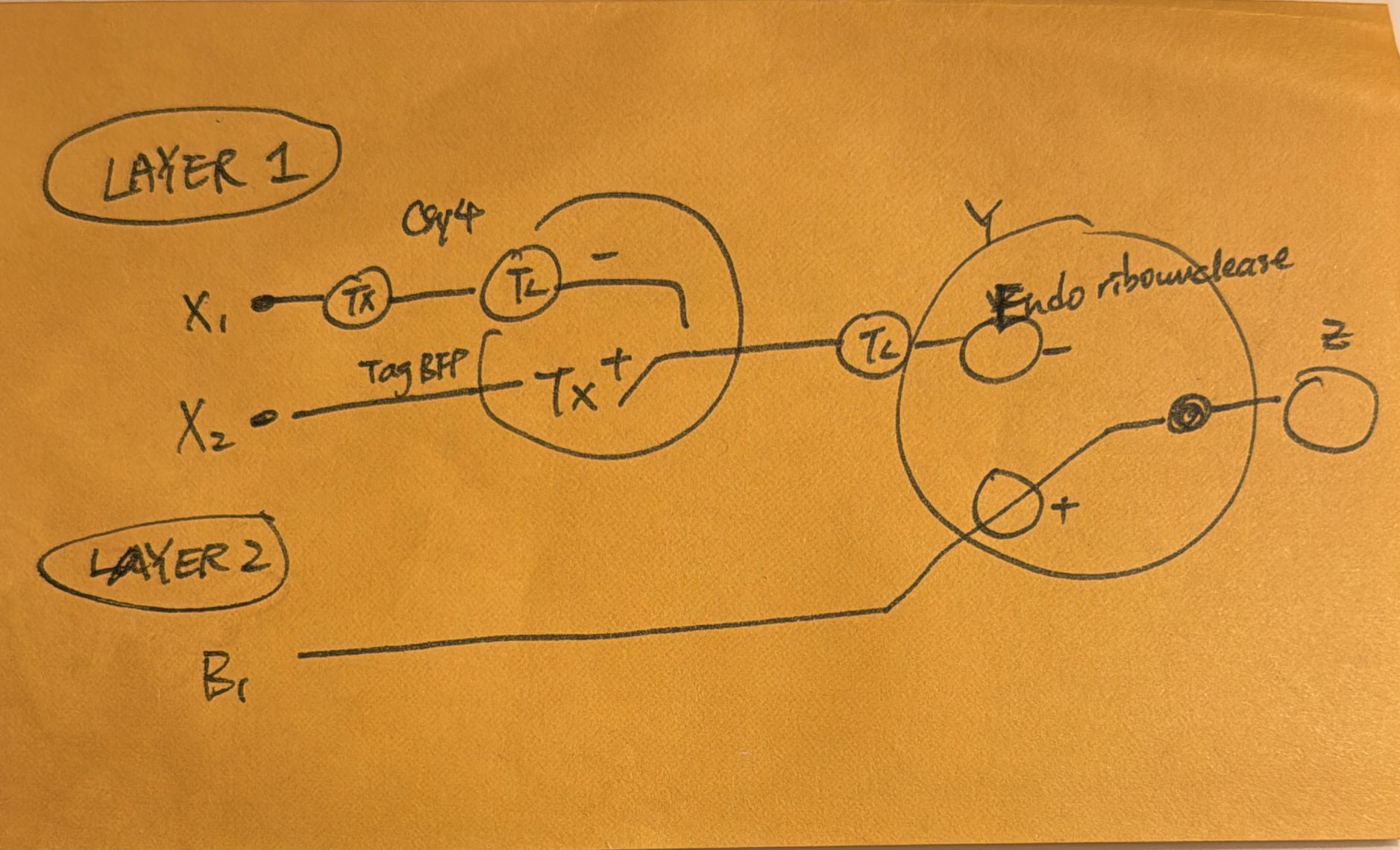
Draw a diagram for an intracellular multilayer perceptron where layer 1 outputs an endoribonuclease that regulates a fluorescent protein output in layer 2.

Assignment Part 2: Fungal Materials
- What are some examples of existing fungal materials and what are they used for? What are their advantages and disadvantages over traditional counterparts?
There’s lots of use of mycelium leather, that is currently being used to manifacture into fashion products. They have used agricultural waste as feedstock and with enough coating, can develop great strength and malleability. There’s much less waste and resources needed to create mycelium leather and will help us with better animal welfare.
- What might you want to genetically engineer fungi to do and why? What are the advantages of doing synthetic biology in fungi as opposed to bacteria?
It might be possible to create mycelium electronics which I’m super interested in. I am interested to genetically engineer mycelium to have better conductivity or bioelectric activity, or to use bioglue to stick to sensors for activity readout. Fungi is adaptable and can respond to environments quickly and act as good living sensors. They ’learn’ quickly and can be interfaced with EMG sensors without needing to submerge in many culture mediums or inject antibiotics.
Assignment Part 3: First DNA Twist Order
- Review the Individual Final Project documentation guidelines.
- Submit this Google Form with your draft Aim 1, final project summary, HTGAA industry council selections, and shared folder for DNA designs. DUE MARCH 20 FOR MIT/HARVARD/WELLESLEY STUDENTS
Done this!
- Review Part 3: DNA Design Challenge of the week 2 homework. Design at least 1 insert sequence and place it into the Benchling/Kernel/Other folder you shared in the Google Form above. Document the backbone vector it will be synthesized in on your website.
Reading & Resources
- The perceptron, the basis of artificial neural networks: https://www.geeksforgeeks.org/deep-learning/what-is-perceptron-the-simplest-artificial-neural-network/
- Many examples of artificial neural networks made using biomolecules: https://doi.org/10.1016/j.biosystems.2024.105164