Projects

Final projects
Fitness-constrained design of a probiotic sentinel: a quantitative framework for circuit governability under evolutionary pressure.
HTGAA Group Project: Engineering the MS2 Bacteriophage L Protein

Fitness-constrained design of a probiotic sentinel: a quantitative framework for circuit governability under evolutionary pressure.
HTGAA Group Project: Engineering the MS2 Bacteriophage L Protein
ÌṢỌ — Yoruba: to be well; to recover. A fitness-aware engineered probiotic designed to sense gut context, respond with targeted antimicrobials, and remain governable by design.
Childhood diarrhoeal disease kills roughly half a million children under five every year, and the majority of those deaths happen in sub-Saharan Africa. During clinical training in Osogbo, the treatment options were ORS, zinc, and empirical antibiotics. Effective, but blunt. The gap is not a shortage of therapeutics. It is a precision problem.
How do we design microbial circuits that remain governable under the evolutionary and ecological pressures of a real gut environment?
The existing engineered probiotic literature optimises for peak performance under ideal conditions. ÌṢỌ maps design regimes: what works, what breaks, and what stays stable as conditions shift.
ÌṢỌ is a four-module sense-respond-contain system built on E. coli Nissle 1917 (EcN):
| Module | Component | Role |
|---|---|---|
| Biosensor | TtrS/TtrR two-component system | Detects tetrathionate, a pathogen-associated signal produced during gut inflammation by Salmonella and E. coli O157:H7 |
| Regulator | Thresholded Hill-function promoter | Gates activation; suppresses leaky expression and reduces fitness cost at homeostatic baseline |
| Effector | Microcin H47 (MccH47) | Narrow-spectrum antimicrobial; ATP synthase inhibition; active against Salmonella, Shigella, pathogenic E. coli; endogenous immunity protein MchI in EcN chassis |
| Containment | deltaDAPA auxotrophy | DAP absent from mammalian gut; deletion is lethal without exogenous supply; escape frequency ~10^-8 per generation |
Optimise for stability, not just performance. The output is not a single optimal construct. It is a map of design regimes: parameter regions where the circuit functions, where it fails under burden, and where containment holds under selection pressure.
Computational modelling only. Wet-lab validation, full microbiome simulation, and clinical deployment are explicitly out of scope for this phase.
The first aim of my final project is to build and simulate a genome-scale metabolic and circuit-level ODE model of the four-module ÌṢỌ architecture by utilising Tellurium/libroadrunner for time-course simulation, SALib for global sensitivity analysis, and NumPy-based Moran process modelling for evolutionary containment stability, generating a Pareto-resolved fitness-efficacy landscape and a ranked parameter influence analysis as the primary computational output.
Following a successful Aim 1, the top-ranked parameter regimes from the Pareto landscape will guide assembly and transformation of the sense-respond-contain circuit into EcN. The engineered sentinel will be tested in co-culture assays against Salmonella Typhimurium and E. coli O157:H7, validating both the tetrathionate-sensing threshold and MccH47-mediated kill kinetics experimentally. Discrepancies between model predictions and wet-lab data will feed back into model refinement.
The long-term goal is a rugged, orally delivered live biotherapeutic that operates autonomously in the gut, activates only in the presence of pathogen-associated tetrathionate, kills narrow-spectrum without collateral microbiome disruption, and cannot persist outside the host. If the fitness-governability framework holds, ÌṢỌ becomes a design methodology applicable beyond this specific pathogen set, with direct relevance for AMR management, inflammatory bowel disease, and cancer immunotherapy in low-resource clinical settings.
Palmer et al. (2017, ACS Infectious Diseases) demonstrated that EcN can be engineered to sense gut-luminal tetrathionate via the TtrS/TtrR two-component system and produce Microcin H47 in response, achieving measurable Salmonella inhibition in a mouse colonisation model. Critically for ÌṢỌ, this paper provides experimentally validated, ODE-parameterisable values for sensor activation kinetics, MccH47 production rates, and pathogen kill constants, making it the direct quantitative predecessor to this project rather than simply a conceptual reference.
Stritzker et al. (2007, International Journal of Medical Microbiology) characterised deltaDAPA auxotrophy in EcN in detail, reporting an escape frequency of approximately 10^-8 per generation under DAP-free conditions. That specific number is what makes the containment module computationally tractable: escape probability can be directly parameterised in the Moran process model rather than estimated from first principles.
What ÌṢỌ does that neither paper does is treat fitness cost as a first-class design variable rather than a post-hoc observation. Every published EcN engineering study acknowledges metabolic burden; none model it explicitly as a design input alongside efficacy. ÌṢỌ builds a Pareto frontier that makes the tradeoff navigable rather than anecdotal. The containment module also moves from binary characterisation (auxotrophy is present or absent) to a dynamic system property, asking how quickly a loss-of-function mutant fixes in a finite population over evolutionary time.
Diarrhoeal disease causes approximately 1.6 million deaths per year globally, with the under-five burden concentrated in West and East Africa. Nigeria alone accounts for a disproportionate share of this mortality. Existing interventions reduce severity but do not prevent recurrence in high-transmission settings, and empirical antibiotic use is accelerating resistance emergence in the pathogens most responsible for paediatric deaths: Salmonella, enterotoxigenic E. coli, and Shigella. A sentinel probiotic that activates conditionally, kills narrow-spectrum, and cannot persist outside the host addresses this without adding to AMR pressure.
Beyond the immediate clinical problem, the fitness-governability framework ÌṢỌ develops has broader implications. Any engineered living therapeutic faces the same core question: will the circuit hold under the evolutionary pressure of a real biological environment? Current regulatory frameworks for live biotherapeutics have no standardised computational tool for answering this before a clinical trial. ÌṢỌ begins building one. Nigerian and broader West African epidemiological data (Egbewale 2022; Gayawan 2024) are used to parameterise disease burden and clinical context from the start, not as a framing afterthought.
Two principles are directly engaged here: beneficence and justice. A precision antimicrobial that spares the commensal microbiome and cannot persist outside the host is strictly better than empirical broad-spectrum antibiotics for the patient, for the microbiome, and for the resistance landscape. Research that addresses paediatric mortality in West Africa while remaining computationally grounded in West African epidemiology represents a genuine departure from the default of developing interventions for high-income contexts and adapting them downstream.
The risks require honesty. A single deltaDAPA deletion is probably not sufficient for any real-world deployment. The current model assumes a closed population and does not account for horizontal gene transfer of the dapA gene from environmental bacteria. The Moran process also excludes commensal competition dynamics, so estimates of circuit persistence are optimistic. These are known limitations, explicitly scope-bounded to this computational phase. Non-maleficence requires that these caveats travel with any communication of the results. Open-source model release via GitHub (MIT licensed) is a deliberate act toward equitable access to the methodology.
Chosen: tetrathionate via TtrS/TtrR two-component system. Pathogen-specific: Salmonella and E. coli O157:H7 produce tetrathionate during gut inflammation via reactive oxygen species. Experimentally validated in EcN (Palmer et al. 2017). Signal is absent under homeostatic conditions, directly minimising leaky expression burden at baseline.
Chosen: Microcin H47 (MccH47). Naturally produced by EcN. Narrow-spectrum: E. coli, Salmonella, Shigella. Mechanism is ATP synthase inhibition, a well-characterised mode of action enabling direct ODE kill-kinetics parameterisation. Immunity protein MchI is endogenous to the EcN chassis. Palmer 2017 provides benchmarked production and kill-rate values for exactly this design.
Chosen: deltaDAPA auxotrophy (diaminopimelic acid / DAP). DapA is essential for lysine and peptidoglycan synthesis. DAP is absent from the mammalian gut: no dietary source, no commensal production. Deletion is lethal without exogenous supply. Validated in EcN (Stritzker et al. 2007). Published escape frequency ~10^-8 per generation is directly parameterisable for the containment escape model.
Chosen: Tellurium + libroadrunner (SBML/Antimony).
Purpose-built for systems biology ODE modelling. Antimony syntax maps directly onto circuit topology (promoter to mRNA to protein). libroadrunner’s stiff CVODE solver handles fast mRNA turnover and slow protein accumulation dynamics without manual configuration. SBML export makes every model citable and reproducible. SciPy solve_ivp (LSODA flag) runs in parallel for parameter sweeps and Pareto grid computation.
Primary: PRCC via SALib (Marino et al. 2008). Designed for nonlinear, monotonic systems, exactly what Hill-function gene circuits produce. 500 to 2000 Latin hypercube samples sufficient for 6 to 8 parameters.
Supplementary: Sobol total-order indices. Captures interaction effects (Hill coefficient n and KD interact in the sensor module). 5000 to 10000 samples, tractable on a laptop in minutes.
Chosen: Moran process with fitness-weighted selection.
Two competing types: functional circuit (fitness 1 minus delta) and loss-of-function mutant (fitness 1). Fixation probability computed analytically (Nowak 2006), then 1000 stochastic trajectories via numpy.random.choice() with fitness-weighted birth-death events. Directly answers: how long does the circuit remain functional under selection pressure?
Tellurium + libroadrunner — all four-module ODE construction and time-course simulation written in Antimony syntax. SBML export for reproducibility and citability.
SciPy solve_ivp (LSODA) — parameter sweeps and Pareto grid computation. LSODA auto-switches between stiff and non-stiff regimes.
SALib — PRCC for main figures, Sobol as supplementary. Canonical citation: Marino et al. 2008, J. Theor. Biol.
NumPy random.choice() — Moran process. Fitness-weighted birth-death events across 1000 independent trajectories. No additional dependencies.
Matplotlib + Seaborn (seaborn-whitegrid). Pareto scatter, PRCC tornado chart, containment escape semi-log, Moran fixation fan. Exported at 300 dpi PNG and SVG.
GitHub + venv + requirements.txt (pinned exact versions). One command regenerates all figures. MIT licensed. CITATION.cff included.
Install and configure the full modelling stack: Tellurium, libroadrunner, SALib, NumPy, Matplotlib, Seaborn, SciPy, all pinned in a venv with requirements.txt. Build the biosensor module as a two-ODE Hill-function model encoding the TtrS/TtrR tetrathionate-to-promoter activation pathway. Fit activation threshold KD and Hill coefficient n against Palmer 2017 time-course data.
Expected result: Simulated sensor activation curve matches digitised Palmer 2017 experimental data within 20% across the measured tetrathionate concentration range.
Extend the biosensor ODE to include the regulator module (thresholded Hill-function promoter gating effector expression), the effector module (MccH47 production and pathogen kill kinetics), and the containment module (deltaDAPA escape probability). Write all models in Antimony syntax within Tellurium. Export validated models to SBML and commit to GitHub.
Expected result: Stable steady-state solutions for all four modules under both homeostatic and pathogen-present conditions. Leaky expression at baseline should approach zero.
Use SciPy solve_ivp with LSODA flag to sweep burden parameter delta and effector output rate k_M across a 50 x 50 parameter grid. Record steady-state growth rate and pathogen suppression ratio for each grid point. Plot Pareto frontier, colour-coded by regulator variant (linear vs. thresholded).
Expected result: A visible Pareto frontier separating viable design space from over-burdened and under-effective regions. The thresholded regulator variant should dominate the frontier.
Run PRCC analysis via SALib using Latin hypercube sampling across 6 to 8 parameters: Hill coefficient n, signal threshold KD, burden delta, MccH47 production rate k_M, pathogen kill rate k_kill, mRNA degradation rate, and protein dilution rate. Generate ranked tornado chart. Run supplementary Sobol total-order index analysis.
Expected result: n and KD rank as the top two PRCC drivers of sensor module output. Sobol indices confirm a significant interaction effect between the two.
Implement the Moran process in NumPy. Define two competing cell types (functional circuit, fitness 1 minus delta; loss-of-function mutant, fitness 1). Compute analytical fixation probability from Nowak 2006. Run 1000 stochastic trajectories. Vary delta across the Pareto-viable range; plot fixation probability across three population sizes with analytical solution overlaid.
Expected result: Fixation probability of the loss-of-function mutant increases sharply above delta = 0.1. This is the quantitative argument for why the thresholded regulator module is not optional.
| Company | Role |
|---|---|
| Asimov (Kernel) | Validate Pareto landscape and containment circuit architecture; independent cross-check against Tellurium ODE results |
| SecureDNA | Screen all DNA sequences (mccH47, deltaDAPA cassette) before synthesis |
| Cultivarium | EcN-specific transformation protocols and characterised parts for Aim 2 |
| Twist Biosciences | Codon-optimised construct synthesis for Aim 2 |
| Opentrons | Co-culture assay automation for Aim 2 parallel screening |
Burden parameter delta and effector output k_M swept across a 50 x 50 parameter grid. Each point represents steady-state growth rate and pathogen suppression ratio. Pareto frontier overlaid. Colour-coded by circuit variant (linear vs. thresholded regulator). This figure makes the design regime concept concrete: the viable parameter space, the over-burdened region, and the under-effective region visible in a single plot. No equivalent exists in the published EcN engineering literature.
PRCC bar chart ranked by absolute influence on steady-state pathogen suppression. Parameters: Hill coefficient n, signal threshold KD, burden delta, microcin production rate k_M, pathogen kill rate k_kill. Sobol indices shown as supplementary to capture n-KD interaction effects. Identifies which parameters drive circuit behaviour most strongly, informing which design variables to constrain first in any future experimental build.
Semi-log plot of escape frequency vs. generations. Single deltaDAPA vs. dual deltaDAPA + deltaThyA auxotrophy compared. Analytical curve overlaid on stochastic simulation trajectories. Anchored to published escape frequency ~10^-8 per generation (Stritzker 2007).
Fixation probability of loss-of-function mutant as a function of burden delta, across three population sizes. 1000-trajectory stochastic fan with analytical Nowak 2006 solution overlaid. Demonstrates that the thresholded regulator (Module 2) extends functional circuit half-life under selection relative to constitutive expression: the quantitative argument for why the regulator module is not optional.
The primary validation for this phase is the biosensor module ODE fitted to Palmer 2017 experimental data, and the resulting Pareto landscape generated by the four-module parameter sweep. Together, these demonstrate that the computational framework is grounded in real parameterisation, not arbitrary simulation.
curve_fit with residual sum of squares minimisation. Report fitted values with 95% confidence intervals.The main challenge so far is parameter identifiability in the effector module. The MccH47 kill-rate constant k_kill is not directly reported in Palmer 2017; it is inferred from pathogen viable-count time-courses. The current approach uses a range of literature-sourced bacteriocin kill-rate values (106 to 108 cells per micromolar per hour) and propagates this uncertainty explicitly through the sensitivity analysis rather than selecting a single point estimate. This produces confidence intervals on the Pareto frontier that reflect real parametric uncertainty rather than false precision.
A second known limitation is the absence of commensal competition in the current Moran process model. Current fixation probability estimates are therefore optimistic by design. This is flagged explicitly in every figure caption involving evolutionary stability output. The Lotka-Volterra competition extension planned for post-course development is the right fix; it is out of scope here and is stated as such.
The engineered probiotic field asks: can we build a circuit that works?
ÌṢỌ asks: across what design regimes does a circuit remain both functional and governable under the pressures that will actually be present?
The models, figures, and write-up constitute the core of a bioRxiv preprint. Abstract, introduction, and discussion sections bring it to a citable first-author computational biology paper.
Add a simplified Lotka-Volterra competition term for commensal species. Explores how microbiome density affects EcN colonisation stability and circuit persistence under realistic ecological conditions.
Apply the PepMLM/moPPIt peptide generation pipeline (HTGAA Week 5) to propose microcin-analog sequences with improved target specificity. AlphaFold3 structural prediction of microcin-pathogen outer membrane protein complexes bridges the computational peptide design and engineered probiotic work.
Parameterise the pathogen kill model with AMR prevalence data from Nigerian clinical isolates (WHONET/GLASS). Grounds the model in Sub-Saharan African epidemiology and connects to the planned AMR West Africa genomic data paper.
All model code, SBML files, and figures. MIT licensed. CITATION.cff included.
Palmer, J. D., Piattelli, E., McCormick, B. A., Silby, M. W., Brigham, C. J., and Bucci, V. (2017). Engineered probiotic for the inhibition of Salmonella via tetrathionate-induced production of microcin H47. ACS Infectious Diseases, 4(1), 39-45. https://doi.org/10.1021/acsinfecdis.7b00114
Weibel, N., Curcio, M., Schreiber, A., et al. (2024). Engineering a novel probiotic toolkit in Escherichia coli Nissle 1917 for sensing and mitigating gut inflammatory diseases. ACS Synthetic Biology, 13(8), 2376-2390. https://doi.org/10.1021/acssynbio.4c00036
Lynch, J. P., Goers, L., and Lesser, C. F. (2022). Emerging strategies for engineering Escherichia coli Nissle 1917-based therapeutics. Trends in Pharmacological Sciences, 43(9). https://doi.org/10.1016/j.tips.2022.02.002
Ba, F., Zhang, Y., Ji, X., Liu, W.-Q., Ling, S., and Li, J. (2023). Expanding the toolbox of probiotic Escherichia coli Nissle 1917 for synthetic biology. bioRxiv. https://doi.org/10.1101/2023.06.05.543671
Stritzker, J., Weibel, S., Hill, P. J., Oelschlaeger, T. A., Goebel, W., and Szalay, A. A. (2007). Tumor-specific colonisation, tissue distribution, and gene induction by probiotic E. coli Nissle 1917 in live mice. International Journal of Medical Microbiology, 297(3), 151-162.
Marino, S., Hogue, I. B., Ray, C. J., and Kirschner, D. E. (2008). A methodology for performing global uncertainty and sensitivity analysis in systems biology. Journal of Theoretical Biology, 254(1), 178-196.
Nowak, M. A. (2006). Evolutionary Dynamics: Exploring the Equations of Life. Harvard University Press.
Moran, P. A. P. (1958). Random processes in genetics. Mathematical Proceedings of the Cambridge Philosophical Society, 54(1), 60-71.
Egbewale, B. E., Karlsson, O., and Sudfeld, C. R. (2022). Childhood diarrhea prevalence and uptake of oral rehydration solution and zinc treatment in Nigeria. Children, 9(11), 1722.
Gayawan, E., Cameron, E., Okitika, T., Egbon, O. A., and Gething, P. (2024). A situational assessment of treatments received for childhood diarrhoea in the Federal Republic of Nigeria. PLOS ONE, 19(5), e0303963.
World Health Organization. (2024). Diarrhoeal disease. https://www.who.int/news-room/fact-sheets/detail/diarrhoeal-disease
Authored and reviewed by:
This document captures the full scope of our group work within the Genspace node focused on engineering the MS2 bacteriophage L protein. Group 2 formed around a shared interest in improving the toxicity, stability, and tunability of the L protein through computational design.
Our early brainstorming sessions centered on three broad goals:
After several meetings and independent exploration, the group converged on two main computational directions. The first centered on systematic truncation and mutagenesis of the N-terminal regulatory domain. The second focused on point mutations within conserved regions that could alter electrostatic interactions while preserving structure.
Two major pipelines emerged from that work. John’s pipeline explored N-terminal truncations, DnaJ disruption, sequence redesign, codon optimization, and sequencing validation. Eric’s pipeline focused on charge-based mutations, conservation mapping, structural modeling, ORF overlap analysis, and cross-referencing with experimental lysis data.
Both approaches identified strong but distinct candidates for improving L protein function.
The MS2 lysis protein L is a 75 amino acid single-pass transmembrane protein whose N-terminal region acts as a regulatory brake on lysis. Rather than directly participating in membrane disruption, this region delays insertion and oligomerization of the transmembrane domain.
My pipeline focused on systematically removing portions of that inhibitory region while preserving the membrane-spanning lytic core. The central hypothesis was simple: if the N-terminal domain slows lysis, then partial removal should release that inhibition and produce earlier, stronger lytic activity.
The strongest candidate to emerge from the analysis was L_trunc30, which removes the first 30 amino acids while preserving the entire transmembrane domain.
Confirmed L protein sequence:
Confirmed DNA sequence:
Three ideas guided the pipeline:
| Stage | Tool | Purpose |
|---|---|---|
| 1 | ESM2 | Mutational scanning across all 75 residues |
| 2 | ESMFold | Structural prediction of truncation variants |
| 3 | AlphaFold-Multimer | Modeling interaction with DnaJ |
| 4 | GROMACS | Molecular dynamics and RMSF analysis |
| 5 | ProteinMPNN | Junction redesign and charge reduction |
| 6 | Codon optimization | Prepare E. coli expression constructs |
| 7 | Synthetic construct design | Assemble expression cassette |
| 8 | Bowtie2 + BCFtools | Variant calling and sequencing validation |
| 9 | IGV | Manual inspection of called variants |
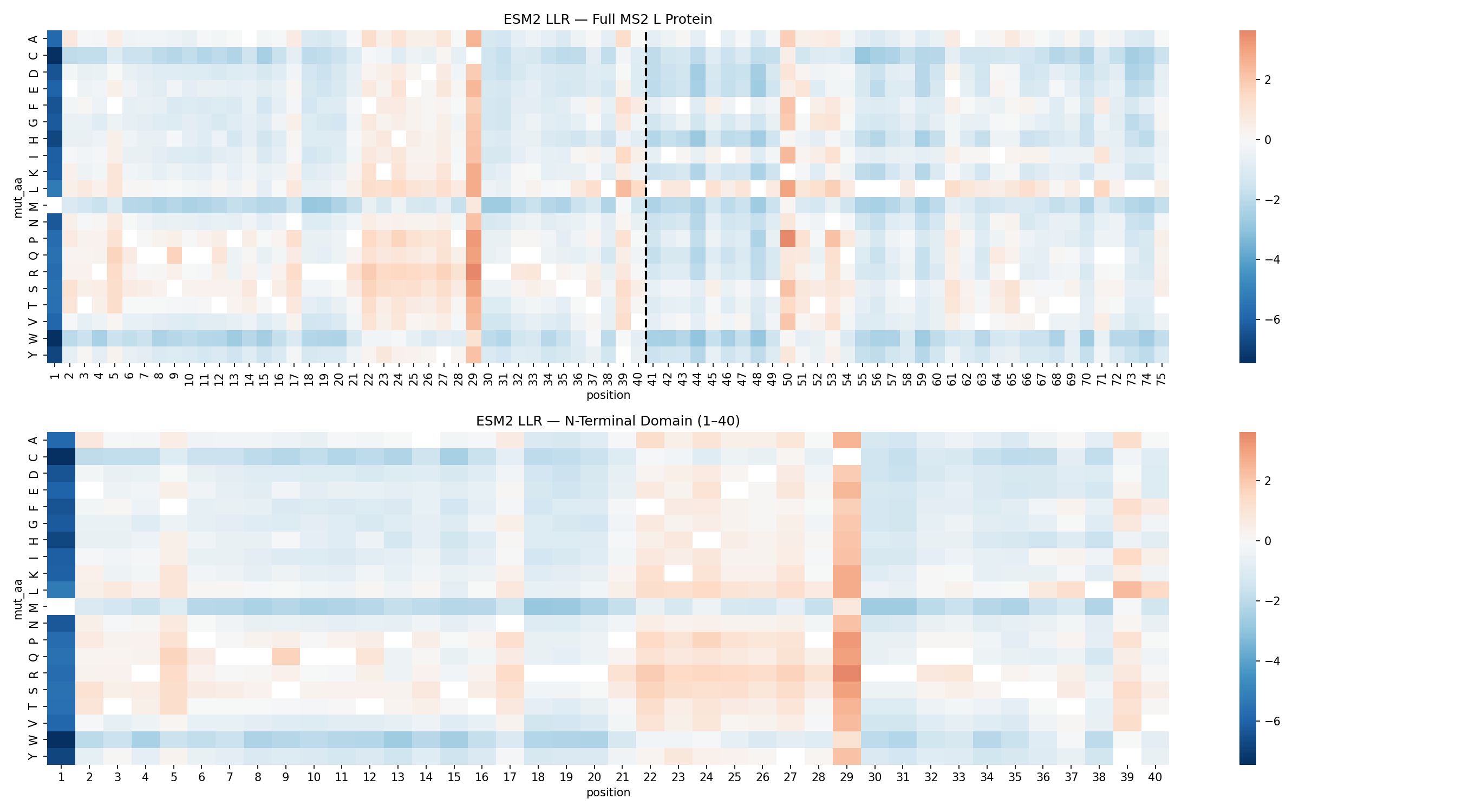

The ESM2 scan identified position C29 as the dominant mutational hotspot in the N-terminal domain.
| Mutation | LLR | Notes |
|---|---|---|
| C29R | 3.64 | Top-ranked substitution |
| C29P | 3.17 | Strong helix-disrupting mutation |
| C29Q | 3.06 | Conservative but highly favored |
| F22R | 1.86 | Introduces basic charge |
| S9Q | 1.69 | Recovered independently in prior work |
C29 accounted for 12 of the top 20 substitutions. That concentration strongly suggested that the wild-type residue at this site is not ideal for maximizing toxicity outside the native viral context.
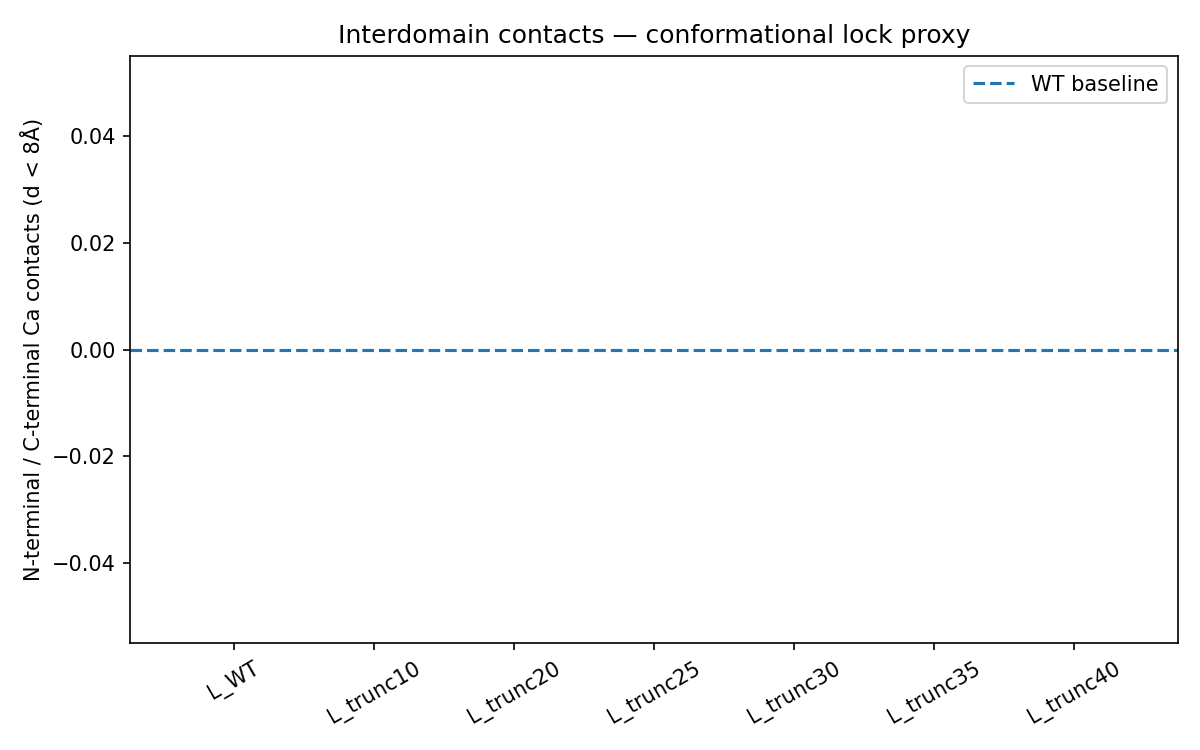

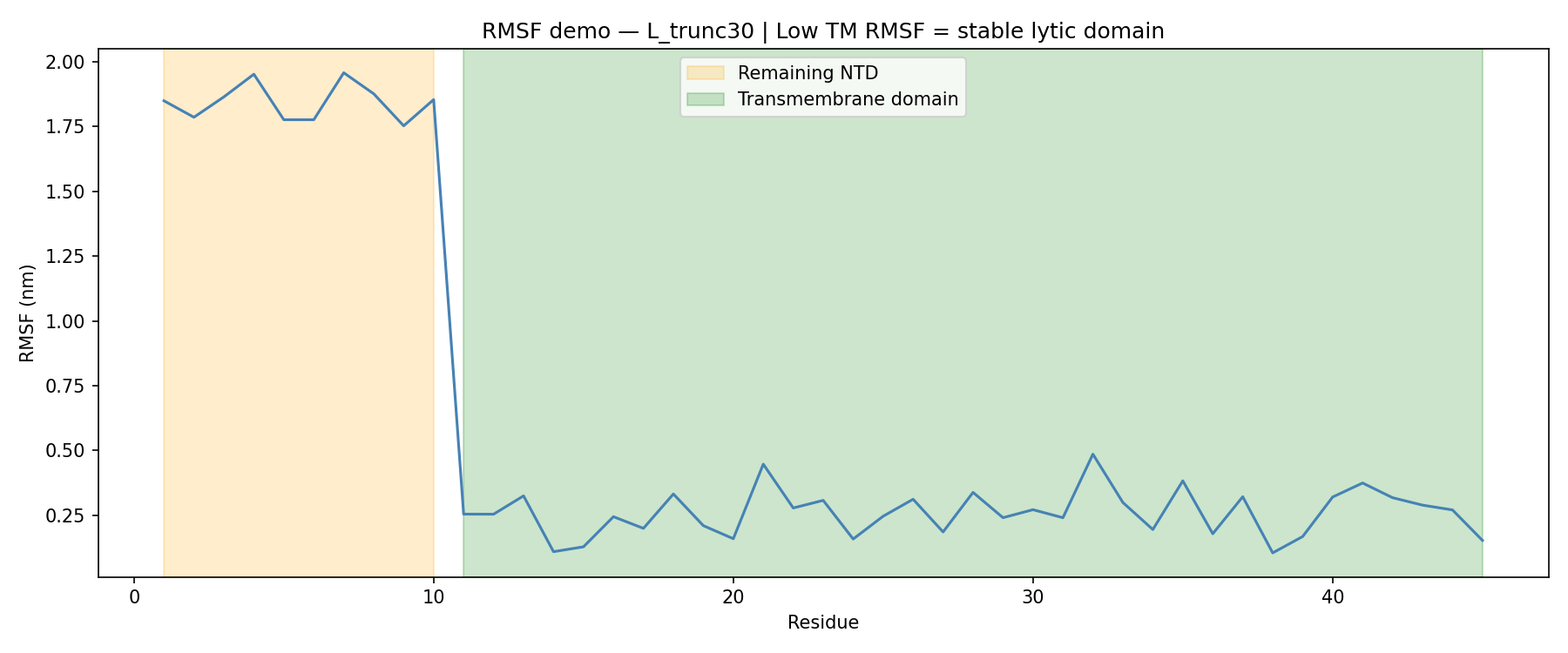

ESMFold predictions for all truncation variants suggested that the N-terminal domain is highly disordered in solution. Interdomain contact analysis returned essentially zero contacts across all variants, which fits with the known biology of the L protein.
The more useful signal came from molecular dynamics.
For L_trunc30:
That sharp drop in flexibility confirmed that the transmembrane region remains stable even after removing 30 amino acids from the N-terminus.
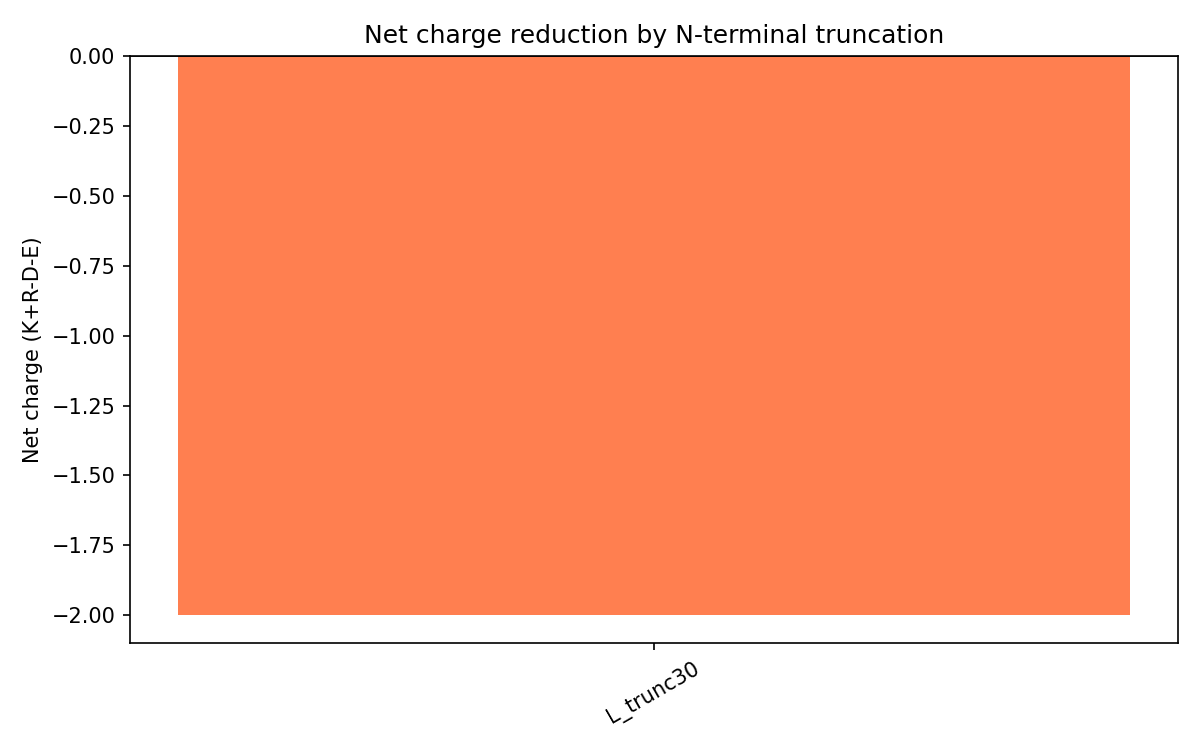

The wild-type N-terminal region is strongly basic due to motifs like RRRPFK and RRQQR.
L_trunc30 reverses the overall charge profile:
| Variant | Net charge | Interpretation |
|---|---|---|
| Wild-type L | Approximately +8 | Strong DnaJ interaction expected |
| L_trunc30 | -2 | Reduced DnaJ binding and earlier lysis expected |
This was important mechanistically because DnaJ binding depends heavily on electrostatic interactions with the positively charged N-terminal region.
All major truncation variants were codon-optimized for E. coli K-12.
The lead construct, L_trunc30, preserved the essential LS motif and was assembled into a complete 230 bp expression cassette with:
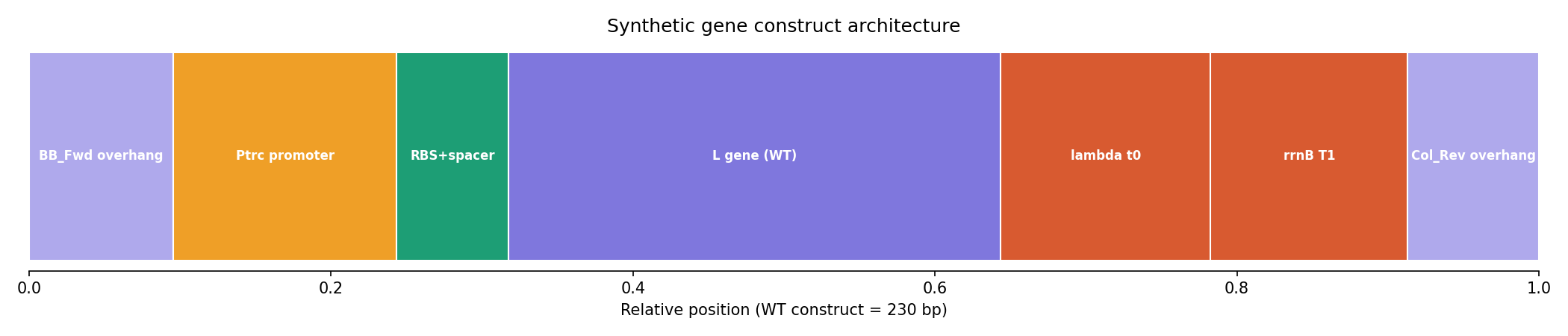

| Candidate | Key Feature | Reason |
|---|---|---|
| L_trunc30 | Removes aa 1-30 | Strongest balance of toxicity, structural stability, and DnaJ disruption |
| Candidate | Reason for Inclusion |
|---|---|
| C29R | Highest ESM2 score overall |
| F22R | Adds positive charge in N-terminal region |
| S9Q | Recovered independently in previous scans |
| L_trunc40 | Most aggressive truncation, likely strongest toxicity |
Eric approached the same problem from a different angle. Instead of removing large sections of the N-terminus, he focused on identifying individual amino acid substitutions that could improve toxicity while preserving the overall structure of the protein.
His strongest candidate was P13L, a single amino acid change in the N-terminal region.
| Stage | Tool | Purpose |
|---|---|---|
| 1 | UniProt + BLAST | Sequence retrieval and homolog identification |
| 2 | Clustal Omega | Conservation mapping |
| 3 | AlphaFold-Multimer | Oligomer modeling |
| 4 | ESM2 | Mutation scoring |
| 5 | ESMFold | Structural confidence and pTM analysis |
| 6 | ChimeraX | Electrostatic visualization |
| 7 | Benchling | ORF overlap analysis |
Eric identified a relatively unconstrained region between amino acids 16 and 28 that could tolerate mutation without damaging essential structure.
| Position | Wild-type residue | Interpretation |
|---|---|---|
| 18 | R | Fully conserved, avoid |
| 21 | P | Fully conserved, avoid |
| 23 | K | Fully conserved, avoid |
| 26 | D | Variable, strong candidate |
| 13 | P | Weakly conserved, potentially safe |
P13L produced the strongest ESMFold result among all variants tested.
| Variant | pTM | Change vs WT |
|---|---|---|
| Wild-type | 0.273 | Reference |
| D26R | 0.267 | Slight decrease |
| P13L | 0.420 | Strong increase |
The jump from 0.273 to 0.420 made P13L the most structurally favorable point mutation in Eric’s pipeline.
Unlike my pipeline, Eric cross-referenced computational candidates with available lysis data.
| Mutation | Replicate A | Replicate B | Result |
|---|---|---|---|
| P13L | 1 | 1 | Confirmed lytic |
| D26G | 1 | 0 | Mixed |
| K23E | 1 | 0 | Mixed |
| E25G | 1 | 0 | Mixed |
P13L was the only candidate to remain consistently positive across both replicates.
One of the more interesting parts of Eric’s work was the DNA-level overlap analysis.
P13L falls within the overlap region between the coat protein and the L protein, which initially made it look risky. After codon-level analysis, though, the mutation turned out to be safe.
| Gene | WT codon | Mutant codon | Result |
|---|---|---|---|
| L protein | CCG | CTG | Pro → Leu |
| Coat protein | TCC | TCT | Ser → Ser |
That synonymous change in the coat protein meant the mutation could proceed without disrupting the overlapping reading frame.
| Candidate | Key Feature | Reason |
|---|---|---|
| P13L | Single amino acid substitution | Best structural score and strongest experimental support |
| Candidate | Status |
|---|---|
| D26R | Untested but promising |
| D26G | Mixed experimental results |
| N17R | Open candidate |
| H24R | Open candidate |
Albert focused primarily on structural stability.
His workflow emphasized:
His key concern was preserving structure while introducing beneficial mutations.
He also pointed out an important limitation that kept showing up across the project: membrane proteins are underrepresented in both structural databases and protein language model training sets. That means even high-scoring mutations should still be interpreted cautiously.
Tehseen’s approach aligned closely with my truncation-based strategy but focused more on identifying the smallest regulatory segment required for precise control over lysis timing.
The central idea was not simply to remove the N-terminal region, but to identify exactly which residues are responsible for slowing lysis.
That led to three closely related hypotheses:
| Aspect | John’s Pipeline | Eric’s Pipeline |
|---|---|---|
| Main strategy | Progressive N-terminal truncation | Point mutation design |
| Lead candidate | L_trunc30 | P13L |
| Core hypothesis | Remove inhibitory domain | Increase local electrostatic effects |
| ESM2 scope | Full 1,425-substitution scan | Single-site targeted analysis |
| Structural analysis | ESMFold + GROMACS RMSF | ESMFold + ChimeraX |
| DnaJ interaction | Central to model | Considered indirectly |
| Experimental validation | Not yet completed | P13L confirmed experimentally |
| Construct design | Fully assembled | Still planned |
| Sequencing workflow | Fully designed with Bowtie2, BCFtools, IGV | Listed as future step |
The project ended up producing two very different but complementary engineering directions.
L_trunc30 represents the stronger systems-level redesign. It removes the inhibitory N-terminal region, reduces DnaJ engagement, preserves the transmembrane core, and provides a fully buildable expression construct ready for synthesis and sequencing validation.
P13L represents the cleaner minimal-change strategy. It preserves the full-length protein, improves structural confidence, survives ORF overlap analysis, and already has positive experimental support.
If the goal is maximum disruption of the native regulatory system, L_trunc30 is the stronger candidate.
If the goal is a simpler mutation with lower engineering risk and existing wet lab support, P13L is the better starting point.
The most practical next step would be to synthesize and compare both side by side.