Week 3 HW: Lab Automation
Contents
Python Script for Opentrons Artwork
Note: This is due before the Victoria node does its Opentrons artwork lab, at a future date TBD. This homework assignment is still in-progress because the due date is not yet established.
Post-lab questions
- Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
A paper published this month in ACS Synthetic Biology details a new workflow for automating MoClo plasmid assembly and transformation, with a semi-automated colony PCR on an Opentrons OT-2 and Opentrons Flex. These workflows are designed to be user-friendly and output the Opentrons protocol from user-supplied CSV files, which provided README files describe how to produce.
Alternatively, the authors also developed a graphical user interface which requires no coding ability. This is a novel application because it is only the second automation of MoClo/Golden Gate cloning for Opentrons system (as opposed to advanced high-throughput liquid handling systems), and this new workflow does not require Python ability as the previously published AssemblyTron workflow.
These workflows were validated by assembling plasmids with the MoClo Yeast Toolkit and MoClo SubtiToolKit, and transforming these plasmids into Saccharomyces cerevisiae and sequentially Escherichia coli and Bacillus subtilis, respectively. With both toolkits, the automated procedure achieved efficiency comparable to the manual procedures (> 90% and 60%, respectively).
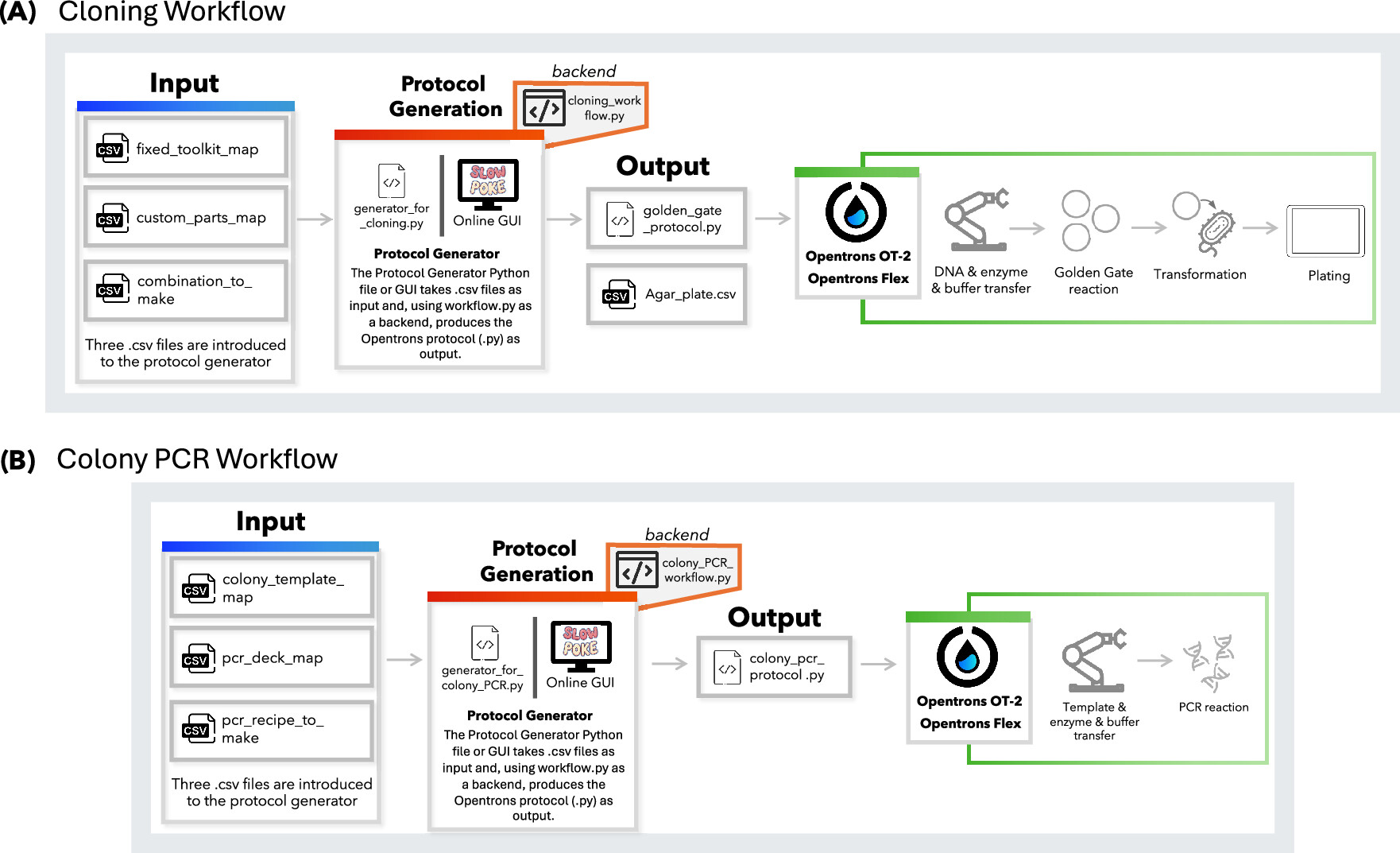

Figure 1: Schematic overview of the protocol design workflows developed for the Opentrons platform. Protocols can be generated using either the generator.py Python script via the command line or the online Slowpoke tool, which features a user-friendly GUI. Both tools run the workflow.py files in the backend. (A) Workflow for Golden Gate-based cloning, where users define genetic part layouts and assembly combinations. (B) Workflow for colony PCR, including colony selection, reagent layout, and reaction recipe input.
- Malci, K; Meng, F; Galez, H; et al. Slowpoke: An Automated Golden Gate Cloning Workflow for Opentrons OT-2 and Flex. 2026. ACS Synthetic Biology, 15(2): 511-521. DOI: 10.1021/acssynbio.5c00629
- Write a description about what you intend to do with automation tools for your final project.
I’d want to utilize the Opentrons set-up in the Victoria node to enable the possible execution of my medium-term aim with as little scientist benchtime as possible. I don’t know the exact make and model of all modules that the Victoria Opentrons has, but below is a series of possible steps that might be automatable (best use of automation would be medium or high throughput, depending on the number of designs we are able to test):- Gibson Assembly or MoClo plasmid assembly
- Transfer reaction components into wells
- Heat block for digestion/ligation/PCR steps
- Transformation of expression plasmid
- Transfer plasmids and competent cells into wells
- Heat block for heat shock
- Transfer media into wells
- Heated shaker for recovery
- Incubator for overnight growth
- Stamp onto new plate or pick into multiple liquid cultures for culturing
- Incubator or heated shaker for overnight growth
- Readout
- Transfer cells (and reagents) into wells
- Plate reader for fluorescent or colorimetric output
- Gibson Assembly or MoClo plasmid assembly
Final project ideas
Brainstorming:
- Identification of PhaC analog in Cyanobacterium aponium UTEX 3222 and overproducing or engineering for increased efficiency
- BLAST/align with known PHA-synthases
- Compare efficiency / mutations that improved turnover in other PhaC - test analogous mutations (aligned location, similar or different AAs). improved substrate specificity?
- Site-specific saturation mutagenesis? Would be good use for automation
- Quorum sensing based killswitch (i.e. cell dies if it escapes bioreactor)
- Has to have some kind of inducible element or won’t grow after initial transformation
- What’s good at quorum sensing already?
- Something else??? Something in E coli that can be done on Opentron
- Because it’s more convenient for a final project to be executed in Victoria remotely
- Cyanobacterial expression plasmid across multiple cyano species
- needs to include E coli machinery for manipulation and production (and conjugation, for relevant species)
Ideas:
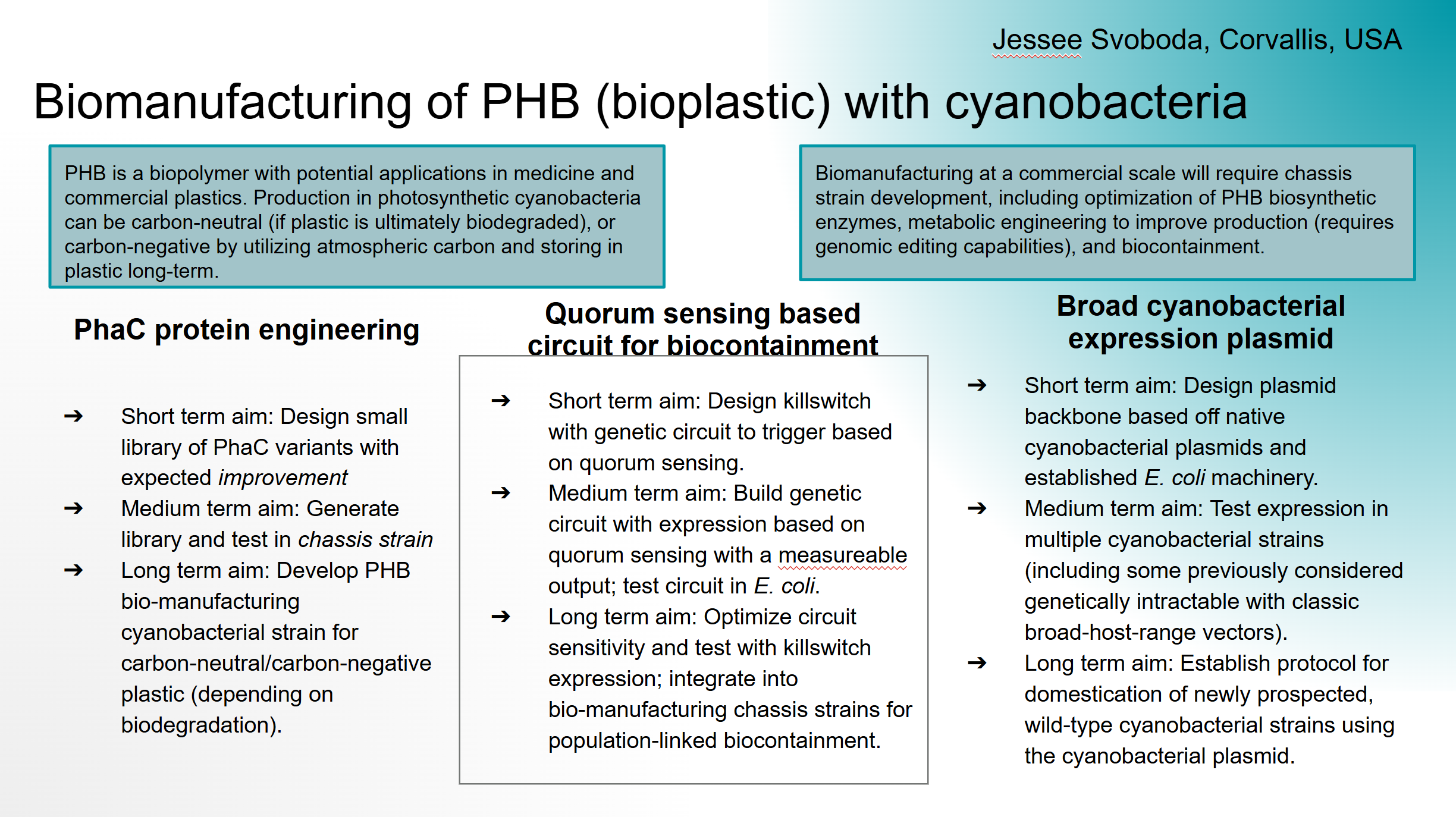
- PhaC protein engineering
- Short term aim: Design small library of PhaC variants with expected improvement
- Medium term aim: Generate library and test in chassis strain
- Long term aim: Develop PHB bio-manufacturing cyanobacterial strain for carbon-neutral/carbon-negative plastic (depending on biodegradation).
- Quorum sensing based circuit for biocontainment
- Short term aim: Design killswitch with genetic circuit to trigger based on quorum sensing.
- Medium term aim: Build genetic circuit with expression based on quorum sensing with a measureable output; test circuit in E. coli.
- Long term aim: Optimize circuit sensitivity and test with killswitch expression; integrate into bio-manufacturing chassis strains for population-linked biocontainment.
- Broad cyanobacterial expression plasmid
- Short term aim: Design plasmid backbone based off native cyanobacterial plasmids and established E. coli machinery.
- Medium term aim: Test expression in multiple cyanobacterial strains (including some previously considered genetically intractable with classic broad-host-range vectors).
- Long term aim: Establish protocol for domestication of newly prospected, wild-type cyanobacterial strains using the cyanobacterial plasmid.