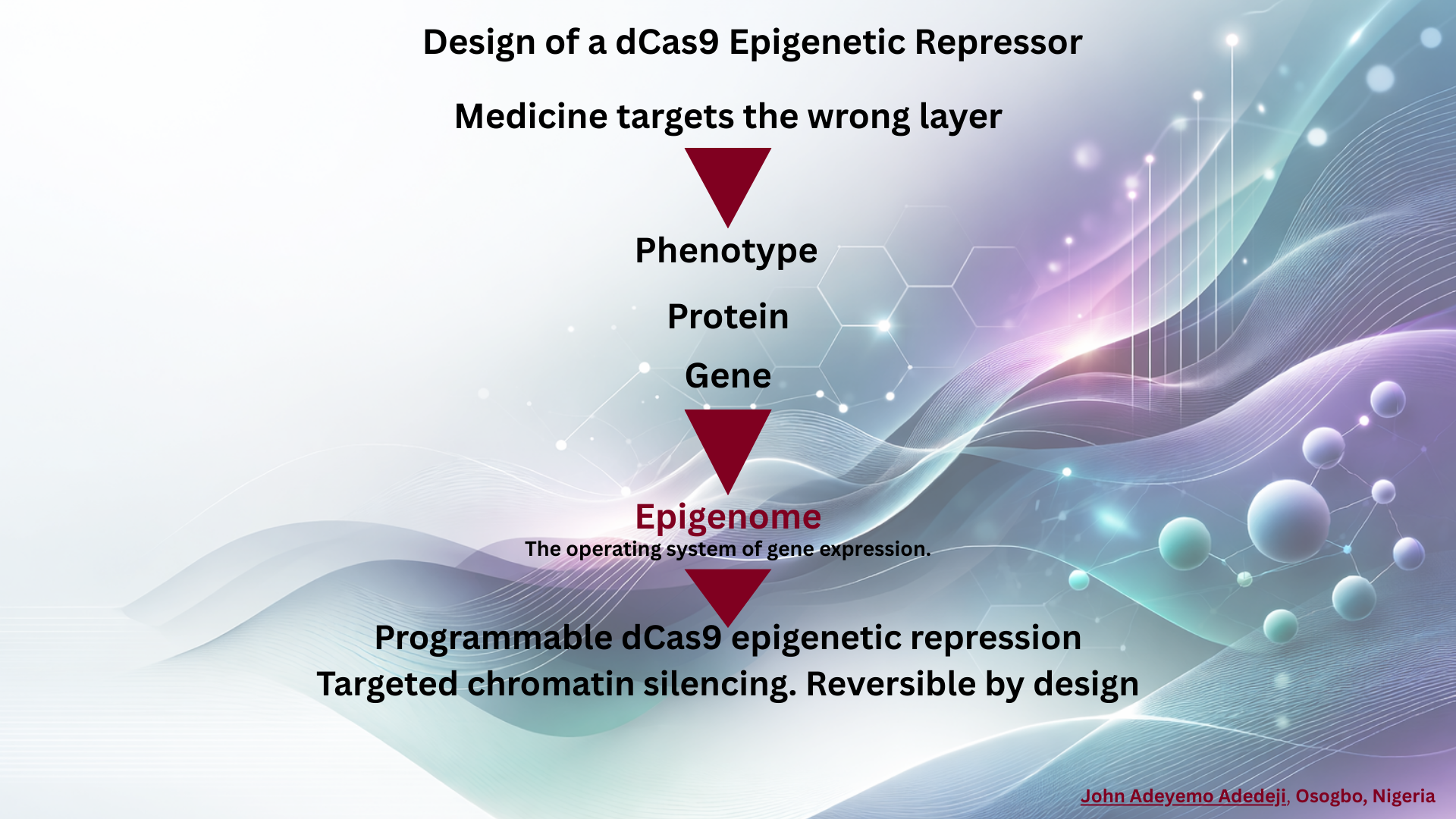
Week 3
Class Assignment — Week 3
1) Opentrons Artwork
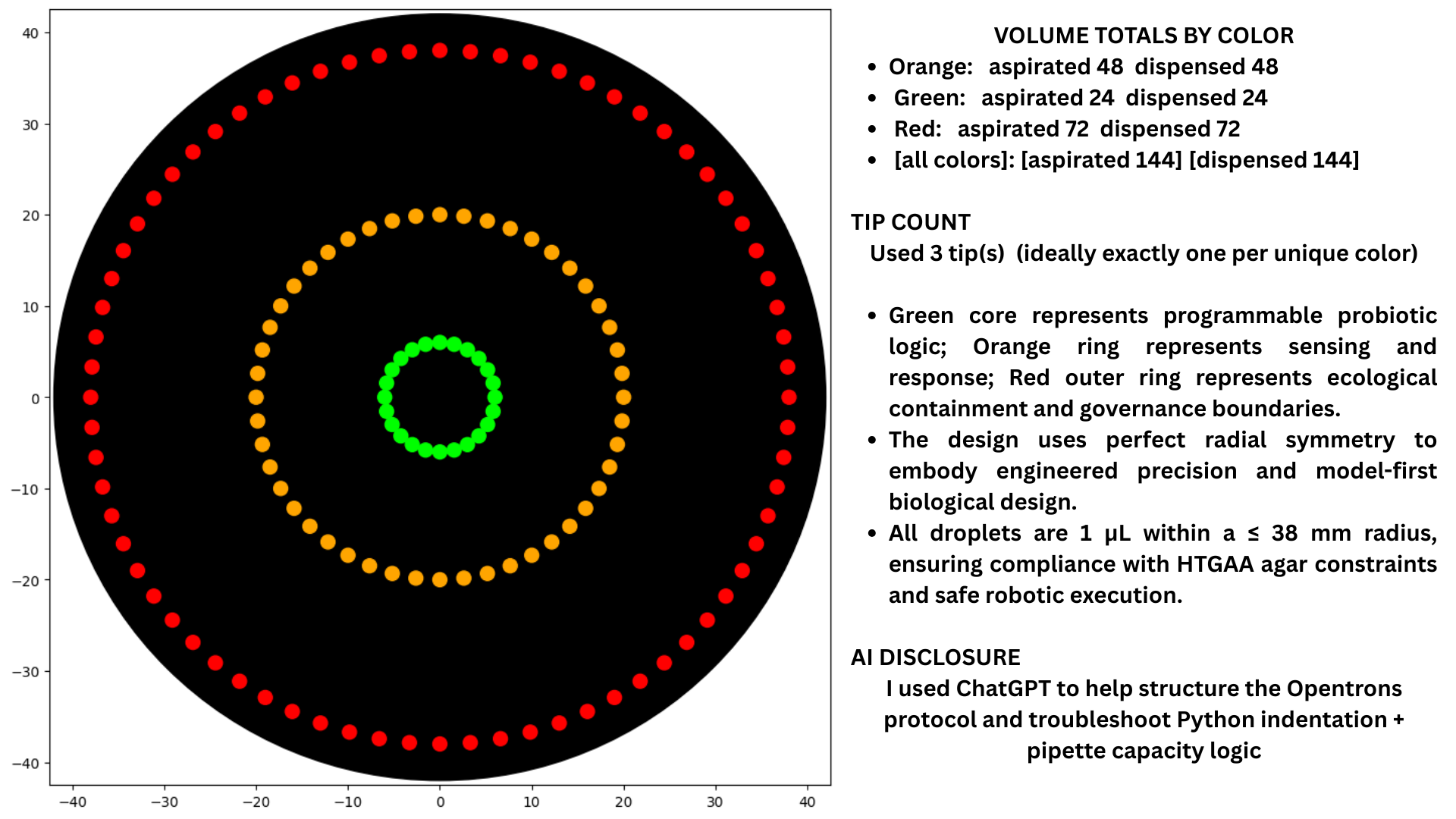

2) Published Papers Utilizing Automation
LabscriptAI — Autonomous Liquid-Handling Robotics Scripting
Gao et al., 2025 introduce LabscriptAI, a multi-agent framework that translates natural language experimental descriptions into validated Python scripts for heterogeneous liquid-handling robots, including Opentrons platforms.
The system integrates:
- Hierarchical task planning
- Platform-specific simulation validation
- A precise refactoring engine for targeted debugging
- Domain-specific knowledge retrieval
- Human-in-the-loop safety checkpoints
Experimental validation included:
- Cross-platform fluorescence calibration
- Automated cell-free expression and screening of 298 GFP variants
- Distributed enzyme engineering involving hazardous substrates
The central contribution is not pipetting precision alone. It is structured experimental execution with embedded validation and safety logic. Automation becomes reproducible, cross-platform, and governable.
Active Learning Directed Evolution (ALDE)
Active Learning Directed Evolution which integrates machine learning uncertainty estimation with iterative experimental screening to guide protein engineering efficiently was introduced by Yang, Lal, Arnold, et al. 2025.
ALDE automates experimental decision-making by:
- Training predictive sequence–function models
- Quantifying uncertainty across unexplored sequence space
- Selecting optimal next-round variants
- Iteratively refining search trajectories
Rather than brute-force screening, ALDE navigates design space intelligently, minimizing experimental waste while maximizing functional discovery.
Together, these systems represent complementary layers:
- ALDE enables intelligent experimental proposal
- Robotic scripting platforms enable validated execution
Automation becomes both cognitive and mechanical.
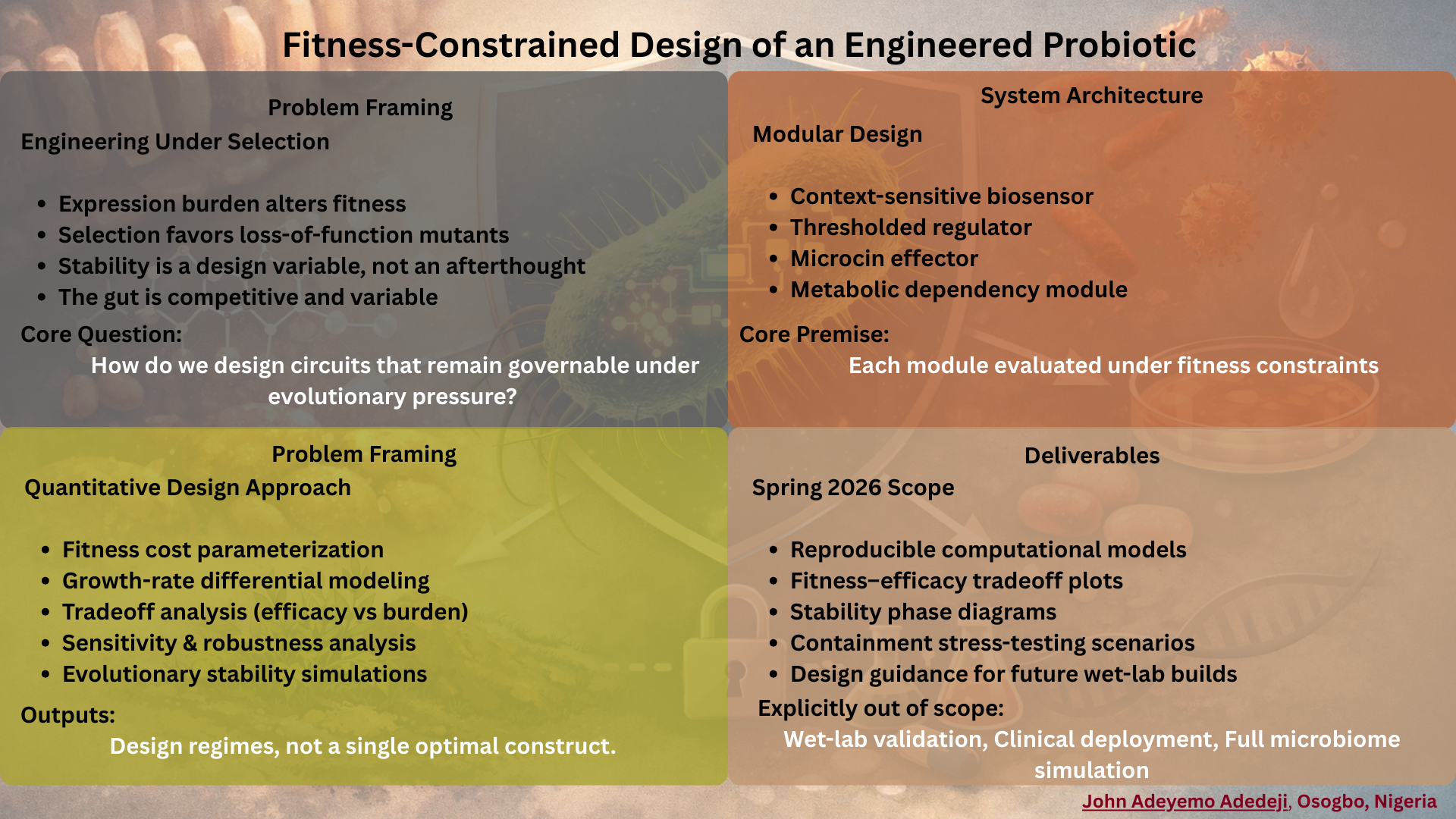
3) Automation Architecture for ÌṢỌ — Sentinel EcN
ÌṢỌ is a fitness-aware engineered probiotic system designed to sense gut context, produce targeted antimicrobial responses, and remain bounded through intrinsic containment.
Automation enables a structured Design–Build–Test–Learn loop.
A) Combinatorial Genetic Circuit Screening (requires automation)
Objective: Evaluate sensor–effector variants under growth constraints.
Automated workflow:
- Dispense transformation master mix into 96-well plate
- Add plasmid constructs into defined coordinates
- Perform serial dilution plating
- Inoculate colonies into induction gradient
- Measure OD600 for growth
- Measure fluorescence for reporter output
- Normalize fluorescence by growth to assess fitness-aware performance
Example Opentrons pseudocode:
This enables reproducible and remotely deployable transformation workflows.
B) Cell-Free Circuit Screening
To decouple metabolic burden from host growth:
- Echo transfer DNA constructs into 384-well plate
- Stamp CFPS master mix
- Dispense lysate to initiate expression
- Incubate at 37°C
- Measure fluorescence
This permits rapid high-throughput screening prior to in vivo validation.
C) Active Learning Integration
After first-round screening:
- Fit sequence–function predictive model
- Quantify uncertainty across design space
- Propose next construct library
- Upload variants for synthesis or robotic cloning
- Repeat screening
This reduces combinatorial explosion and focuses experimentation where information gain is highest.
D) 3D Printed Hardware Integration (requires automation)
To approximate ecological realism:
- Custom 96-well anaerobic incubation adapter
- Microfluidic gradient diffusion holder
- Plate alignment fixtures for reproducible layout
These hardware additions introduce environmental constraint into automated pipelines rather than assuming ideal laboratory conditions.
E) Use of Ginkgo Nebula
For larger combinatorial libraries:
- Upload sequence designs
- Automated synthesis and cloning
- High-throughput transformation
- Automated phenotyping
- Structured dataset return
Cloud laboratories enable distributed execution while preserving structured feedback into the design loop.
Summary
Automation within ÌṢỌ operates at two levels:
- Cognitive layer: uncertainty-aware experimental selection
- Execution layer: validated robotic implementation
Together, they form a closed-loop, governable engineering system that prioritizes stability under ecological pressure rather than maximal output under ideal conditions.
Works Cited
Yang, J., Lal, R. G., Bowden, J. C., et al. (2025). Active learning-assisted directed evolution. Nature Communications, 16, 714. https://doi.org/10.1038/s41467-025-55987-8
Gao, Y., Luo, Y., Li, W., Lan, Y., Jiang, H., Chen, Y., Yi, X., Li, B., Alinejad-Rokny, H., Wang, T., Fu, L., Yang, M., & Si, T. (2025). Autonomous liquid-handling robotics scripting for accessible and responsible protein engineering. bioRxiv. https://doi.org/10.1101/2025.09.30.679666
Proposed Final Project Ideas
Process Reflections
This week shifted my understanding of automation from technical convenience to systems architecture.
Initially, I approached the assignment by identifying a strong automation framework in LabscriptAI. However, as I explored complementary tools such as ALDE, it became clear that robotic precision alone is insufficient. Scalable biological engineering requires structured exploration, specifically uncertainty-aware active learning to navigate sequence and design space intelligently.
The key insight was recognizing that automation operates on two layers:
- Cognitive layer deciding what experiment to run next
- Execution layer safely and reproducibly running it
By combining both, my thinking moved beyond pipetting workflows toward a closed-loop, governable Design–Build–Test–Learn system. This reframing aligns directly with ÌṢỌ, which requires ecological realism, fitness awareness, and safety constraints.
Another important shift was recognizing the role of governance. Automation increases capability, but without structured safety checkpoints, biosecurity screening, and human oversight, it becomes fragile or irresponsible. Designing the automation architecture required explicit consideration of containment, ecological competition, and reproducibility.
This process strengthened three core skills:
- Systems-level integration rather than tool-level selection
- Designing for constraint rather than brute-force optimization
- Framing automation as a platform rather than a procedure
Ultimately, I realized that my final project is not only an engineered probiotic. It is a structured, uncertainty-aware engineering pipeline for responsible biological deployment.
AI Prompts Employed
- Compare ALDE and LabscriptAI to see if they work well together as a system
- Design a closed-loop setup where AI chooses experiments and robots run them
- List what I would automate for ÌṢỌ (Sentinel EcN)
- Draft simple Opentrons-style pseudocode for running reactions
- Integrate 3D printed tools, cloud labs, and governance into the automation workflow