John Adedeji — HTGAA Spring 2026
Weeks
- Class Assignment — Week 1 1) Biological Engineering Application I aim to develop a computational and experimental platform for engineering metabolically constrained microbial systems designed for responsible real-world use. Inspired by clinical exposure to preventable infectious disease and my research at the intersection of microbiology and computational biology, the platform integrates genomic design rules, programmed auxotrophies, and environmental sensing circuits that couple microbial survival to defined ecological contexts.
- Class Assignment — Week 2 Part 1 — Sequence Retrieval and Design Workflow 1) Sequence Retrieval and Benchling Initialization The process began with obtaining a Lambda GenBank file from New England Biolabs. After confirming the correct format, I imported the file into Benchling as a DNA sequence. Care was taken to ensure that the file was not mistakenly uploaded as RNA and that annotations displayed properly within the platform.
- Class Assignment — Week 3 1) Opentrons Artwork 2) Published Papers Utilizing Automation LabscriptAI — Autonomous Liquid-Handling Robotics Scripting Gao et al., 2025 introduce LabscriptAI, a multi-agent framework that translates natural language experimental descriptions into validated Python scripts for heterogeneous liquid-handling robots, including Opentrons platforms.
- Class Assignment — Week 4 Part A. Conceptual Questions 1) How many molecules of amino acids do you take with a piece of 500 grams of meat? Assumptions: lean meat is ~20% protein by mass, average amino acid residue ~100 Da (≈100 g/mol).
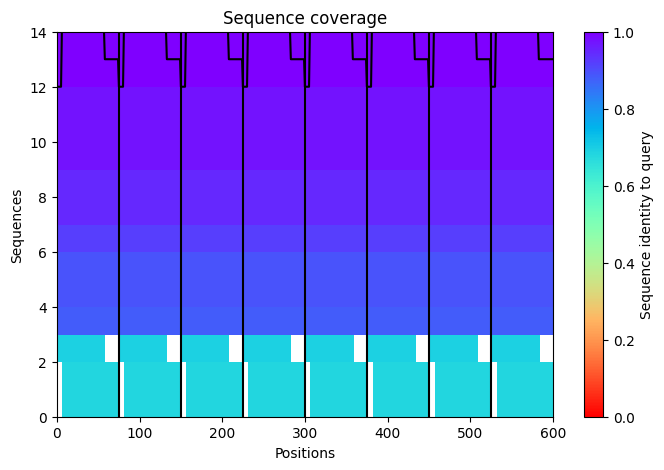
- Class Assignment — Week 5 Part A. SOD1 Binder Peptide Design Background ALS remains one of the more intractable neurodegenerative diseases partly because its genetic architecture is well-defined but hard to drug. The A4V mutation in SOD1 - a single alanine-to-valine substitution at residue 4 - is one of the most aggressive familial variants, accelerating disease progression significantly compared to other SOD1 mutations. The aggregation-prone nature of the A4V protein makes it an interesting peptide-binding target: if you can design a peptide that engages the misfolded or oligomerizing form, you potentially disrupt a key early step in motor neuron toxicity.
- Class Assignment — Week 6 Part A. DNA Assembly 1. Components of Phusion High-Fidelity PCR Master Mix A) Phusion DNA Polymerase A DNA-binding protein subunit that ensures higher template processivity, speed, and accuracy/fidelity alongside 5´→3´ polymerase activity and 3´→5´ exonuclease activity for proofreading.
- Class Assignment — Week 7 Part A. Intracellular Artificial Neural Networks (IANNs) 1. Advantages of IANNs over Boolean Genetic Circuits Boolean genetic circuits are fundamentally limited by their design logic: every input gets collapsed into a binary state, and the circuit operates on those discrete values. That works for simple switch-like decisions, but most physiologically relevant signals (metabolite concentrations, osmotic gradients, and quorum sensing molecule titres), exist on a continuum, and forcing them through a hard threshold discards information. IANNs avoid this by processing analog inputs directly, generating graded outputs that reflect the actual magnitude of the input rather than just which side of a threshold it fell on.
- Class Assignment — Week 9 Part A. General and Lecturer-Specific Questions 1. General homework questions 1. Advantages of Cell-Free Protein Synthesis Over In Vivo Methods Cell-free systems decouple protein production from cell viability, giving you direct control over reaction composition, temperature, redox state, and cofactor concentrations, none of which are easily tunable in living cells.
- Class Assignment — Week 10 Homework: Final Project ÌṢỌ is currently computational, so the “measurements” in scope are model outputs rather than physical assays. The key quantities I track are: steady-state pathogen kill rate as a function of MccH47 production, growth rate as a function of expression burden δ, biosensor activation ratio across tetrathionate concentrations, and containment escape probability over generational time. These are computed from ODE integration and Moran process simulation rather than physical instruments, but they map directly onto measurable biological quantities that would need experimental validation in a future phase of the project.
- Class Assignment — Week 11 Part A. Community Bioart Reflections | The 1,536 Pixel Artwork Canvas I contributed to the “Love” apple-shaped yellow sign at the mid-bottom of the artwork, working on the DNA assembly for that section of the plate.
- Homework + course notes; project constraints memo start.
- Homework + course notes; project constraints memo start.
- Homework + course notes; project constraints memo start.
Project
ÌṢỌ (Sentinel EcN)
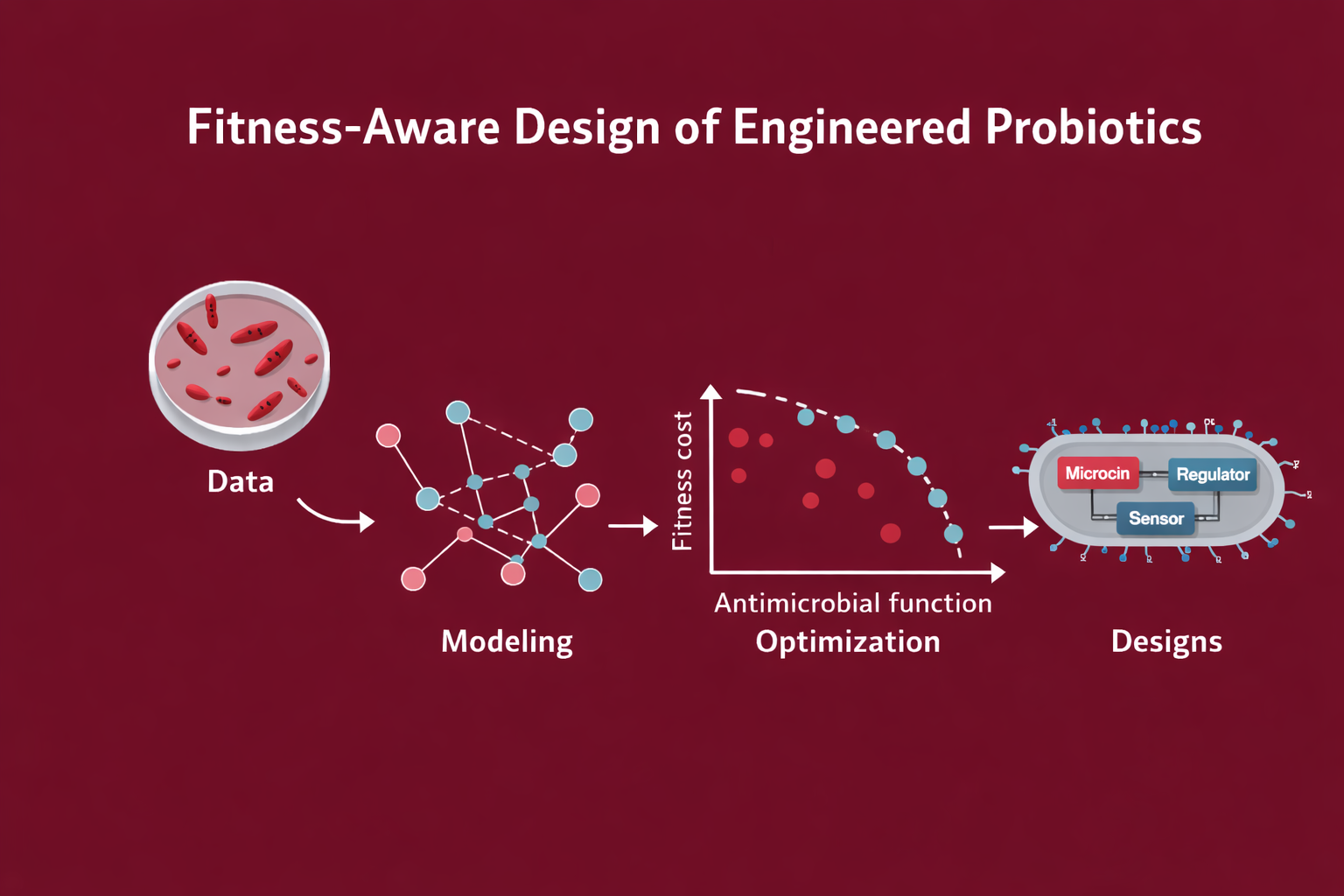
Fitness-aware design of engineered probiotics under ecological and evolutionary constraints.
This project is a model-first, constraint-aware approach to engineering E. coli Nissle 1917 (EcN) as a gut sentinel: sensing context, responding with targeted antimicrobials, and remaining governable through built-in containment.

Inspiration
During my medical training in Osogbo, diarrheal admissions became a rhythm I could not ignore. Children arrived dehydrated, eyes sunken, mothers anxious yet composed in that uniquely Nigerian way, strong because they had to be. We gave ORS, zinc, fluids. Sometimes antibiotics “just in case.” Sometimes it worked. Sometimes the silence afterward stayed with me longer than the ward round.
In microbiology, I encountered E. coli again, this time not only as culprit but as chassis. That shift lingered. What if the organism we blamed could be redesigned as a responder—quiet in health and active only when toxin or inflammatory signals rise—constrained and context bound, unable to persist beyond intention?
The idea was not dramatic. It was patterned. Repetition in the pediatric ward met ecological thinking in the lab. If microbes shape disease landscapes, perhaps they can also stabilize them—precisely, intelligently, and safely—within the same environments where I first learned to treat the consequences.
Why this matters
Childhood diarrhoeal disease remains high-burden with persistent treatment gaps, despite well-known interventions. The ambition here is not spectacle—it is reliable behavior under pressure: a responder that stays quiet in health, activates only under risk signals, and remains bounded by design.
Core design stance
Optimize for stability, not just performance.
I’m not chasing one “best construct.” I’m mapping design regimes: what works, what breaks, and what stays governable as conditions shift—fitness cost vs efficacy, signal vs noise, activation vs survivability.
System overview
ÌṢỌ is designed as a three-layer system:
- Detection: a biosensor tuned to a pathogen-associated signal or inflammation-linked marker
- Response: context-dependent expression of targeted antimicrobials (microcins)
- Containment: survival becomes conditional via metabolic dependency (“metabolic contract”)
Modeling assumptions & constraints
- Burden matters: expression cost is a first-class design variable, not a footnote
- Selection is always running: anything that reduces fitness will be negotiated by evolution
- The gut isn’t a flask: competition and variability are the setting, not edge-cases
- Outputs are design guidance: models inform what to build next, not clinical claims
- Containment is a system property: not only “does it exist,” but “does it hold under pressure?”
Out of scope (Spring 2026)
- Wet-lab validation
- Full microbiome ecosystem simulation
- Inventing novel antimicrobials
- Clinical deployment trials
- Regulatory implementation
Pipeline
Model → explore → optimize → stress-test.
The goal is to produce:
- reproducible computational models
- tradeoff plots (fitness vs efficacy)
- robustness/sensitivity analyses
- design regimes rather than a single “optimal” construct
Circuit modules
- Module 1 — Biosensor: reads a context signal and gates activation to reduce unnecessary burden
- Module 2 — Regulator: thresholded activation to limit leaky expression and improve stability under selection
- Module 3 — Effector (microcin): narrow-spectrum antimicrobial peptides aiming to pressure pathogens while minimizing broader disruption
- Module 4 — Containment: metabolic dependency to embed governance in biology
Governance & biosafety
Metabolic Dependency: if the engineered organism is made dependent on an externally supplied essential metabolite, it becomes non-viable without deliberate human-provided support.
Ecological Firewall: escapees cannot persist in nature, reducing ecological risk.
Human-Controlled Survival (“metabolic contract”): survival is coupled to oversight and supply chains, embedding accountability into the organism’s survival logic.
References
- Ba, F., Zhang, Y., Ji, X., Liu, W.-Q., Ling, S., & Li, J. (2023). Expanding the toolbox of probiotic Escherichia coli Nissle 1917 for synthetic biology. bioRxiv. https://doi.org/10.1101/2023.06.05.543671
- Egbewale, B. E., Karlsson, O., & Sudfeld, C. R. (2022). Childhood Diarrhea Prevalence and Uptake of Oral Rehydration Solution and Zinc Treatment in Nigeria. Children, 9(11), 1722. https://doi.org/10.3390/children9111722
- Gayawan, E., Cameron, E., Okitika, T., Egbon, O. A., & Gething, P. (2024). A situational assessment of treatments received for childhood diarrhea in the Federal Republic of Nigeria. PLOS ONE, 19(5), e0303963. https://doi.org/10.1371/journal.pone.0303963
- Lynch, J. P., Goers, L., & Lesser, C. F. (2022). Emerging strategies for engineering Escherichia coli Nissle 1917-based therapeutics. Trends in Pharmacological Sciences, 43(9). https://doi.org/10.1016/j.tips.2022.02.002
- Palmer, J. D., Piattelli, E., McCormick, B. A., Silby, M. W., Brigham, C. J., & Bucci, V. (2017). Engineered Probiotic for the Inhibition of Salmonella via Tetrathionate-Induced Production of Microcin H47. ACS Infectious Diseases, 4(1), 39–45. https://doi.org/10.1021/acsinfecdis.7b00114
- Weibel, N., Curcio, M., Schreiber, A., et al. (2024). Engineering a Novel Probiotic Toolkit in Escherichia coli Nissle 1917 for Sensing and Mitigating Gut Inflammatory Diseases. ACS Synthetic Biology, 13(8), 2376–2390. https://doi.org/10.1021/acssynbio.4c00036
- World Health Organization. (2024, March 7). Diarrhoeal Disease. https://www.who.int/news-room/fact-sheets/detail/diarrhoeal-disease