Week 7 HW: Genetic Circuits Part II
Contents:
Intracellular Artificial Neural Networks
- What advantages do IANNs have over traditional genetic circuits, whose input/output behaviors are Boolean functions?
IANNs do analog computing instead of digital. So functions are additive (positive or negative) rather than just present/absent. This means that they can respond to an input that’s beyond (over or under) a certain threshold, instead of just is the input present or not. Non-digital dosage. IANNs can also stack with multiple layers for multiple inputs as well. - Describe a useful application for an IANN; include a detailed description of input/output behavior, as well as any limitations an IANN might face to achieve your goal.
IANNs can be used to identify cell types, such as cancer cells by differentiating them from the surrounding healthy cells. The cancer cells might not have a single unique signal to use as an identifier, but it might have a few different metabolites (or other signals) present in different amounts from the healthy cells. So an IANN can be used to recognize multiple inputs, and how much of those inputs are present (is it more/less than the baseline amount present in the healthy cells). The output might be fluorescence to tag tumor locations for a surgeon to excise, or maybe the output could be a medication for specific targeted release. - Draw a diagram for an intracellular multilayer perceptron where layer 1 outputs an endoribonuclease that regulates a fluorescent protein output in layer 2.


Fungal Materials
- What are some examples of existing fungal materials and what are they used for? What are their advantages and disadvantages over traditional counterparts?
Some existing fungal materials include fungal leather and fabrics for clothing, primarily of mycelium or cellulose; biocement, which uses bacteria or fungi to produce calcium carbonate around gravel; and fungal composite materials, which uses a fungal mycelium around an organic or agricultural substrate. Fungal composite materials can be leather-like fabrics, packaging, acoustic insulation, thermal insulation, and hard particle board or brick-like building materials for furniture or architecture. - What might you want to genetically engineer fungi to do and why? What are the advantages of doing synthetic biology in fungi as opposed to bacteria?
Engineered fungi might form mycelium materials that can produce different colors; contain biosensors; have different material properties like hardness/flexibility; or be able to actively bioremediate the location that the mycelium-made object is placed in. Fungi are eukaryotic instead of prokaryotic like bacteria, which means there is more diversity both within the cell (organelles) and on a cell-to-cell level (cell differentiation). This complexity both increases the difficulty of synthetic biology in fungi over bacteria, but also allows for engineering that complexity (such as only having the bioremediation turned on in the fruiting bodies).
First DNA Twist Order
Design at least 1 insert sequence and place it into the Benchling/Kernel/Other folder you shared in the Google Form above. Document the backbone vector it will be synthesized in on your website.
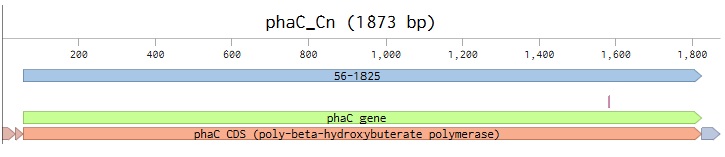
My first sequence is the wild-type Cupriavidus necator PhaC. For cell-free synthesis, it will be transcribed by T7 polymerase, so it needs to have those components. I designed it in Kernel, using the parts from the iGEM repository and the PhaC_Cnecator gene from the Uniprot repository.
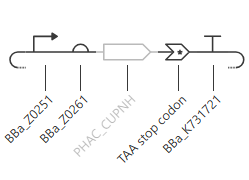

The promoter, Bba_Z0251 is the T7 promoter with the consensus sequence. The RBS, Bba_Z0261 is a wild-type T7 RBS that has been characterized as a strong RBS by an iGEM team. The terminator, Bba_K731721 is a wild-type T7 terminator that has been characterized by an iGEM team. The Uniprot PhaC_Cnecator part has no DNA sequnce in Kernel, so I remade this circuit in Benchling, using the PhaC_Cnecator sequence that I had previously codon optimized for E. coli expression in homework2; and copied the regulatory elements from Kernel.
This will be synthesized into a Twist cloning vector. Ronan suggested a chloramphenicol marker for constructs at the Ginkgo Nebula facility, so I’ll use pTwist-Chlor-HighCopy.
