Week 2
Class Assignment — Week 2
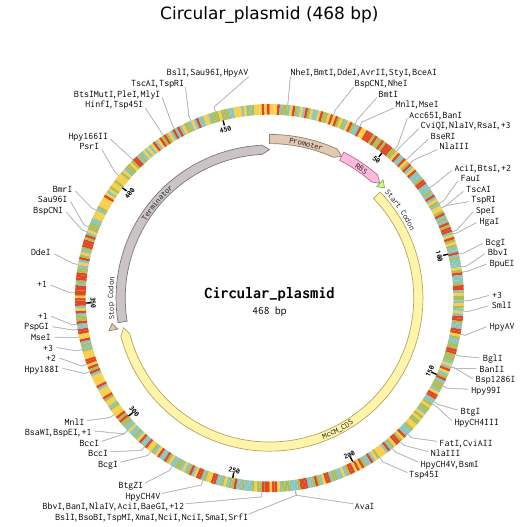



Part 1 — Sequence Retrieval and Design Workflow
1) Sequence Retrieval and Benchling Initialization
The process began with obtaining a Lambda GenBank file from New England Biolabs. After confirming the correct format, I imported the file into Benchling as a DNA sequence. Care was taken to ensure that the file was not mistakenly uploaded as RNA and that annotations displayed properly within the platform.
This step established a stable working environment before any design modifications were introduced. Confirming correct topology and annotation structure prevented downstream formatting or visualization issues.
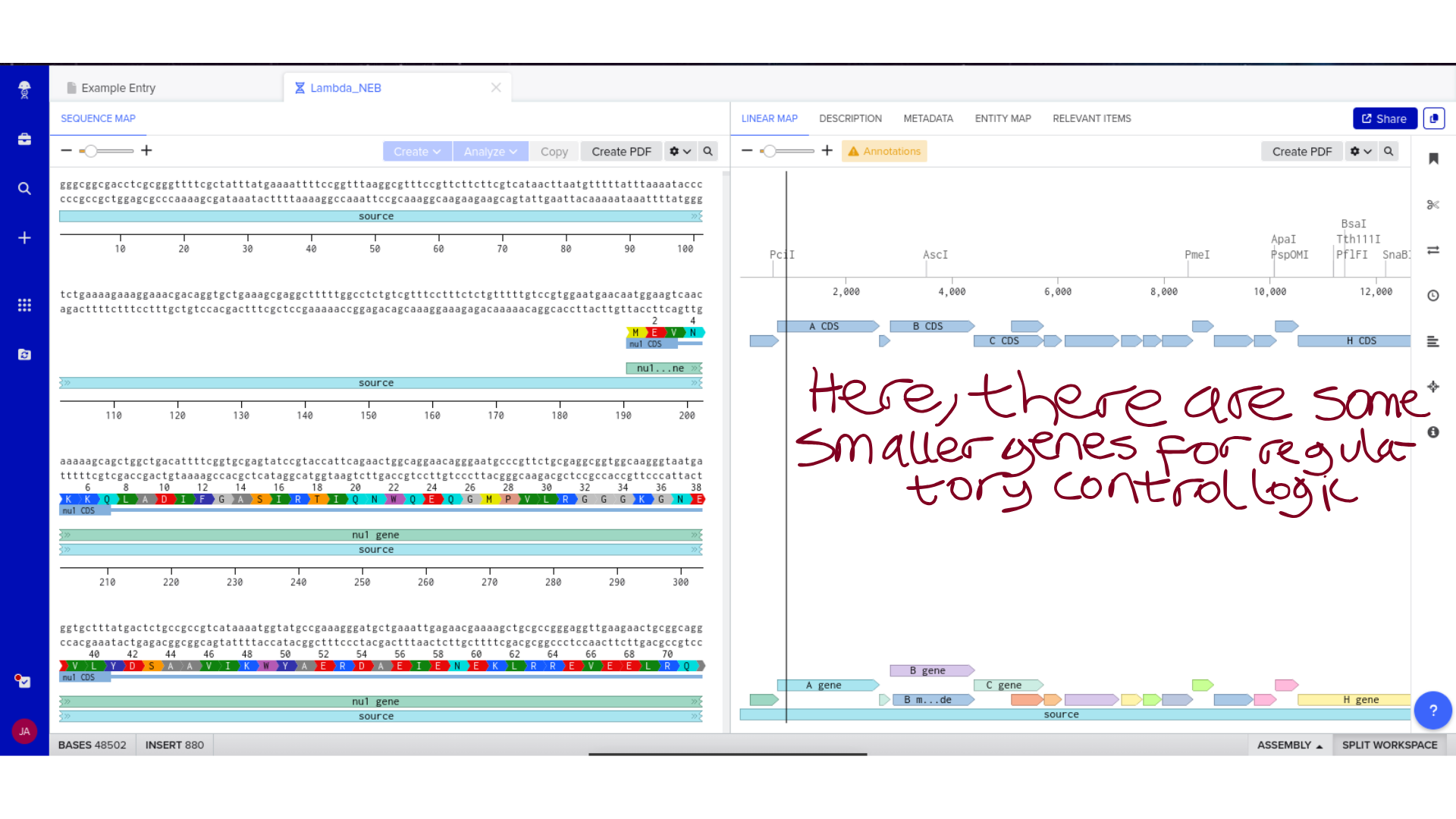
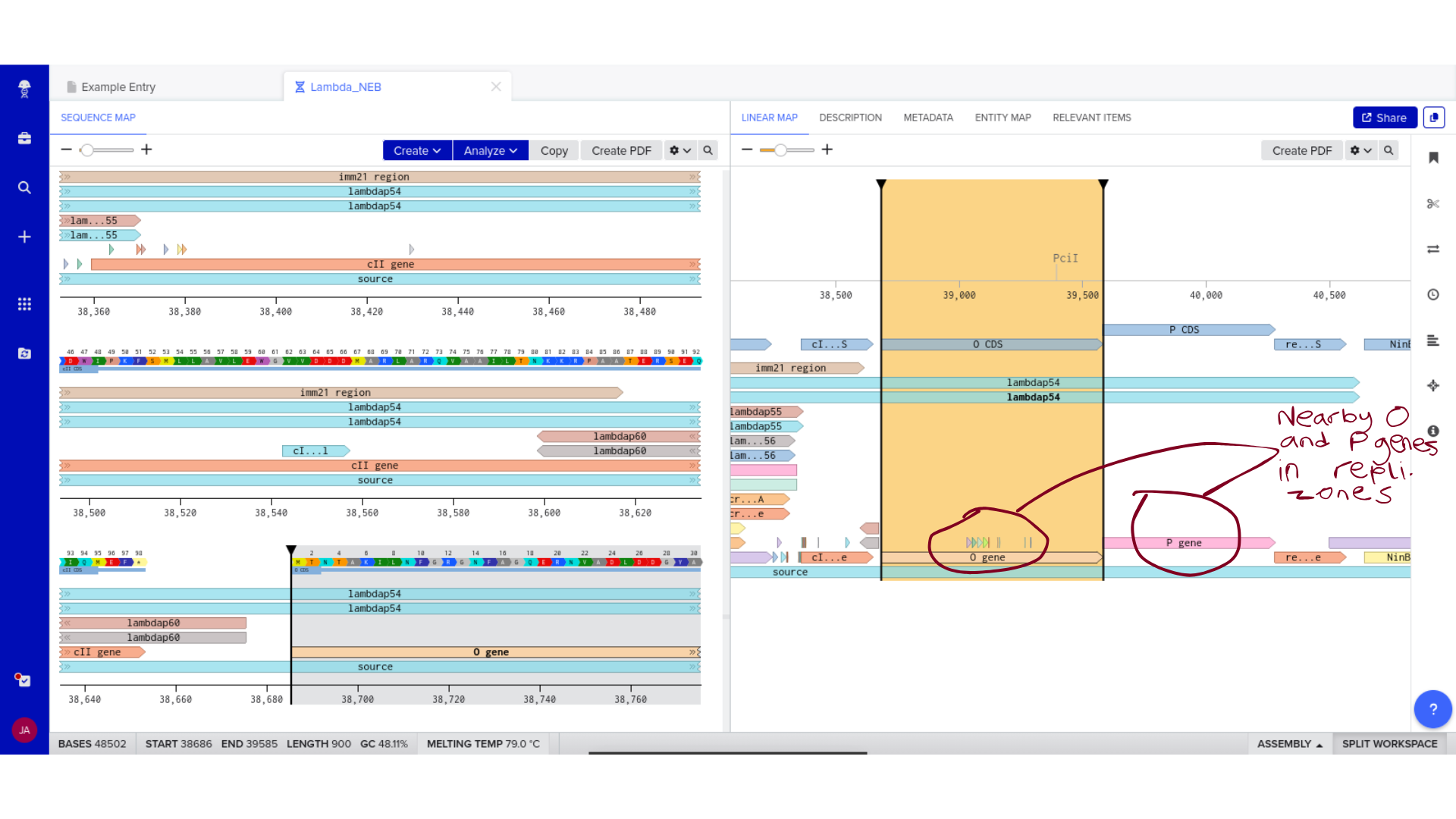
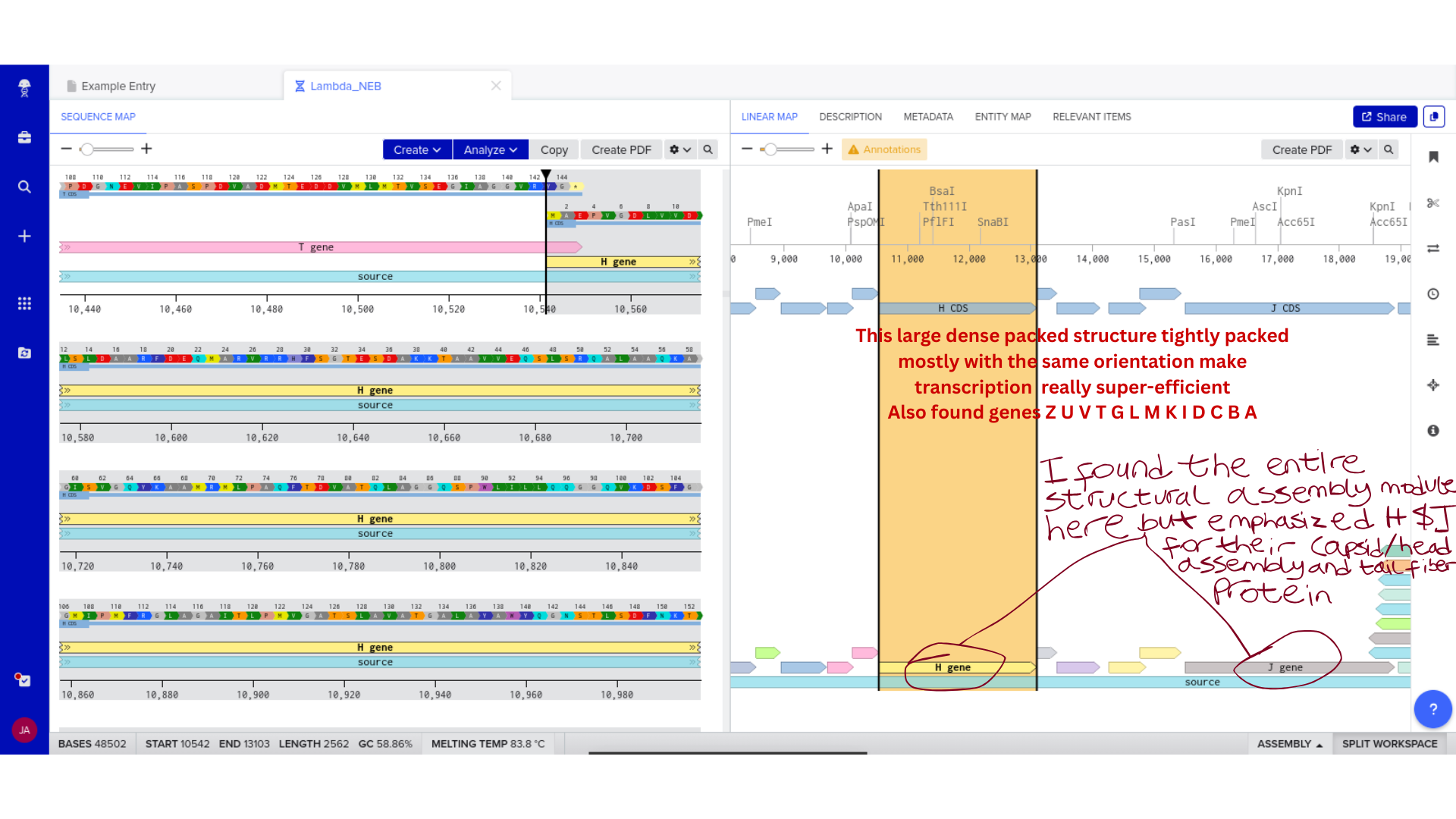
2) Genomic Exploration and Annotation Familiarization
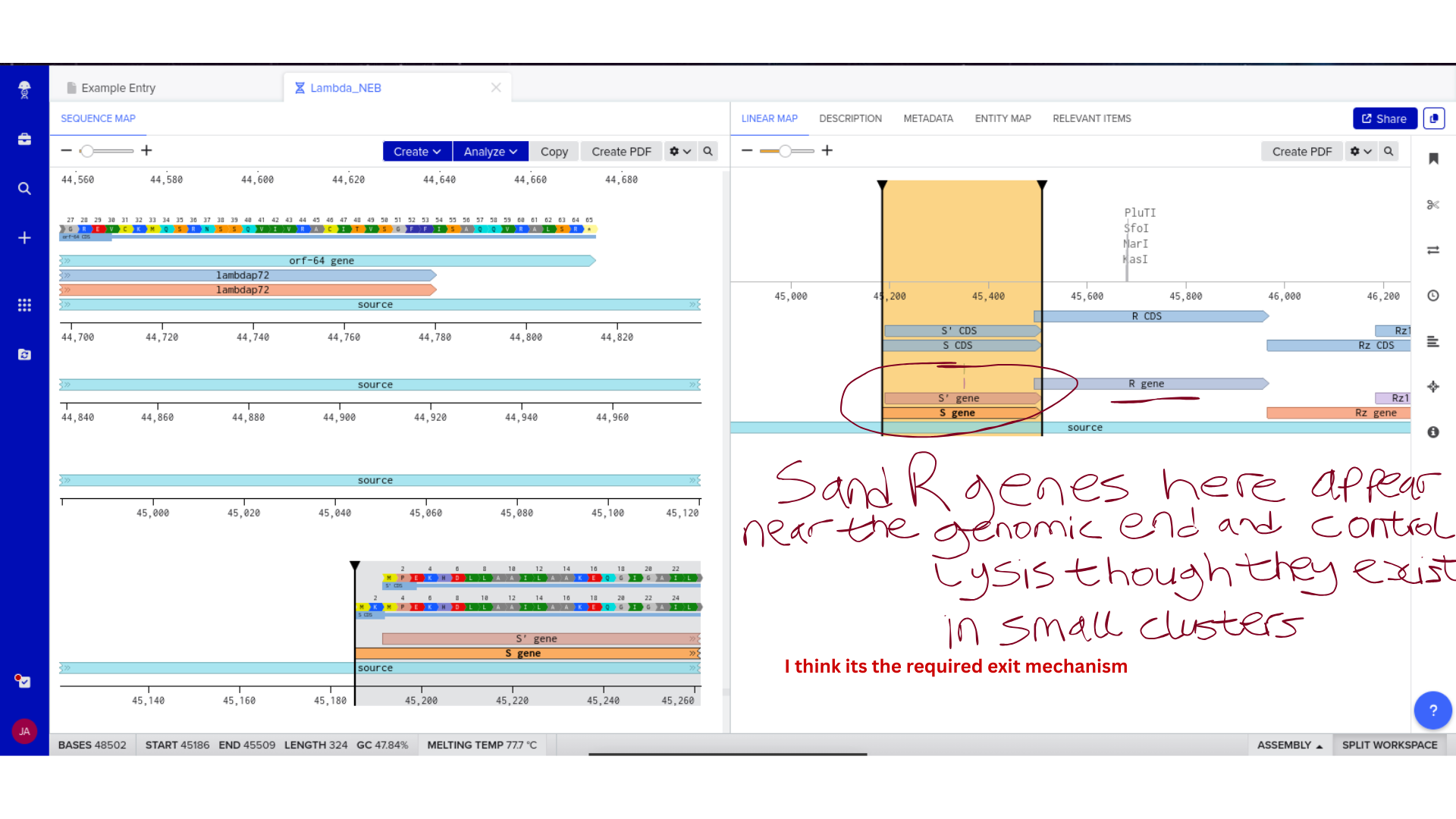
Once imported, I explored the annotated regions of the Lambda genome within Benchling. This involved confirming gene orientation, identifying labeled regions, and understanding the graphical interface for both linear and circular visualization.
Although exploratory, this step reinforced familiarity with the design environment. It ensured that I could distinguish between expected gene clusters and annotation artifacts, and that I could confidently navigate the interface for subsequent editing.
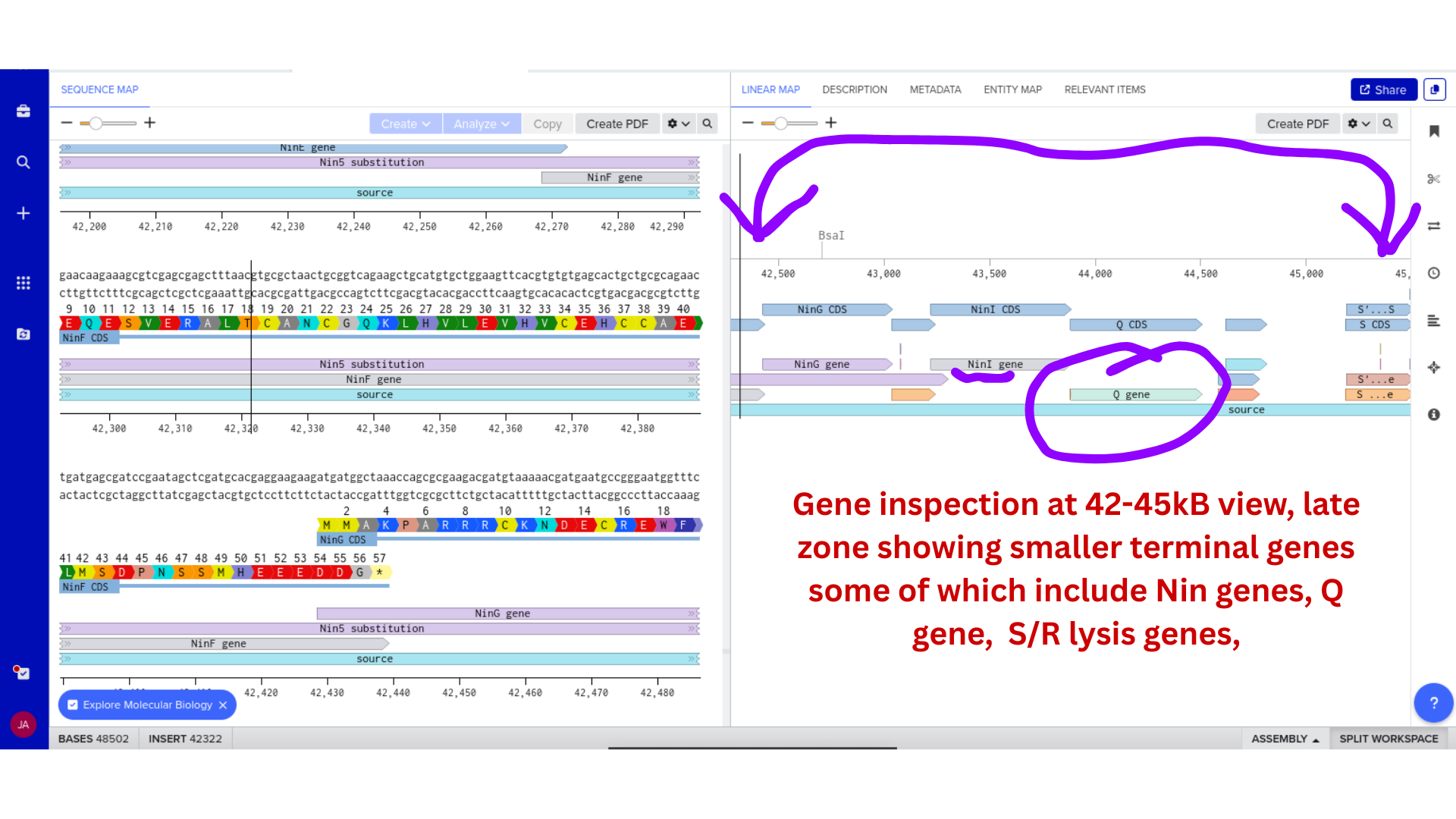

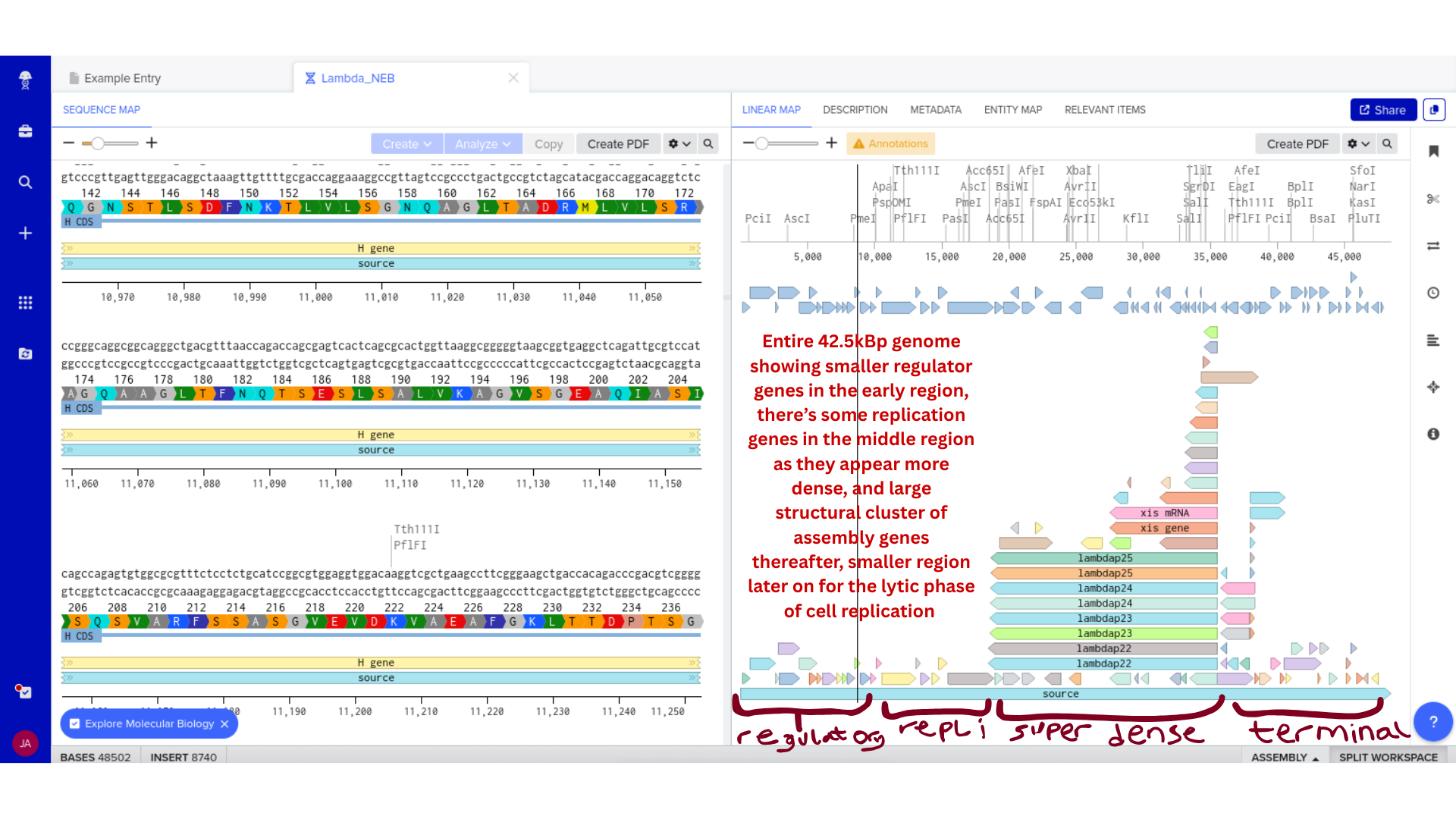

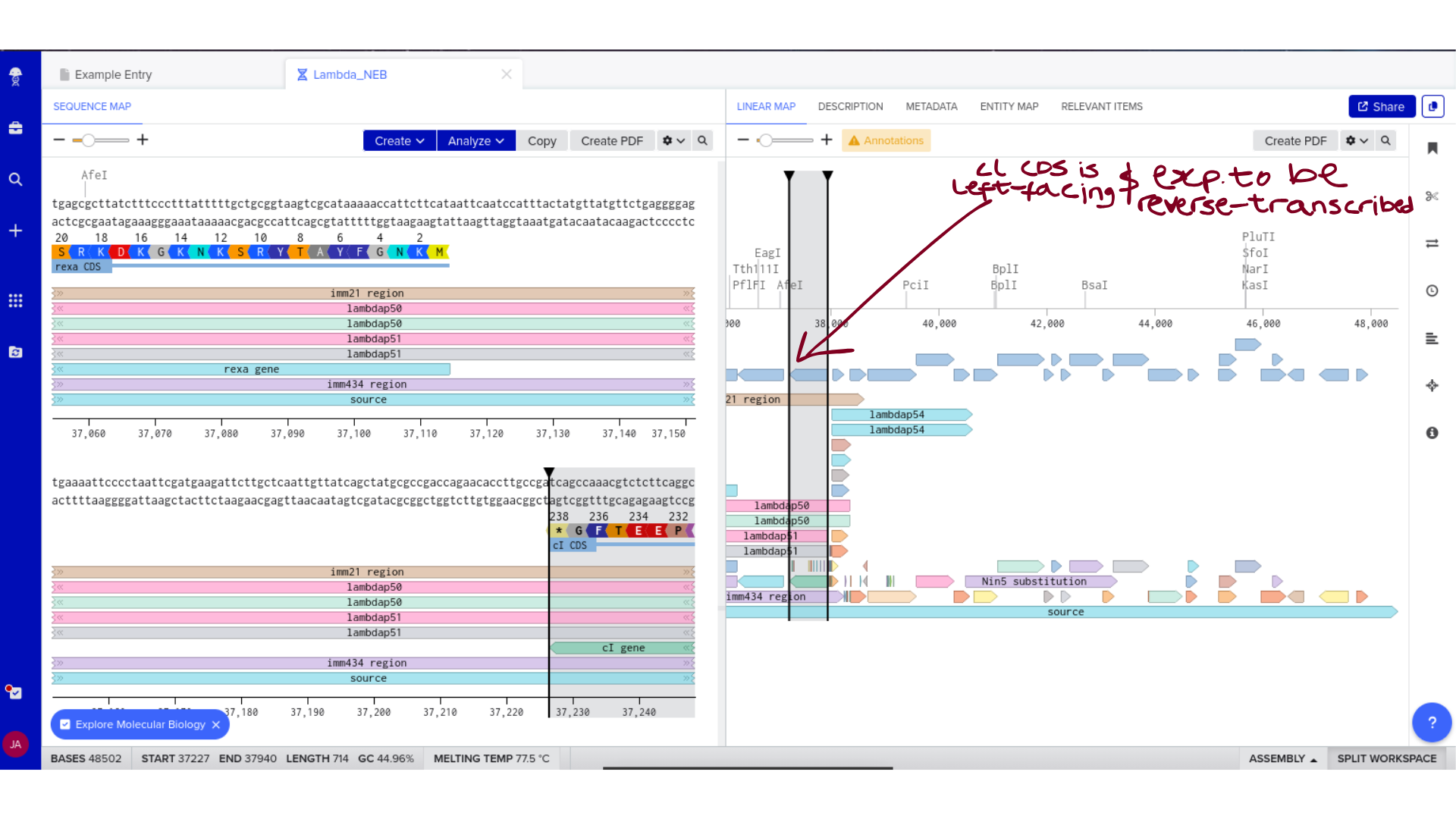

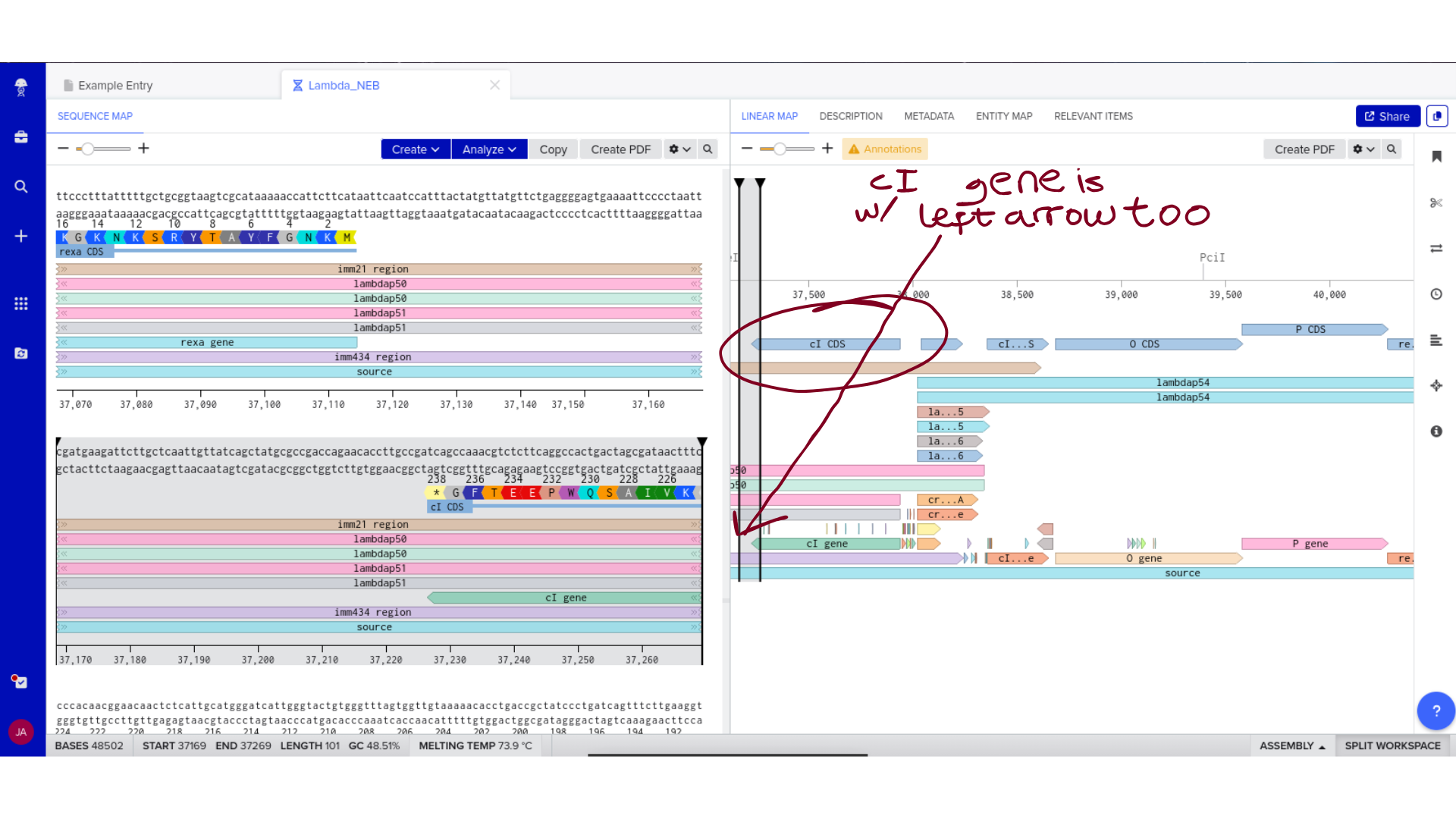

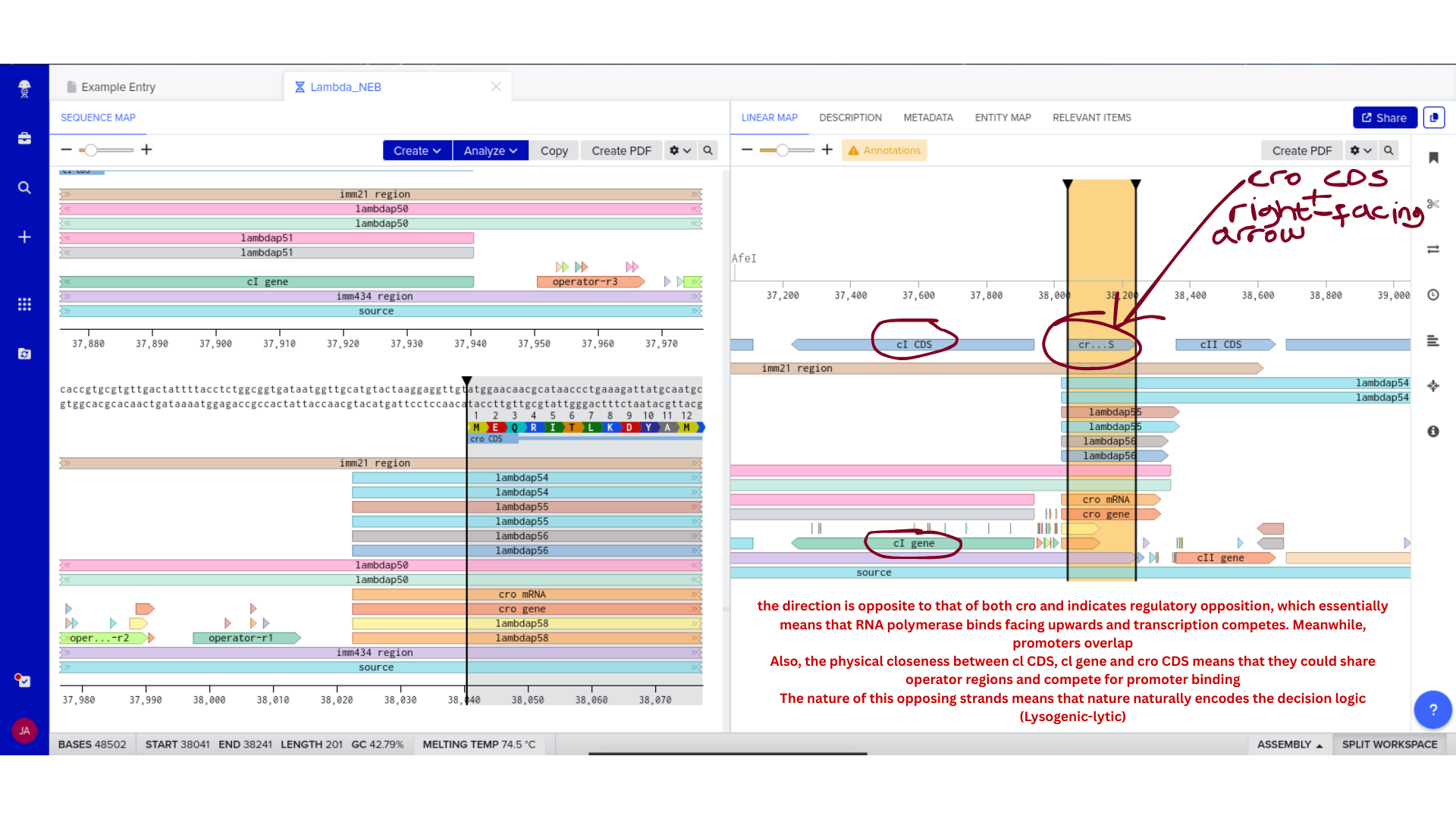

3) Protein Selection and Sequence Acquisition
Furthermore, I selected Microcin M as the protein of interest. The choice aligned with my project, ÌṢỌ, which focuses on context-sensitive antimicrobial response within the gut ecosystem.
The selection criteria included:
- Narrow-spectrum antimicrobial activity
- Relevance to microbial competition
- Compatibility with a governed probiotic chassis
The amino acid sequence was retrieved in FASTA format from a reliable database (NCBI GenBank: CAE55705.1). I verified the header structure and ensured that the sequence corresponded exactly to the intended protein.
4) Reverse Translation
Using Benchling’s reverse translation functionality, I converted the amino acid sequence into a nucleotide sequence suitable for expression in Escherichia coli.
Key considerations included:
- Maintaining correct reading frame
- Ensuring inclusion of a start codon
- Confirming appropriate stop codon placement
- Selecting E. coli codon usage
The output DNA sequence was checked to ensure it translated back to the original protein sequence without truncation or frame shift.
5) Codon Optimization
Following reverse translation, codon optimization was performed for expression in E. coli. This step aimed to improve translational efficiency while minimizing expression burden and avoiding rare codons.
Optimization included:
- Aligning codon usage with host bias
- Avoiding problematic restriction sites
- Preserving protein sequence integrity
This stage reinforced that codon choice influences not only protein yield but also metabolic load and evolutionary stability.
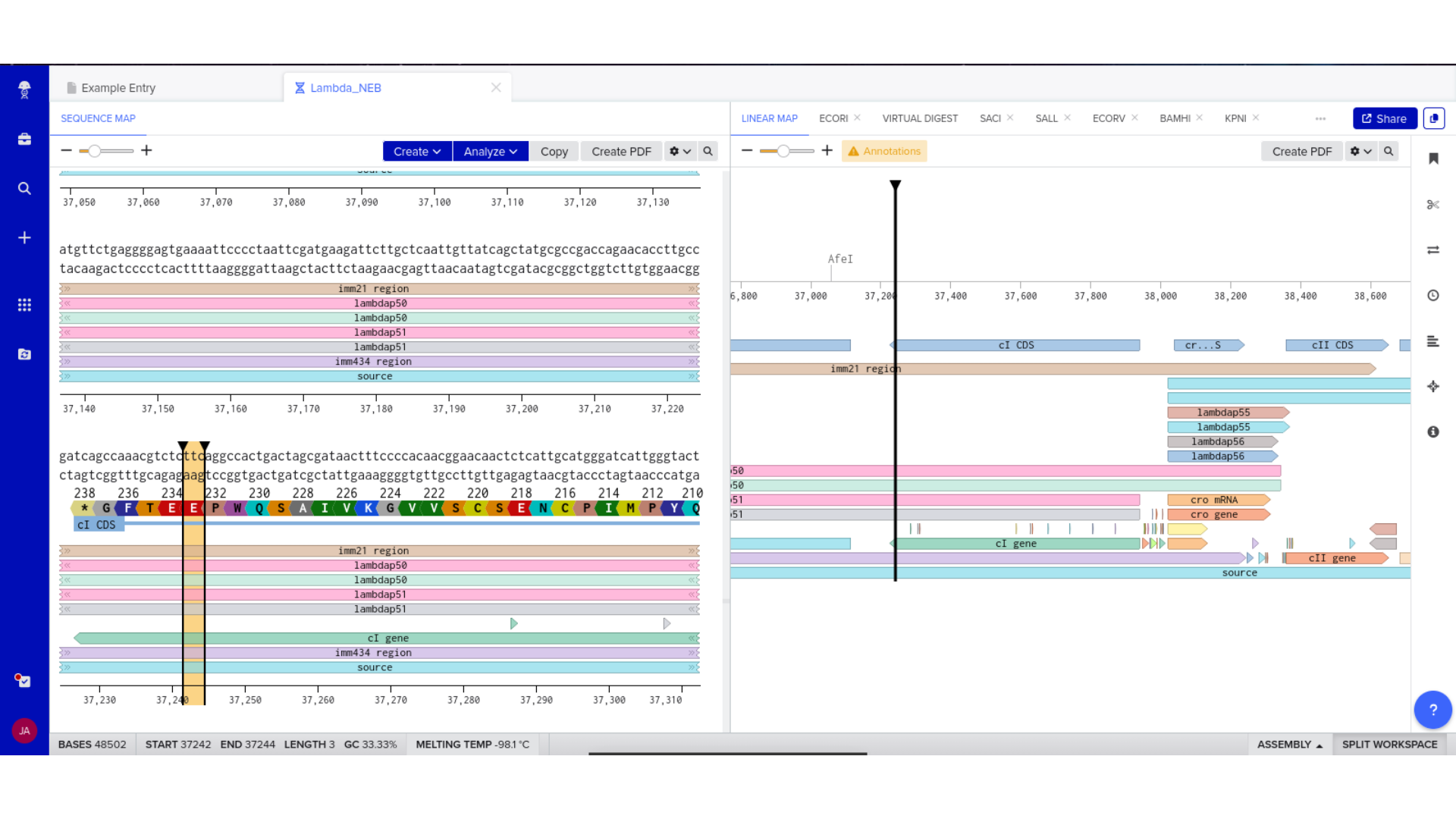

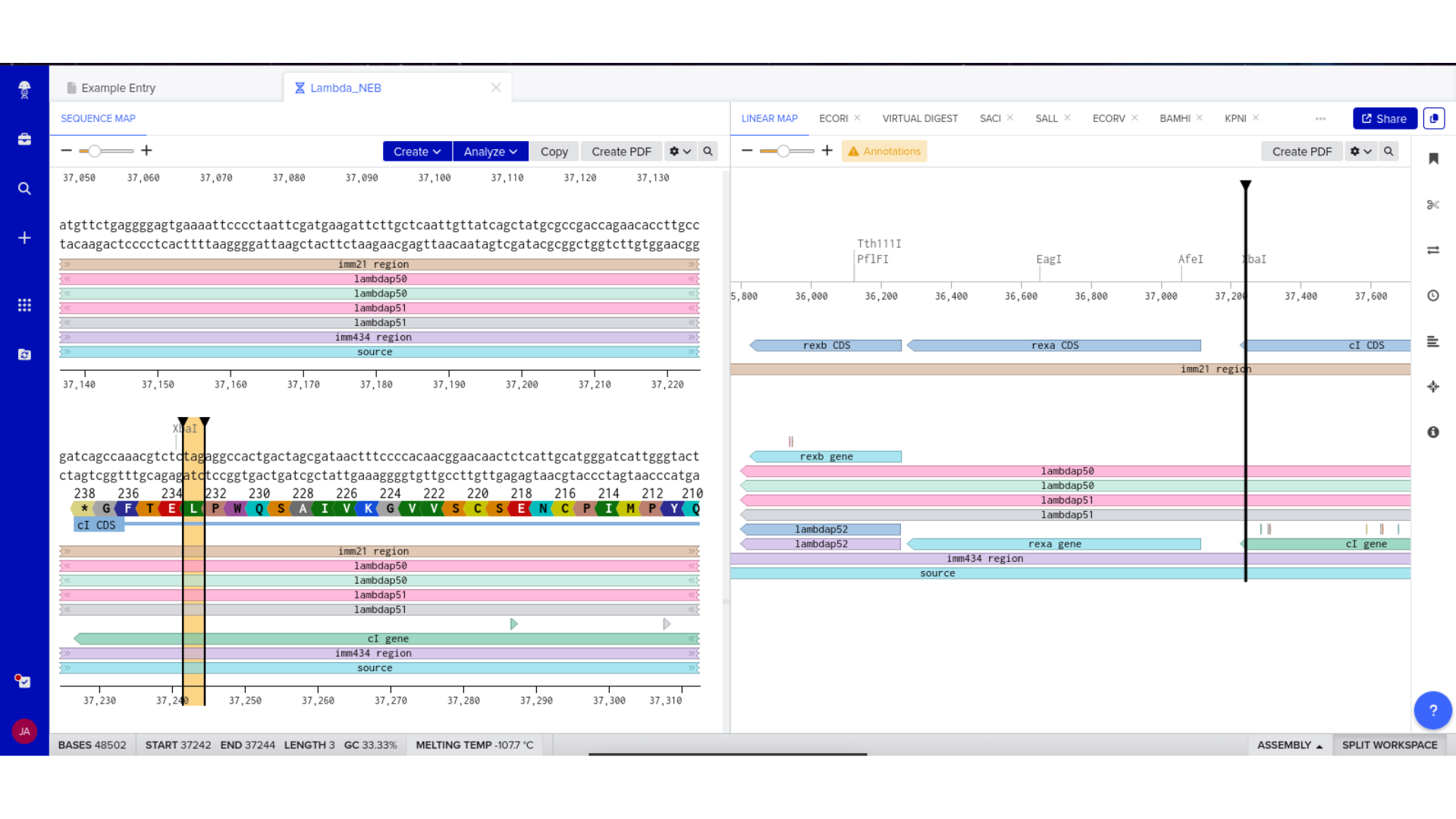

Part 2 — Construct Assembly and Validation
6) Expression Cassette Assembly
The optimized coding sequence was integrated into a complete expression cassette using the assignment’s structural framework:
Promoter → Ribosome Binding Site → Start Codon → Codon-Optimized CDS → Optional His Tag → Stop Codon → Terminator
Each component was manually inserted and annotated within Benchling. Particular care was taken to ensure that the coding region replaced the example scaffold sequence rather than being appended to it.
Linear and circular map views were used to confirm structural continuity, annotation accuracy, and absence of unintended sequence artifacts.
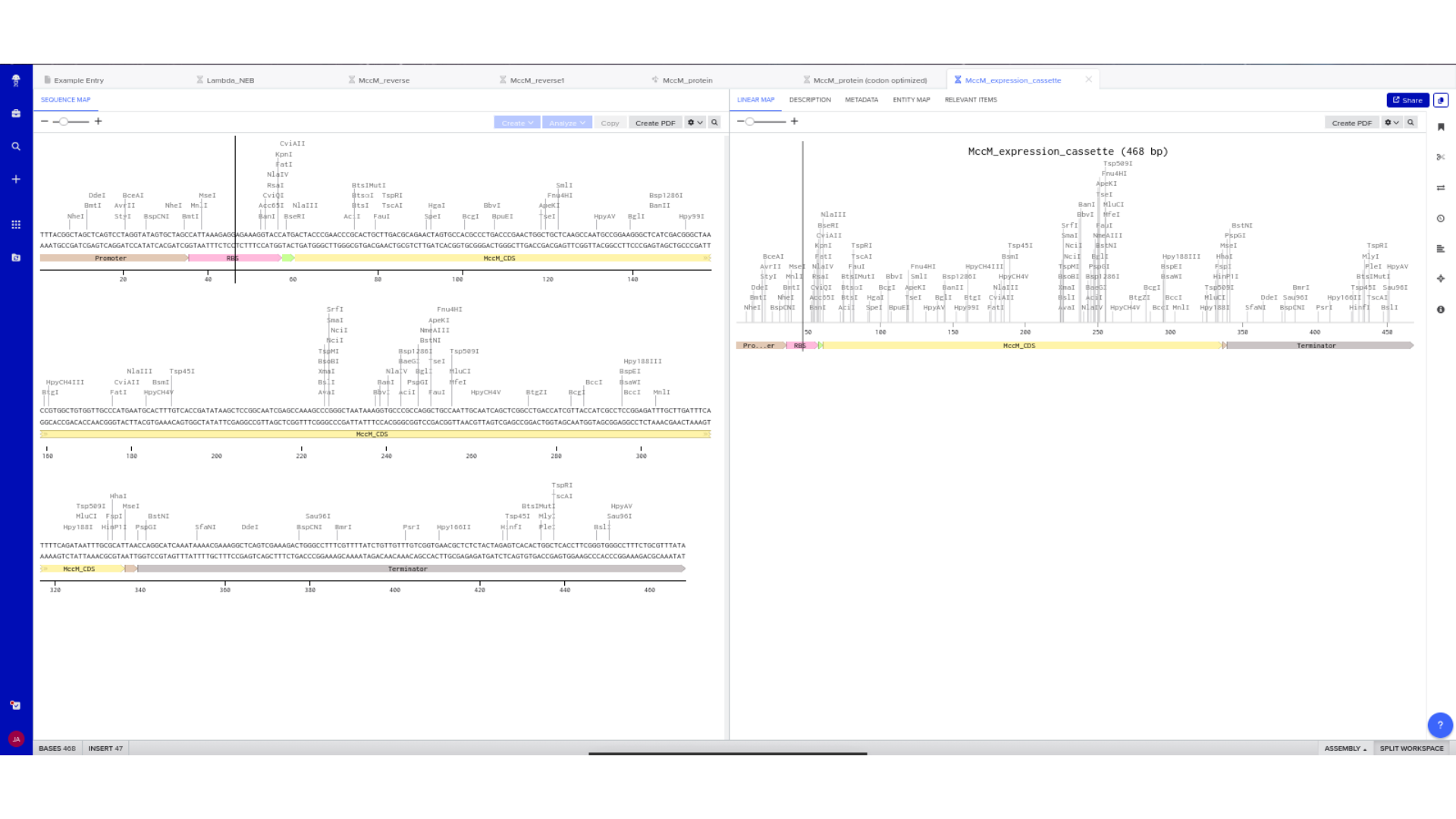

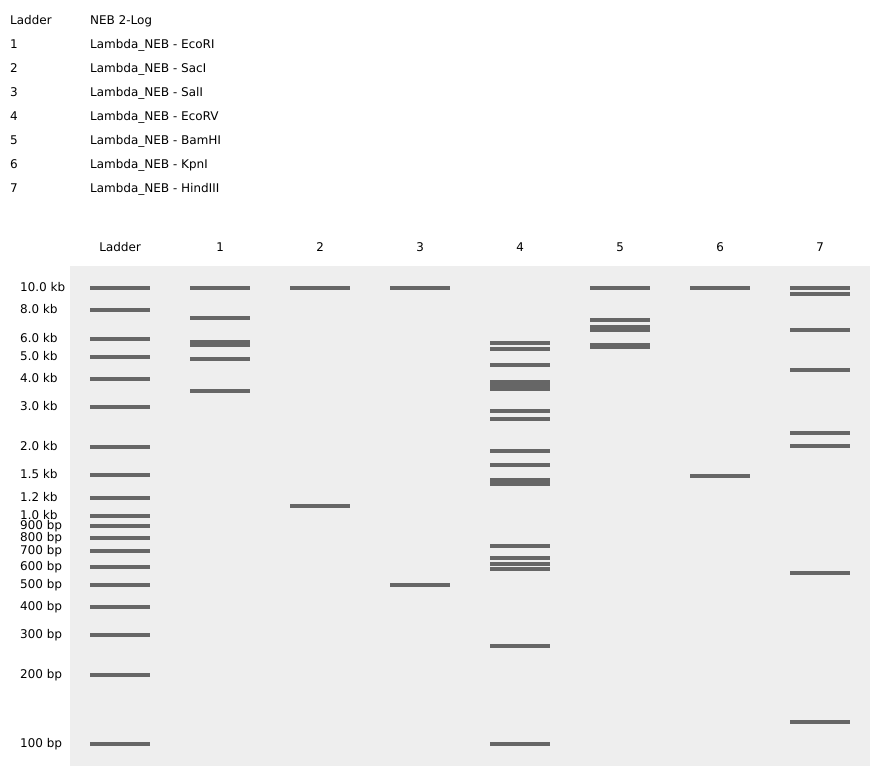
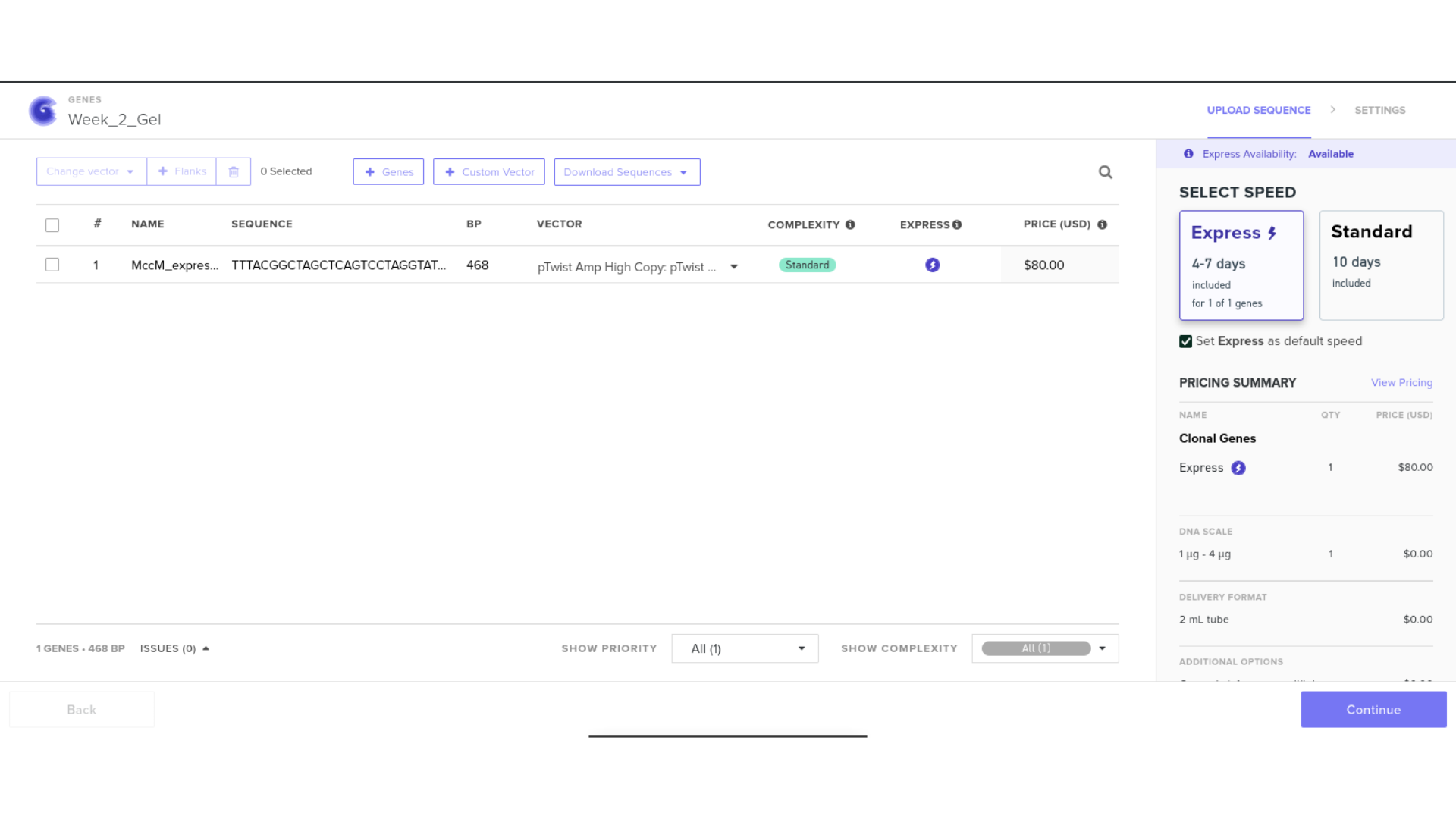
7) Virtual Digest and Gel Simulation
To validate construct integrity, I performed a virtual digest within Benchling and obtained predicted fragment sizes. These fragment sizes were then visualized using an external gel simulation tool.
This step confirmed that the construct behaved as expected under restriction enzyme analysis and reinforced my understanding of plasmid verification workflows.
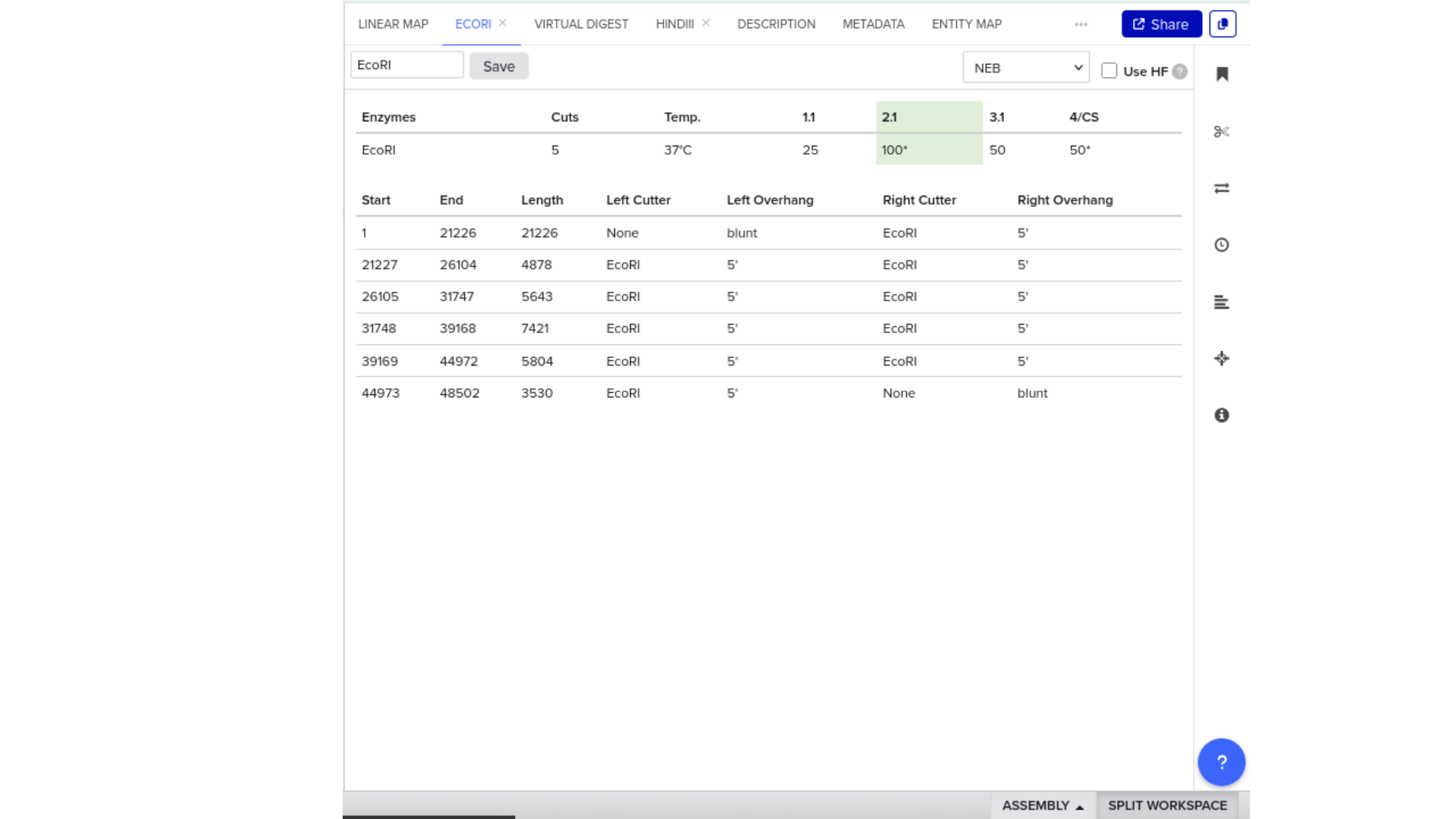

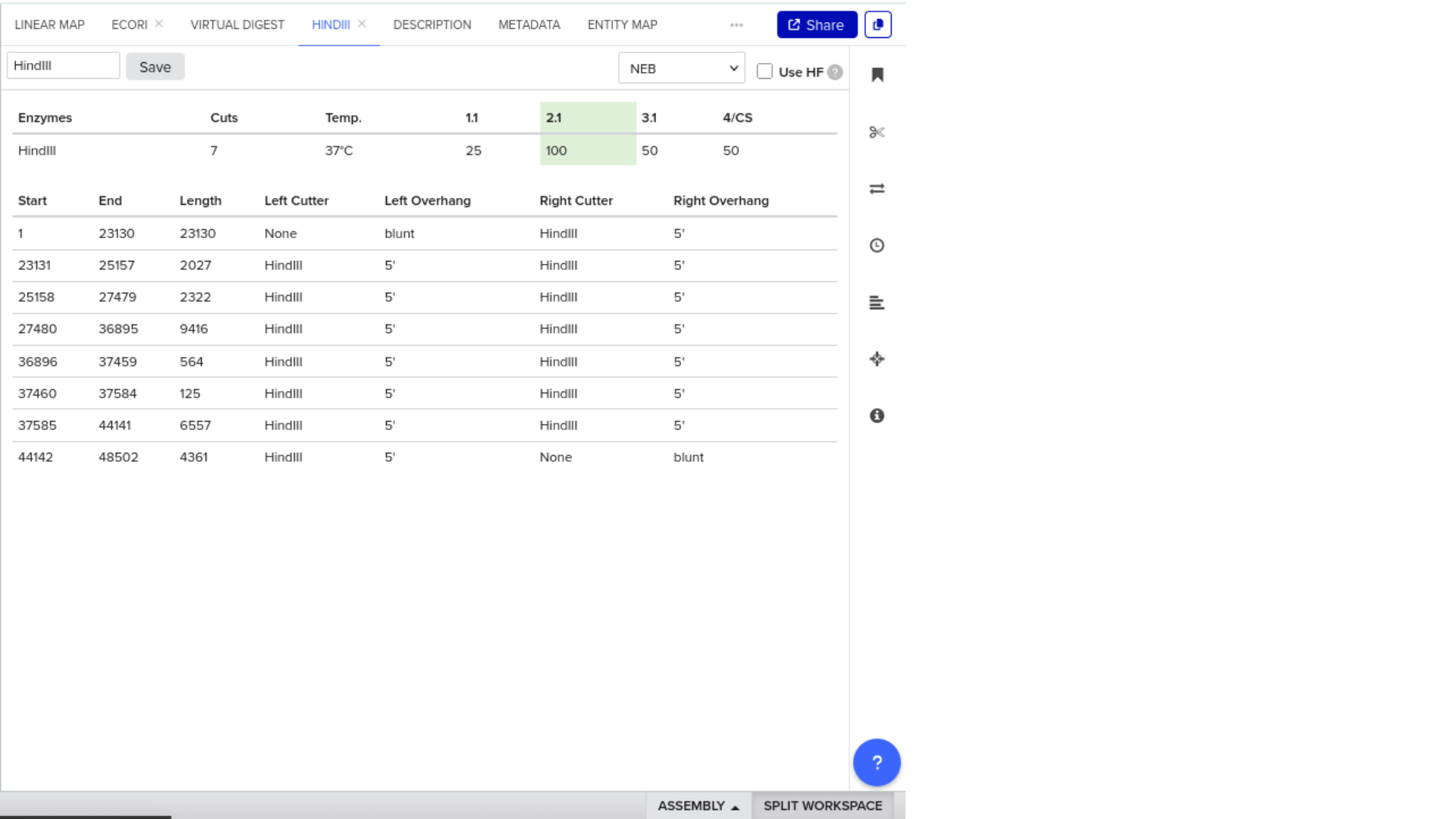

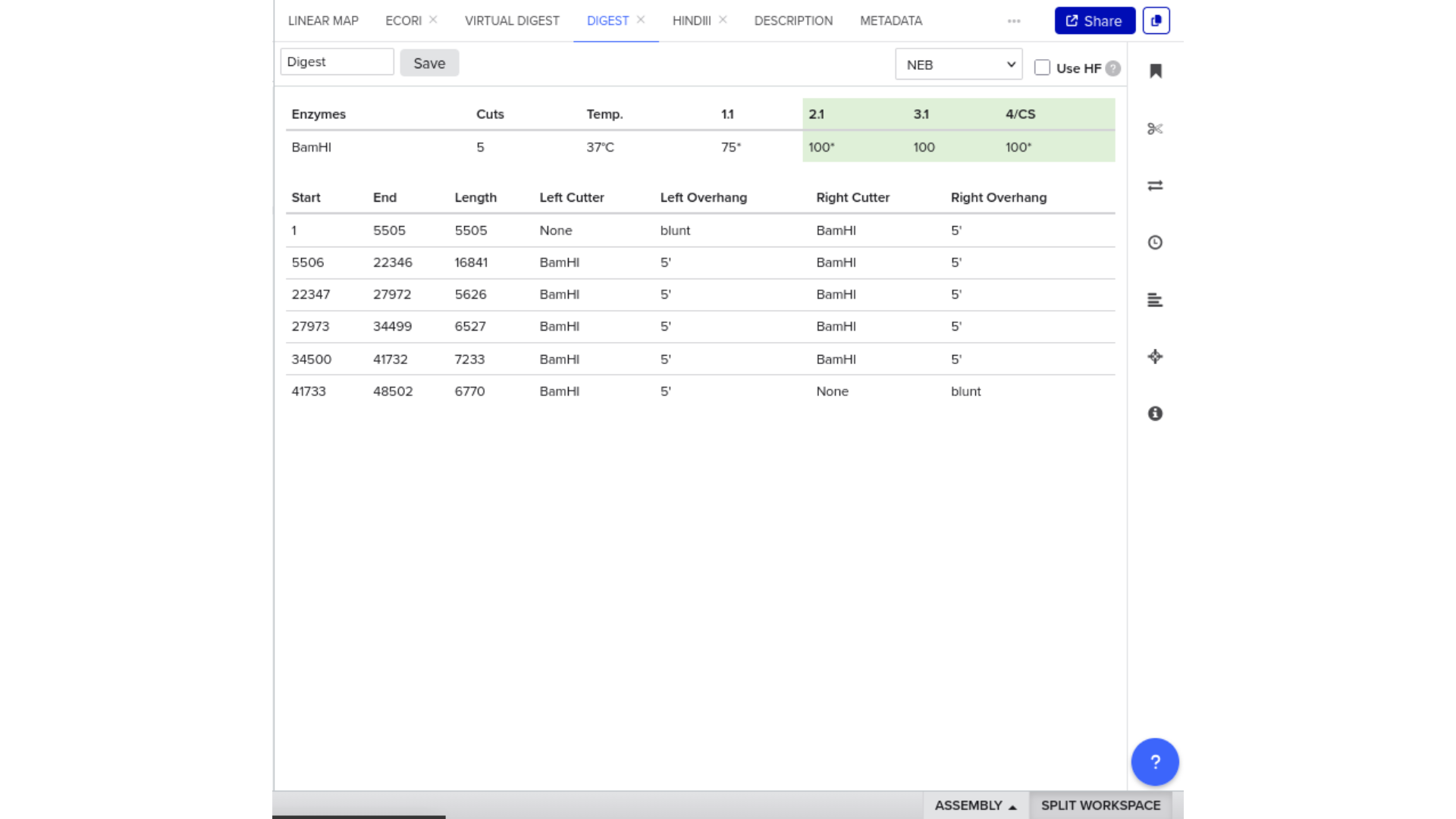

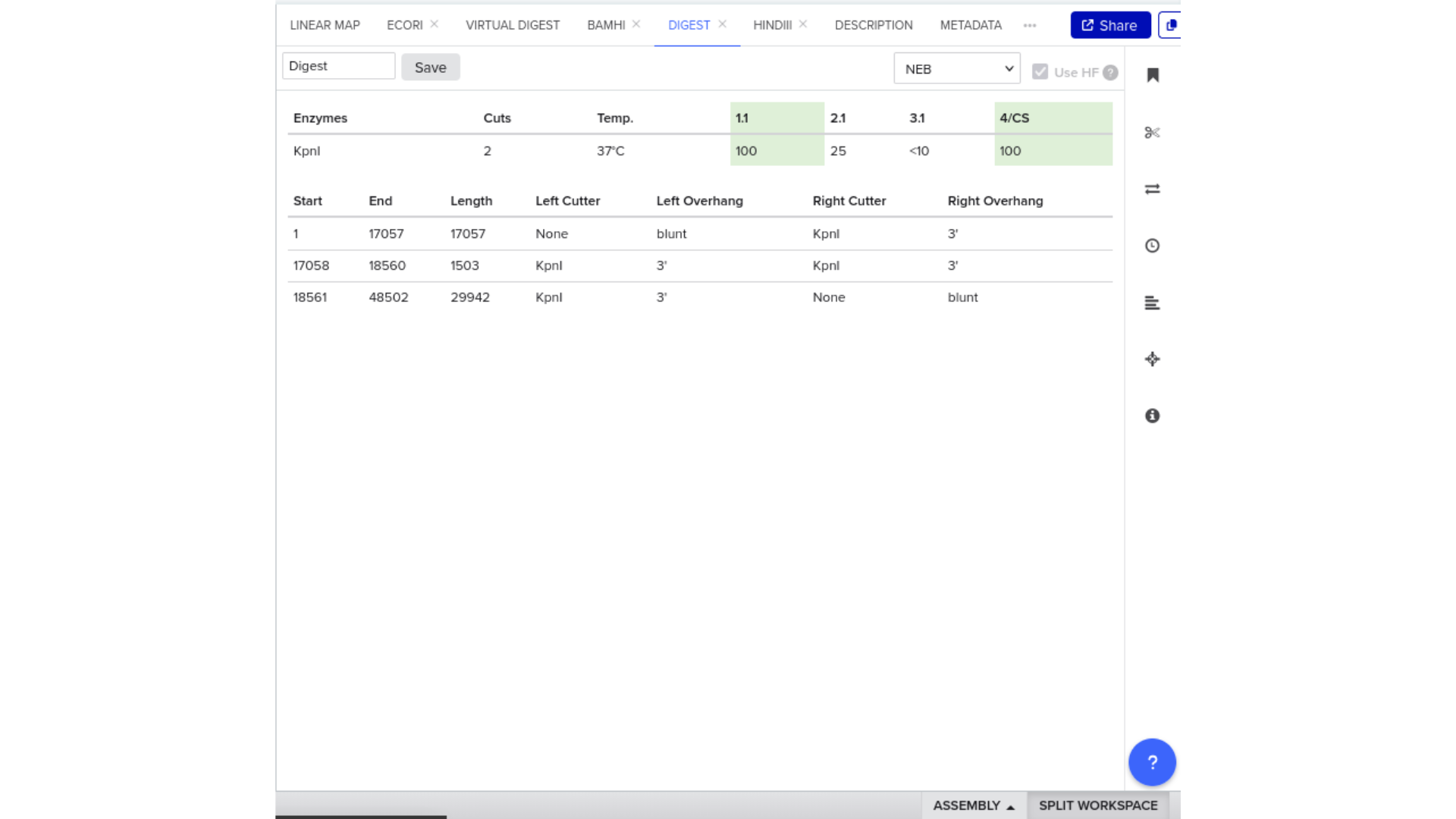

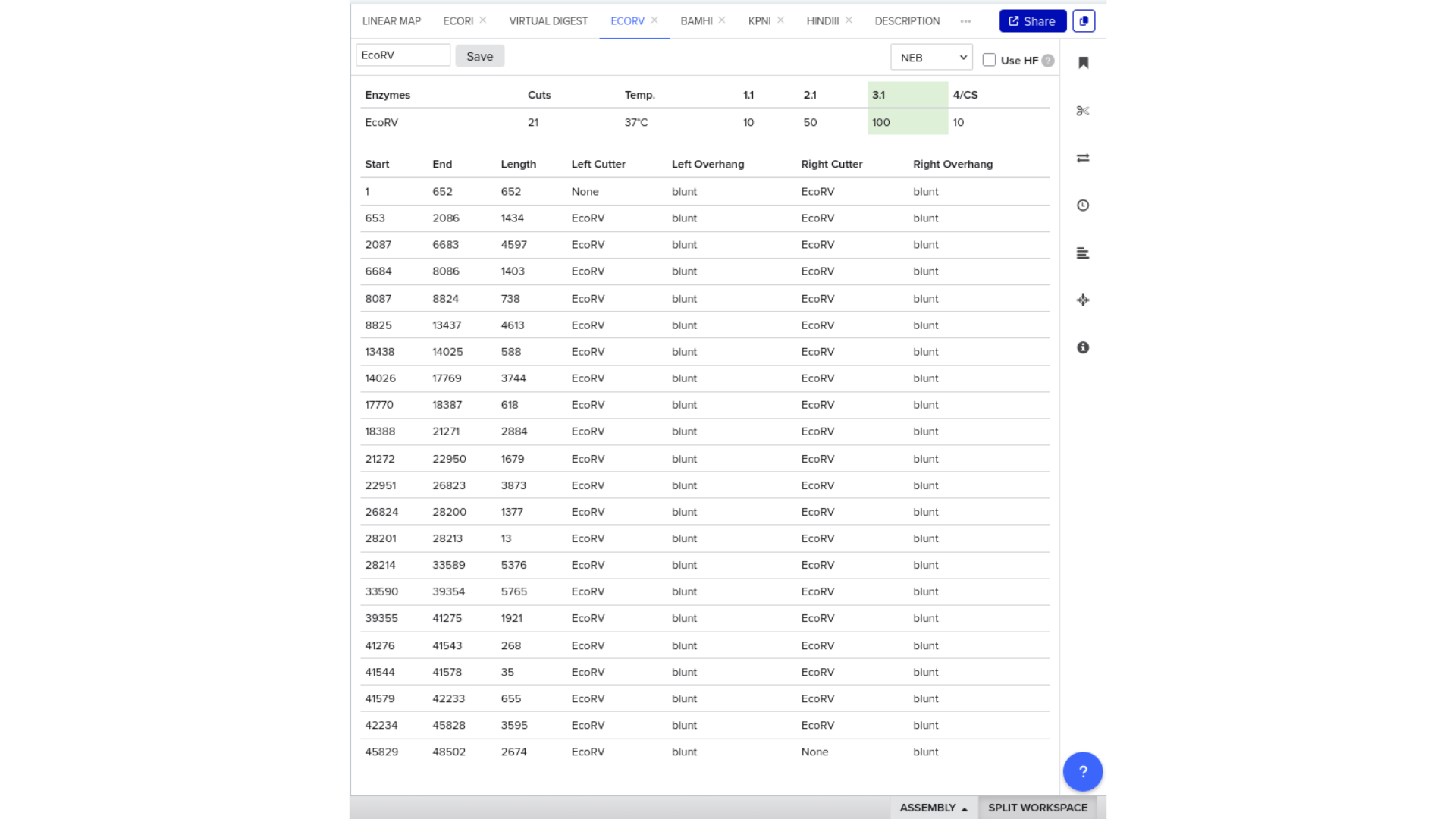

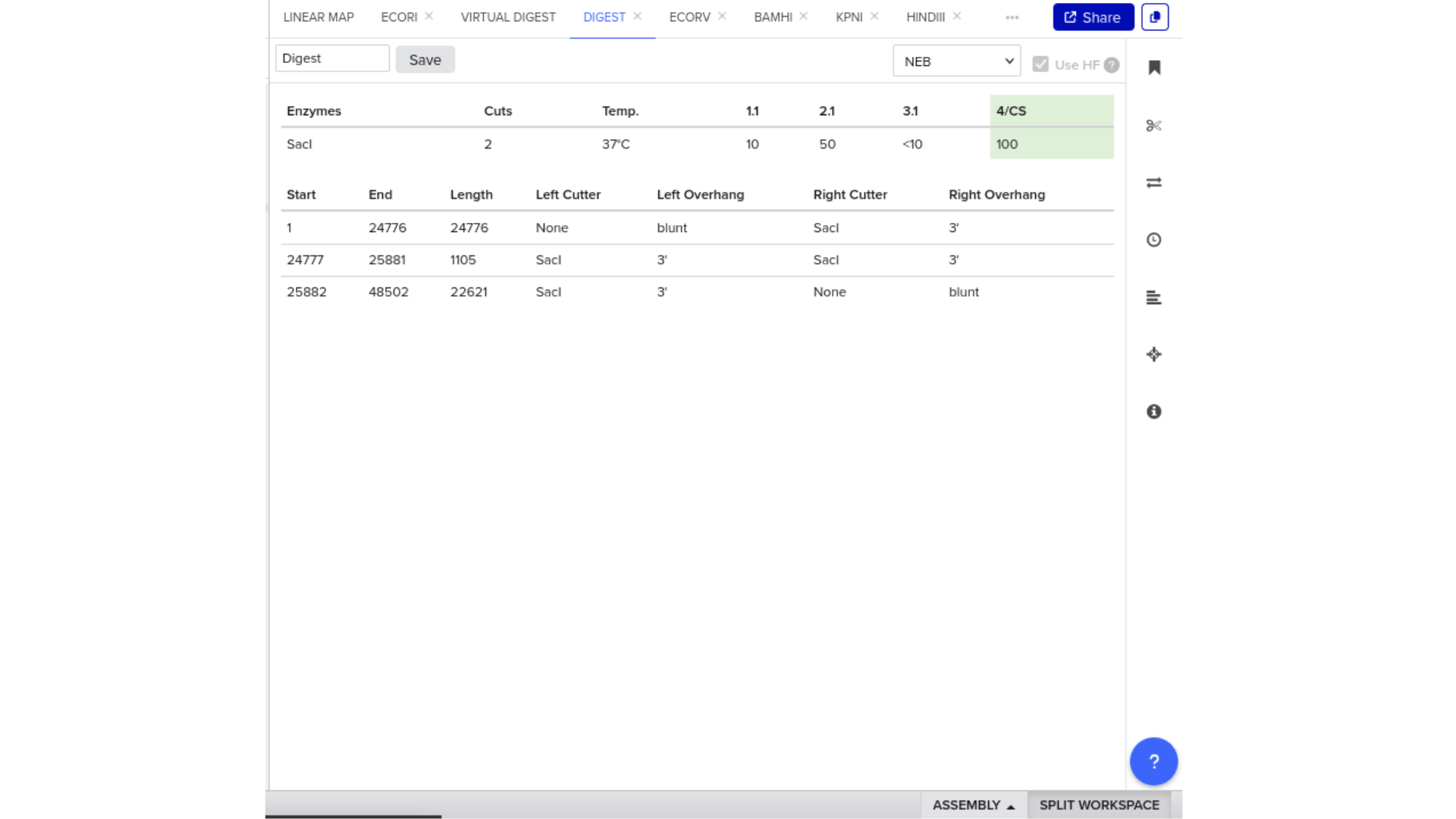

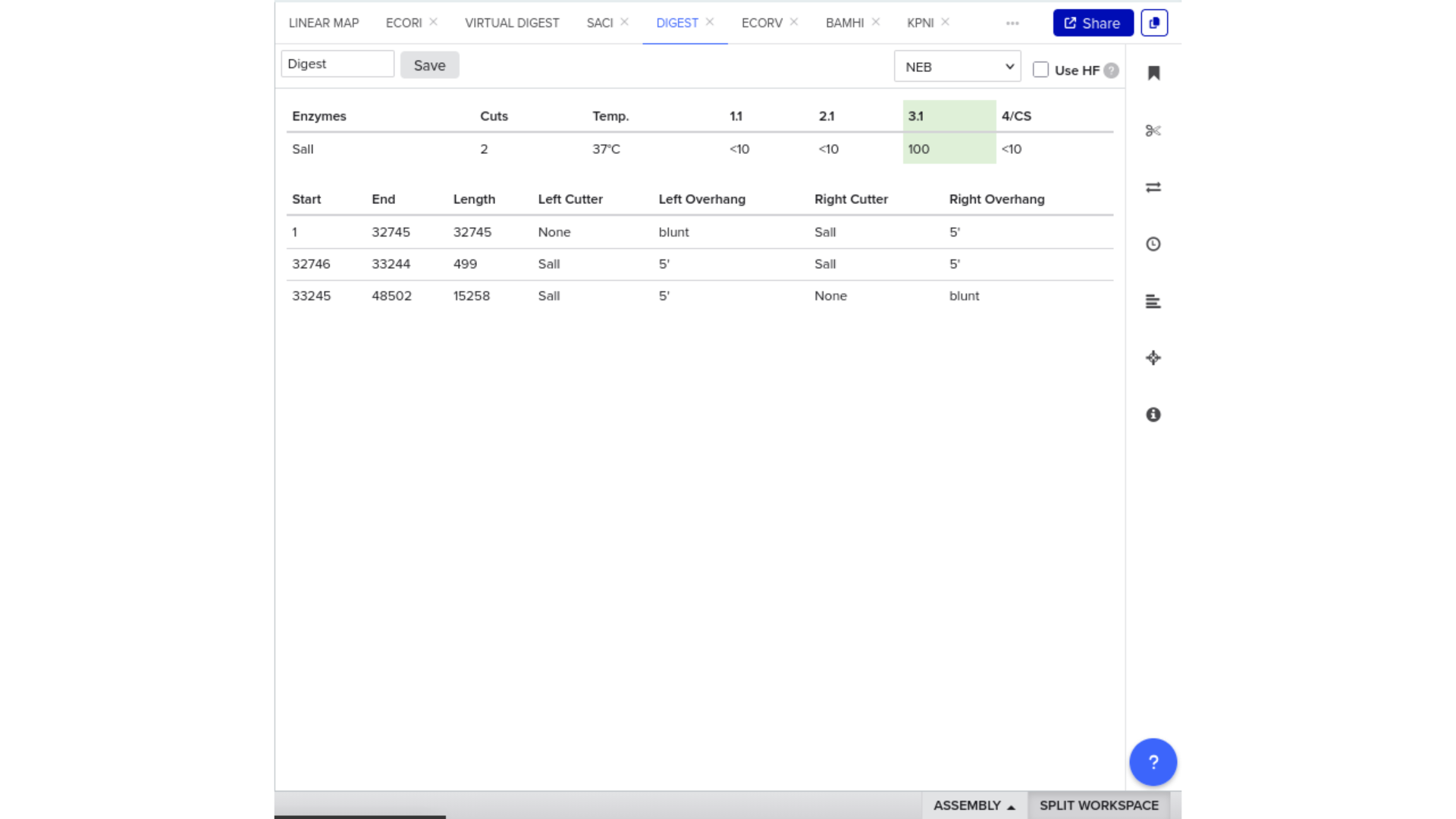

8) FASTA Export and Synthesis Preparation
The completed expression cassette was exported in FASTA format for potential synthesis ordering. Care was taken to ensure:
- Correct header formatting beginning with the greater-than symbol
- No extraneous spaces or formatting characters
- Proper file extension
Although synthesis ordering through Twist was initiated, access limitations prevented full completion. Instead of halting progress, I pivoted toward generating a complete plasmid visualization within Benchling.

9) Plasmid Map Generation
To simulate a complete plasmid construct, the sequence topology was converted to circular within Benchling. Circular map visualization confirmed clear annotation of promoter, ribosome binding site, coding sequence, and terminator.
This produced a plasmid map without requiring external synthesis confirmation. The visualization ensured structural coherence and clear representation of the engineered construct.


Technical Milestones Achieved
- Successful import and annotation of GenBank files
- Accurate reverse translation from protein to DNA
- Codon optimization aligned with host expression
- Proper construction of an annotated expression cassette
- Verified FASTA export formatting
- Simulated plasmid visualization in circular topology
- Integration of molecular workflow with ecological design philosophy




Backbone Vector Documentation
The Microcin M expression cassette was designed for cloning into pUC19, a high-copy ColE1-origin plasmid carrying ampicillin resistance. pUC19 was selected primarily for its well-characterised cloning sites and broad compatibility with standard E. coli transformation protocols — practical considerations given that the immediate goal is sequence verification rather than stable expression. The MccH47 insert is flanked by EcoRI and HindIII sites for directional cloning into the multiple cloning site. The complete annotated construct is deposited in the class Benchling folder as MccH47_pUC19_EcN_construct.
For downstream ÌṢỌ deployment, the cassette would need migration to a lower-copy backbone — pSC101 or a chromosomal integration vector — to reduce metabolic burden on the EcN chassis and improve evolutionary stability under selection.
Referenced from Week 7, Part 3
Design Integration
Throughout the experience, I maintained alignment with the core principles of ÌṢỌ:
- Fitness cost is a primary design variable
- Selection operates continuously
- Expression burden affects evolutionary stability
- Containment must be intrinsic to architecture
- Models inform design boundaries
This reframed it for me from a cloning exercise into a constraint-aware engineering process.
Process Reflections
The workflow required iterative verification at each stage. Formatting, reading frame integrity, codon usage, annotation accuracy, and topology conversion each presented potential points of error and addressing them incrementally reduced compounding mistakes.
More importantly, it reinforced that biological engineering is not simply about inserting genes. It requires contextual awareness, ecological humility, and structural foresight.
Sequence design is only the beginning. Stability under pressure determines whether a system is viable outside controlled conditions.
This process strengthened both my technical fluency and design discipline, linking molecular implementation to ecological responsibility.