Week 3 HW: Lab Automation
Python Script for Opentrons Artwork
Opentrons Art
I tried to play around with math functions to create a design, like the Mathematical Heart sample. I drew up a cute fox in Desmos graphing calculator using the following functions, making sure to scale them to the 40 mm limit. Transferring that to Colab was a bit more difficult, and I had to play around with the functions, ranges, and dispense volume to find something that looked good.
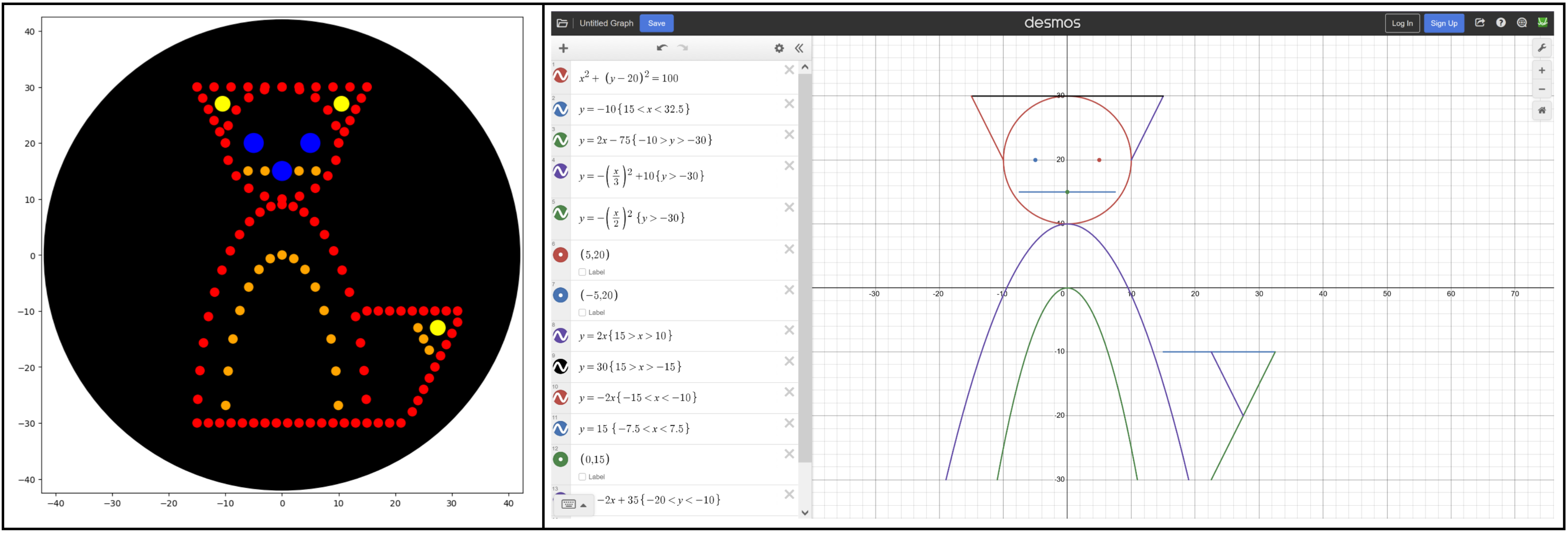

Some notes from the process:
- Polar functions are ideal for spacing out the points. I used polynomial equations (y=x2) for the body but ended up switching to ellipses (y2 + x2 = r2).
- I had trouble picking up/dropping tips, especially with the color changes in each part of the fox. So my program appends all points to lists sorted by color and then dispenses each list in a
draw()function (like the Mathematical Heart). - I had a lot of trouble tracking volume to aspirate, especially as some for loops go into the negatives, so my refill is based on the actual volume of the pipette (
pipette_20ul.current_volume) and is handled all at once in thedraw()function.
I also used https://ginkgoartworks.com/ to draw a mushroom and imported the program into Colab. Since the bacteria names don’t register as RGB colors, I had to “color-correct” well_colors to get the visualization to show up (but I assume both versions will work as long as the PCR tubes are physically in order).
Colab link for both projects here, including the color-corrected version for the mushroom.
Extra: Nebula art
I designed some artwork for the 1536-well plates on the Nebula, which were made during the Saturday 2-6 pm Cloud Lab session. The first one was a firefly squid, inspired by bioluminescent photos I’ve seen of them underwater. Link to gallery image here.
I also made a second one resembling the Chinese jianzhi for Lunar New Year. I experimented with two color sets, to see how bacteria with similar coloring would contrast against each other. Link to gallery images here and here.
Post-Lab Questions
- Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
This paper explores “dyeing” bacterial cellulose, a bioplastic alternative to leather, which I found pretty interesting. Instead of applying an independent biodegradable dye, the researchers engineered Komagataeibacter rhaeticus to develop eumelanin (dark melanin), which gives a range of shades seen in the photo above. Opentrons is used in the production of the eumelanin development buffer, in which the K. rhaeticus pellicle is incubated to “dye” itself. The robot mixes the buffer, cells, and supernatant within a reaction plate while maintaining a constant low temperature to prevent initial eumelanin growth.
Walker, K.T., Li, I.S., Keane, J. et al. Self-pigmenting textiles grown from cellulose-producing bacteria with engineered tyrosinase expression. Nat Biotechnol 43, 345–354 (2025). https://doi.org/10.1038/s41587-024-02194-3
- Write a description about what you intend to do with automation tools for your final project.
More research is needed on my part for this, but I’d like to explore scaling up or going in depth with the range of results for my project (e.g. if self-pigmenting, then trials to develop swatches of colors). This would require a lot of samples, and liquid handlers like the Opentrons would be necessary for producing all the samples indentically.
Final Project Ideas
Coloring Bioplastic/Biotextiles (with an art-focused approach)
a. Self-pigmenting Bacterial Cellulose: Building on the above paper, further development with dyed bacterial cellulose using pheomelanin instead of eumelanin for a different color range. Likely this is already being explored, so as an ambitious goal Komagataeibacter rhaeticus could be edited to express both pheomelanin and eumelanin, allowing you a 2D range of colors.
b. Structural Color on Textiles as Biopigment: Naturally occuring structural color is tied to the genome, so we could intentionally induce colors as a natural, biodegradable dyes for textiles. This paper highlights a bacteria that naturally form to output structural color, and this paper explores gene knockout to change the color expressed by one of the bacterias Flavobacterium IR1. A potential project could explore different colors on these bacteria, or find a way to introduce the bacteria to textiles without affecting its formation.
Environmental Sensors (Algae biosensors?)
- I’d like to explore fluorescence through engineering algae to detect toxic pollutants. I’d be interested in getting a biosensor to detect a singular type of pollutant (e.g. presence of heavy methods), which would require multiple strains of microalgae versus just one general strain that reacts uniformly to all pollutants. So part of this work would probably also include characterizing different responses to find the one that points to that pollutant.
Polyester Biodegradation
a. Integration with Byproduct Biodegradation: Ideonella sakaiensis is a bacteria that can break down PET plastic through a two step process involving PETase and MHETase enzymes. The end products include carbon dioxide, water, and methane, which in itself a pollutant. Methane itself is a subject of research, with methanotrophs being a type of bacteria that metabolize methane. I’m wondering if the bacteria used for breaking down plastic can be somehow integrated with the added function of breaking down methane through gene engineering.
b. Polyester-Eating Enzymes: This is a less familiar topic for me, but current work on enzyme degradation focuses on improving the performance of natural enzymes, e.g. its thermostability, pH control, etc. Since there’s such a wide range of work being done, I’m sure there’s some further testing that could be done on an underfocused bacteria/performance metric/modification method.