Week 2 HW: Read, Write, Edit DNA Version2
Contents
Part 1: Benchling and In-Silico Gel Art
Simulated lambda DNA digestions:
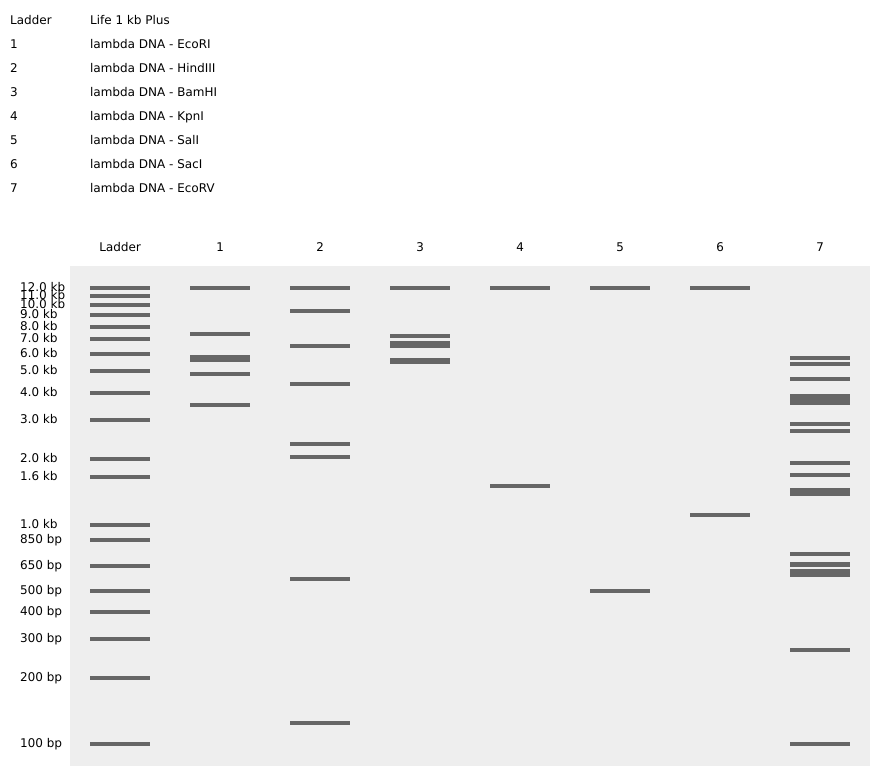


I couldn’t figure out how to use Ronan’s website other than the randomization button unfortunately. As a result, I went with a pretty simple smiley face design for my in-silico art.
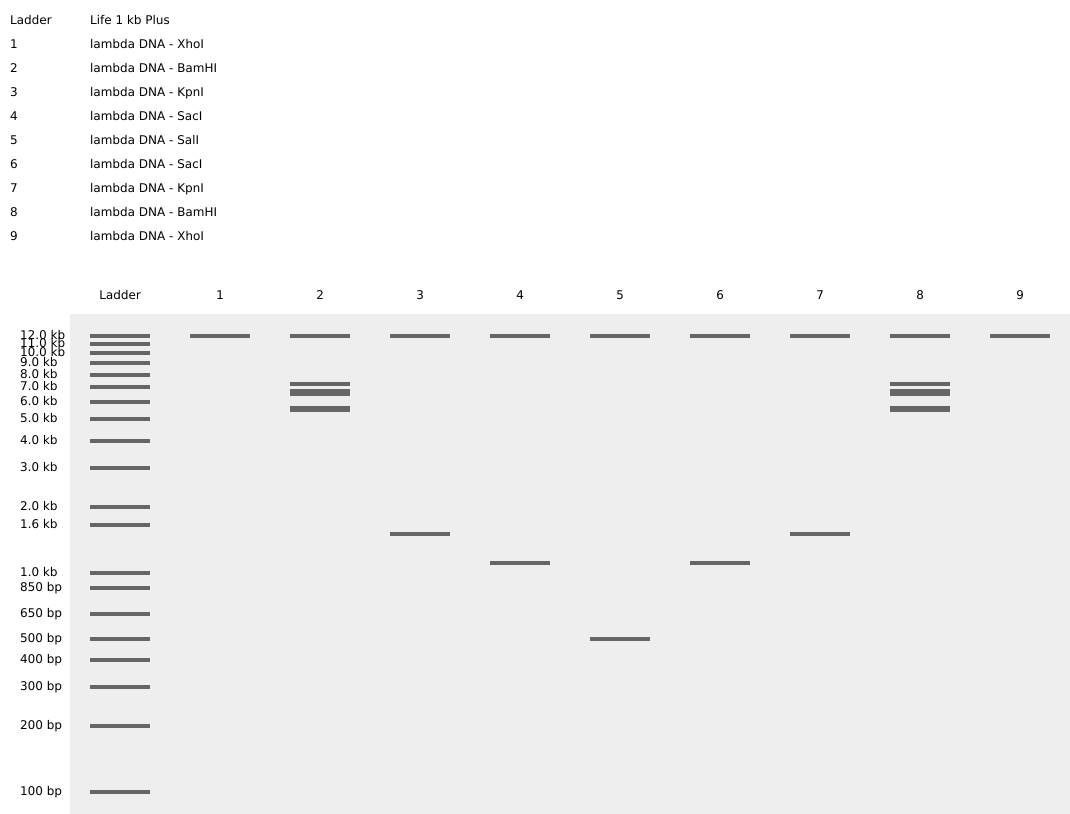


Part 2: Gel Art - Restriction Digests and Gel Electrophoresis
See Week 2 lab for details.
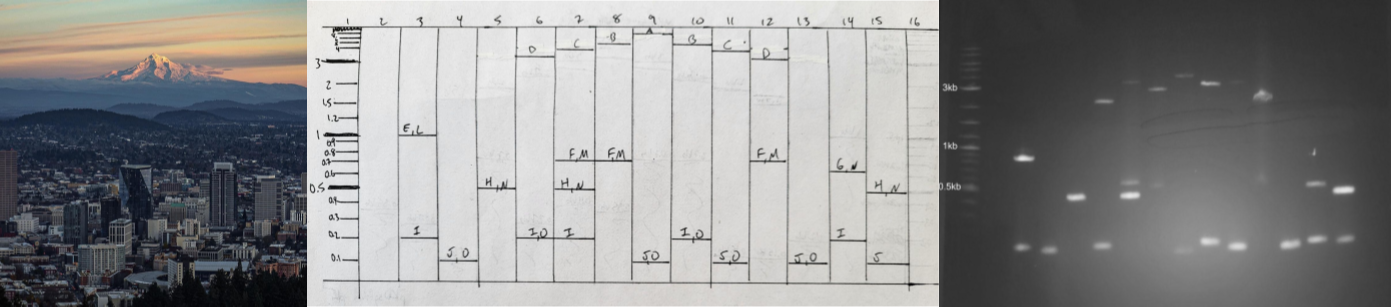

Part 3: DNA Design Challenge
3.1 Choose protein
I’m interested in PhaC, a PHA synthase. This is an enzyme involved in the synthesis of polyhydroxyalkanoates (PHAs), a class of biopolymer that is considered a potential non-petroleum-derived thermoplastic. PHAs are also of interest for possible medical uses as biodegradable polymers. PhaC is the enzyme that catalyzes the polymerization step, adding on monomers to the chain.
I selected PhaC from Cupriavidus necator H16 whose primary product is poly(3-hydroxybutyurate). From UniProt, the accession number is P23608 · PHAC_CUPNH.
MATGKGAAASTQEGKSQPFKVTPGPFDPATWLEWSRQWQGTEGNGHAAASGIPGLDALAGVKIAPAQLGDIQQRYMKDFSALWQAMAEGKAEATGPLHDRRFAGDAWRTNLPYRFAAAFYLLNARALTELADAVEADAKTRQRIRFAISQWVDAMSPANFLATNPEAQRLLIESGGESLRAGVRNMMEDLTRGKISQTDESAFEVGRNVAVTEGAVVFENEYFQLLQYKPLTDKVHARPLLMVPPCINKYYILDLQPESSLVRHVVEQGHTVFLVSWRNPDASMAGSTWDDYIEHAAIRAIEVARDISGQDKINVLGFCVGGTIVSTALAVLAARGEHPAASVTLLTTLLDFADTGILDVFVDEGHVQLREATLGGGAGAPCALLRGLELANTFSFLRPNDLVWNYVVDNYLKGNTPVPFDLLFWNGDATNLPGPWYCWYLRHTYLQNELKVPGKLTVCGVPVDLASIDVPTYIYGSREDHIVPWTAAYASTALLANKLRFVLGASGHIAGVINPPAKNKRSHWTNDALPESPQQWLAGAIEHHGSWWPDWTAWLAGQAGAKRAAPANYGNARYRAIEPAPGRYVKAKA
3.2 Reverse translate
I used the Benchling back-translate tool set to match Escherichia coli K-12 naturally occuring codon usage because it didn’t have the native host C. necator as an option. They are in the same phylum (Pseudomonadota), so maybe it will be similar.
ATGGCAACTGGAAAGGGTGCGGCCGCGAGCACACAGGAAGGTAAATCACAGCCGTTTAAGGTAACCCCGGGCCCCTTCGATCCTGCCACGTGGCTCGAGTGGTCGCGTCAGTGGCAAGGCACTGAAGGTAATGGGCACGCAGCCGCCTCTGGCATCCCGGGTCTTGATGCCCTGGCAGGCGTGAAGATTGCCCCAGCCCAATTAGGTGACATTCAGCAACGTTACATGAAAGACTTTAGTGCACTATGGCAGGCCATGGCGGAAGGTAAAGCGGAGGCGACGGGGCCTCTGCATGATCGTCGCTTCGCCGGCGATGCGTGGCGTACCAACCTGCCGTATCGCTTCGCAGCGGCGTTTTATCTGCTCAACGCGCGTGCACTTACCGAGCTGGCTGACGCAGTAGAAGCCGACGCCAAAACCAGGCAACGCATCCGTTTTGCGATTAGCCAGTGGGTGGATGCCATGAGTCCGGCTAACTTTCTGGCGACCAACCCGGAAGCCCAGCGCCTCCTGATTGAATCCGGTGGCGAAAGTCTTCGCGCGGGAGTGCGAAACATGATGGAAGATCTGACGCGAGGTAAGATCAGCCAGACGGATGAAAGCGCATTCGAAGTCGGGCGTAATGTTGCCGTTACGGAGGGTGCGGTTGTGTTTGAGAACGAATATTTCCAGTTGTTACAGTATAAGCCGCTGACCGATAAAGTGCATGCCCGCCCACTTCTCATGGTACCTCCGTGCATCAACAAATACTACATTCTGGATCTTCAGCCTGAGAGCTCATTGGTACGCCATGTGGTAGAGCAAGGCCACACAGTGTTTCTAGTCTCATGGCGCAATCCGGACGCATCCATGGCCGGCTCGACGTGGGACGATTATATCGAACACGCGGCAATAAGAGCGATTGAGGTCGCGCGTGATATCAGCGGTCAGGACAAAATTAATGTGTTAGGTTTCTGCGTAGGCGGTACTATCGTGAGTACCGCCCTGGCGGTTTTGGCAGCTCGCGGCGAACATCCGGCCGCTTCAGTTACTCTTCTGACTACCCTGCTGGATTTTGCGGACACCGGCATTCTGGATGTCTTCGTAGATGAAGGACATGTTCAGTTGCGCGAAGCAACCTTAGGCGGGGGGGCGGGTGCCCCGTGTGCCTTACTGCGGGGCCTGGAACTCGCTAACACCTTTTCGTTCCTGCGCCCAAACGATCTGGTTTGGAATTACGTGGTCGATAACTATCTGAAAGGCAACACCCCGGTGCCGTTTGATCTGCTGTTTTGGAATGGCGACGCGACCAACCTGCCGGGCCCGTGGTATTGCTGGTACCTCCGCCACACATACCTGCAAAATGAACTAAAAGTGCCAGGCAAACTGACAGTTTGTGGCGTGCCTGTGGATTTGGCTTCCATTGACGTGCCGACGTACATTTACGGTTCGCGCGAAGATCACATCGTCCCGTGGACCGCTGCCTACGCTTCTACGGCGTTGTTAGCAAATAAACTTCGGTTCGTTTTAGGCGCATCTGGCCATATTGCGGGAGTTATTAATCCACCCGCGAAAAATAAGCGTAGCCATTGGACCAATGACGCGTTGCCTGAAAGCCCCCAGCAATGGCTGGCAGGCGCGATAGAGCATCACGGCAGCTGGTGGCCGGATTGGACCGCATGGTTAGCCGGCCAGGCCGGAGCGAAACGTGCTGCGCCCGCGAATTATGGAAACGCGCGTTATCGTGCCATTGAACCCGCCCCGGGGCGCTATGTCAAAGCGAAAGCA
They are not that similar, it turns out; although that may have less to do with codon usage frequency and more to do with when the reverse translate tool used which codons. Here’s the DNA sequence alignment comparing the genomic sequence from C. necator with the E. coli optimized reverse translation. This sequence alignment was performed in Benchling, using MAFFT with pre-set parameters.
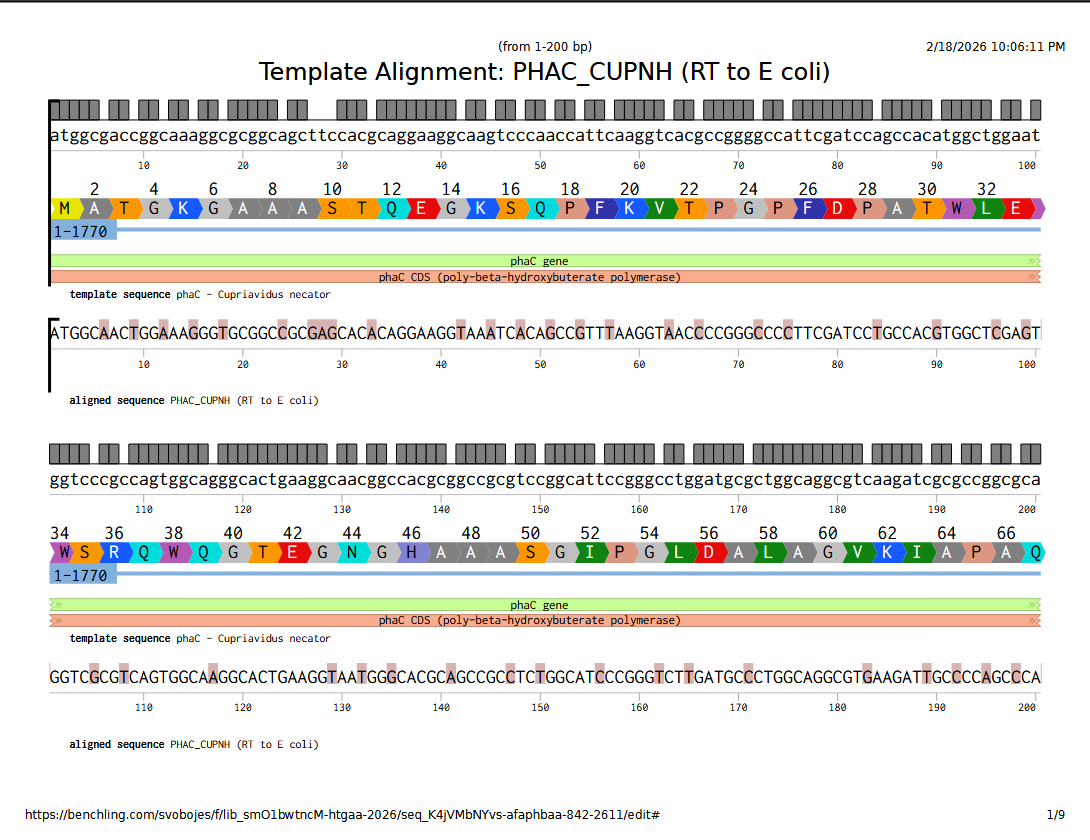
 Full alignment viewable here.
Full alignment viewable here.
3.3 Codon optimize
I once again used the Benchling tool to codon optimize for E. coli K-12, but this time, I selected the Best Codon option in Benchling, and this was performed off the original C. necator phaC DNA sequence - although it should produce the same sequence if it was done as a reverse translate from the amino acid sequence too (since i confirmed that the phaC sequence does translate to the PhaC sequence with 100% identity).
ATGGCAACTGGAAAGGGTGCGGCCGCGAGCACACAGGAAGGTAAATCACAGCCGTTTAAGGTAACCCCGGGCCCCTTCGATCCTGCCACGTGGCTCGAGTGGTCGCGTCAGTGGCAAGGCACTGAAGGTAATGGGCACGCAGCCGCCTCTGGCATCCCGGGTCTTGATGCCCTGGCAGGCGTGAAGATTGCCCCAGCCCAATTAGGTGACATTCAGCAACGTTACATGAAAGACTTTAGTGCACTATGGCAGGCCATGGCGGAAGGTAAAGCGGAGGCGACGGGGCCTCTGCATGATCGTCGCTTCGCCGGCGATGCGTGGCGTACCAACCTGCCGTATCGCTTCGCAGCGGCGTTTTATCTGCTCAACGCGCGTGCACTTACCGAGCTGGCTGACGCAGTAGAAGCCGACGCCAAAACCAGGCAACGCATCCGTTTTGCGATTAGCCAGTGGGTGGATGCCATGAGTCCGGCTAACTTTCTGGCGACCAACCCGGAAGCCCAGCGCCTCCTGATTGAATCCGGTGGCGAAAGTCTTCGCGCGGGAGTGCGAAACATGATGGAAGATCTGACGCGAGGTAAGATCAGCCAGACGGATGAAAGCGCATTCGAAGTCGGGCGTAATGTTGCCGTTACGGAGGGTGCGGTTGTGTTTGAGAACGAATATTTCCAGTTGTTACAGTATAAGCCGCTGACCGATAAAGTGCATGCCCGCCCACTTCTCATGGTACCTCCGTGCATCAACAAATACTACATTCTGGATCTTCAGCCTGAGAGCTCATTGGTACGCCATGTGGTAGAGCAAGGCCACACAGTGTTTCTAGTCTCATGGCGCAATCCGGACGCATCCATGGCCGGCTCGACGTGGGACGATTATATCGAACACGCGGCAATAAGAGCGATTGAGGTCGCGCGTGATATCAGCGGTCAGGACAAAATTAATGTGTTAGGTTTCTGCGTAGGCGGTACTATCGTGAGTACCGCCCTGGCGGTTTTGGCAGCTCGCGGCGAACATCCGGCCGCTTCAGTTACTCTTCTGACTACCCTGCTGGATTTTGCGGACACCGGCATTCTGGATGTCTTCGTAGATGAAGGACATGTTCAGTTGCGCGAAGCAACCTTAGGCGGGGGGGCGGGTGCCCCGTGTGCCTTACTGCGGGGCCTGGAACTCGCTAACACCTTTTCGTTCCTGCGCCCAAACGATCTGGTTTGGAATTACGTGGTCGATAACTATCTGAAAGGCAACACCCCGGTGCCGTTTGATCTGCTGTTTTGGAATGGCGACGCGACCAACCTGCCGGGCCCGTGGTATTGCTGGTACCTCCGCCACACATACCTGCAAAATGAACTAAAAGTGCCAGGCAAACTGACAGTTTGTGGCGTGCCTGTGGATTTGGCTTCCATTGACGTGCCGACGTACATTTACGGTTCGCGCGAAGATCACATCGTCCCGTGGACCGCTGCCTACGCTTCTACGGCGTTGTTAGCAAATAAACTTCGGTTCGTTTTAGGCGCATCTGGCCATATTGCGGGAGTTATTAATCCACCCGCGAAAAATAAGCGTAGCCATTGGACCAATGACGCGTTGCCTGAAAGCCCCCAGCAATGGCTGGCAGGCGCGATAGAGCATCACGGCAGCTGGTGGCCGGATTGGACCGCATGGTTAGCCGGCCAGGCCGGAGCGAAACGTGCTGCGCCCGCGAATTATGGAAACGCGCGTTATCGTGCCATTGAACCCGCCCCGGGGCGCTATGTCAAAGCGAAAGCA
3.4 Now what?
This sequence could be used to express PhaC in E. coli. I would probably put the gene onto an expression plasmid, under a strong constitutive promoter, just to ensure it works. After transforming E. coli with the plasmid, I would test expression by looking at protein production with a Western blot, and looking at cells under a microscope to look for PHA granules. I need to do a little more literature searching on heterologous expression of PhaC in E. coli - I think maybe other enzymes are needed for PHB synthesis.
3.5 Optional - how does it work in natural biological systems?
- Describe how a single gene codes for multiple proteins at the transcriptional level.
Different reading frames on the same string of DNA bases gives different codons that are off-set by which base (1-3) starts it. In this way, genes for multiple proteins can overlap on the same sequence of DNA. - Try aligning the DNA sequence, the transcribed RNA, and also the resulting translated Protein!
I created the transcript by using Benchling to create a new RNA sequence off the reverse of my coodon-optimized sequence. I kept the annotations, so the translation should still be visible. Then I made a new alignment in Benchling using MAFFT with the automatic parameters. Again, the sequences match perfectly - although it’s not 100% identity because technically the T/U difference between DNA and RNA are considered mismatches, but we can see visually across the bottom of the screenshot that we don’t have any actual mismatches.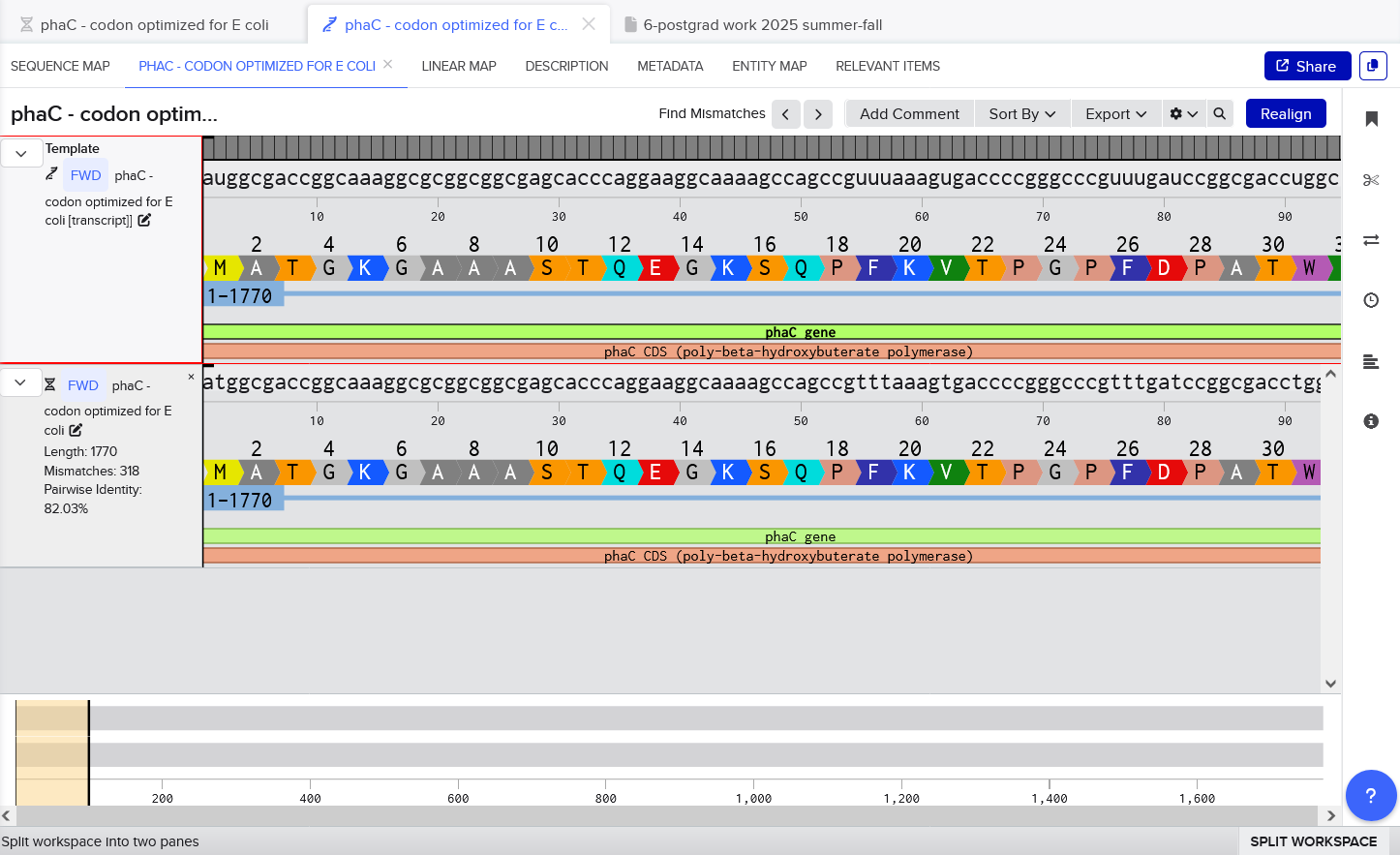

Part 4: Prepare a Twist DNA Synthesis Order
Following the instructions in the Week2 Homework, I added the J23106 promoter and an RBS at the beginning of my codon-optimized phaC sequence. My coding sequence already had a start and stop codon, so I didn’t need to add those. I inserted the 7x-His tag just before the stop codon, and then I put the terminator after the stop codon at the end.
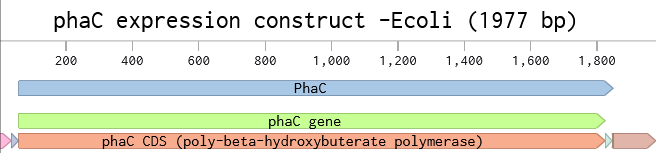

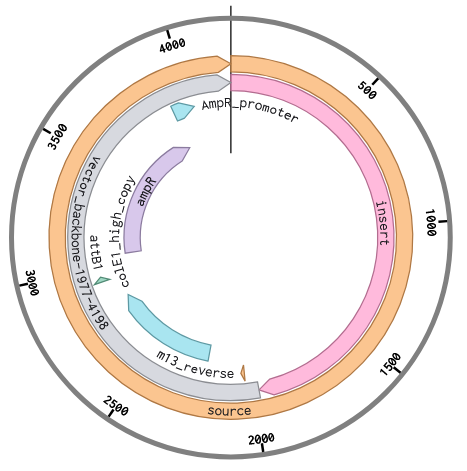
I then set up the Twist order, as if I was going to order this cassette to be synthesized. Again, following the instructions for upload, I chose cloning vector pTwist Amp High Copy to make a full plasmid. My sequence was high complexity, so I went through the Twist codon optimization process to improve the sequence for easier synthesis. I chose E. coli as my host strain again, and selected the ORF that matched my gene. I chose the promoter and RBS, and terminator regions as regions to preserve during the codon optimization process so that it kept the sequences for the genetic parts that I chose. The optimized sequence was no longer high complexity as the regions of high GC% and repeats were changed.

Part 5: DNA Read/Write/Edit
5.1 Read
- What DNA would you want to sequence (e.g., read) and why?
I’d like to sequence the genomes of all cyanobacterial strains known to produce PHAs or specifically PHB (some already are sequenced, I think). I want to align all the known cyanobacterial PHA-synthases, and then align with the assembled genomes of the cyanobacterial strains known to produce PHAs that maybe aren’t annotated yet to try to find the PHA-synthases and add those to my comparisons. - In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
I would use third-generation sequencing on an Oxford nanopore. By using long-read technology, I would get much longer contigs, to make genomic assembly easier.
5.2 Write
- What DNA would you want to synthesize (e.g., write) and why?
I’d like to get a CRISPR-Cas12a multiplexed gRNA cassette synthesized. This would allow multiple genomic edits to occur simultaneously, if the appropriate repair templates are included (one for gRNA target). - What technology or technologies would you use to perform this DNA synthesis and why?
I would submit an order to Twist to get this synthesized because it has multiple internal repeats because of the CRISPR region, which means traditional DNA synthesis technologies would struggle with this sequence.
5.3 Edit
- What DNA would you want to edit and why?
I’d like to improve PHA-synthase expression in my cyanobacterial chassis strain of choice (specific strain yet to be determined). This could be accomplished through promoter replacement if we’re staying in the genome rather than adding a plasmid, but I’d also be interested in knocking out other biosynthetic pathways to improve carbon flux towards PHA synthesis. So I’d want to edit the genomic DNA of a cyanobacterial chassis. - What technology or technologies would you use to perform these DNA edits and why?
I’d use a CRISPR-Cas12a vector because it allows for multiplexed targeting, so I could make multiple genomic edits. Cas12a both processes the CRISPR-gRNA cassette and makes the cuts, so it requires fewer components than Cas9. Additionally, there’s some evidence suggesting Cas12a shows less off-target effects than Cas9.