Week 7
Class Assignment — Week 7
Part A. Intracellular Artificial Neural Networks (IANNs)
1. Advantages of IANNs over Boolean Genetic Circuits
Boolean genetic circuits are fundamentally limited by their design logic: every input gets collapsed into a binary state, and the circuit operates on those discrete values. That works for simple switch-like decisions, but most physiologically relevant signals (metabolite concentrations, osmotic gradients, and quorum sensing molecule titres), exist on a continuum, and forcing them through a hard threshold discards information. IANNs avoid this by processing analog inputs directly, generating graded outputs that reflect the actual magnitude of the input rather than just which side of a threshold it fell on.
The deeper advantage is function approximation capacity. A sufficiently wide or deep network of gene-regulatory elements functioning as weighted summing nodes can approximate arbitrary continuous input-output relationships, which means you can in principle encode complex multi-factor decisions (that respond strongly when signal A is high and signal B is moderate and signal C is low, but not when all three are high) without the combinatorial explosion of logic gates that an equivalent Boolean circuit would require. Practically, this also reduces the parameterisation burden: you train the network on data rather than manually calibrating each gate’s individual threshold and transfer function, which for complex Boolean circuits is a significant experimental cost.
Noise robustness is the third real advantage. Biological systems are stochastic, and Boolean circuits that depend on clean thresholding behave poorly when input signals are noisy or when component expression varies between cells. Analog processing distributes the computation across multiple nodes, so no single component’s noise dominates the output.
2. IANN Application — ÌṢỌ / Gut Sentinel Context
The continuous modelling capacity of an IANN is directly relevant to the gut sentinel problem. The challenge with engineering E. coli Nissle 1917 as a therapeutic probiotic is that its fitness and output behaviour depend on a genuinely continuous environmental landscape — luminal pH, competing commensal species densities, pathogen metabolite concentrations, mucus layer thickness, transit rate. A Boolean circuit could in principle be designed to activate effector expression above some threshold concentration of a target metabolite, but that assumes a single clean input drives the decision. Real gut ecology doesn’t work that way.
An IANN implemented in EcN could integrate multiple continuous environmental inputs simultaneously, tetrathionate concentration, competing species quorum signals, local oxygen tension, and produce a graded effector output proportional to the true threat level rather than a binary kill switch. This is particularly relevant to the evolutionary stability question in the ÌṢỌ framework: a cell population making graded decisions about resource allocation to effector production versus growth will, under selection, behave more like a stable evolutionarily stable strategy than one operating a hard switch that either maximally expresses a costly effector or doesn’t express it at all.
The limitations are substantial though. Implementing an IANN in a living cell requires physical instantiation of weighted connections as actual molecular interactions (protein-protein binding affinities, RNA regulatory elements, transcription factor binding strengths), all of which drift under evolutionary pressure, are sensitive to cellular metabolic state, and cannot be reconfigured in situ once the cell is deployed. Training the network computationally is achievable; translating the learned weights into specific DNA sequences encoding the required regulatory strengths is not straightforward, and verifying that the implemented network actually computes what you intended in a complex in vivo environment like the gut is a significant experimental challenge. There is also a metabolic cost argument: implementing even a shallow network requires expressing multiple non-native regulatory proteins simultaneously, which imposes a fitness burden that selection will work against over time.
3. Intracellular Multilayer Perceptron
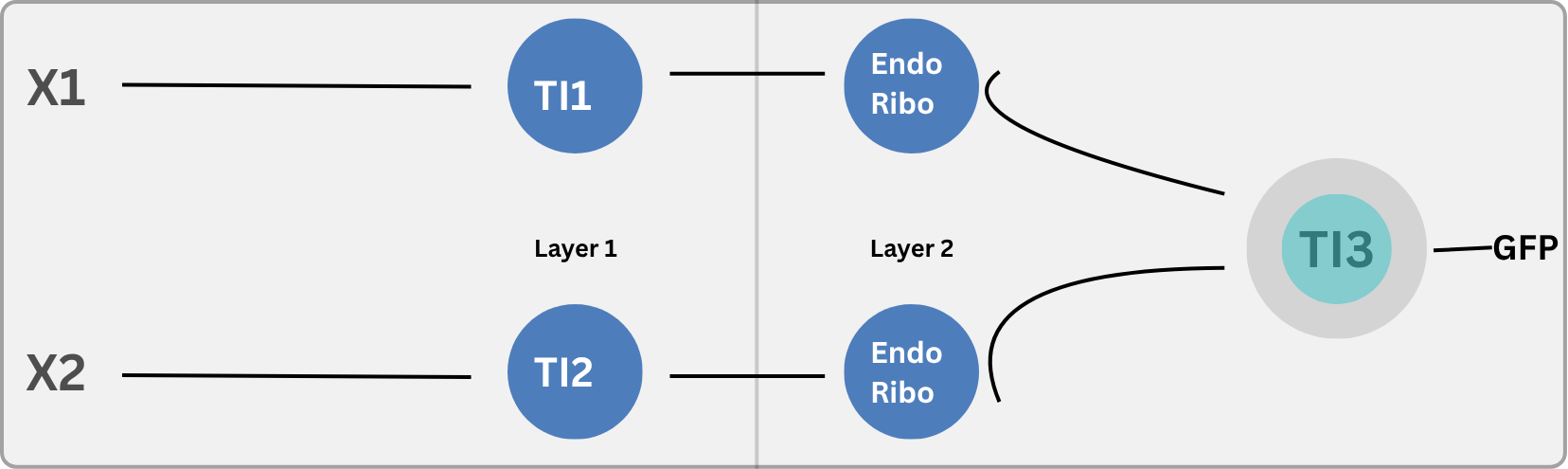

Part B. Fungal Materials
1. Examples of Existing Fungal Materials and Their Applications
The most commercially visible fungal materials are mycelium-based composites — mycelial networks grown through agricultural waste substrates like hemp hurds or corn stalks, then heat-treated to halt growth and pressed into rigid forms. Companies like Ecovative have used this to produce packaging, acoustic panels, and leather-like textiles. In construction contexts, mycelium composites offer comparable compressive strength to expanded polystyrene at a fraction of the carbon cost, with full biodegradability at end of life.
In the medical context specifically, fungal-derived materials have a longer history than the mycelium-composite trend might suggest. Chitin and its deacetylated derivative chitosan (both derived from fungal cell walls) have been extensively evaluated as wound dressings, drug delivery scaffolds, and haemostatic agents. Chitosan’s cationic character at physiological pH allows it to interact electrostatically with bacterial membranes and negatively-charged wound exudate, giving it both antimicrobial and pro-coagulant properties without the immunogenicity concerns associated with animal-derived alternatives like collagen. For biosecurity and field-medicine applications, chitosan-based haemostatic dressings are already in clinical and military deployment, HemCon dressings were among the first to translate this directly into combat casualty care.
The disadvantages are real though. Batch-to-batch consistency in fungal-derived biomaterials is harder to control than synthetic polymer manufacturing: chitin extraction yields vary with growth conditions, and residual endotoxin or beta-glucan contamination from fungal cell wall debris poses immunogenicity risks in any implantable or injectable application. Regulatory classification is also still unsettled in many jurisdictions: a mycelium-derived scaffold sits awkwardly between a device and a biological, which complicates approval pathways considerably.
For biofabrication purposes, the more interesting frontier is using fungal hyphal networks as living scaffolds for tissue engineering — mycelial architecture naturally produces interconnected porous networks at scales relevant to vascularisation, something genuinely difficult to replicate by synthetic additive manufacturing. The limitation here is that you are working with a eukaryotic organism that has its own growth agenda, and getting predictable pore geometry without precise genetic intervention remains challenging.
2. Genetic Engineering in Fungi for Biopharmaceuticals and Protein Therapeutics
The application I find most compelling is using engineered Pichia pastoris (now reclassified as Komagataella phaffii) or Saccharomyces cerevisiae as chassis for producing complex glycosylated therapeutic proteins, biologics that bacteria fundamentally cannot make correctly.
This is where the core advantage of fungal synthetic biology over bacterial systems becomes concrete: post-translational modification. Bacteria lack the endoplasmic reticulum machinery for N-linked glycosylation, disulfide bond formation in a controlled oxidising environment, and proper signal peptide processing for secretion. A therapeutic antibody fragment, a vaccine antigen, or a receptor-binding protein domain that depends on correct glycosylation for receptor recognition, serum half-life, or effector function simply cannot be produced functionally in E. coli without extensive refolding steps that introduce batch variability and reduce yield. Yeast do all of this co-translationally in a compartmentalised secretory pathway that is genuinely homologous to mammalian cells.
For vaccinology specifically, yeast-expressed virus-like particles are already an established platform, the hepatitis B surface antigen in Engerix-B is produced in S. cerevisiae, and the HPV L1 capsid proteins in Gardasil are produced in the same host. The self-assembly capacity of these proteins into immunogenic particles in a yeast secretory environment is something a bacterial chassis would struggle with. Engineering Pichia further, humanising its N-glycosylation pathway to reduce the hypermannose patterns that drive immunogenicity in native yeast glycoproteins, moves the output closer to what a mammalian CHO cell would produce, but at fermentation costs that are orders of magnitude lower.
The limitations worth being honest about: yeast genetic toolkits are less mature than bacterial ones. CRISPR-based genome editing in S. cerevisiae is well-established, but in non-model yeasts the efficiency drops sharply. Promoter libraries, ribosome binding site tuning, and the kind of fine transcriptional control you take for granted in E. coli requires considerably more development effort in a fungal host. Secretion titres for complex proteins also remain lower than CHO cells for the most demanding biologics, and hypermannose glycosylation, even with humanisation efforts, is still not identical to human-type glycans, which matters for Fc-mediated effector functions in therapeutic antibody applications.
Part C. First DNA Twist Order
The Microcin M expression cassette was designed for cloning into pUC19, a high-copy ColE1-origin plasmid carrying ampicillin resistance. pUC19 was selected primarily for its well-characterised cloning sites and broad compatibility with standard E. coli transformation protocols, practical considerations given that the immediate goal is sequence verification rather than stable expression. The MccH47 insert is flanked by EcoRI and HindIII sites for directional cloning into the multiple cloning site. The complete annotated construct is deposited in the class Benchling folder as MccH47_pUC19_EcN_construct.
For downstream ÌṢỌ deployment, the cassette would need migration to a lower-copy backbone — pSC101 or a chromosomal integration vector — to reduce metabolic burden on the EcN chassis and improve evolutionary stability under selection.