Week 2 HW: DNA Read, Write, Edit
Week 2 Pre-Lecure Homework:
Homework Questions from Professor Jacobson:
- Nature’s machinery for copying DNA is called polymerase. What is the error rate of polymerase? How does this compare to the length of the human genome. How does biology deal with that discrepancy?
DNA polymerase makes about 1 error per 10⁴ bases during replication. Since the human genome is about 3 billion base pairs, this would result in roughly 300,000 errors per replication if left uncorrected. Biology handles this through proofreading and DNA repair systems, which dramatically reduce the final number of mistakes.
- How many different ways are there to code (DNA nucleotide code) for an average human protein? In practice what are some of the reasons that all of these different codes don’t work to code for the protein of interest?
Because 61 codons code for 20 amino acids, there are many possible DNA sequences that can encode the same protein. In reality, not all work well due to factors like tRNA availability, mRNA structure, and protein folding during translation, which affect how efficiently and accurately a protein is made.
Sources: ChatGPT Prompt: “Detail the most common different factors in which variant amino acid codes may fail and explain each”
ChatGPT Prompt: “How does biology deal with the error rate of DNA polymerase on a molecular level?”
Homework Questions from Dr. LeProust:
Most common oligo synthesis method: The most widely used method is solid-phase synthesis using phosphoramidite chemistry.
Why are oligos longer than ~200 nt hard to make? Errors build up during synthesis, making long sequences increasingly inaccurate.
Why can’t a 2000 bp gene be made directly? The error rate becomes too high, so genes are instead built from shorter oligos that are assembled together.
Homework Question from George Church:
The ten essential amino acids in animals are phenylalanine, valine, threonine, tryptophan, isoleucine, methionine, histidine, arginine, leucine, and lysine.
Because animals must get all of these from their diet, the “lysine contingency” proposed in Jurassic Park is not a strong safety mechanism as limiting lysine alone would not effectively control an organism.
Week 2 Homework
1. Benchling & In-silico Gel Art
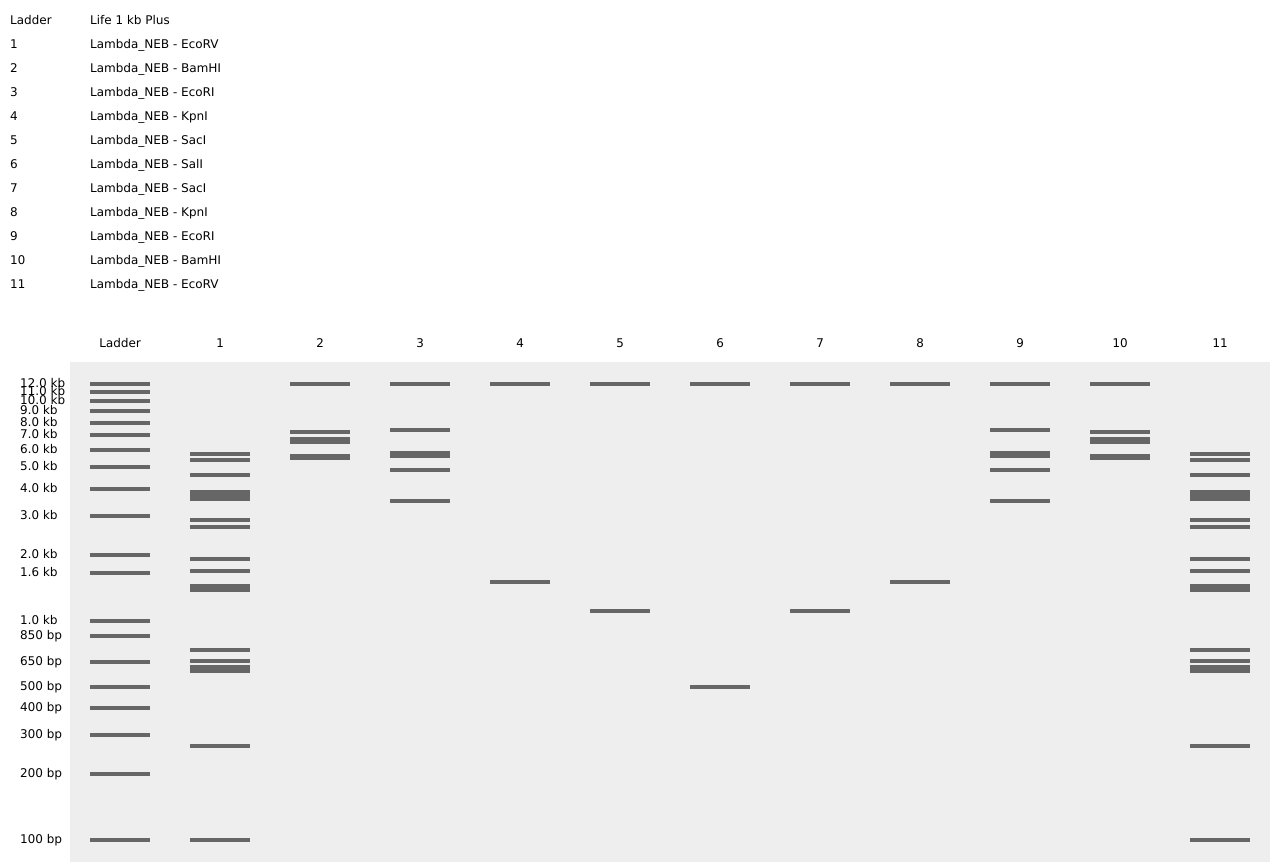


I made the letter M! (I grew up in Michigan, Go Blue!)
2. Gel Art - Restriction Digests and Gel Electrophoresis
I ran a gel in lab! I pipetted DNA into 10 wells, only 1 band shows up clearly besides the ladder, but there are a couple of fainter bands as well.
3. DNA Design Challenge
3.1: I chose the protein Magainin-2, an antimicrobial peptide isolated from the skin of the African Clawed Frog, Xenopus Laevis. I thought this would be an interesting protein to study because of its antibiotic properties and possible function as a resistance to chytrid fungus in amphibians.
Amino Acid Sequence: GIGKFLHSAKKFGKAFVGEIMNS
3.2:
reverse translation of Untitled to a 69 base sequence of most likely codons. ggcattggcaaatttctgcatagcgcgaaaaaatttggcaaagcgtttgtgggcgaaatt atgaacagc
3.3: Codon Optomization
Here is the optomized sequence using Xenopus Laevis as my chosen model organism.
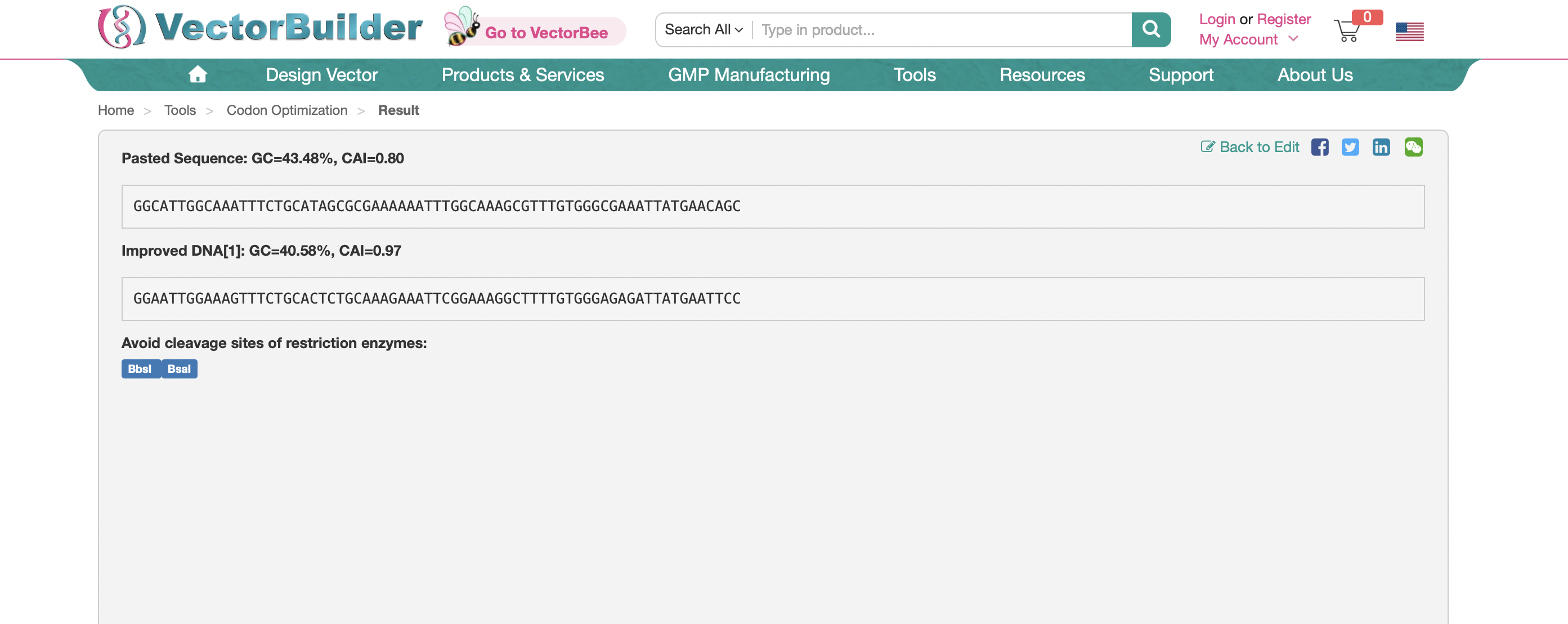

Here is the sequence using E.coli as the model organism for codon optomization: GGCATTGGCAAATTTCTGCATAGCGCGAAAAAATTTGGCAAAGCGTTTGTGGGCGAAATTATGAACAGC
E.Coli would be preferable to create this protein in for ease-of-use, widespread availability, and fast growth.
3.4 You Have A Sequence! Now What?
To synthesize a protein from this sequence, a common method used is cell-dependent, where I create a plasmid vector that includes a promoter, ribosome-binding sequence, my sequence, a terminator, and possibly a selection marker that helps verify that the plasmid is being expressed (such as an antibiotic resistance sequence). The plasmid can then be transformed into a host organism such as a bacteria or yeast, via electroporation or heat shock. The sequence is now able to express my protein!
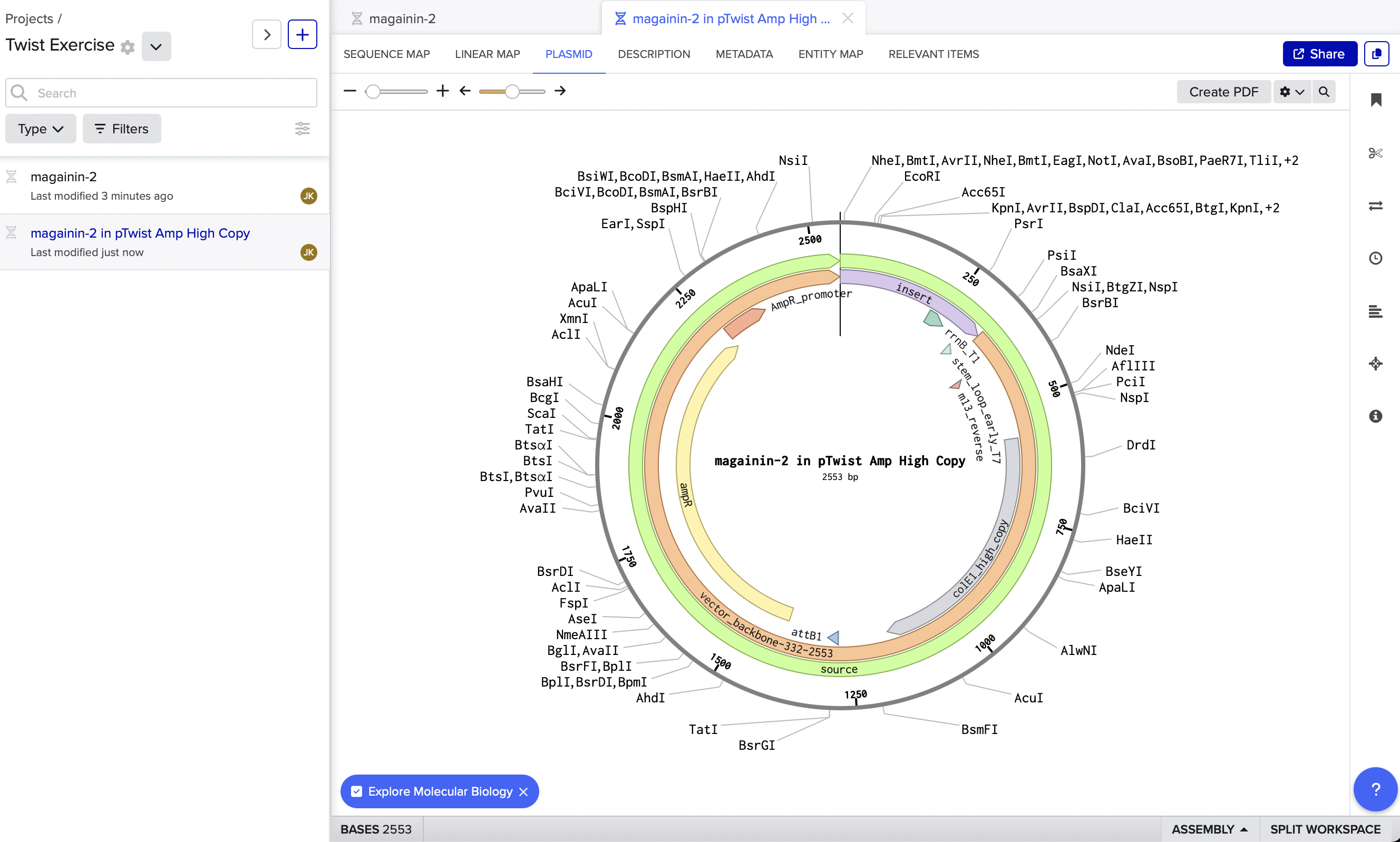
4. Prepare a Twist DNA Synthesis Order
Here is my Twist - generated plasmid in Benchling. My coding sequence was short, so I had to add in a neutral regulatory element upstream of my coding region so I had enough BP for Twist to generate the rest of the plasmid.

5. DNA Read/Write/Edit
5.1: If I were to sequence and read any DNA, I would love to read a full genome of an extinct species that still has available tissue for ancient DNA sequencing. It would be interesting to compare the genome of these extinct species to their closest relatives and see what genes are conserved and how they differ genetically. This would give some insight into the evolutionary history behind the speciation of that particular lineage and could give clues on how to safely and ethically bring back the species from extinction if it would help a current ecosystem ecologically.
5.2: