Week 1 HW: Principles and Practices
Class Assignment
Describe a biological engineering application or tool you want to develop and why.
Aplication title: De novo design of proteins binders for neutralizing Bothrops venom toxins
Antivenoms are a mix of immunoglobulins produced traditionally by the hyperimmunization of large animals with crude venom obtained from clinically-relevant snakes (Ratanabangkoon, K., 2023). Novel alternatives have emerged to neutralize venom toxins without the use of animals. For example, Torres and collaborators (2025) designed proteins with high affinity for important regions of cytotoxins from the 3FTx family. These proteins showed great neutralizating capacity in vitro and great protective capacity in vivo .
“Omics” strategies applied to snake venoms have been developed as Venomics, these strategies allows the characterization of whole venoms building protein profiles. Similar estrategies can also be used for studying antibody-toxin complexes as Antivenomics (Lomonte, 2017). The information obtained through venomics and antivenomics give us the ability to build databases with structural and functional information of snake toxins; this information is important for disigning de novo proteins
Bothrops genus is one of the most relevant in South America and has been studied broadly. Many venomics and antivenomics studies has been developed giving high amounts of information that can be used for the design of proteins with high affinity to their important regions.
The following proposal aims to create a system that involves venomics and antivenomics estudies of Bothrops venoms to create a database that can be used to identify key regions with high impact in their toxic activities. Through these regions I propose the design of proteins using artificial intelligence. These candidates could be useful to explore the possibility of designing synthetic antivenoms that don’t depend on the use of animals for their production (Figure 1)
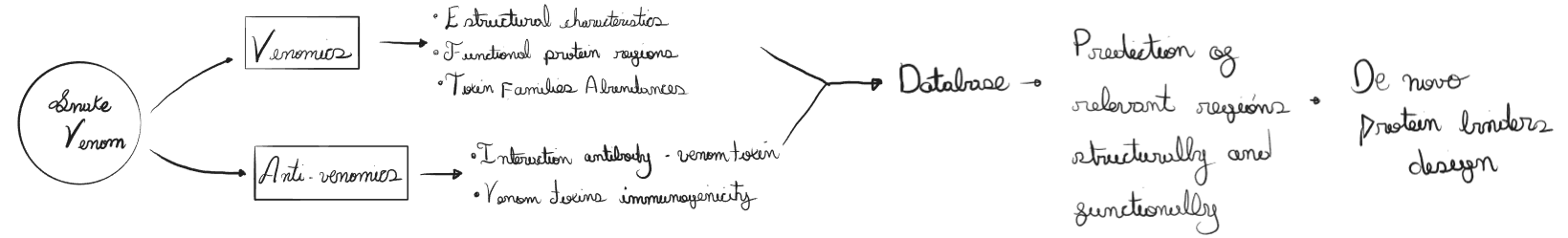
Justification: Snakebites are classified as Neglected Tropical Diseases by the World Health Organization (WHO) affecting low-and middle-income countries from Africa, Asia, and South America (World Health Organization, 2019).
Antivenoms are the only approved treatment against snakebites. Antivenoms show several limitations in their efficacy and production. Snake venoms present variability in their composition. This could lead to antivenoms with different efficacies depending on the venom used for their production. Additionally, antibodies present in the antivenom can cause adverse reactions when administered to the patient.
Antivenom production is technologically complex with high costs, resulting in a limitation for low-income countries (Alangode et al., 2020). Antivenoms not only present but also numerous challenges in their production but also in their requisites at different levels to be used safely (Figure 2, Potet et al, 2021)
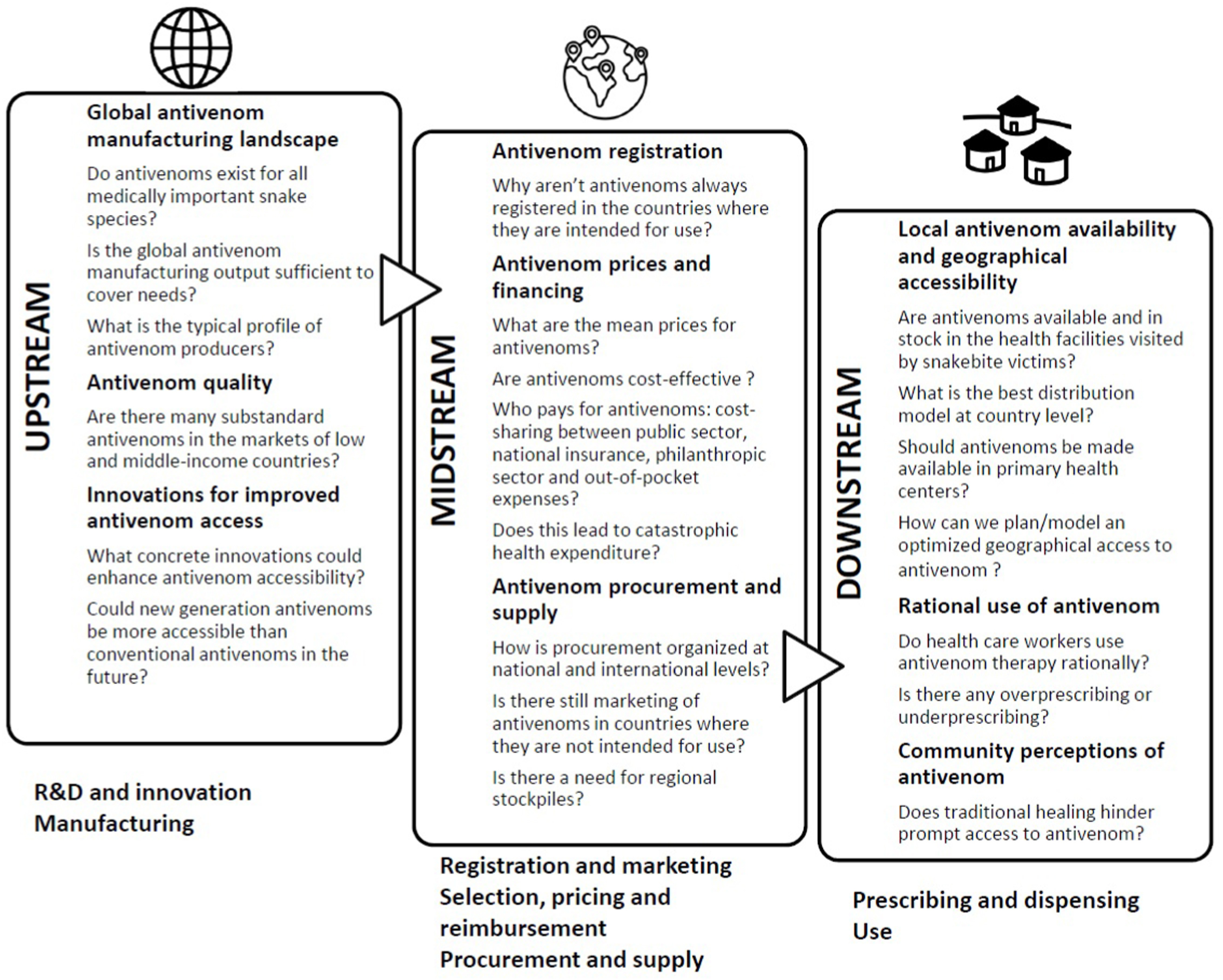
The limitations observed in antivenoms produced traditionally supports the necessity of novel alternatives that can be produced safely and with low cost in their design. The use of artificial intelligence with the information provided by venomics and antivenomics opens the possibility of creating synthetic alternatives for the neutralization of venom toxins, and their design could also be optimized to a production at large scale increasing their availabilty and reducing their cost.
Analysis of Protein Binders for Governance Goals and Actions
The World Health Organization has established a programme to evaluate the safety and effectivenes of current antivenoms intended for their use in different countries. This programme led to the recruit of several world experts, forming the Working Group on Snakebite Envenoming. Through this group, the WHO has established a goal of reducing the mortality and disabiluty of snakebite envenomings by 2030. (World Health Organization, 2019)
To accomplish this goal, the working group has developed a road map with objetives at different scales (Figure 3, Williams et al, 2019)
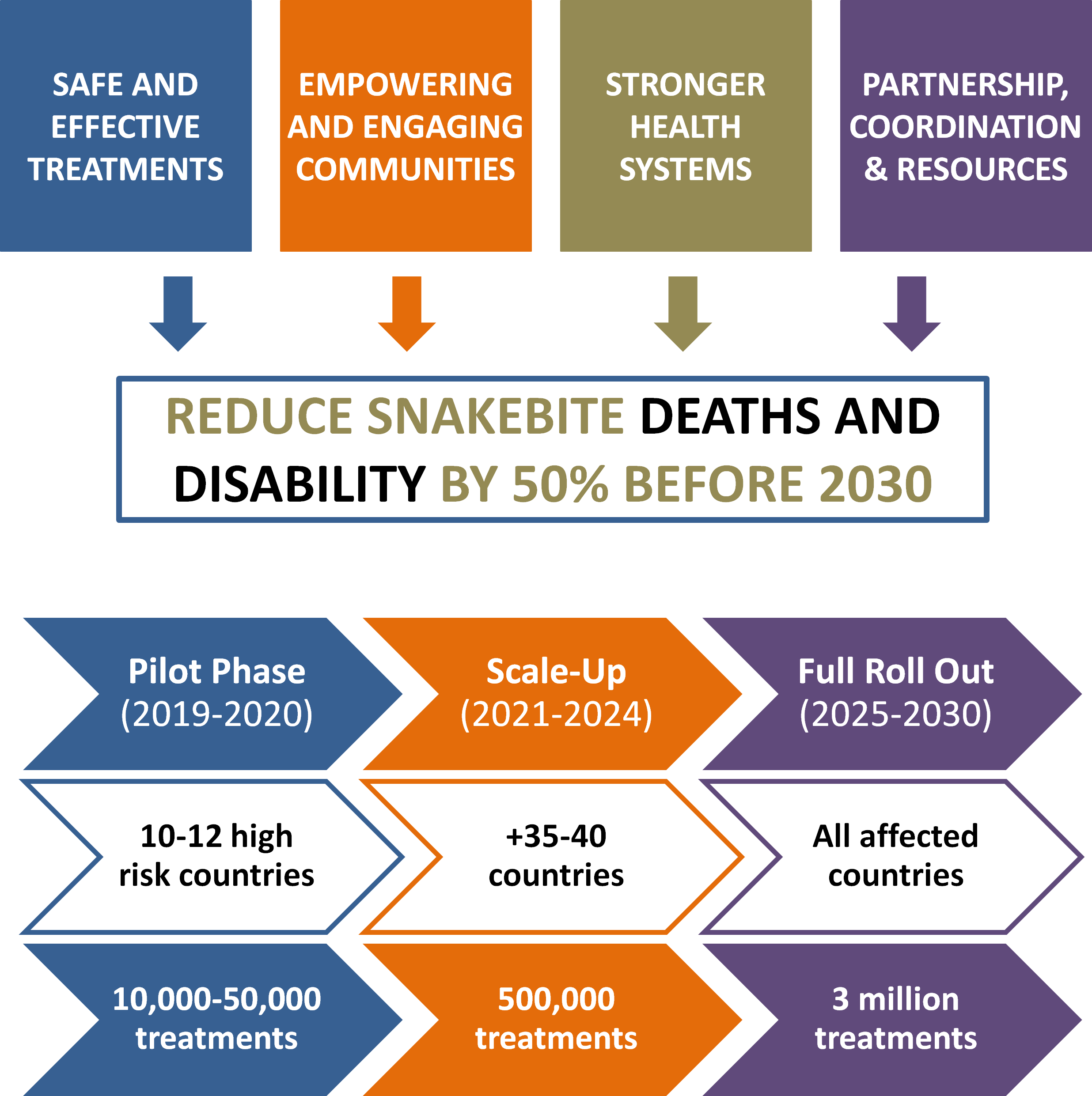
Designing protein binders De novo fits with the objective “Safe and Effective Treatment” from the WHO roadmap. This objective proposes 5 key activities:
- Make safe, effective antivenoms available, and affordable to all
- Better control and regulation of antivenoms
- Prequalification of antivenoms
- Integrated health worker training and education
- Improving clinical decision making, treatment, recovery and rehabilitation
- Investing in innovative research on new therapeutics
The implementation of these protein binders as an alternative to traditionally-produced antivenoms should meet with these 5 key activites. The image below analysis how portein binders could contributo to these key activities and proposes 4 potential governance actions according to the objetive and key activites proposes by the WHO (Figure 4)
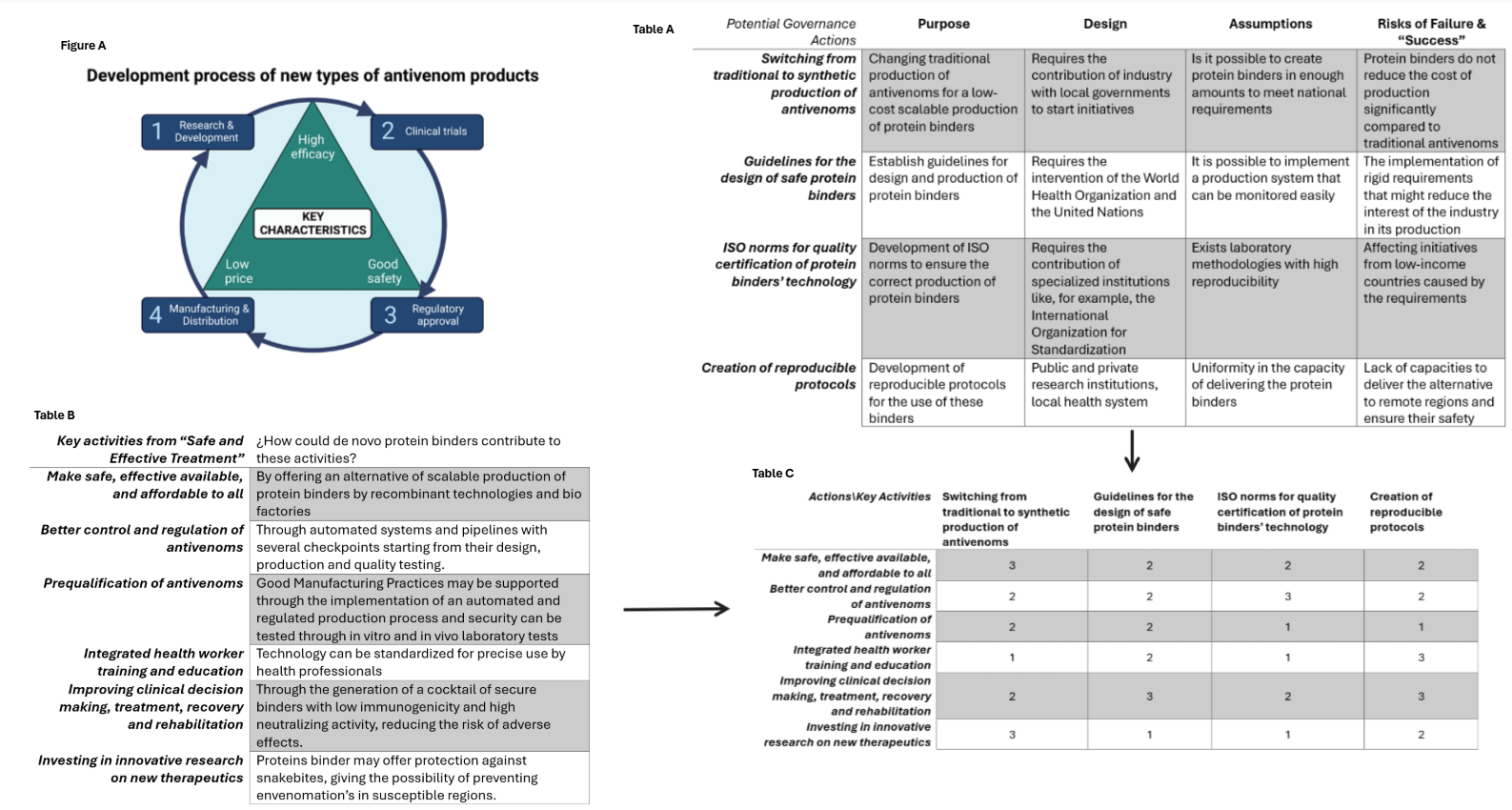
One of the most important governance actions that I would prioritize is the development of reproducible protocols for the design and use of protein binders against snake venoms. Reproducible protocols require the participation of public and private research institutions and involves the development of clear and highly reproducible strategies for de novo prediction of these protein binders, recombinant production and scalation. This action may contribute to other actions like the creation of guidelines promoted by the WHO using these protocols. In Peru, the National Health Institute is in charge of antivenom production, the development of reproducible protocols requires the association of research laboratories with this institute. A pilot program can also be created using different species of the genus Bothrops to design and test the efficacy of protein binders.
References
- Alangode, A., Rajan, K. & Nair, B. G. (2020). Snake antivenom: Challenges and alternate approaches.. Biochemical Pharmacology, 181. https://doi.org/10.1016/J.BCP.2020.114135
- Lomonte, B. and Calvete, J. J. (2017). Strategies in ‘snake venomics’ aiming at an integrative view of compositional, functional, and immunological characteristics of venoms. Journal of Venomous Animals and Toxins including Tropical Diseases, 23(1). https://doi.org/10.1186/S40409-017-0117-8
- Potet, J., Beran, D., Ray, N., Alcoba, G., Habib, A. G., Iliyasu, G., Waldmann, B., Ralph, R., Faiz, M. A., Monteiro, W. M., Sachett, J. d. A. G., Di Fábio, J. L., Cortés, M. d. l. Á., Brown, N. & Williams, D. (2021). Access to antivenoms in the developing world: A multidisciplinary analysis.. Toxicon: X, 12. https://doi.org/10.1016/J.TOXCX.2021.100086
- Ratanabanangkoon, K. (2023). Polyvalent Snake Antivenoms: Production Strategy and Their Therapeutic Benefits. Toxins, 15. https://doi.org/10.3390/TOXINS15090517
- Thumtecho, S., Burlet, N. J., Ljungars, A. & Laustsen, A. H. (2023). Towards better antivenoms: navigating the road to new types of snakebite envenoming therapies. Journal of Venomous Animals and Toxins including Tropical Diseases, 29.
- Torres, S. V., Valle, M. B., Mackessy, S., Menzies, S. K., Casewell, N. R., Ahmadi, S., Muratspahić, E., Sappington, I., Overath, M., Rivera-de-Torre, E., Ledergerber, J., Laustsen, A. H., Boddum, K., Bera, A. K., Kang, A., Brackenbrough, E., Cardoso, I. A., Crittenden, E., Edge, R. & Decarreau, J. (2025). De novo designed proteins neutralize lethal snake venom toxins.. Nature, 639. https://doi.org/10.1038/S41586-024-08393-X
- Williams, D., Faiz, M. A., Abela-Ridder, B., Ainsworth, S., Bulfone, T. C., Nickerson, A., Habib, A. G., Junghanss, T., Wen, F. H., Turner, M. J., Harrison, R. A. & Warrell, D. A. (2019). Strategy for a globally coordinated response to a priority neglected tropical disease: Snakebite envenoming.. PLoS Neglected Tropical Diseases, 13. https://doi.org/10.1371/JOURNAL.PNTD.0007059
- World Health Organisation. (2019, April 8). Snakebite Envenoming. Who.int; World Health Organization: WHO. https://www.who.int/news-room/fact-sheets/detail/snakebite-envenoming
- OpenAI (2026). CHATGPT(GTP-5-based-model). Used for conceptual discussion and feedback on project development. https://chat.openai.com/
Week 2 Lecture Prep
Professor Jacobson Questions:
Nature’s machinery for copying DNA is called polymerase. What is the error rate of polymerase? How does this compare to the length of the human genome. How does biology deal with that discrepancy?
During DNA replication our cells use DNA polymerases for DNA synthesis, these polymerases can have an error rate of 1 bp per every 100,00 nucleotides. As the human genome is composed of 6 billion bp per diploid cell, every time a cell divides DNA polymerases will make about 120,000 errors (Pray, 2008). While these errors may become mutations that could lead to new adaptations, it is important to correct these errors since they could lead to many dangerous effects on the organism’s life. To correct these errors some DNA polymerases come with an extra exonuclease 3’-5’ activity that serves as proofreading. For example, PolƐ is a DNA polymerase that is involved in the process of DNA replication of the leading strand. PolƐ is a holoenzyme compose of many subunits, when a mismatch is detected in the pol site of PolƐ the proteins arrest the pol activity and the protein moves away from the mismatched 3’end preventing additional base incorporation. Then, the proofreading region generates a change in the DNA conformation. This takes the mismatched base to the exo site of the polymerase generating the excision, after that the polymerase resumes its activity after correcting the mistake (Wang et al, 2025). Proofreading mechanisms help to reduce errors induced by the replication process, for that reason, polymerases with proofreading activity are highly important in different applications. To design complex synthetics systems, it is necessary to reduce the possibility of bp mismatches caused by the polymerase, for that reason, high fidelity polymerase with proofreading activity is available commercially
How many different ways are there to code (DNA nucleotide code) for an average human protein? In practice what are some of the reasons that all of these different codes don’t work to code for the protein of interest?
Human genetic code is a set of three RNA bases called codon; every codon decodes a specific amino acid. The genetic code shows codon degeneracy, in other word codons that can be used to decode the same amino acid. In average most amino acids correspond to three codons, with some exceptions like Methionine and Tryptophan that only belong to a single codon. While codon degeneracy allows the use of different codons to produce an amino acid, different organisms have different preferences for the codons they use. Codon preference may occur for different reasons like metabolic pressures where some specific tRNAs are used instead of a wide variety of tRNAs for every codon available. Similarly, protein characteristics may influence the preference for some tRNAs than others (Ford, n.d).
Dr. LeProust Questions:
- What’s the most used method for oligo synthesis currently? The most used method for DNA synthesis is through Phosphoramidite chemistry. This technology consists of the use of Nucleoside Phosphoramidites, a type of modified nucleosides that allows the sequential addition of new bases in a cyclic manner. These modified nucleosides are protected in a way that chemists can control the reaction of oligonucleotide synthesis by exposing only the regions of the nucleotide they desire.
- Why is it difficult to make oligos longer than 200nt via direct synthesis? One of the reasons why synthesis of oligos longer than 200 nt is the increase of errors caused by the natural DNA polymerases error rate or fidelity of nucleoside phosphoramidite thecnology. Another reason could be the limitations of Quality Control, since oligos require MALDI spectrometry to test their quality, this method limits the length to 10-50 nucleotides.
- Why can’t you make a 2000bp gene via direct oligo synthesis? Production of long oligos faces the main challenge of accumulating errors in their formation making it difficult to obtain high yield of oligos with high quality (Yin et al, 2024)
George Church Questions:
What are the 10 essential amino acids in all animals and how does this affect your view of the “Lysine Contingency”?
While many organisms are capable to synthesize all these 20 amino acids, some groups like ours (Metazoa) have lost the capability to synthetize nine EAAs. These amino acids are histidine, isoleucine, leucine, lysine, methionine, phenylalanine, threonine, tryptophan, and valine. Additionally, arginine can also be considered essential because of the incapacity of the body to synthetize it under special periods of growth (Lopez & Mohiuddin, 2024). One explanation for the loss of the synthetic capacity of these essential amino acids is because of energetic efficiency. Estimates suggest that essential amino acids have high energetic costs in their synthesis. Selective pressures towards energy efficiency may contribute to the loss of capacity to produce essential amino acids and relying on them by direct consumption (Kasalo et al., 2026). Considering that in Jurassic Park movies the lysine contingency consists in limiting the expansion of dinosaurs by creating to them the incapacity of lysine production. This contingency now seems futile because animals have more essential amino acids than dinosaurs in that case. Animals have overcome this limitation through the ingest of these amino acids in their diets, so in consequence dinosaurs can also survive by consuming other living things that produce lysine either from animal or plant sources.
References:
- Ford, T. (n.d.). Plasmids 101: Codon usage bias. Available at https://blog.addgene.org/plasmis-101-codon-usage-bias?
- Kasalo, N., Domazet-Lošo, M., & Domazet-Lošo, T. (2026). Outsourcing of energetically costly amino acids at the origin of animals. Nature Communications. https://doi.org/10.1038/s41467-026-68724-6
- Lopez, M. J., & Mohiuddin, S. S. (2024, April 30). Biochemistry, essential amino acids. StatPearls - NCBI Bookshelf. https://www.ncbi.nlm.nih.gov/books/NBK557845/
- Pray, L. (2008) DNA Replication and Causes of Mutation. Nature Education 1(1):214
- Wang, F., He, Q., O’Donnell, M. E., & Li, H. (2025). The proofreading mechanism of the human leading-strand DNA polymerase ε holoenzyme. Proceedings of the National Academy of Sciences, 122(22), e2507232122. https://doi.org/10.1073/pnas.2507232122
- Yin, Y., Arneson, R., Yuan, Y., & Fang, S. (2024). Long oligos: direct chemical synthesis of genes with up to 1728 nucleotides. Chemical Science, 16(4), 1966–1973. https://doi.org/10.1039/d4sc06958g